
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ZMIZ1-CPEB3 (FusionGDB2 ID:101300) |
Fusion Gene Summary for ZMIZ1-CPEB3 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ZMIZ1-CPEB3 | Fusion gene ID: 101300 | Hgene | Tgene | Gene symbol | ZMIZ1 | CPEB3 | Gene ID | 57178 | 22849 |
| Gene name | zinc finger MIZ-type containing 1 | cytoplasmic polyadenylation element binding protein 3 | |
| Synonyms | MIZ|NEDDFSA|RAI17|TRAFIP10|ZIMP10 | - | |
| Cytomap | 10q22.3 | 10q23.32 | |
| Type of gene | protein-coding | protein-coding | |
| Description | zinc finger MIZ domain-containing protein 1retinoic acid induced 17zinc finger-containing, Miz1, PIAS-like protein on chromosome 10 | cytoplasmic polyadenylation element-binding protein 3CPE-BP3CPE-binding protein 3hCPEB-3 | |
| Modification date | 20200315 | 20200322 | |
| UniProtAcc | . | Q8NE35 | |
| Ensembl transtripts involved in fusion gene | ENST00000334512, ENST00000446377, ENST00000478357, | ENST00000265997, ENST00000412050, | |
| Fusion gene scores | * DoF score | 20 X 11 X 11=2420 | 8 X 8 X 2=128 |
| # samples | 25 | 8 | |
| ** MAII score | log2(25/2420*10)=-3.27500704749987 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(8/128*10)=-0.678071905112638 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: ZMIZ1 [Title/Abstract] AND CPEB3 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ZMIZ1(80976031)-CPEB3(93904869), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | ZMIZ1 | GO:0007179 | transforming growth factor beta receptor signaling pathway | 16777850 |
| Hgene | ZMIZ1 | GO:0030521 | androgen receptor signaling pathway | 14609956 |
| Hgene | ZMIZ1 | GO:0033233 | regulation of protein sumoylation | 14609956 |
| Hgene | ZMIZ1 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 26522984 |
| Hgene | ZMIZ1 | GO:0060395 | SMAD protein signal transduction | 16777850 |
| Hgene | ZMIZ1 | GO:1903508 | positive regulation of nucleic acid-templated transcription | 14609956|16777850 |
| Tgene | CPEB3 | GO:0000122 | negative regulation of transcription by RNA polymerase II | 20639532 |
| Tgene | CPEB3 | GO:0017148 | negative regulation of translation | 21336257|22711986 |
| Tgene | CPEB3 | GO:0060213 | positive regulation of nuclear-transcribed mRNA poly(A) tail shortening | 21336257 |
| Tgene | CPEB3 | GO:0061158 | 3'-UTR-mediated mRNA destabilization | 21336257 |
| Tgene | CPEB3 | GO:0071230 | cellular response to amino acid stimulus | 20639532|22730302 |
