
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:YME1L1-SLC35D2 (FusionGDB2 ID:100045) |
Fusion Gene Summary for YME1L1-SLC35D2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: YME1L1-SLC35D2 | Fusion gene ID: 100045 | Hgene | Tgene | Gene symbol | YME1L1 | SLC35D2 | Gene ID | 10730 | 11046 |
| Gene name | YME1 like 1 ATPase | solute carrier family 35 member D2 | |
| Synonyms | FTSH|MEG4|OPA11|PAMP|YME1L | HFRC1|SQV7L|UGTrel8|hfrc | |
| Cytomap | 10p12.1 | 9q22.32 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ATP-dependent zinc metalloprotease YME1L1ATP-dependent metalloprotease FtsH1 homologYME1-like protein 1meg-4presenilin-associated metalloprotease | UDP-N-acetylglucosamine/UDP-glucose/GDP-mannose transporterSQV7-like proteinUDP-N-acetylglucosamine transporterUDP-galactose transporter-related protein 8fringe connectionhomolog of Fringe connection protein 1solute carrier family 35 (UDP-GlcNAc/UDP | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000326799, ENST00000375972, ENST00000376016, ENST00000477432, ENST00000463270, | ENST00000482643, ENST00000375257, ENST00000375259, ENST00000253270, | |
| Fusion gene scores | * DoF score | 7 X 6 X 5=210 | 3 X 4 X 3=36 |
| # samples | 7 | 3 | |
| ** MAII score | log2(7/210*10)=-1.58496250072116 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(3/36*10)=-0.263034405833794 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: YME1L1 [Title/Abstract] AND SLC35D2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | YME1L1(27437835)-SLC35D2(99122503), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | YME1L1-SLC35D2 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. YME1L1-SLC35D2 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | YME1L1 | GO:0034214 | protein hexamerization | 27786171 |
 Fusion gene breakpoints across YME1L1 (5'-gene) Fusion gene breakpoints across YME1L1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
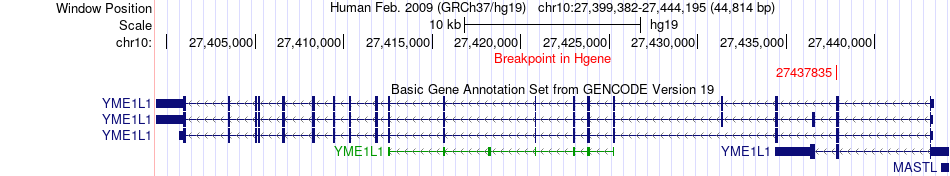 |
 Fusion gene breakpoints across SLC35D2 (3'-gene) Fusion gene breakpoints across SLC35D2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
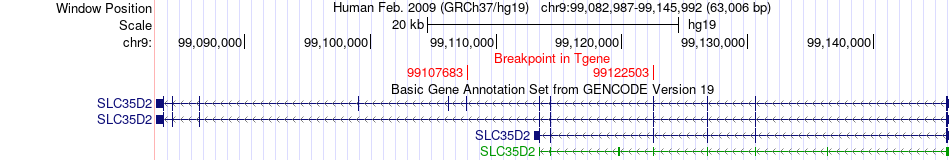 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-E2-A1IH-01A | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| ChimerDB4 | BRCA | TCGA-E2-A1IH-01A | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
Top |
Fusion Gene ORF analysis for YME1L1-SLC35D2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000326799 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| 5CDS-5UTR | ENST00000375972 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| 5CDS-5UTR | ENST00000376016 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| 5CDS-5UTR | ENST00000477432 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| 5CDS-intron | ENST00000326799 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000326799 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000326799 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000375972 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000375972 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000375972 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000376016 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000376016 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000376016 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000477432 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000477432 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| 5CDS-intron | ENST00000477432 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| Frame-shift | ENST00000326799 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| Frame-shift | ENST00000375972 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| Frame-shift | ENST00000376016 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| Frame-shift | ENST00000477432 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| In-frame | ENST00000326799 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000326799 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000326799 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000375972 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000375972 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000375972 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000376016 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000376016 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000376016 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000477432 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000477432 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| In-frame | ENST00000477432 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| intron-3CDS | ENST00000463270 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| intron-3CDS | ENST00000463270 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| intron-3CDS | ENST00000463270 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| intron-3CDS | ENST00000463270 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| intron-5UTR | ENST00000463270 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - |
| intron-intron | ENST00000463270 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| intron-intron | ENST00000463270 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
| intron-intron | ENST00000463270 | ENST00000482643 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99107683 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000326799 | YME1L1 | chr10 | 27437835 | - | ENST00000253270 | SLC35D2 | chr9 | 99122503 | - | 1588 | 317 | 149 | 1051 | 300 |
| ENST00000326799 | YME1L1 | chr10 | 27437835 | - | ENST00000375259 | SLC35D2 | chr9 | 99122503 | - | 1323 | 317 | 149 | 787 | 212 |
| ENST00000326799 | YME1L1 | chr10 | 27437835 | - | ENST00000375257 | SLC35D2 | chr9 | 99122503 | - | 879 | 317 | 149 | 529 | 126 |
| ENST00000376016 | YME1L1 | chr10 | 27437835 | - | ENST00000253270 | SLC35D2 | chr9 | 99122503 | - | 1621 | 350 | 182 | 1084 | 300 |
| ENST00000376016 | YME1L1 | chr10 | 27437835 | - | ENST00000375259 | SLC35D2 | chr9 | 99122503 | - | 1356 | 350 | 182 | 820 | 212 |
| ENST00000376016 | YME1L1 | chr10 | 27437835 | - | ENST00000375257 | SLC35D2 | chr9 | 99122503 | - | 912 | 350 | 182 | 562 | 126 |
| ENST00000375972 | YME1L1 | chr10 | 27437835 | - | ENST00000253270 | SLC35D2 | chr9 | 99122503 | - | 1613 | 342 | 174 | 1076 | 300 |
| ENST00000375972 | YME1L1 | chr10 | 27437835 | - | ENST00000375259 | SLC35D2 | chr9 | 99122503 | - | 1348 | 342 | 174 | 812 | 212 |
| ENST00000375972 | YME1L1 | chr10 | 27437835 | - | ENST00000375257 | SLC35D2 | chr9 | 99122503 | - | 904 | 342 | 174 | 554 | 126 |
| ENST00000477432 | YME1L1 | chr10 | 27437835 | - | ENST00000253270 | SLC35D2 | chr9 | 99122503 | - | 2495 | 1224 | 813 | 1958 | 381 |
| ENST00000477432 | YME1L1 | chr10 | 27437835 | - | ENST00000375259 | SLC35D2 | chr9 | 99122503 | - | 2230 | 1224 | 813 | 1694 | 293 |
| ENST00000477432 | YME1L1 | chr10 | 27437835 | - | ENST00000375257 | SLC35D2 | chr9 | 99122503 | - | 1786 | 1224 | 813 | 1436 | 207 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000326799 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.00593634 | 0.9940637 |
| ENST00000326799 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.03406683 | 0.96593314 |
| ENST00000326799 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.3092277 | 0.69077235 |
| ENST00000376016 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.00516626 | 0.9948337 |
| ENST00000376016 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.02779859 | 0.9722014 |
| ENST00000376016 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.1789389 | 0.8210611 |
| ENST00000375972 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.005258682 | 0.9947413 |
| ENST00000375972 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.028896626 | 0.9711033 |
| ENST00000375972 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.1910921 | 0.8089079 |
| ENST00000477432 | ENST00000253270 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.00898627 | 0.9910137 |
| ENST00000477432 | ENST00000375259 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.06287518 | 0.9371248 |
| ENST00000477432 | ENST00000375257 | YME1L1 | chr10 | 27437835 | - | SLC35D2 | chr9 | 99122503 | - | 0.41519707 | 0.584803 |
Top |
Fusion Genomic Features for YME1L1-SLC35D2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
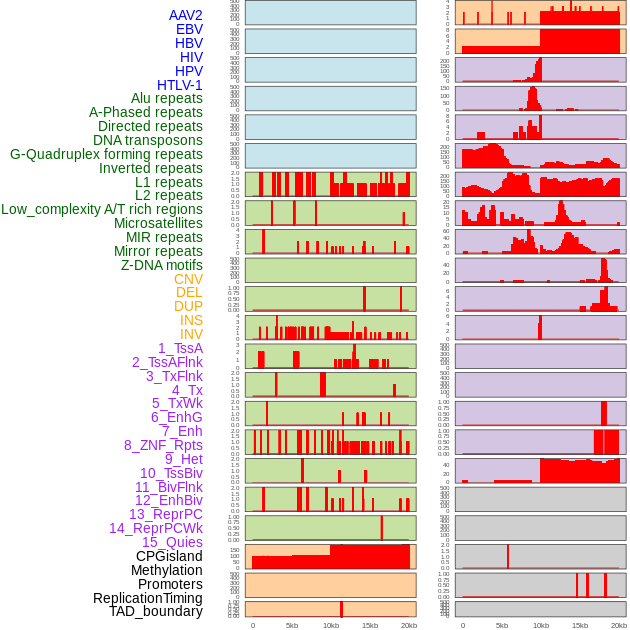 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for YME1L1-SLC35D2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr10:27437835/chr9:99122503) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 189_201 | 93 | 338.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 223_237 | 93 | 338.0 | Topological domain | Extracellular | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 259_265 | 93 | 338.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 289_292 | 93 | 338.0 | Topological domain | Extracellular | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 316_337 | 93 | 338.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 189_201 | 93 | 250.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 223_237 | 93 | 250.0 | Topological domain | Extracellular | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 259_265 | 93 | 250.