
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:BRCA1-BECN1 (FusionGDB2 ID:10149) |
Fusion Gene Summary for BRCA1-BECN1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: BRCA1-BECN1 | Fusion gene ID: 10149 | Hgene | Tgene | Gene symbol | BRCA1 | BECN1 | Gene ID | 672 | 8678 |
| Gene name | BRCA1 DNA repair associated | beclin 1 | |
| Synonyms | BRCAI|BRCC1|BROVCA1|FANCS|IRIS|PNCA4|PPP1R53|PSCP|RNF53 | ATG6|VPS30|beclin1 | |
| Cytomap | 17q21.31 | 17q21.31 | |
| Type of gene | protein-coding | protein-coding | |
| Description | breast cancer type 1 susceptibility proteinBRCA1/BRCA2-containing complex, subunit 1Fanconi anemia, complementation group SRING finger protein 53breast and ovarian cancer susceptibility protein 1breast cancer 1, early onsetearly onset breast cancer | beclin-1ATG6 autophagy related 6 homologbeclin 1 (coiled-coil, moesin-like BCL2-interacting protein)beclin 1, autophagy relatedcoiled-coil myosin-like BCL2-interacting proteintestis secretory sperm-binding protein Li 215e | |
| Modification date | 20200329 | 20200327 | |
| UniProtAcc | P38398 | Q14457 | |
| Ensembl transtripts involved in fusion gene | ENST00000309486, ENST00000346315, ENST00000351666, ENST00000352993, ENST00000354071, ENST00000357654, ENST00000468300, ENST00000471181, ENST00000491747, ENST00000493795, ENST00000586385, ENST00000591534, ENST00000591849, | ENST00000361523, ENST00000438274, ENST00000590099, | |
| Fusion gene scores | * DoF score | 9 X 8 X 7=504 | 4 X 3 X 4=48 |
| # samples | 9 | 4 | |
| ** MAII score | log2(9/504*10)=-2.48542682717024 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(4/48*10)=-0.263034405833794 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: BRCA1 [Title/Abstract] AND BECN1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | BRCA1(41209069)-BECN1(40968072), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | BRCA1-BECN1 seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. BRCA1-BECN1 seems lost the major protein functional domain in Hgene partner, which is a epigenetic factor due to the frame-shifted ORF. BRCA1-BECN1 seems lost the major protein functional domain in Hgene partner, which is a tumor suppressor due to the frame-shifted ORF. BRCA1-BECN1 seems lost the major protein functional domain in Tgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | BRCA1 | GO:0000724 | double-strand break repair via homologous recombination | 17349954 |
| Hgene | BRCA1 | GO:0006301 | postreplication repair | 17349954 |
| Hgene | BRCA1 | GO:0006302 | double-strand break repair | 22186889 |
| Hgene | BRCA1 | GO:0008630 | intrinsic apoptotic signaling pathway in response to DNA damage | 14654789 |
| Hgene | BRCA1 | GO:0016567 | protein ubiquitination | 17349954 |
| Hgene | BRCA1 | GO:0031398 | positive regulation of protein ubiquitination | 15965487 |
| Hgene | BRCA1 | GO:0035066 | positive regulation of histone acetylation | 20820192 |