| Tgene | CPEB3 | GO:1900153 | positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay | 21336257 |
 Fusion gene breakpoints across ZMIZ1 (5'-gene) Fusion gene breakpoints across ZMIZ1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
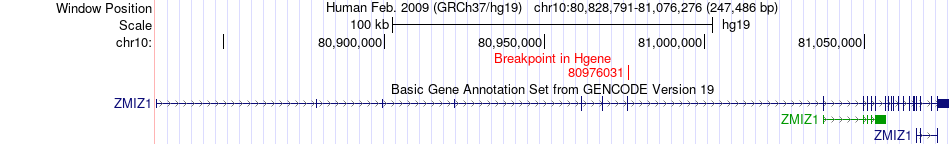 |
 Fusion gene breakpoints across CPEB3 (3'-gene) Fusion gene breakpoints across CPEB3 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
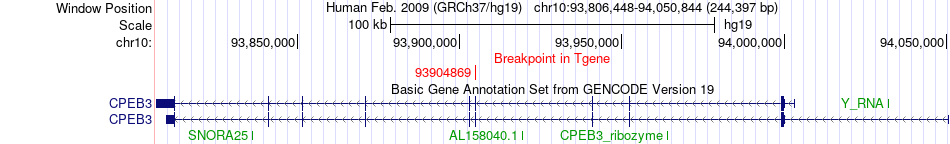 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | UCEC | TCGA-5B-A90C-01A | ZMIZ1 | chr10 | 80976031 | - | CPEB3 | chr10 | 93904869 | - |
| ChimerDB4 | UCEC | TCGA-5B-A90C-01A | ZMIZ1 | chr10 | 80976031 | + | CPEB3 | chr10 | 93904869 | - |
| ChimerDB4 | UCEC | TCGA-5B-A90C | ZMIZ1 | chr10 | 80976031 | + | CPEB3 | chr10 | 93904869 | - |
Top |
Fusion Gene ORF analysis for ZMIZ1-CPEB3 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000334512 | ENST00000265997 | ZMIZ1 | chr10 | 80976031 | + | CPEB3 | chr10 | 93904869 | - |
| In-frame | ENST00000334512 | ENST00000412050 | ZMIZ1 | chr10 | 80976031 | + | CPEB3 | chr10 | 93904869 | - |
| intron-3CDS | ENST00000446377 | ENST00000412050 | ZMIZ1 | chr10 | 80976031 | + | CPEB3 | chr10 | 93904869 | - |
| intron-3CDS | ENST00000478357 | ENST00000412050 | ZMIZ1 | chr10 | 80976031 | + | CPEB3 | chr10 | 93904869 | - |
| intron-intron | ENST00000446377 | ENST00000265997 | ZMIZ1 | chr10 | 80976031 | + | CPEB3 | chr10 | 93904869 | - |
| intron-intron | ENST00000478357 | ENST00000265997 | ZMIZ1 | chr10 | 80976031 | + | CPEB3 | chr10 | 93904869 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000334512 | ZMIZ1 | chr10 | 80976031 | + | ENST00000412050 | CPEB3 | chr10 | 93904869 | - | 7274 | 852 | 500 | 1753 | 417 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000334512 | ENST00000412050 | ZMIZ1 | chr10 | 80976031 | + | CPEB3 | chr10 | 93904869 | - | 0.000115289 | 0.9998847 |
Top |
Fusion Genomic Features for ZMIZ1-CPEB3 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
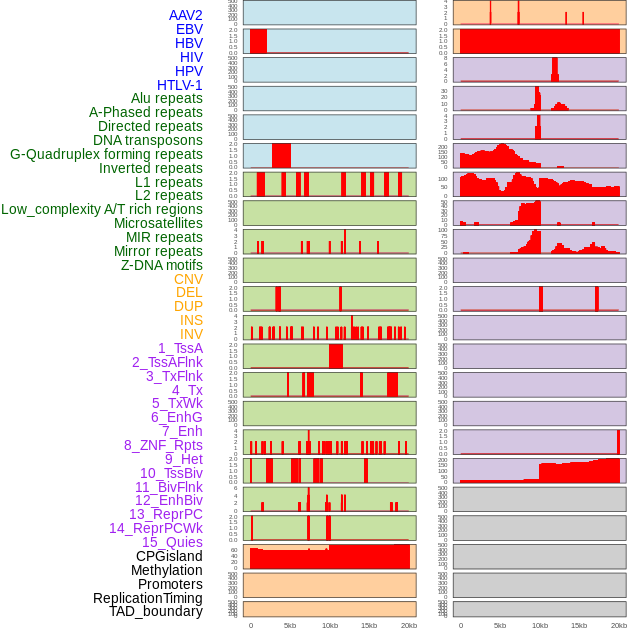 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for ZMIZ1-CPEB3 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr10:80976031/chr10:93904869) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | CPEB3 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Sequence-specific RNA-binding protein which acts as a translational repressor in the