
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ZNF83-PPP2R1A (FusionGDB2 ID:103051) |
Fusion Gene Summary for ZNF83-PPP2R1A |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ZNF83-PPP2R1A | Fusion gene ID: 103051 | Hgene | Tgene | Gene symbol | ZNF83 | PPP2R1A | Gene ID | 55769 | 5518 |
| Gene name | zinc finger protein 83 | protein phosphatase 2 scaffold subunit Aalpha | |
| Synonyms | HPF1|ZNF816B | MRD36|PP2A-Aalpha|PP2AA|PP2AAALPHA|PR65A | |
| Cytomap | 19q13.41 | 19q13.41 | |
| Type of gene | protein-coding | protein-coding | |
| Description | zinc finger protein 83zinc finger protein 816Bzinc finger protein HPF1 | serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoformPP2A subunit A isoform PR65-alphaPP2A subunit A isoform R1-alphamedium tumor antigen-associated 61 KDA proteinprotein phosphatase 2 (formerly 2A), regulatory subunit A (P | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | . | |
| Ensembl transtripts involved in fusion gene | ENST00000594682, ENST00000596930, ENST00000597161, ENST00000598536, ENST00000600714, ENST00000601257, ENST00000301096, ENST00000536937, ENST00000544146, ENST00000545872, ENST00000597597, ENST00000391789, ENST00000541777, ENST00000600457, | ENST00000473455, ENST00000462990, ENST00000477989, ENST00000322088, ENST00000444322, | |
| Fusion gene scores | * DoF score | 11 X 9 X 5=495 | 10 X 12 X 4=480 |
| # samples | 12 | 12 | |
| ** MAII score | log2(12/495*10)=-2.04439411935845 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(12/480*10)=-2 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: ZNF83 [Title/Abstract] AND PPP2R1A [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ZNF83(53138318)-PPP2R1A(52705196), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | ZNF83-PPP2R1A seems lost the major protein functional domain in Hgene partner, which is a transcription factor due to the frame-shifted ORF. ZNF83-PPP2R1A seems lost the major protein functional domain in Tgene partner, which is a cell metabolism gene due to the frame-shifted ORF. ZNF83-PPP2R1A seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. ZNF83-PPP2R1A seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | PPP2R1A | GO:0007059 | chromosome segregation | 16580887 |
 Fusion gene breakpoints across ZNF83 (5'-gene) Fusion gene breakpoints across ZNF83 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
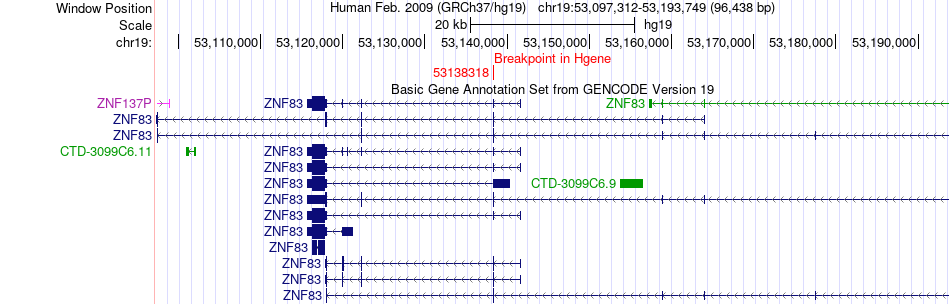 |
 Fusion gene breakpoints across PPP2R1A (3'-gene) Fusion gene breakpoints across PPP2R1A (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
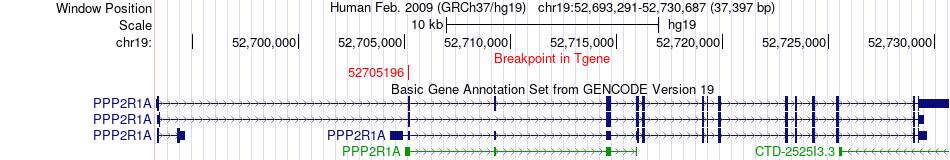 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-BR-A4PD | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
Top |
Fusion Gene ORF analysis for ZNF83-PPP2R1A |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000594682 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-3UTR | ENST00000596930 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-3UTR | ENST00000597161 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-3UTR | ENST00000598536 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-3UTR | ENST00000600714 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-3UTR | ENST00000601257 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-5UTR | ENST00000594682 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-5UTR | ENST00000596930 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-5UTR | ENST00000597161 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-5UTR | ENST00000598536 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-5UTR | ENST00000600714 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-5UTR | ENST00000601257 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-intron | ENST00000594682 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-intron | ENST00000596930 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-intron | ENST00000597161 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-intron | ENST00000598536 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-intron | ENST00000600714 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5CDS-intron | ENST00000601257 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000301096 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000301096 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000536937 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000536937 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000544146 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000544146 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000545872 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000545872 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000597597 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3CDS | ENST00000597597 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3UTR | ENST00000301096 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3UTR | ENST00000536937 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3UTR | ENST00000544146 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3UTR | ENST00000545872 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-3UTR | ENST00000597597 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-5UTR | ENST00000301096 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-5UTR | ENST00000536937 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-5UTR | ENST00000544146 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-5UTR | ENST00000545872 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-5UTR | ENST00000597597 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-intron | ENST00000301096 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-intron | ENST00000536937 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-intron | ENST00000544146 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-intron | ENST00000545872 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| 5UTR-intron | ENST00000597597 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| Frame-shift | ENST00000594682 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| Frame-shift | ENST00000596930 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| Frame-shift | ENST00000596930 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| Frame-shift | ENST00000597161 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| Frame-shift | ENST00000597161 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| Frame-shift | ENST00000598536 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| Frame-shift | ENST00000600714 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| Frame-shift | ENST00000600714 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| Frame-shift | ENST00000601257 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| In-frame | ENST00000594682 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| In-frame | ENST00000598536 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| In-frame | ENST00000601257 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-3CDS | ENST00000391789 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-3CDS | ENST00000391789 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-3CDS | ENST00000541777 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-3CDS | ENST00000541777 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-3CDS | ENST00000600457 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-3CDS | ENST00000600457 | ENST00000444322 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-3UTR | ENST00000391789 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-3UTR | ENST00000541777 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-3UTR | ENST00000600457 | ENST00000473455 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-5UTR | ENST00000391789 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-5UTR | ENST00000541777 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-5UTR | ENST00000600457 | ENST00000462990 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-intron | ENST00000391789 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-intron | ENST00000541777 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
| intron-intron | ENST00000600457 | ENST00000477989 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000601257 | ZNF83 | chr19 | 53138318 | - | ENST00000322088 | PPP2R1A | chr19 | 52705196 | + | 3568 | 423 | 354 | 2114 | 586 |
| ENST00000594682 | ZNF83 | chr19 | 53138318 | - | ENST00000322088 | PPP2R1A | chr19 | 52705196 | + | 3488 | 343 | 274 | 2034 | 586 |
| ENST00000598536 | ZNF83 | chr19 | 53138318 | - | ENST00000322088 | PPP2R1A | chr19 | 52705196 | + | 3558 | 413 | 344 | 2104 | 586 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000601257 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + | 0.001589841 | 0.9984101 |
| ENST00000594682 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + | 0.001672826 | 0.9983272 |
| ENST00000598536 | ENST00000322088 | ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + | 0.001568646 | 0.9984314 |
Top |
Fusion Genomic Features for ZNF83-PPP2R1A |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + | 0.003720762 | 0.9962792 |
| ZNF83 | chr19 | 53138318 | - | PPP2R1A | chr19 | 52705196 | + | 0.003720762 | 0.9962792 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
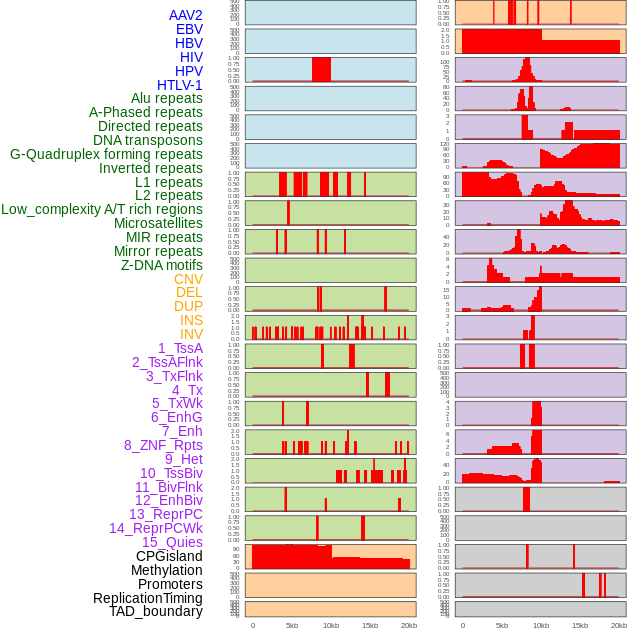 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
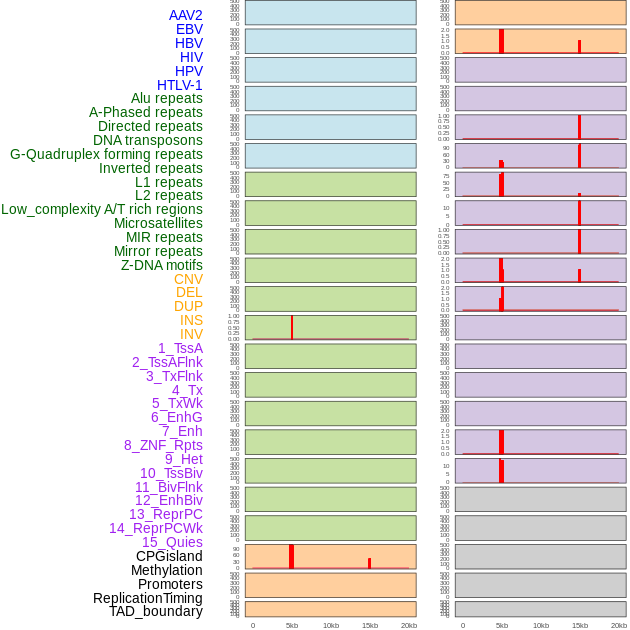 |
Top |