| Hgene | BRCA1 | GO:0043627 | response to estrogen | 8895509 |
| Hgene | BRCA1 | GO:0045892 | negative regulation of transcription, DNA-templated | 16288014 |
| Hgene | BRCA1 | GO:0045893 | positive regulation of transcription, DNA-templated | 20160719 |
| Hgene | BRCA1 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 16331276 |
| Hgene | BRCA1 | GO:0051571 | positive regulation of histone H3-K4 methylation | 20820192 |
| Hgene | BRCA1 | GO:0051573 | negative regulation of histone H3-K9 methylation | 20820192 |
| Hgene | BRCA1 | GO:0051865 | protein autoubiquitination | 12890688|20351172 |
| Hgene | BRCA1 | GO:0070512 | positive regulation of histone H4-K20 methylation | 20820192 |
| Hgene | BRCA1 | GO:0071158 | positive regulation of cell cycle arrest | 21102443 |
| Hgene | BRCA1 | GO:0071681 | cellular response to indole-3-methanol | 10868478 |
| Hgene | BRCA1 | GO:0085020 | protein K6-linked ubiquitination | 12890688|20351172 |
| Hgene | BRCA1 | GO:2000617 | positive regulation of histone H3-K9 acetylation | 20820192 |
| Hgene | BRCA1 | GO:2000620 | positive regulation of histone H4-K16 acetylation | 20820192 |
| Tgene | BECN1 | GO:0006914 | autophagy | 23629966 |
 Fusion gene breakpoints across BRCA1 (5'-gene) Fusion gene breakpoints across BRCA1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
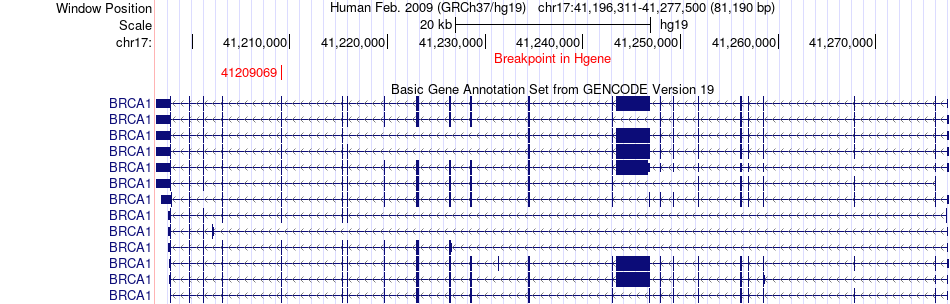 |
 Fusion gene breakpoints across BECN1 (3'-gene) Fusion gene breakpoints across BECN1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
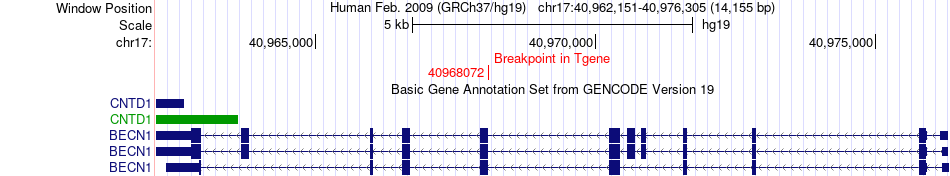 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LIHC | TCGA-CC-5263-01A | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
Top |
Fusion Gene ORF analysis for BRCA1-BECN1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000309486 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000309486 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000309486 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000346315 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000346315 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000346315 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000351666 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000351666 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000351666 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000352993 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000352993 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000352993 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000354071 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000354071 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000354071 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000357654 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000357654 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000357654 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000468300 