basal unstimulated state but, following neuronal stimulation, acts as a translational activator (By similarity). In contrast to CPEB1, does not bind to the cytoplasmic polyadenylation element (CPE), a uridine-rich sequence element within the mRNA 3'-UTR, but binds to a U-rich loop within a stem-loop structure (By similarity). Required for the consolidation and maintenance of hippocampal-based long term memory (By similarity). In the basal state, binds to the mRNA 3'-UTR of the glutamate receptors GRIA2/GLUR2 mRNA and negatively regulates their translation (By similarity). Also represses the translation of DLG4, GRIN1, GRIN2A and GRIN2B (By similarity). When activated, acts as a translational activator of GRIA1 and GRIA2 (By similarity). In the basal state, suppresses SUMO2 translation but activates it following neuronal stimulation (By similarity). Binds to the 3'-UTR of TRPV1 mRNA and represses TRPV1 translation which is required to maintain normal thermoception (By similarity). Binds actin mRNA, leading to actin translational repression in the basal state and to translational activation following neuronal stimulation (By similarity). Negatively regulates target mRNA levels by binding to TOB1 which recruits CNOT7/CAF1 to a ternary complex and this leads to target mRNA deadenylation and decay (PubMed:21336257). In addition to its role in translation, binds to and inhibits the transcriptional activation activity of STAT5B without affecting its dimerization or DNA-binding activity. This, in turn, represses transcription of the STAT5B target gene EGFR which has been shown to play a role in enhancing learning and memory performance (PubMed:20639532). In contrast to CPEB1, CPEB2 and CPEB4, not required for cell cycle progression (PubMed:26398195). {ECO:0000250|UniProtKB:Q7TN99, ECO:0000269|PubMed:20639532, ECO:0000269|PubMed:21336257, ECO:0000269|PubMed:26398195}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000265997 | 0 | 10 | 13_27 | 0 | 699.0 | Compositional bias | Note=Gln-rich | |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000265997 | 0 | 10 | 170_238 | 0 | 699.0 | Compositional bias | Note=Ala-rich | |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000265997 | 0 | 10 | 26_200 | 0 | 699.0 | Compositional bias | Note=Pro-rich | |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000265997 | 0 | 10 | 441_532 | 0 | 699.0 | Domain | RRM 1 | |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000265997 | 0 | 10 | 549_631 | 0 | 699.0 | Domain | RRM 2 | |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000412050 | 3 | 10 | 441_532 | 384 | 685.0 | Domain | RRM 1 | |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000412050 | 3 | 10 | 549_631 | 384 | 685.0 | Domain | RRM 2 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ZMIZ1 | chr10:80976031 | chr10:93904869 | ENST00000334512 | + | 7 | 25 | 280_305 | 93 | 1068.0 | Compositional bias | Note=Ala-rich |
| Hgene | ZMIZ1 | chr10:80976031 | chr10:93904869 | ENST00000334512 | + | 7 | 25 | 334_555 | 93 | 1068.0 | Compositional bias | Note=Pro-rich |
| Hgene | ZMIZ1 | chr10:80976031 | chr10:93904869 | ENST00000334512 | + | 7 | 25 | 867_1002 | 93 | 1068.0 | Compositional bias | Note=Pro-rich |
| Hgene | ZMIZ1 | chr10:80976031 | chr10:93904869 | ENST00000334512 | + | 7 | 25 | 837_1067 | 93 | 1068.0 | Region | Transactivation domain |
| Hgene | ZMIZ1 | chr10:80976031 | chr10:93904869 | ENST00000334512 | + | 7 | 25 | 727_804 | 93 | 1068.0 | Zinc finger | SP-RING-type |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000412050 | 3 | 10 | 13_27 | 384 | 685.0 | Compositional bias | Note=Gln-rich | |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000412050 | 3 | 10 | 170_238 | 384 | 685.0 | Compositional bias | Note=Ala-rich | |
| Tgene | CPEB3 | chr10:80976031 | chr10:93904869 | ENST00000412050 | 3 | 10 | 26_200 | 384 | 685.0 | Compositional bias | Note=Pro-rich |
Top |
Fusion Gene Sequence for ZMIZ1-CPEB3 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >101300_101300_1_ZMIZ1-CPEB3_ZMIZ1_chr10_80976031_ENST00000334512_CPEB3_chr10_93904869_ENST00000412050_length(transcript)=7274nt_BP=852nt CGGGCGCCGGGGAGAGCGGGCGGCCGGGCGGCAGGCGGGCGAGCAGCGATCGGGCGGCCGAGCGAGCGAGCAACGCCGGCGCAGCGCGGT GACCCCAGCCCCAGCCGGCGCGGAGCAGGAGCCGGAGCCGAGCGGATCTCGGCGCCCTCGCTGCGCTCCTCCCGGCCCGAGCCTGCCCTA CCCGGCGGTGGCGGCGGCGCGTCCTCCAGCGGCGGCAGCGGCGCTCGCAGCGCCCGGACTCACGGCAGCTGTGTTCTGATTTCGTACTAC TGCTGGGGCTGCCACCTCCTCCTCCAGACGCTCTCAGCAGACTTGAGTCCTGGTCCTTCTGCAGAGGCCTGAGCAGGAGGAAGAGGAGGA GGCCCGTTGGCGTCGGACCAATGCTGCAAGGGGTGTGAGGAGAGGAGCCGCTGTTTTTCACTGAGCTGCCATACCCCGAAAGCAGGATGG AGCTGGAGTGAGGTGGAGGGGCCGCAAGCTGCTGACCGGCGTGTGGAACACTGGTGGTTTGCAGATCACTGAGGCTGGACAACGTTCATG GCTCTCGGGTAGAACCTAGTGAAACGGCCAGAATGAATTCTATGGACAGGCACATCCAGCAGACCAATGACCGACTGCAGTGCATCAAGC AGCACTTACAGAATCCTGCCAACTTCCACAATGCCGCCACGGAGCTGCTGGACTGGTGCGGAGACCCACGGGCCTTCCAGCGGCCCTTCG