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 289_292 | 93 | 250.0 | Topological domain | Extracellular | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 316_337 | 93 | 250.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 147_167 | 93 | 338.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 168_188 | 93 | 338.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 202_222 | 93 | 338.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 238_258 | 93 | 338.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 266_288 | 93 | 338.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 293_315 | 93 | 338.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 147_167 | 93 | 250.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 168_188 | 93 | 250.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 202_222 | 93 | 250.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 238_258 | 93 | 250.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 266_288 | 93 | 250.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 293_315 | 93 | 250.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000326799 | - | 2 | 20 | 379_386 | 56 | 774.0 | Nucleotide binding | ATP |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000375972 | - | 2 | 18 | 379_386 | 56 | 684.0 | Nucleotide binding | ATP |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000376016 | - | 2 | 19 | 379_386 | 56 | 717.0 | Nucleotide binding | ATP |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000326799 | - | 2 | 20 | 1_295 | 56 | 774.0 | Topological domain | Mitochondrial matrix |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000326799 | - | 2 | 20 | 317_773 | 56 | 774.0 | Topological domain | Mitochondrial intermembrane |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000375972 | - | 2 | 18 | 1_295 | 56 | 684.0 | Topological domain | Mitochondrial matrix |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000375972 | - | 2 | 18 | 317_773 | 56 | 684.0 | Topological domain | Mitochondrial intermembrane |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000376016 | - | 2 | 19 | 1_295 | 56 | 717.0 | Topological domain | Mitochondrial matrix |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000376016 | - | 2 | 19 | 317_773 | 56 | 717.0 | Topological domain | Mitochondrial intermembrane |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000326799 | - | 2 | 20 | 296_316 | 56 | 774.0 | Transmembrane | Helical |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000375972 | - | 2 | 18 | 296_316 | 56 | 684.0 | Transmembrane | Helical |
| Hgene | YME1L1 | chr10:27437835 | chr9:99122503 | ENST00000376016 | - | 2 | 19 | 296_316 | 56 | 717.0 | Transmembrane | Helical |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 1_27 | 93 | 338.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 49_53 | 93 | 338.0 | Topological domain | Extracellular | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 75_146 | 93 | 338.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 1_27 | 93 | 250.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 49_53 | 93 | 250.0 | Topological domain | Extracellular | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 75_146 | 93 | 250.0 | Topological domain | Cytoplasmic | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 28_48 | 93 | 338.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000253270 | 2 | 12 | 54_74 | 93 | 338.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 28_48 | 93 | 250.0 | Transmembrane | Helical | |
| Tgene | SLC35D2 | chr10:27437835 | chr9:99122503 | ENST00000375259 | 2 | 9 | 54_74 | 93 | 250.0 | Transmembrane | Helical |
Top |
Fusion Gene Sequence for YME1L1-SLC35D2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >100045_100045_1_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000326799_SLC35D2_chr9_99122503_ENST00000253270_length(transcript)=1588nt_BP=317nt AGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAGGAAAAAAGAAGGGCTCAGCGCCTCCCCGCCGGG CCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGGAGATGTTTTCCTTGTCGAGCACGGTGCAACCCC AGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTTCTGTTTCTCTCAGTGGAGTGTCAGTTTCTCAAA ACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTCTGCCTCTCCTCTACGTTGGAAACCACATAAGTG GATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCACCATTCCACTTACCTTACTTCTGGAAACCATCA TACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCGGGGCTTTCATAGCAGCTGGGTCTGACCTTGCTT TTAACTTAGAAGGCTATATTTTTGTATTCCTGAATGATATCTTCACAGCAGCAAATGGAGTTTATACCAAACAGAAAATGGACCCAAAGG AGCTAGGGAAATACGGAGTACTTTTCTACAATGCCTGCTTCATGATTATCCCAACTCTTATTATTAGTGTCTCCACTGGAGACCTGCAAC AGGCTACTGAATTCAACCAATGGAAGAATGTTGTGTTTATCCTACAGTTTCTTCTTTCCTGTTTTTTGGGGTTTCTGCTGATGTACTCCA