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000468300 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000468300 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000471181 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000471181 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000471181 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000491747 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000491747 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000491747 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000493795 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000493795 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000493795 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000586385 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000586385 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| Frame-shift | ENST00000586385 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| In-frame | ENST00000591534 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| In-frame | ENST00000591534 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| In-frame | ENST00000591534 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| intron-3CDS | ENST00000591849 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| intron-3CDS | ENST00000591849 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
| intron-3CDS | ENST00000591849 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000591534 | BRCA1 | chr17 | 41209069 | - | ENST00000361523 | BECN1 | chr17 | 40968072 | - | 2148 | 852 | 102 | 872 | 256 |
| ENST00000591534 | BRCA1 | chr17 | 41209069 | - | ENST00000590099 | BECN1 | chr17 | 40968072 | - | 2145 | 852 | 102 | 872 | 256 |
| ENST00000591534 | BRCA1 | chr17 | 41209069 | - | ENST00000438274 | BECN1 | chr17 | 40968072 | - | 1831 | 852 | 102 | 872 | 256 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000591534 | ENST00000361523 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - | 0.001018873 | 0.99898106 |
| ENST00000591534 | ENST00000590099 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - | 0.001017529 | 0.9989825 |
| ENST00000591534 | ENST00000438274 | BRCA1 | chr17 | 41209069 | - | BECN1 | chr17 | 40968072 | - | 0.001025217 | 0.9989748 |
Top |
Fusion Genomic Features for BRCA1-BECN1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
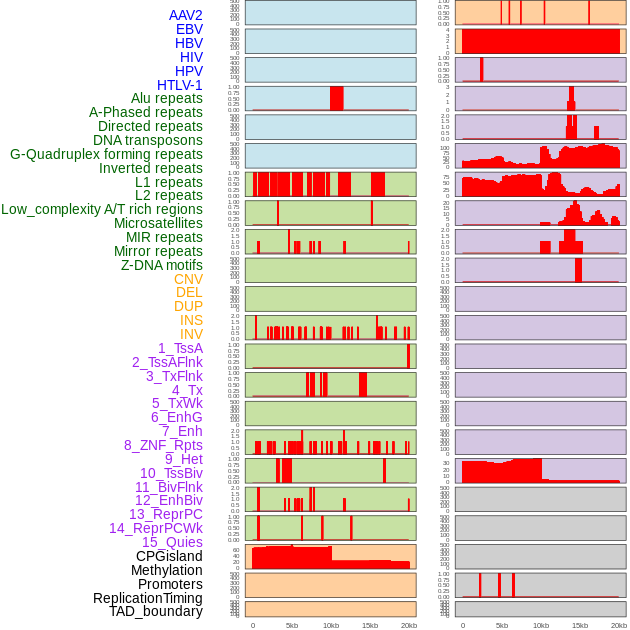 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for BRCA1-BECN1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr17:41209069/chr17:40968072) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| BRCA1 | BECN1 |