AGCAGAGCCTGATGGGCTGTTTGACGGTGGTCAGTCGGGTGGCAGCCCAGCAAGGCTTTGACCTGGACCTCGGCTACAGACTGCTGGCTG TGTGTGCTGCAAACCGAGACAAGTTCACCCCGAAGTCTGCCGGTCGATCTTCTCTCTTTCCCTTTGAAGATGCCTTCCTGGATGATAGCC ATGGTGATCAAGCCTTGTCATCTGGCTTAAGTTCTCCCACTCGCTGTCAAAATGGGGAACGAGTAGAACGCTACTCTAGAAAGGTGTTTG TTGGAGGACTTCCTCCTGATATTGATGAAGATGAGATCACTGCCAGCTTTCGCAGGTTTGGACCTCTCGTAGTAGACTGGCCTCACAAAG CTGAAAGCAAGTCTTATTTTCCTCCTAAAGGCTATGCCTTTCTGCTGTTCCAAGAGGAAAGCTCAGTACAAGCTTTGATAGATGCCTGCC TAGAAGAAGATGGGAAACTCTACCTGTGTGTGTCAAGCCCCACCATCAAGGACAAGCCAGTGCAAATTCGACCATGGAACCTAAGTGACA GTGACTTTGTAATGGATGGTTCTCAGCCTTTGGACCCCAGAAAAACTATCTTTGTTGGGGGAGTTCCACGACCCCTTCGAGCTGTTGAAC TGGCAATGATCATGGACCGTTTGTATGGTGGTGTCTGCTATGCTGGCATTGATACGGACCCAGAGCTGAAGTACCCCAAAGGTGCTGGCC GCGTGGCATTCTCCAATCAGCAGAGTTACATTGCAGCCATCAGCGCTCGTTTTGTGCAGCTTCAGCACAATGACATTGACAAACGGGTTG AAGTAAAGCCATATGTGCTGGATGATCAGATGTGTGATGAGTGCCAGGGCACACGCTGTGGTGGGAAGTTTGCCCCGTTCTTCTGTGCCA ACGTCACCTGTCTGCAGTATTACTGTGAATACTGCTGGGCGAGCATACATTCCCGAGCCGGGCGGGAGTTCCACAAACCGCTGGTGAAGG AGGGAGGCGACCGCCCTCGTCACGTCCCGTTCCGCTGGAGCTGAGCCACGGGCCGACCCAGCCACAAGTACTGGAGTAGATAACAAGGAG GGAAAAGAGAGGGCACTGCCTCAGGGCTTACAGTGTTCTGGAAATCTGTGCATTCGTTCTCGATTTTTAAAGAAGAATTGGGTCTTTTTA TTATTATTATTATTATTTACAGTCATATTACTGAACTCAGTGTCCAGTATAATGCAGAATTGCAGAGTGTAGAATGGTACTGATTTGCAC TTTGATTCACACCTTTGTCTGCTTGGTACCTTAATTTTTTATACCTGATTTGTCACTCGTGACAGTATGTAGACTGCCATTCTCATCAGC GGCCGTCAGGCTACTGTCTGTGATTCTTTTGTTTGTTGTTGTGGTGTTGTTCTGATTGTTTTGAACTGGAGCATGCACTTATTTTCAGAA ACTTAGAGAGTTGCCCCTAGGACAAGACTTACCAAAGGTAATACTCGTGAGTAGGTGGCAGATGTAGCAAAAACTTCACAACAATTCAGC AATCATTAGCTCGGGTCACTTAGAGTAAGGATAACCCTTCATGAAAATAAGCCTTTAGTTGCTCTCTATTTTGACTTGTAAGGATACCTC ATAAGCTGGATCTTTTATTTTTCCTTCATGTTTGATTTTTGTTTTGCCGAGGATTTTGGTTAGATCTTTTAGTATTATGTAAATTTATGC TGCAATAGCTTTTGCTGCTATTAAAATGGTATTATTAATACCATAATACTCTATTCAGGCTAGGATATCAATTAAACACAAAAACCCAAC AACACACTTTTGTGATTTTAGGGATAGAGTCAGCTGCCGAACGGAGATCAGAATAGACTTCAGAAAGCTTCGGTGGAAGGTCACTATTTC TGACTGCATGTTTACTTAATCAGGTTGCTGCATGCAATAAATTTTATATCCAATGTCTAGTATAAAACGTTCCCACAATGCACAAATTAA GAATCTATATAAGCTATAAAAATACTCATTTTTTAAAAAAAGTTTTTTTCATTCAAAGAATTGGTAATTGACCGGAGGAAGGCATGCCTC AATAAATGGAATGCTCTGAATGGGAAGAGCCCATAACAGAATGACCTATATTACAACCGATATAAAACCTAGGTGTAGGATATTCTTCAA GCAAATTTGTGGATGGTAGATGAAGTACTTTGCGGTGCTCTGTTATATATTGTGAAATTTATTTTATATAGTAAAGTGATTATGTCTAAC TGTGATCAAACTTAACTGTTTAAAGCCTCTGTCAGACATTAATGAACTTGACAGTTCATAACTGATATATTATAAACTAACACATGAGCT CTTAGTTACAATCTCTTAGTGCTTGCACCATTTTACATAGCTTCAGACCTTGTCCAGAAAACCAAAGAGCTCATCTTAGCTTACAAACAC