Fusion Protein Features for ZNF83-PPP2R1A |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:53138318/chr19:52705196) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | . |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 400_589 | 26 | 590.0 | Region | Note=PP2A subunit C binding | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 47_321 | 26 | 590.0 | Region | Note=Polyoma small and medium T antigens Binding | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 85_239 | 26 | 590.0 | Region | Note=SV40 small T antigen binding | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 124_161 | 26 | 590.0 | Repeat | Note=HEAT 4 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 162_200 | 26 | 590.0 | Repeat | Note=HEAT 5 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 201_239 | 26 | 590.0 | Repeat | Note=HEAT 6 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 240_278 | 26 | 590.0 | Repeat | Note=HEAT 7 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 279_321 | 26 | 590.0 | Repeat | Note=HEAT 8 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 322_360 | 26 | 590.0 | Repeat | Note=HEAT 9 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 361_399 | 26 | 590.0 | Repeat | Note=HEAT 10 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 400_438 | 26 | 590.0 | Repeat | Note=HEAT 11 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 439_477 | 26 | 590.0 | Repeat | Note=HEAT 12 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 478_516 | 26 | 590.0 | Repeat | Note=HEAT 13 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 47_84 | 26 | 590.0 | Repeat | Note=HEAT 2 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 517_555 | 26 | 590.0 | Repeat | Note=HEAT 14 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 556_589 | 26 | 590.0 | Repeat | Note=HEAT 15 | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 85_123 | 26 | 590.0 | Repeat | Note=HEAT 3 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 121_143 | 0 | 517.0 | Zinc finger | C2H2-type 2 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 149_171 | 0 | 517.0 | Zinc finger | C2H2-type 3 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 177_199 | 0 | 517.0 | Zinc finger | C2H2-type 4 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 205_227 | 0 | 517.0 | Zinc finger | C2H2-type 5 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 233_255 | 0 | 517.0 | Zinc finger | C2H2-type 6 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 261_283 | 0 | 517.0 | Zinc finger | C2H2-type 7 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 289_311 | 0 | 517.0 | Zinc finger | C2H2-type 8 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 317_339 | 0 | 517.0 | Zinc finger | C2H2-type 9 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 345_367 | 0 | 517.0 | Zinc finger | C2H2-type 10 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 373_395 | 0 | 517.0 | Zinc finger | C2H2-type 11 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 401_423 | 0 | 517.0 | Zinc finger | C2H2-type 12 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 429_451 | 0 | 517.0 | Zinc finger | C2H2-type 13 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 457_479 | 0 | 517.0 | Zinc finger | C2H2-type 14 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 485_507 | 0 | 517.0 | Zinc finger | C2H2-type 15 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000301096 | - | 2 | 3 | 93_115 | 0 | 517.0 | Zinc finger | C2H2-type 1%3B degenerate |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 121_143 | 0 | 489.0 | Zinc finger | C2H2-type 2 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 149_171 | 0 | 489.0 | Zinc finger | C2H2-type 3 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 177_199 | 0 | 489.0 | Zinc finger | C2H2-type 4 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 205_227 | 0 | 489.0 | Zinc finger | C2H2-type 5 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 233_255 | 0 | 489.0 | Zinc finger | C2H2-type 6 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 261_283 | 0 | 489.0 | Zinc finger | C2H2-type 7 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 289_311 | 0 | 489.0 | Zinc finger | C2H2-type 8 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 317_339 | 0 | 489.0 | Zinc finger | C2H2-type 9 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 345_367 | 0 | 489.0 | Zinc finger | C2H2-type 10 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 373_395 | 0 | 489.0 | Zinc finger | C2H2-type 11 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 401_423 | 0 | 489.0 | Zinc finger | C2H2-type 12 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 429_451 | 0 | 489.0 | Zinc finger | C2H2-type 13 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 457_479 | 0 | 489.0 | Zinc finger | C2H2-type 14 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 485_507 | 0 | 489.0 | Zinc finger | C2H2-type 15 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000391789 | - | 1 | 2 | 93_115 | 0 | 489.0 | Zinc finger | C2H2-type 1%3B degenerate |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 121_143 | 0 | 517.0 | Zinc finger | C2H2-type 2 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 149_171 | 0 | 517.0 | Zinc finger | C2H2-type 3 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 177_199 | 0 | 517.0 | Zinc finger | C2H2-type 4 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 205_227 | 0 | 517.0 | Zinc finger | C2H2-type 5 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 233_255 | 0 | 517.0 | Zinc finger | C2H2-type 6 