| FUNCTION: E3 ubiquitin-protein ligase that specifically mediates the formation of 'Lys-6'-linked polyubiquitin chains and plays a central role in DNA repair by facilitating cellular responses to DNA damage (PubMed:12890688, PubMed:14976165, PubMed:16818604, PubMed:17525340, PubMed:12887909, PubMed:10500182, PubMed:19261748). It is unclear whether it also mediates the formation of other types of polyubiquitin chains (PubMed:12890688). The BRCA1-BARD1 heterodimer coordinates a diverse range of cellular pathways such as DNA damage repair, ubiquitination and transcriptional regulation to maintain genomic stability (PubMed:12890688, PubMed:14976165, PubMed:20351172). Regulates centrosomal microtubule nucleation (PubMed:18056443). Required for appropriate cell cycle arrests after ionizing irradiation in both the S-phase and the G2 phase of the cell cycle (PubMed:10724175, PubMed:12183412, PubMed:11836499, PubMed:19261748). Required for FANCD2 targeting to sites of DNA damage (PubMed:12887909). Inhibits lipid synthesis by binding to inactive phosphorylated ACACA and preventing its dephosphorylation (PubMed:16326698). Contributes to homologous recombination repair (HRR) via its direct interaction with PALB2, fine-tunes recombinational repair partly through its modulatory role in the PALB2-dependent loading of BRCA2-RAD51 repair machinery at DNA breaks (PubMed:19369211). Component of the BRCA1-RBBP8 complex which regulates CHEK1 activation and controls cell cycle G2/M checkpoints on DNA damage via BRCA1-mediated ubiquitination of RBBP8 (PubMed:16818604). Acts as a transcriptional activator (PubMed:20160719). {ECO:0000269|PubMed:10500182, ECO:0000269|PubMed:10724175, ECO:0000269|PubMed:11836499, ECO:0000269|PubMed:12183412, ECO:0000269|PubMed:12887909, ECO:0000269|PubMed:12890688, ECO:0000269|PubMed:14976165, ECO:0000269|PubMed:16326698, ECO:0000269|PubMed:16818604, ECO:0000269|PubMed:17525340, ECO:0000269|PubMed:18056443, ECO:0000269|PubMed:19261748, ECO:0000269|PubMed:19369211, ECO:0000269|PubMed:20160719, ECO:0000269|PubMed:20351172}. | FUNCTION: Plays a central role in autophagy (PubMed:23184933, PubMed:28445460). Acts as core subunit of the PI3K complex that mediates formation of phosphatidylinositol 3-phosphate; different complex forms are believed to play a role in multiple membrane trafficking pathways: PI3KC3-C1 is involved in initiation of autophagosomes and PI3KC3-C2 in maturation of autophagosomes and endocytosis. Involved in regulation of degradative endocytic trafficking and required for the abcission step in cytokinesis, probably in the context of PI3KC3-C2 (PubMed:20643123, PubMed:20208530, PubMed:26783301). Essential for the formation of PI3KC3-C2 but not PI3KC3-C1 PI3K complex forms. Involved in endocytosis (PubMed:25275521). Protects against infection by a neurovirulent strain of Sindbis virus (PubMed:9765397). May play a role in antiviral host defense. {ECO:0000269|PubMed:20208530, ECO:0000269|PubMed:20643123, ECO:0000269|PubMed:23184933, ECO:0000269|PubMed:25275521, ECO:0000269|PubMed:26783301, ECO:0000269|PubMed:28445460, ECO:0000269|PubMed:9765397, ECO:0000305}.; FUNCTION: Beclin-1-C 35 kDa localized to mitochondria can promote apoptosis; it induces the mitochondrial translocation of BAX and the