TAAGAATATATACAGGATATAGAGATAAGTTGTCAGATTGACAAACTAGGCACATTTTTAAGAAAGTTGTGTAATGGTGATGCTAAAGAA ATGTCTGTACAAAATAAGATGGCCTGGGCAGAGTTGGTTGTCTGAGTGAGAAACTAACTACTTAATTCCCTGGATTTATCCTGTTAAGCT TGAGTAGAATGAAGGCCTGCTAGCACTAGCGTGGTCTAGAATGAAGGCCTGCTAGCACTAGCGTGGACATTTTTCTAGCATTCACCCTTG GGCTGCAGATAGTAATATATAAACACACACAATGATTGCAAGACTATGTAAATCTTTTTTTAATCTTCTTCCTATGTTTTGGAATTCTGG GGACAATCTGACTGGATATGACACTGCAGAATAGATTTAAGTCGCTGTGTTTTCTGTGCTGTGATGTCCTTGTACTGTCTTAAGTTAGTG TTGCTTGTTTTTGTTTTGTTCTCCCATGTTTAGTTGCAATTACTGGCTCTCAAGCTATAGTTTGTTTTCAAAAAGGAAAAACAAAATAGT TTTTTACTGGATTAAAAATGCTTATTTTAGAATCCTCCCCTTTTTGAGGCTTTTTGGTAATAATTGCTTTTTTCCAGCCCAGGTGTGGTT TTGTTTGTTTTTTTCTATGTTTGTGGGTTTTTTTTCATCTGTACAAGAAATTTTGTTATGCACTACTACTTTTTGTATTCTAGACAGTTT TCAAAAGTTGGCAGATTTTTTTTTTTGTTTTTAAGGAAAAAGGCATGCTTTCTGACTGACATGATCTTTTGCAATACCTCAATTTTGAAA TATAGGAAAGAAAATGATATTTTCTAAGTAAGTTTATTCAGAAAAATATATATTAATTTAATTCTGCAAGAAAATGATTTAACAGATGGG TAACTTTATTTTTTTCATGCTTATTTGTGTTTTTTGAAAATGAAGAAGTTAGAGCTTTATAACTATAATATTAAAGATCATATCTAACCT ACAGAAATCTACCTAATTATGTGAAAGAAGCAAAACTTTTTGAAGAAGCACCCCTCTTTCATGTATTAAAATAACACTGAGATTTTTTTA AAAGCTGGAAGATTGTACAGCATAAAGCTAAAGTGAAGGCAAAAAACATTGTCCTGAAGAATAGGTTACAAAAAATAAACTCCATTTGGT GCTGAAAGTTGTGAATAGATGGTGGAAACTACTTTCCCTTAATTTGTATATTTCTAATCAAGTGTTGATTGTGCGCGACTGACCAGGATT GTGAAAGAGTTCTTCTGTTTGTATTTTTTTAAAGAGAACTTTTTTAGTTTATTAAATTATAACATGTTCCCTTTTGTTTTGTAAATAAAT GTGCCCCCAAGTTGTTAATGTTATATTAAAAGAGAGGGTCCTTCAAAAAAATTATTTTTTAAAACACAGAGGTGTTCTGACCTGCAACTT GACTGGGAAGCCCCTGGGTAACACAGGTCCTGCTGTGGCCTTTCCTTGGTGTGACACTTGAACTAGTCGATTTTTTGAAGGCTATAACCT AACCTCGACTATTTGTTGTTATGTGCAATACTGTTTGTACTGTTTGTATAAAAAATAGAAAAAAAAAAGAAAAGAAAAAAGGCATAAATC TTTATTGCAGATTTATTTTCATTGCAAACTTTAGACATTGTGATTCACTGAAATCATTTTTCTACTGTCAAATATGTACCTTTTCTTCAT GCTGGTATTTTTTCAATAGTAATGTAGCCTTTGTACTGAGACAAATTTTCATTGTTATTTTTAGAAAGATTTTTTGCTAAGAAAAAAATT AAACATATTTACATATTTTTCATTTTATTTCGTATTTTCTTTAAGGAGTGCTTTCTCAATCATTAATAGAGTTGTTTCTGGGAGGGACTT TCATATCTGGTCAATGTCATAATATTATTTTGTATTTTTATTGGAATAAATTAATTTTATGATGCTAATTCTTTAAAAGTTTATTTTTTT CATTTTCAGTTATTTTTGAAGCTAAAACCTTTGTAATATTGTTACTAAAGTGTCTAGTTTTTTGAAATGCAAAGCTTTATGTATTAACTT GCTGATGTGGTTTACAATAGTGTGACAACGATGAAAATGCTTGGTTATGTAAAGTGATTGGAAATGTAATGGATAATTGGGTAATAAAAT GACAGAATGAGCAGAACATGTGAGATTTTGAGAAGCCTTTTCCTTTATCACAGTGGTTTAGTGTAAGACTAGGGATGTCACAGTGCAGTT CTCTAGGGATGTCACGGTGGATTTATACAACACCAGGTGCAGCCAAACCTGTGGCCCTGAGAATGAAGTCACCCAACCGAAGTACCACTG GGTTTAAAATCTGGCTAAGTCAACAGAATGTTTCATTGCTTTTTCCCCTCTAACTCGTCATTATGTCGTGTGTGTGTGTGTGTGTGTGAG