CGGTTCTGTGCAGCTATTACAATTCAGCCCTGACGACAGCAGTGGTTGGAGCCATCAAGAATGTATCCGTTGCCTACATTGGGATATTAA TCGGTGGAGACTACATTTTCTCTTTGTTAAACTTTGTAGGGTTAAATATTTGCATGGCAGGGGGCTTGAGATATTCCTTTTTAACACTGA GCAGCCAGTTAAAACCTAAACCTGTGGGTGAAGAAAACATCTGTTTGGATTTGAAGAGCTAAAGAGTCTGCAGCAGGATTGGAGACTGAC TTGTGACTGCGGGCTGGGGGGGCATTCCCAGTAGGAATGTGAAGCCAGAGGTTTCGGATTCGTGACATCCACCCCCTGGGCAAGTGAGAG CATCTGCAAAATGCAAAGAGAACTACCTCATATGCAGGATGAGCCAATGGCAGTCTCAAGAAATGTACTCGGGCGACACCTTACCTGTGG AAAGCAAATCTTTTCAAAATAAGCCACTGGGACTCGGTAGGTGGAGCCCCAGCTGCTCTTCTAGGGACCTATGGGGCCTTCGTGGCATCT CTGTGCTGTGTGCTGGGGAGGAGGTTGATGTAATGGTGACTCTTTTCTGATCAGCACCTTGGCCGTGATTCCCAAGGTCCCAGCCAAAGC AAAGGGCCAGTTGTTTCAGTTTAAACAGACATGTCTTTAGTCTAATAAAATTAGTTAACTGCCAGTAAAGTTATTTGTTAGCTTTGATGA >100045_100045_1_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000326799_SLC35D2_chr9_99122503_ENST00000253270_length(amino acids)=300AA_BP=56 MFSLSSTVQPQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFT IPLTLLLETIILGKQYSLNIILSVFAIILGAFIAAGSDLAFNLEGYIFVFLNDIFTAANGVYTKQKMDPKELGKYGVLFYNACFMIIPTL IISVSTGDLQQATEFNQWKNVVFILQFLLSCFLGFLLMYSTVLCSYYNSALTTAVVGAIKNVSVAYIGILIGGDYIFSLLNFVGLNICMA -------------------------------------------------------------- >100045_100045_2_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000326799_SLC35D2_chr9_99122503_ENST00000375257_length(transcript)=879nt_BP=317nt AGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAGGAAAAAAGAAGGGCTCAGCGCCTCCCCGCCGGG CCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGGAGATGTTTTCCTTGTCGAGCACGGTGCAACCCC AGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTTCTGTTTCTCTCAGTGGAGTGTCAGTTTCTCAAA ACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTCTGCCTCTCCTCTACGTTGGAAACCACATAAGTG GATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCACCATTCCACTTACCTTACTTCTGGAAACCATCA TACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCGGGGCTTTCATAGCAGCTGGGTAAGACTTTGGAC TGTTTCTGCTCAATAACTTCTTTGTGTCTTTGACAATAGGCCCAGCCCCATCCTCAGTTTTTCATTTTCCATCTGGCTGCTGTTGAATTG AGCAGGCAAGAGACAAATCTGTTGAAGTTAGTGGAATCCCCATGTGTTCCTATGAAAGTCAAAGTGGGTCCCACATTGGCCAGAAAATTT CTGAGAATATATGAAGAGAGCATTGCAAGATATTAATATAGCTTTTCTTCACAAAAAGCAGTGAACTGTTTACATTTCTTACTGAATGTT >100045_100045_2_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000326799_SLC35D2_chr9_99122503_ENST00000375257_length(amino acids)=126AA_BP=56 MFSLSSTVQPQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFT -------------------------------------------------------------- >100045_100045_3_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000326799_SLC35D2_chr9_99122503_ENST00000375259_length(transcript)=1323nt_BP=317nt AGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAGGAAAAAAGAAGGGCTCAGCGCCTCCCCGCCGGG CCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGGAGATGTTTTCCTTGTCGAGCACGGTGCAACCCC AGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTTCTGTTTCTCTCAGTGGAGTGTCAGTTTCTCAAA ACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTCTGCCTCTCCTCTACGTTGGAAACCACATAAGTG GATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCACCATTCCACTTACCTTACTTCTGGAAACCATCA TACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCGGGGCTTTCATAGCAGCTGGGTTTCTGCTGATGT ACTCCACGGTTCTGTGCAGCTATTACAATTCAGCCCTGACGACAGCAGTGGTTGGAGCCATCAAGAATGTATCCGTTGCCTACATTGGGA TATTAATCGGTGGAGACTACATTTTCTCTTTGTTAAACTTTGTAGGGTTAAATATTTGCATGGCAGGGGGCTTGAGATATTCCTTTTTAA CACTGAGCAGCCAGTTAAAACCTAAACCTGTGGGTGAAGAAAACATCTGTTTGGATTTGAAGAGCTAAAGAGTCTGCAGCAGGATTGGAG ACTGACTTGTGACTGCGGGCTGGGGGGGCATTCCCAGTAGGAATGTGAAGCCAGAGGTTTCGGATTCGTGACATCCACCCCCTGGGCAAG TGAGAGCATCTGCAAAATGCAAAGAGAACTACCTCATATGCAGGATGAGCCAATGGCAGTCTCAAGAAATGTACTCGGGCGACACCTTAC CTGTGGAAAGCAAATCTTTTCAAAATAAGCCACTGGGACTCGGTAGGTGGAGCCCCAGCTGCTCTTCTAGGGACCTATGGGGCCTTCGTG GCATCTCTGTGCTGTGTGCTGGGGAGGAGGTTGATGTAATGGTGACTCTTTTCTGATCAGCACCTTGGCCGTGATTCCCAAGGTCCCAGC CAAAGCAAAGGGCCAGTTGTTTCAGTTTAAACAGACATGTCTTTAGTCTAATAAAATTAGTTAACTGCCAGTAAAGTTATTTGTTAGCTT >100045_100045_3_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000326799_SLC35D2_chr9_99122503_ENST00000375259_length(amino acids)=212AA_BP=56 MFSLSSTVQPQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFT IPLTLLLETIILGKQYSLNIILSVFAIILGAFIAAGFLLMYSTVLCSYYNSALTTAVVGAIKNVSVAYIGILIGGDYIFSLLNFVGLNIC -------------------------------------------------------------- >100045_100045_4_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000375972_SLC35D2_chr9_99122503_ENST00000253270_length(transcript)=1613nt_BP=342nt GTCTGTCGTTGCAGTAGCTGTAGGAAGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAGGAAAAAAG AAGGGCTCAGCGCCTCCCCGCCGGGCCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGGAGATGTTT TCCTTGTCGAGCACGGTGCAACCCCAGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTTCTGTTTCT CTCAGTGGAGTGTCAGTTTCTCAAAACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTCTGCCTCTC CTCTACGTTGGAAACCACATAAGTGGATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCACCATTCCA CTTACCTTACTTCTGGAAACCATCATACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCGGGGCTTTC ATAGCAGCTGGGTCTGACCTTGCTTTTAACTTAGAAGGCTATATTTTTGTATTCCTGAATGATATCTTCACAGCAGCAAATGGAGTTTAT ACCAAACAGAAAATGGACCCAAAGGAGCTAGGGAAATACGGAGTACTTTTCTACAATGCCTGCTTCATGATTATCCCAACTCTTATTATT AGTGTCTCCACTGGAGACCTGCAACAGGCTACTGAATTCAACCAATGGAAGAATGTTGTGTTTATCCTACAGTTTCTTCTTTCCTGTTTT TTGGGGTTTCTGCTGATGTACTCCACGGTTCTGTGCAGCTATTACAATTCAGCCCTGACGACAGCAGTGGTTGGAGCCATCAAGAATGTA TCCGTTGCCTACATTGGGATATTAATCGGTGGAGACTACATTTTCTCTTTGTTAAACTTTGTAGGGTTAAATATTTGCATGGCAGGGGGC TTGAGATATTCCTTTTTAACACTGAGCAGCCAGTTAAAACCTAAACCTGTGGGTGAAGAAAACATCTGTTTGGATTTGAAGAGCTAAAGA GTCTGCAGCAGGATTGGAGACTGACTTGTGACTGCGGGCTGGGGGGGCATTCCCAGTAGGAATGTGAAGCCAGAGGTTTCGGATTCGTGA CATCCACCCCCTGGGCAAGTGAGAGCATCTGCAAAATGCAAAGAGAACTACCTCATATGCAGGATGAGCCAATGGCAGTCTCAAGAAATG TACTCGGGCGACACCTTACCTGTGGAAAGCAAATCTTTTCAAAATAAGCCACTGGGACTCGGTAGGTGGAGCCCCAGCTGCTCTTCTAGG GACCTATGGGGCCTTCGTGGCATCTCTGTGCTGTGTGCTGGGGAGGAGGTTGATGTAATGGTGACTCTTTTCTGATCAGCACCTTGGCCG TGATTCCCAAGGTCCCAGCCAAAGCAAAGGGCCAGTTGTTTCAGTTTAAACAGACATGTCTTTAGTCTAATAAAATTAGTTAACTGCCAG >100045_100045_4_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000375972_SLC35D2_chr9_99122503_ENST00000253270_length(amino acids)=300AA_BP=56 MFSLSSTVQPQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFT IPLTLLLETIILGKQYSLNIILSVFAIILGAFIAAGSDLAFNLEGYIFVFLNDIFTAANGVYTKQKMDPKELGKYGVLFYNACFMIIPTL IISVSTGDLQQATEFNQWKNVVFILQFLLSCFLGFLLMYSTVLCSYYNSALTTAVVGAIKNVSVAYIGILIGGDYIFSLLNFVGLNICMA -------------------------------------------------------------- >100045_100045_5_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000375972_SLC35D2_chr9_99122503_ENST00000375257_length(transcript)=904nt_BP=342nt GTCTGTCGTTGCAGTAGCTGTAGGAAGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAGGAAAAAAG AAGGGCTCAGCGCCTCCCCGCCGGGCCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGGAGATGTTT TCCTTGTCGAGCACGGTGCAACCCCAGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTTCTGTTTCT CTCAGTGGAGTGTCAGTTTCTCAAAACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTCTGCCTCTC CTCTACGTTGGAAACCACATAAGTGGATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCACCATTCCA CTTACCTTACTTCTGGAAACCATCATACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCGGGGCTTTC ATAGCAGCTGGGTAAGACTTTGGACTGTTTCTGCTCAATAACTTCTTTGTGTCTTTGACAATAGGCCCAGCCCCATCCTCAGTTTTTCAT TTTCCATCTGGCTGCTGTTGAATTGAGCAGGCAAGAGACAAATCTGTTGAAGTTAGTGGAATCCCCATGTGTTCCTATGAAAGTCAAAGT GGGTCCCACATTGGCCAGAAAATTTCTGAGAATATATGAAGAGAGCATTGCAAGATATTAATATAGCTTTTCTTCACAAAAAGCAGTGAA CTGTTTACATTTCTTACTGAATGTTTGTCTTCATTATAACAACTTTACTTGAGAAGTTCTTAGGCATTTGATTAATAATAACAGACTTAC >100045_100045_5_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000375972_SLC35D2_chr9_99122503_ENST00000375257_length(amino acids)=126AA_BP=56 MFSLSSTVQPQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFT -------------------------------------------------------------- >100045_100045_6_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000375972_SLC35D2_chr9_99122503_ENST00000375259_length(transcript)=1348nt_BP=342nt GTCTGTCGTTGCAGTAGCTGTAGGAAGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAGGAAAAAAG AAGGGCTCAGCGCCTCCCCGCCGGGCCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGGAGATGTTT TCCTTGTCGAGCACGGTGCAACCCCAGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTTCTGTTTCT CTCAGTGGAGTGTCAGTTTCTCAAAACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTCTGCCTCTC CTCTACGTTGGAAACCACATAAGTGGATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCACCATTCCA CTTACCTTACTTCTGGAAACCATCATACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCGGGGCTTTC ATAGCAGCTGGGTTTCTGCTGATGTACTCCACGGTTCTGTGCAGCTATTACAATTCAGCCCTGACGACAGCAGTGGTTGGAGCCATCAAG AATGTATCCGTTGCCTACATTGGGATATTAATCGGTGGAGACTACATTTTCTCTTTGTTAAACTTTGTAGGGTTAAATATTTGCATGGCA GGGGGCTTGAGATATTCCTTTTTAACACTGAGCAGCCAGTTAAAACCTAAACCTGTGGGTGAAGAAAACATCTGTTTGGATTTGAAGAGC TAAAGAGTCTGCAGCAGGATTGGAGACTGACTTGTGACTGCGGGCTGGGGGGGCATTCCCAGTAGGAATGTGAAGCCAGAGGTTTCGGAT TCGTGACATCCACCCCCTGGGCAAGTGAGAGCATCTGCAAAATGCAAAGAGAACTACCTCATATGCAGGATGAGCCAATGGCAGTCTCAA GAAATGTACTCGGGCGACACCTTACCTGTGGAAAGCAAATCTTTTCAAAATAAGCCACTGGGACTCGGTAGGTGGAGCCCCAGCTGCTCT TCTAGGGACCTATGGGGCCTTCGTGGCATCTCTGTGCTGTGTGCTGGGGAGGAGGTTGATGTAATGGTGACTCTTTTCTGATCAGCACCT TGGCCGTGATTCCCAAGGTCCCAGCCAAAGCAAAGGGCCAGTTGTTTCAGTTTAAACAGACATGTCTTTAGTCTAATAAAATTAGTTAAC >100045_100045_6_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000375972_SLC35D2_chr9_99122503_ENST00000375259_length(amino acids)=212AA_BP=56 MFSLSSTVQPQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFT IPLTLLLETIILGKQYSLNIILSVFAIILGAFIAAGFLLMYSTVLCSYYNSALTTAVVGAIKNVSVAYIGILIGGDYIFSLLNFVGLNIC -------------------------------------------------------------- >100045_100045_7_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000376016_SLC35D2_chr9_99122503_ENST00000253270_length(transcript)=1621nt_BP=350nt GGCGGATTGTCTGTCGTTGCAGTAGCTGTAGGAAGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAG GAAAAAAGAAGGGCTCAGCGCCTCCCCGCCGGGCCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGG AGATGTTTTCCTTGTCGAGCACGGTGCAACCCCAGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTT CTGTTTCTCTCAGTGGAGTGTCAGTTTCTCAAAACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTC TGCCTCTCCTCTACGTTGGAAACCACATAAGTGGATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCA CCATTCCACTTACCTTACTTCTGGAAACCATCATACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCG GGGCTTTCATAGCAGCTGGGTCTGACCTTGCTTTTAACTTAGAAGGCTATATTTTTGTATTCCTGAATGATATCTTCACAGCAGCAAATG GAGTTTATACCAAACAGAAAATGGACCCAAAGGAGCTAGGGAAATACGGAGTACTTTTCTACAATGCCTGCTTCATGATTATCCCAACTC TTATTATTAGTGTCTCCACTGGAGACCTGCAACAGGCTACTGAATTCAACCAATGGAAGAATGTTGTGTTTATCCTACAGTTTCTTCTTT CCTGTTTTTTGGGGTTTCTGCTGATGTACTCCACGGTTCTGTGCAGCTATTACAATTCAGCCCTGACGACAGCAGTGGTTGGAGCCATCA AGAATGTATCCGTTGCCTACATTGGGATATTAATCGGTGGAGACTACATTTTCTCTTTGTTAAACTTTGTAGGGTTAAATATTTGCATGG CAGGGGGCTTGAGATATTCCTTTTTAACACTGAGCAGCCAGTTAAAACCTAAACCTGTGGGTGAAGAAAACATCTGTTTGGATTTGAAGA GCTAAAGAGTCTGCAGCAGGATTGGAGACTGACTTGTGACTGCGGGCTGGGGGGGCATTCCCAGTAGGAATGTGAAGCCAGAGGTTTCGG ATTCGTGACATCCACCCCCTGGGCAAGTGAGAGCATCTGCAAAATGCAAAGAGAACTACCTCATATGCAGGATGAGCCAATGGCAGTCTC AAGAAATGTACTCGGGCGACACCTTACCTGTGGAAAGCAAATCTTTTCAAAATAAGCCACTGGGACTCGGTAGGTGGAGCCCCAGCTGCT CTTCTAGGGACCTATGGGGCCTTCGTGGCATCTCTGTGCTGTGTGCTGGGGAGGAGGTTGATGTAATGGTGACTCTTTTCTGATCAGCAC CTTGGCCGTGATTCCCAAGGTCCCAGCCAAAGCAAAGGGCCAGTTGTTTCAGTTTAAACAGACATGTCTTTAGTCTAATAAAATTAGTTA ACTGCCAGTAAAGTTATTTGTTAGCTTTGATGAAAGCTATGTTGGTATCTTTCCCTAATCATCAAAGTAAATAAAAAATCATTTCTATGT >100045_100045_7_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000376016_SLC35D2_chr9_99122503_ENST00000253270_length(amino acids)=300AA_BP=56 MFSLSSTVQPQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFT IPLTLLLETIILGKQYSLNIILSVFAIILGAFIAAGSDLAFNLEGYIFVFLNDIFTAANGVYTKQKMDPKELGKYGVLFYNACFMIIPTL IISVSTGDLQQATEFNQWKNVVFILQFLLSCFLGFLLMYSTVLCSYYNSALTTAVVGAIKNVSVAYIGILIGGDYIFSLLNFVGLNICMA -------------------------------------------------------------- >100045_100045_8_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000376016_SLC35D2_chr9_99122503_ENST00000375257_length(transcript)=912nt_BP=350nt GGCGGATTGTCTGTCGTTGCAGTAGCTGTAGGAAGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAG GAAAAAAGAAGGGCTCAGCGCCTCCCCGCCGGGCCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGG AGATGTTTTCCTTGTCGAGCACGGTGCAACCCCAGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTT CTGTTTCTCTCAGTGGAGTGTCAGTTTCTCAAAACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTC TGCCTCTCCTCTACGTTGGAAACCACATAAGTGGATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCA CCATTCCACTTACCTTACTTCTGGAAACCATCATACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCG GGGCTTTCATAGCAGCTGGGTAAGACTTTGGACTGTTTCTGCTCAATAACTTCTTTGTGTCTTTGACAATAGGCCCAGCCCCATCCTCAG TTTTTCATTTTCCATCTGGCTGCTGTTGAATTGAGCAGGCAAGAGACAAATCTGTTGAAGTTAGTGGAATCCCCATGTGTTCCTATGAAA GTCAAAGTGGGTCCCACATTGGCCAGAAAATTTCTGAGAATATATGAAGAGAGCATTGCAAGATATTAATATAGCTTTTCTTCACAAAAA GCAGTGAACTGTTTACATTTCTTACTGAATGTTTGTCTTCATTATAACAACTTTACTTGAGAAGTTCTTAGGCATTTGATTAATAATAAC >100045_100045_8_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000376016_SLC35D2_chr9_99122503_ENST00000375257_length(amino acids)=126AA_BP=56 MFSLSSTVQPQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFT -------------------------------------------------------------- >100045_100045_9_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000376016_SLC35D2_chr9_99122503_ENST00000375259_length(transcript)=1356nt_BP=350nt GGCGGATTGTCTGTCGTTGCAGTAGCTGTAGGAAGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAG GAAAAAAGAAGGGCTCAGCGCCTCCCCGCCGGGCCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGG AGATGTTTTCCTTGTCGAGCACGGTGCAACCCCAGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTT CTGTTTCTCTCAGTGGAGTGTCAGTTTCTCAAAACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTC TGCCTCTCCTCTACGTTGGAAACCACATAAGTGGATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCA CCATTCCACTTACCTTACTTCTGGAAACCATCATACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCG GGGCTTTCATAGCAGCTGGGTTTCTGCTGATGTACTCCACGGTTCTGTGCAGCTATTACAATTCAGCCCTGACGACAGCAGTGGTTGGAG CCATCAAGAATGTATCCGTTGCCTACATTGGGATATTAATCGGTGGAGACTACATTTTCTCTTTGTTAAACTTTGTAGGGTTAAATATTT GCATGGCAGGGGGCTTGAGATATTCCTTTTTAACACTGAGCAGCCAGTTAAAACCTAAACCTGTGGGTGAAGAAAACATCTGTTTGGATT