release of proapoptotic factors. {ECO:0000269|PubMed:21364619, ECO:0000269|PubMed:26263979}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000357654 | - | 19 | 23 | 651_654 | 1759 | 1864.0 | Compositional bias | Note=Poly-Lys |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000468300 | - | 19 | 22 | 651_654 | 655 | 700.0 | Compositional bias | Note=Poly-Lys |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000491747 | - | 19 | 23 | 651_654 | 655 | 760.0 | Compositional bias | Note=Poly-Lys |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000357654 | - | 19 | 23 | 1642_1736 | 1759 | 1864.0 | Domain | BRCT 1 |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000352993 | - | 18 | 22 | 24_65 | 617 | 722.0 | Zinc finger | RING-type |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000357654 | - | 19 | 23 | 24_65 | 1759 | 1864.0 | Zinc finger | RING-type |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000468300 | - | 19 | 22 | 24_65 | 655 | 700.0 | Zinc finger | RING-type |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000491747 | - | 19 | 23 | 24_65 | 655 | 760.0 | Zinc finger | RING-type |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000361523 | 6 | 12 | 245_450 | 227 | 451.0 | Region | Evolutionary conserved domain (ECD) | |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000361523 | 6 | 12 | 425_450 | 227 | 451.0 | Region | Required for membrane-association | |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000590099 | 6 | 12 | 245_450 | 227 | 451.0 | Region | Evolutionary conserved domain (ECD) | |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000590099 | 6 | 12 | 425_450 | 227 | 451.0 | Region | Required for membrane-association |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000352993 | - | 18 | 22 | 651_654 | 617 | 722.0 | Compositional bias | Note=Poly-Lys |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000352993 | - | 18 | 22 | 1642_1736 | 617 | 722.0 | Domain | BRCT 1 |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000352993 | - | 18 | 22 | 1756_1855 | 617 | 722.0 | Domain | BRCT 2 |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000357654 | - | 19 | 23 | 1756_1855 | 1759 | 1864.0 | Domain | BRCT 2 |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000468300 | - | 19 | 22 | 1642_1736 | 655 | 700.0 | Domain | BRCT 1 |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000468300 | - | 19 | 22 | 1756_1855 | 655 | 700.0 | Domain | BRCT 2 |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000491747 | - | 19 | 23 | 1642_1736 | 655 | 760.0 | Domain | BRCT 1 |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000491747 | - | 19 | 23 | 1756_1855 | 655 | 760.0 | Domain | BRCT 2 |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000361523 | 6 | 12 | 142_270 | 227 | 451.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000590099 | 6 | 12 | 142_270 | 227 | 451.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000361523 | 6 | 12 | 108_127 | 227 | 451.0 | Motif | Note=BH3 | |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000590099 | 6 | 12 | 108_127 | 227 | 451.0 | Motif | Note=BH3 |