AGAGAGAGAGAGAGAGAGAGAGAAAAACTGCCACCAACTGCTGTCCTTGTGTGAGACAGTGCCTTTCTCCCAGGCCGAATCTTTTATGTG CACTGACAGTGGTGATATCGGAGTCATACAGGAAGTTGCTAAGAGGACACGGCTTTCGTGGTGTACTGCAGACCTCACCTGCAGGGTCTC ACTCATGCTGCCTCCCTGTAAGGCTCAGTTGGTGGCACTGAGGAAGCACACTCGGATGTCACCACTGTCTGTGGACCCTGACTATGTGTC GGGTTGCATCTTCCTCCACACCCATGCCTGCTTCCTGTTCTGGTGCCCCTTGCCTTTCATTGTGGCAACTGCGTGCATGATCTGTCTGCC TTTCTGCTGCTGCTTTTCCAACACCCTCCTCACCTGGAAGCTGGGCAGTTAAGGCACAGTCATTAATGTGTCATTGCAATAGCACCTACA GTCAAAGCAAAAACTACAACTGCAAAACTGGTGTTGCGTAAACAGAACAGCCAATGACTTGATCCAGATAGATGATATGTTTCTACTAGT GGTTGATGGCCTCTTTCAGCTCCATCGCTATGGAGGCGAGGTTCATCAATAACTCAGAGAAACTTGTGTCCAGAGTTGGTCTTGGGTATG CTCTATGTATTCAGTTTTGTAAATAATATATAGATTAGATCAAGTTGTGATGTAAAATTTTAACTCAAATTGGGTACTACTGGTTGGAAT TTTACATTAACTACAAATAAATGATTAGGAACAGAAGAAAAGTGGAAAATTCTTGCTTAGTGGTATAAAGATTATGCTTAGATTTCTGCC TATATCTTGTGTTTTATAGCTTGTTAGTGAGGCATGGGCTAATGCAGAATGTACTGCATTTATGCAATTAGCAAATAACTGCTAGTTTTC ATTTTATCTATTATATTAGCAACAAAAATGGCATTTAACTGGGTAAAGCTCCTACCTAAATATTTGAGTATTGTGACATCTCTACTTGCA AGGTTTGAGTGAAAGATTAGTCACAGATTAGTGAAAAAAATGTATCTTTTTGTTAAAACTGGCTTATTTGAATGGATTGTTGCTTTTCAC TAGCTTGTATGGTATGTCAGCTAACATTTGTTTTTTTTTTTCTTTTATAATAACCAAACTTCTCTATCTCAGCTGCAATAGTGTTCCATC TTACCACAGAGTGTTTTATAATTAATGCTGTTGACTTTATACCCCAAAATGGGTCAAGAAGTGGCTACATGATTAGGGGATGTTTAAATT TGTATGACGACAAAGTTAATTGCAACAAGTAGAAAACTTGGGTGCTAACACAGGATAACTTCTGCTCTTTTTCTTTCCTGGATATAGTTC TGTTGGGGTATAAAGTATTAGGTATTTGTATTTTTTGCTGTCAAAATAATGGACTTAACTTTTAGAGTGTTCACAGCTAGGATCTCTGTA TTTGTTTTTCAACTAACAACTAATTTTAAATGTATTGTATCTTTTATTACCTGTTCTCTTAGATTGTTTTTTGCTAGTAACCCTTACTCA GCTTCTGCCTTTGGGGCTTGGACTAATATTGCTTGTATGGAACTACAGTACTGTGACTAAACATGGAGCTCAATGCAGCTACTGCTTACA >101300_101300_1_ZMIZ1-CPEB3_ZMIZ1_chr10_80976031_ENST00000334512_CPEB3_chr10_93904869_ENST00000412050_length(amino acids)=417AA_BP=117 MVVCRSLRLDNVHGSRVEPSETARMNSMDRHIQQTNDRLQCIKQHLQNPANFHNAATELLDWCGDPRAFQRPFEQSLMGCLTVVSRVAAQ QGFDLDLGYRLLAVCAANRDKFTPKSAGRSSLFPFEDAFLDDSHGDQALSSGLSSPTRCQNGERVERYSRKVFVGGLPPDIDEDEITASF RRFGPLVVDWPHKAESKSYFPPKGYAFLLFQEESSVQALIDACLEEDGKLYLCVSSPTIKDKPVQIRPWNLSDSDFVMDGSQPLDPRKTI FVGGVPRPLRAVELAMIMDRLYGGVCYAGIDTDPELKYPKGAGRVAFSNQQSYIAAISARFVQLQHNDIDKRVEVKPYVLDDQMCDECQG -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for ZMIZ1-CPEB3 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ZMIZ1-CPEB3 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ZMIZ1-CPEB3 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