TGAAGAGCTAAAGAGTCTGCAGCAGGATTGGAGACTGACTTGTGACTGCGGGCTGGGGGGGCATTCCCAGTAGGAATGTGAAGCCAGAGG TTTCGGATTCGTGACATCCACCCCCTGGGCAAGTGAGAGCATCTGCAAAATGCAAAGAGAACTACCTCATATGCAGGATGAGCCAATGGC AGTCTCAAGAAATGTACTCGGGCGACACCTTACCTGTGGAAAGCAAATCTTTTCAAAATAAGCCACTGGGACTCGGTAGGTGGAGCCCCA GCTGCTCTTCTAGGGACCTATGGGGCCTTCGTGGCATCTCTGTGCTGTGTGCTGGGGAGGAGGTTGATGTAATGGTGACTCTTTTCTGAT CAGCACCTTGGCCGTGATTCCCAAGGTCCCAGCCAAAGCAAAGGGCCAGTTGTTTCAGTTTAAACAGACATGTCTTTAGTCTAATAAAAT TAGTTAACTGCCAGTAAAGTTATTTGTTAGCTTTGATGAAAGCTATGTTGGTATCTTTCCCTAATCATCAAAGTAAATAAAAAATCATTT >100045_100045_9_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000376016_SLC35D2_chr9_99122503_ENST00000375259_length(amino acids)=212AA_BP=56 MFSLSSTVQPQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFT IPLTLLLETIILGKQYSLNIILSVFAIILGAFIAAGFLLMYSTVLCSYYNSALTTAVVGAIKNVSVAYIGILIGGDYIFSLLNFVGLNIC -------------------------------------------------------------- >100045_100045_10_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000477432_SLC35D2_chr9_99122503_ENST00000253270_length(transcript)=2495nt_BP=1224nt GGAGGGGGCGTCGGGAAAGCCCCGACTTCGCAGCCTTACACTCTTCGTGGGCGGCGACCGCGGCCCCACTGACATCATTCCTCATGAGGG AGGAGGCACAAACAGTTCTGGGCCGACCAGAAAAAGGACGACTGGGACTTGACTCTGAATCGCAGGATTTGAAGAGATTTCTCCTGGCTT CCCAACGAGGCTGGTGGGAAGCGGTCCTCCTCCCATACACGACCTCCCACCCTCGCGAGGCGTAGAAACCAGTTCTGACTGTACAGTAAA GCGAGGGCCAGGGCTGAGGTCTGGAAGCTAATGAAAGCACAGAAAGTGTCGAAACTGGATGAGCAGGAAGCGAGTGGCCTCCCCTGTCAT CTGACGTTTTCCCAGGATGTAATTTGCCTGACTGAAACAGATCAGGACCAACAGGGAGAGTTTTCGATTTAGTGTGAGGAAAAGAGCACT AAATTGTAGCAAAAGACCTTATTGCTCAAGGCCCAGTCAGAAGATTTCATAAGGAAGCTGTAGAAAGTCTTAAGAGGAAATCAGCCGGGC GTGGTGGCGGGCACCTGTAATCCCAGCTACTCGGGAGGCTGAGGCAGGAGAATCGCTTGAACCCGGGGAGCAGAGGTTGCAGTGAGCCGA GATCGCGCCACTGTACGCCAGCCTGGGCGAAAGAACGAAACTCCGTCTCCAAAAAAAAAAAAAACGAAGAAAAAGTCCAAAGAGGGTAAA GGCTGTTCTCCCCTTAAAAAACAGCTAAAGACCTTTGGGGGCGCTGCTCCTTGTAAATGTCAACTACTTCCGCGGGAAAGAACGCGCAGG CACTTGGCCTTGTGGGCGCTCACTTGCCCCGGAAGTACTGTTGAGTTAGCGCCTCGCCTTCCGGGGCGGATTGTCTGTCGTTGCAGTAGC TGTAGGAAGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAGGAAAAAAGAAGGGCTCAGCGCCTCCC CGCCGGGCCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGGAGATGTTTTCCTTGTCGAGCACGGTG CAACCCCAGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTTCTGTTTCTCTCAGTGGAGTGTCAGTT TCTCAAAACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTCTGCCTCTCCTCTACGTTGGAAACCAC ATAAGTGGATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCACCATTCCACTTACCTTACTTCTGGAA ACCATCATACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCGGGGCTTTCATAGCAGCTGGGTCTGAC CTTGCTTTTAACTTAGAAGGCTATATTTTTGTATTCCTGAATGATATCTTCACAGCAGCAAATGGAGTTTATACCAAACAGAAAATGGAC CCAAAGGAGCTAGGGAAATACGGAGTACTTTTCTACAATGCCTGCTTCATGATTATCCCAACTCTTATTATTAGTGTCTCCACTGGAGAC CTGCAACAGGCTACTGAATTCAACCAATGGAAGAATGTTGTGTTTATCCTACAGTTTCTTCTTTCCTGTTTTTTGGGGTTTCTGCTGATG TACTCCACGGTTCTGTGCAGCTATTACAATTCAGCCCTGACGACAGCAGTGGTTGGAGCCATCAAGAATGTATCCGTTGCCTACATTGGG ATATTAATCGGTGGAGACTACATTTTCTCTTTGTTAAACTTTGTAGGGTTAAATATTTGCATGGCAGGGGGCTTGAGATATTCCTTTTTA ACACTGAGCAGCCAGTTAAAACCTAAACCTGTGGGTGAAGAAAACATCTGTTTGGATTTGAAGAGCTAAAGAGTCTGCAGCAGGATTGGA GACTGACTTGTGACTGCGGGCTGGGGGGGCATTCCCAGTAGGAATGTGAAGCCAGAGGTTTCGGATTCGTGACATCCACCCCCTGGGCAA GTGAGAGCATCTGCAAAATGCAAAGAGAACTACCTCATATGCAGGATGAGCCAATGGCAGTCTCAAGAAATGTACTCGGGCGACACCTTA CCTGTGGAAAGCAAATCTTTTCAAAATAAGCCACTGGGACTCGGTAGGTGGAGCCCCAGCTGCTCTTCTAGGGACCTATGGGGCCTTCGT GGCATCTCTGTGCTGTGTGCTGGGGAGGAGGTTGATGTAATGGTGACTCTTTTCTGATCAGCACCTTGGCCGTGATTCCCAAGGTCCCAG CCAAAGCAAAGGGCCAGTTGTTTCAGTTTAAACAGACATGTCTTTAGTCTAATAAAATTAGTTAACTGCCAGTAAAGTTATTTGTTAGCT >100045_100045_10_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000477432_SLC35D2_chr9_99122503_ENST00000253270_length(amino acids)=381AA_BP=137 MALWALTCPGSTVELAPRLPGRIVCRCSSCRKGRPFSVSGRSEGQRVGEKGKKKGSAPPRRAVDRGAQFRQAGEVAEGPPEMFSLSSTVQ PQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFTIPLTLLLET IILGKQYSLNIILSVFAIILGAFIAAGSDLAFNLEGYIFVFLNDIFTAANGVYTKQKMDPKELGKYGVLFYNACFMIIPTLIISVSTGDL QQATEFNQWKNVVFILQFLLSCFLGFLLMYSTVLCSYYNSALTTAVVGAIKNVSVAYIGILIGGDYIFSLLNFVGLNICMAGGLRYSFLT -------------------------------------------------------------- >100045_100045_11_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000477432_SLC35D2_chr9_99122503_ENST00000375257_length(transcript)=1786nt_BP=1224nt GGAGGGGGCGTCGGGAAAGCCCCGACTTCGCAGCCTTACACTCTTCGTGGGCGGCGACCGCGGCCCCACTGACATCATTCCTCATGAGGG AGGAGGCACAAACAGTTCTGGGCCGACCAGAAAAAGGACGACTGGGACTTGACTCTGAATCGCAGGATTTGAAGAGATTTCTCCTGGCTT CCCAACGAGGCTGGTGGGAAGCGGTCCTCCTCCCATACACGACCTCCCACCCTCGCGAGGCGTAGAAACCAGTTCTGACTGTACAGTAAA GCGAGGGCCAGGGCTGAGGTCTGGAAGCTAATGAAAGCACAGAAAGTGTCGAAACTGGATGAGCAGGAAGCGAGTGGCCTCCCCTGTCAT CTGACGTTTTCCCAGGATGTAATTTGCCTGACTGAAACAGATCAGGACCAACAGGGAGAGTTTTCGATTTAGTGTGAGGAAAAGAGCACT