Top |
Fusion Gene Sequence for BRCA1-BECN1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >10149_10149_1_BRCA1-BECN1_BRCA1_chr17_41209069_ENST00000591534_BECN1_chr17_40968072_ENST00000361523_length(transcript)=2148nt_BP=852nt TTTCTCAGATAACTGGGCCCCTGCGCTCAGGAGGCCTTCACCCTCTGCTCTGGGTAAAGGTCATCCCCTTCTAAATGCCCATCATTAGAT GATAGGTGGTACATGCACAGTTGCTCTGGGAGTCTTCAGAATAGAAACTACCCATCTCAAGAGGAGCTCATTAAGGTTGTTGATGTGGAG GAGCAACAGCTGGAAGAGTCTGGGCCACACGATTTGACGGAAACATCTTACTTGCCAAGGCAAGATCTAGAGGGAACCCCTTACCTGGAA TCTGGAATCAGCCTCTTCTCTGATGACCCTGAATCTGATCCTTCTGAAGACAGAGCCCCAGAGTCAGCTCGTGTTGGCAACATACCATCT TCAACCTCTGCATTGAAAGTTCCCCAATTGAAAGTTGCAGAATCTGCCCAGAGTCCAGCTGCTGCTCATACTACTGATACTGCTGGGTAT AATGCAATGGAAGAAAGTGTGAGCAGGGAGAAGCCAGAATTGACAGCTTCAACAGAAAGGGTCAACAAAAGAATGTCCATGGTGGTGTCT GGCCTGACCCCAGAAGAATTTATGCTCGTGTACAAGTTTGCCAGAAAACACCACATCACTTTAACTAATCTAATTACTGAAGAGACTACT CATGTTGTTATGAAAACAGATGCTGAGTTTGTGTGTGAACGGACACTGAAATATTTTCTAGGAATTGCGGGAGGAAAATGGGTAGTTAGC TATTTCTGGGTGACCCAGTCTATTAAAGAAAGAAAAATGCTGAATGAGCATGATTTTGAAGTCAGAGGAGATGTGGTCAATGGAAGAAAC CACCAAGGTCCAAAGCGAGCAAGAGAATCCCAGGACAGAAAGGTATCAGAGAGAATACAGTGAATTTAAACGACAGCAGCTGGAGCTGGA TGATGAGCTGAAGAGTGTTGAAAACCAGATGCGTTATGCCCAGACGCAGCTGGATAAGCTGAAGAAAACCAACGTCTTTAATGCAACCTT CCACATCTGGCACAGTGGACAGTTTGGCACAATCAATAACTTCAGGCTGGGTCGCCTGCCCAGTGTTCCCGTGGAATGGAATGAGATTAA TGCTGCTTGGGGCCAGACTGTGTTGCTGCTCCATGCTCTGGCCAATAAGATGGGTCTGAAATTTCAGAGATACCGACTTGTTCCTTACGG AAACCATTCATATCTGGAGTCTCTGACAGACAAATCTAAGGAGCTGCCGTTATACTGTTCTGGGGGGTTGCGGTTTTTCTGGGACAACAA GTTTGACCATGCAATGGTGGCTTTCCTGGACTGTGTGCAGCAGTTCAAAGAAGAGGTTGAGAAAGGCGAGACACGTTTTTGTCTTCCCTA CAGGATGGATGTGGAGAAAGGCAAGATTGAAGACACAGGAGGCAGTGGCGGCTCCTATTCCATCAAAACCCAGTTTAACTCTGAGGAGCA GTGGACAAAAGCTCTCAAGTTCATGCTGACGAATCTTAAGTGGGGTCTTGCTTGGGTGTCCTCACAATTTTATAACAAATGACTTTTTTC CTTAGGGGGAGGTTTGCCTTAAAGGCTTTTAATTTTGTTTTGTTTGCAAACATGTTTTAAATTAAATTCGGGTAATATTAAACAGTACAT GTTTACAATACCAAAAAAGAAAAAATCCACAAAAGCCACTTTATTTTAAAATATCATGTGACAGATACTTTCCAGAGCTACAACATGCCA TCTATAGTTGCCAGCCCTGGTCAGTTTTGATTCTTAACCCCATGGACTCCTTTCCCTTTCTTCTCTGAAAAAAACTAATTTAAATTTGCT TTTCTTTTTTTTAACTGAGTTGAATTGAGATTGATGTGTTTTCACTGGATTTTTATCTCTCTCAACTTCCTGCACTTAACAATATGAAAT AGAAACTTTTGTCTTTACTGAGATGAGGATATGTTTGAGATGCACAGTTGGATAATGTGGGAAAATGACATCTAAGCTTTACCTGGTCAC CATGTGATGTGATCAGATGCTTGAAATTTAACACTTTTCACTTGGTTCTTATACTGAATGCCGACTCTGCTCTGTGTTAGAGATATGAAA >10149_10149_1_BRCA1-BECN1_BRCA1_chr17_41209069_ENST00000591534_BECN1_chr17_40968072_ENST00000361523_length(amino acids)=256AA_BP= MHSCSGSLQNRNYPSQEELIKVVDVEEQQLEESGPHDLTETSYLPRQDLEGTPYLESGISLFSDDPESDPSEDRAPESARVGNIPSSTSA LKVPQLKVAESAQSPAAAHTTDTAGYNAMEESVSREKPELTASTERVNKRMSMVVSGLTPEEFMLVYKFARKHHITLTNLITEETTHVVM -------------------------------------------------------------- >10149_10149_2_BRCA1-BECN1_BRCA1_chr17_41209069_ENST00000591534_BECN1_chr17_40968072_ENST00000438274_length(transcript)=1831nt_BP=852nt TTTCTCAGATAACTGGGCCCCTGCGCTCAGGAGGCCTTCACCCTCTGCTCTGGGTAAAGGTCATCCCCTTCTAAATGCCCATCATTAGAT GATAGGTGGTACATGCACAGTTGCTCTGGGAGTCTTCAGAATAGAAACTACCCATCTCAAGAGGAGCTCATTAAGGTTGTTGATGTGGAG GAGCAACAGCTGGAAGAGTCTGGGCCACACGATTTGACGGAAACATCTTACTTGCCAAGGCAAGATCTAGAGGGAACCCCTTACCTGGAA TCTGGAATCAGCCTCTTCTCTGATGACCCTGAATCTGATCCTTCTGAAGACAGAGCCCCAGAGTCAGCTCGTGTTGGCAACATACCATCT TCAACCTCTGCATTGAAAGTTCCCCAATTGAAAGTTGCAGAATCTGCCCAGAGTCCAGCTGCTGCTCATACTACTGATACTGCTGGGTAT AATGCAATGGAAGAAAGTGTGAGCAGGGAGAAGCCAGAATTGACAGCTTCAACAGAAAGGGTCAACAAAAGAATGTCCATGGTGGTGTCT GGCCTGACCCCAGAAGAATTTATGCTCGTGTACAAGTTTGCCAGAAAACACCACATCACTTTAACTAATCTAATTACTGAAGAGACTACT CATGTTGTTATGAAAACAGATGCTGAGTTTGTGTGTGAACGGACACTGAAATATTTTCTAGGAATTGCGGGAGGAAAATGGGTAGTTAGC TATTTCTGGGTGACCCAGTCTATTAAAGAAAGAAAAATGCTGAATGAGCATGATTTTGAAGTCAGAGGAGATGTGGTCAATGGAAGAAAC CACCAAGGTCCAAAGCGAGCAAGAGAATCCCAGGACAGAAAGGTATCAGAGAGAATACAGTGAATTTAAACGACAGCAGCTGGAGCTGGA TGATGAGCTGAAGAGTGTTGAAAACCAGATGCGTTATGCCCAGACGCAGCTGGATAAGCTGAAGAAAACCAACGTCTTTAATGCAACCTT CCACATCTGGCACAGTGGACAGTTTGGCACAATCAATAACTTCAGGCTGGGTCGCCTGCCCAGTGTTCCCGTGGAATGGAATGAGATTAA TGCTGCTTGGGGCCAGACTGTGTTGCTGCTCCATGCTCTGGCCAATAAGATGGGTCTGAAATTTCAGAGATACCGACTTGTTCCTTACGG AAACCATTCATATCTGGAGTCTCTGACAGACAAATCTAAGGATGGATGTGGAGAAAGGCAAGATTGAAGACACAGGAGGCAGTGGCGGCT CCTATTCCATCAAAACCCAGTTTAACTCTGAGGAGCAGTGGACAAAAGCTCTCAAGTTCATGCTGACGAATCTTAAGTGGGGTCTTGCTT GGGTGTCCTCACAATTTTATAACAAATGACTTTTTTCCTTAGGGGGAGGTTTGCCTTAAAGGCTTTTAATTTTGTTTTGTTTGCAAACAT GTTTTAAATTAAATTCGGGTAATATTAAACAGTACATGTTTACAATACCAAAAAAGAAAAAATCCACAAAAGCCACTTTATTTTAAAATA TCATGTGACAGATACTTTCCAGAGCTACAACATGCCATCTATAGTTGCCAGCCCTGGTCAGTTTTGATTCTTAACCCCATGGACTCCTTT CCCTTTCTTCTCTGAAAAAAACTAATTTAAATTTGCTTTTCTTTTTTTTAACTGAGTTGAATTGAGATTGATGTGTTTTCACTGGATTTT TATCTCTCTCAACTTCCTGCACTTAACAATATGAAATAGAAACTTTTGTCTTTACTGAGATGAGGATATGTTTGAGATGCACAGTTGGAT >10149_10149_2_BRCA1-BECN1_BRCA1_chr17_41209069_ENST00000591534_BECN1_chr17_40968072_ENST00000438274_length(amino acids)=256AA_BP= MHSCSGSLQNRNYPSQEELIKVVDVEEQQLEESGPHDLTETSYLPRQDLEGTPYLESGISLFSDDPESDPSEDRAPESARVGNIPSSTSA LKVPQLKVAESAQSPAAAHTTDTAGYNAMEESVSREKPELTASTERVNKRMSMVVSGLTPEEFMLVYKFARKHHITLTNLITEETTHVVM -------------------------------------------------------------- >10149_10149_3_BRCA1-BECN1_BRCA1_chr17_41209069_ENST00000591534_BECN1_chr17_40968072_ENST00000590099_length(transcript)=2145nt_BP=852nt TTTCTCAGATAACTGGGCCCCTGCGCTCAGGAGGCCTTCACCCTCTGCTCTGGGTAAAGGTCATCCCCTTCTAAATGCCCATCATTAGAT GATAGGTGGTACATGCACAGTTGCTCTGGGAGTCTTCAGAATAGAAACTACCCATCTCAAGAGGAGCTCATTAAGGTTGTTGATGTGGAG GAGCAACAGCTGGAAGAGTCTGGGCCACACGATTTGACGGAAACATCTTACTTGCCAAGGCAAGATCTAGAGGGAACCCCTTACCTGGAA TCTGGAATCAGCCTCTTCTCTGATGACCCTGAATCTGATCCTTCTGAAGACAGAGCCCCAGAGTCAGCTCGTGTTGGCAACATACCATCT TCAACCTCTGCATTGAAAGTTCCCCAATTGAAAGTTGCAGAATCTGCCCAGAGTCCAGCTGCTGCTCATACTACTGATACTGCTGGGTAT