AAATTGTAGCAAAAGACCTTATTGCTCAAGGCCCAGTCAGAAGATTTCATAAGGAAGCTGTAGAAAGTCTTAAGAGGAAATCAGCCGGGC GTGGTGGCGGGCACCTGTAATCCCAGCTACTCGGGAGGCTGAGGCAGGAGAATCGCTTGAACCCGGGGAGCAGAGGTTGCAGTGAGCCGA GATCGCGCCACTGTACGCCAGCCTGGGCGAAAGAACGAAACTCCGTCTCCAAAAAAAAAAAAAACGAAGAAAAAGTCCAAAGAGGGTAAA GGCTGTTCTCCCCTTAAAAAACAGCTAAAGACCTTTGGGGGCGCTGCTCCTTGTAAATGTCAACTACTTCCGCGGGAAAGAACGCGCAGG CACTTGGCCTTGTGGGCGCTCACTTGCCCCGGAAGTACTGTTGAGTTAGCGCCTCGCCTTCCGGGGCGGATTGTCTGTCGTTGCAGTAGC TGTAGGAAGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAGGAAAAAAGAAGGGCTCAGCGCCTCCC CGCCGGGCCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGGAGATGTTTTCCTTGTCGAGCACGGTG CAACCCCAGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTTCTGTTTCTCTCAGTGGAGTGTCAGTT TCTCAAAACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTCTGCCTCTCCTCTACGTTGGAAACCAC ATAAGTGGATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCACCATTCCACTTACCTTACTTCTGGAA ACCATCATACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCGGGGCTTTCATAGCAGCTGGGTAAGAC TTTGGACTGTTTCTGCTCAATAACTTCTTTGTGTCTTTGACAATAGGCCCAGCCCCATCCTCAGTTTTTCATTTTCCATCTGGCTGCTGT TGAATTGAGCAGGCAAGAGACAAATCTGTTGAAGTTAGTGGAATCCCCATGTGTTCCTATGAAAGTCAAAGTGGGTCCCACATTGGCCAG AAAATTTCTGAGAATATATGAAGAGAGCATTGCAAGATATTAATATAGCTTTTCTTCACAAAAAGCAGTGAACTGTTTACATTTCTTACT >100045_100045_11_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000477432_SLC35D2_chr9_99122503_ENST00000375257_length(amino acids)=207AA_BP=137 MALWALTCPGSTVELAPRLPGRIVCRCSSCRKGRPFSVSGRSEGQRVGEKGKKKGSAPPRRAVDRGAQFRQAGEVAEGPPEMFSLSSTVQ PQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFTIPLTLLLET -------------------------------------------------------------- >100045_100045_12_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000477432_SLC35D2_chr9_99122503_ENST00000375259_length(transcript)=2230nt_BP=1224nt GGAGGGGGCGTCGGGAAAGCCCCGACTTCGCAGCCTTACACTCTTCGTGGGCGGCGACCGCGGCCCCACTGACATCATTCCTCATGAGGG AGGAGGCACAAACAGTTCTGGGCCGACCAGAAAAAGGACGACTGGGACTTGACTCTGAATCGCAGGATTTGAAGAGATTTCTCCTGGCTT CCCAACGAGGCTGGTGGGAAGCGGTCCTCCTCCCATACACGACCTCCCACCCTCGCGAGGCGTAGAAACCAGTTCTGACTGTACAGTAAA GCGAGGGCCAGGGCTGAGGTCTGGAAGCTAATGAAAGCACAGAAAGTGTCGAAACTGGATGAGCAGGAAGCGAGTGGCCTCCCCTGTCAT CTGACGTTTTCCCAGGATGTAATTTGCCTGACTGAAACAGATCAGGACCAACAGGGAGAGTTTTCGATTTAGTGTGAGGAAAAGAGCACT AAATTGTAGCAAAAGACCTTATTGCTCAAGGCCCAGTCAGAAGATTTCATAAGGAAGCTGTAGAAAGTCTTAAGAGGAAATCAGCCGGGC GTGGTGGCGGGCACCTGTAATCCCAGCTACTCGGGAGGCTGAGGCAGGAGAATCGCTTGAACCCGGGGAGCAGAGGTTGCAGTGAGCCGA GATCGCGCCACTGTACGCCAGCCTGGGCGAAAGAACGAAACTCCGTCTCCAAAAAAAAAAAAAACGAAGAAAAAGTCCAAAGAGGGTAAA GGCTGTTCTCCCCTTAAAAAACAGCTAAAGACCTTTGGGGGCGCTGCTCCTTGTAAATGTCAACTACTTCCGCGGGAAAGAACGCGCAGG CACTTGGCCTTGTGGGCGCTCACTTGCCCCGGAAGTACTGTTGAGTTAGCGCCTCGCCTTCCGGGGCGGATTGTCTGTCGTTGCAGTAGC TGTAGGAAGGGGAGGCCATTTTCCGTTTCTGGGAGGAGTGAGGGGCAACGGGTCGGAGAAAAAGGAAAAAAGAAGGGCTCAGCGCCTCCC CGCCGGGCCGTGGACAGAGGGGCACAGTTTCGGCAGGCGGGTGAGGTCGCTGAGGGCCCGCCGGAGATGTTTTCCTTGTCGAGCACGGTG CAACCCCAGGTTACAGTTCCTCTGAGTCATCTCATCAATGCCTTCCATACACCAAAAAACACTTCTGTTTCTCTCAGTGGAGTGTCAGTT TCTCAAAACCAGCATCGAGATGTAGTTCCTGAGCATGAGGCTCCCAGCAGTGAGCTGTTTCCTCTGCCTCTCCTCTACGTTGGAAACCAC ATAAGTGGATTATCAAGCACAAGTAAATTAAGCCTACCGATGTTCACCGTGCTCAGGAAATTCACCATTCCACTTACCTTACTTCTGGAA ACCATCATACTTGGGAAGCAGTATTCACTCAACATCATCCTCAGTGTCTTTGCCATTATTCTCGGGGCTTTCATAGCAGCTGGGTTTCTG CTGATGTACTCCACGGTTCTGTGCAGCTATTACAATTCAGCCCTGACGACAGCAGTGGTTGGAGCCATCAAGAATGTATCCGTTGCCTAC ATTGGGATATTAATCGGTGGAGACTACATTTTCTCTTTGTTAAACTTTGTAGGGTTAAATATTTGCATGGCAGGGGGCTTGAGATATTCC TTTTTAACACTGAGCAGCCAGTTAAAACCTAAACCTGTGGGTGAAGAAAACATCTGTTTGGATTTGAAGAGCTAAAGAGTCTGCAGCAGG ATTGGAGACTGACTTGTGACTGCGGGCTGGGGGGGCATTCCCAGTAGGAATGTGAAGCCAGAGGTTTCGGATTCGTGACATCCACCCCCT GGGCAAGTGAGAGCATCTGCAAAATGCAAAGAGAACTACCTCATATGCAGGATGAGCCAATGGCAGTCTCAAGAAATGTACTCGGGCGAC ACCTTACCTGTGGAAAGCAAATCTTTTCAAAATAAGCCACTGGGACTCGGTAGGTGGAGCCCCAGCTGCTCTTCTAGGGACCTATGGGGC CTTCGTGGCATCTCTGTGCTGTGTGCTGGGGAGGAGGTTGATGTAATGGTGACTCTTTTCTGATCAGCACCTTGGCCGTGATTCCCAAGG TCCCAGCCAAAGCAAAGGGCCAGTTGTTTCAGTTTAAACAGACATGTCTTTAGTCTAATAAAATTAGTTAACTGCCAGTAAAGTTATTTG >100045_100045_12_YME1L1-SLC35D2_YME1L1_chr10_27437835_ENST00000477432_SLC35D2_chr9_99122503_ENST00000375259_length(amino acids)=293AA_BP=137 MALWALTCPGSTVELAPRLPGRIVCRCSSCRKGRPFSVSGRSEGQRVGEKGKKKGSAPPRRAVDRGAQFRQAGEVAEGPPEMFSLSSTVQ PQVTVPLSHLINAFHTPKNTSVSLSGVSVSQNQHRDVVPEHEAPSSELFPLPLLYVGNHISGLSSTSKLSLPMFTVLRKFTIPLTLLLET IILGKQYSLNIILSVFAIILGAFIAAGFLLMYSTVLCSYYNSALTTAVVGAIKNVSVAYIGILIGGDYIFSLLNFVGLNICMAGGLRYSF -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for YME1L1-SLC35D2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for YME1L1-SLC35D2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for YME1L1-SLC35D2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