AATGCAATGGAAGAAAGTGTGAGCAGGGAGAAGCCAGAATTGACAGCTTCAACAGAAAGGGTCAACAAAAGAATGTCCATGGTGGTGTCT GGCCTGACCCCAGAAGAATTTATGCTCGTGTACAAGTTTGCCAGAAAACACCACATCACTTTAACTAATCTAATTACTGAAGAGACTACT CATGTTGTTATGAAAACAGATGCTGAGTTTGTGTGTGAACGGACACTGAAATATTTTCTAGGAATTGCGGGAGGAAAATGGGTAGTTAGC TATTTCTGGGTGACCCAGTCTATTAAAGAAAGAAAAATGCTGAATGAGCATGATTTTGAAGTCAGAGGAGATGTGGTCAATGGAAGAAAC CACCAAGGTCCAAAGCGAGCAAGAGAATCCCAGGACAGAAAGGTATCAGAGAGAATACAGTGAATTTAAACGACAGCAGCTGGAGCTGGA TGATGAGCTGAAGAGTGTTGAAAACCAGATGCGTTATGCCCAGACGCAGCTGGATAAGCTGAAGAAAACCAACGTCTTTAATGCAACCTT CCACATCTGGCACAGTGGACAGTTTGGCACAATCAATAACTTCAGGCTGGGTCGCCTGCCCAGTGTTCCCGTGGAATGGAATGAGATTAA TGCTGCTTGGGGCCAGACTGTGTTGCTGCTCCATGCTCTGGCCAATAAGATGGGTCTGAAATTTCAGAGATACCGACTTGTTCCTTACGG AAACCATTCATATCTGGAGTCTCTGACAGACAAATCTAAGGAGCTGCCGTTATACTGTTCTGGGGGGTTGCGGTTTTTCTGGGACAACAA GTTTGACCATGCAATGGTGGCTTTCCTGGACTGTGTGCAGCAGTTCAAAGAAGAGGTTGAGAAAGGCGAGACACGTTTTTGTCTTCCCTA CAGGATGGATGTGGAGAAAGGCAAGATTGAAGACACAGGAGGCAGTGGCGGCTCCTATTCCATCAAAACCCAGTTTAACTCTGAGGAGCA GTGGACAAAAGCTCTCAAGTTCATGCTGACGAATCTTAAGTGGGGTCTTGCTTGGGTGTCCTCACAATTTTATAACAAATGACTTTTTTC CTTAGGGGGAGGTTTGCCTTAAAGGCTTTTAATTTTGTTTTGTTTGCAAACATGTTTTAAATTAAATTCGGGTAATATTAAACAGTACAT GTTTACAATACCAAAAAAGAAAAAATCCACAAAAGCCACTTTATTTTAAAATATCATGTGACAGATACTTTCCAGAGCTACAACATGCCA TCTATAGTTGCCAGCCCTGGTCAGTTTTGATTCTTAACCCCATGGACTCCTTTCCCTTTCTTCTCTGAAAAAAACTAATTTAAATTTGCT TTTCTTTTTTTTAACTGAGTTGAATTGAGATTGATGTGTTTTCACTGGATTTTTATCTCTCTCAACTTCCTGCACTTAACAATATGAAAT AGAAACTTTTGTCTTTACTGAGATGAGGATATGTTTGAGATGCACAGTTGGATAATGTGGGAAAATGACATCTAAGCTTTACCTGGTCAC CATGTGATGTGATCAGATGCTTGAAATTTAACACTTTTCACTTGGTTCTTATACTGAATGCCGACTCTGCTCTGTGTTAGAGATATGAAA >10149_10149_3_BRCA1-BECN1_BRCA1_chr17_41209069_ENST00000591534_BECN1_chr17_40968072_ENST00000590099_length(amino acids)=256AA_BP= MHSCSGSLQNRNYPSQEELIKVVDVEEQQLEESGPHDLTETSYLPRQDLEGTPYLESGISLFSDDPESDPSEDRAPESARVGNIPSSTSA LKVPQLKVAESAQSPAAAHTTDTAGYNAMEESVSREKPELTASTERVNKRMSMVVSGLTPEEFMLVYKFARKHHITLTNLITEETTHVVM -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for BRCA1-BECN1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000357654 | - | 19 | 23 | 1397_1424 | 1759.0 | 1864.0 | PALB2 |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000352993 | - | 18 | 22 | 1397_1424 | 617.0 | 722.0 | PALB2 |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000468300 | - | 19 | 22 | 1397_1424 | 655.0 | 700.0 | PALB2 |
| Hgene | BRCA1 | chr17:41209069 | chr17:40968072 | ENST00000491747 | - | 19 | 23 | 1397_1424 | 655.0 | 760.0 | PALB2 |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000361523 | 6 | 12 | 112_159 | 227.66666666666666 | 451.0 | BCL2 and BCL2L1 isoform Bcl-X(L) | |
| Tgene | BECN1 | chr17:41209069 | chr17:40968072 | ENST00000590099 | 6 | 12 | 112_159 | 227.66666666666666 | 451.0 | BCL2 and BCL2L1 isoform Bcl-X(L) |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for BRCA1-BECN1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for BRCA1-BECN1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