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 261_283 | 0 | 517.0 | Zinc finger | C2H2-type 7 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 289_311 | 0 | 517.0 | Zinc finger | C2H2-type 8 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 317_339 | 0 | 517.0 | Zinc finger | C2H2-type 9 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 345_367 | 0 | 517.0 | Zinc finger | C2H2-type 10 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 373_395 | 0 | 517.0 | Zinc finger | C2H2-type 11 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 401_423 | 0 | 517.0 | Zinc finger | C2H2-type 12 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 429_451 | 0 | 517.0 | Zinc finger | C2H2-type 13 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 457_479 | 0 | 517.0 | Zinc finger | C2H2-type 14 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 485_507 | 0 | 517.0 | Zinc finger | C2H2-type 15 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000536937 | - | 2 | 5 | 93_115 | 0 | 517.0 | Zinc finger | C2H2-type 1%3B degenerate |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 121_143 | 0 | 517.0 | Zinc finger | C2H2-type 2 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 149_171 | 0 | 517.0 | Zinc finger | C2H2-type 3 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 177_199 | 0 | 517.0 | Zinc finger | C2H2-type 4 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 205_227 | 0 | 517.0 | Zinc finger | C2H2-type 5 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 233_255 | 0 | 517.0 | Zinc finger | C2H2-type 6 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 261_283 | 0 | 517.0 | Zinc finger | C2H2-type 7 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 289_311 | 0 | 517.0 | Zinc finger | C2H2-type 8 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 317_339 | 0 | 517.0 | Zinc finger | C2H2-type 9 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 345_367 | 0 | 517.0 | Zinc finger | C2H2-type 10 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 373_395 | 0 | 517.0 | Zinc finger | C2H2-type 11 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 401_423 | 0 | 517.0 | Zinc finger | C2H2-type 12 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 429_451 | 0 | 517.0 | Zinc finger | C2H2-type 13 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 457_479 | 0 | 517.0 | Zinc finger | C2H2-type 14 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 485_507 | 0 | 517.0 | Zinc finger | C2H2-type 15 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000541777 | - | 1 | 2 | 93_115 | 0 | 517.0 | Zinc finger | C2H2-type 1%3B degenerate |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 121_143 | 0 | 517.0 | Zinc finger | C2H2-type 2 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 149_171 | 0 | 517.0 | Zinc finger | C2H2-type 3 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 177_199 | 0 | 517.0 | Zinc finger | C2H2-type 4 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 205_227 | 0 | 517.0 | Zinc finger | C2H2-type 5 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 233_255 | 0 | 517.0 | Zinc finger | C2H2-type 6 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 261_283 | 0 | 517.0 | Zinc finger | C2H2-type 7 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 289_311 | 0 | 517.0 | Zinc finger | C2H2-type 8 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 317_339 | 0 | 517.0 | Zinc finger | C2H2-type 9 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 345_367 | 0 | 517.0 | Zinc finger | C2H2-type 10 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 373_395 | 0 | 517.0 | Zinc finger | C2H2-type 11 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 401_423 | 0 | 517.0 | Zinc finger | C2H2-type 12 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 429_451 | 0 | 517.0 | Zinc finger | C2H2-type 13 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 457_479 | 0 | 517.0 | Zinc finger | C2H2-type 14 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 485_507 | 0 | 517.0 | Zinc finger | C2H2-type 15 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000544146 | - | 2 | 6 | 93_115 | 0 | 517.0 | Zinc finger | C2H2-type 1%3B degenerate |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 121_143 | 0 | 517.0 | Zinc finger | C2H2-type 2 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 149_171 | 0 | 517.0 | Zinc finger | C2H2-type 3 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 177_199 | 0 | 517.0 | Zinc finger | C2H2-type 4 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 205_227 | 0 | 517.0 | Zinc finger | C2H2-type 5 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 233_255 | 0 | 517.0 | Zinc finger | C2H2-type 6 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 261_283 | 0 | 517.0 | Zinc finger | C2H2-type 7 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 289_311 | 0 | 517.0 | Zinc finger | C2H2-type 8 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 317_339 | 0 | 517.0 | Zinc finger | C2H2-type 9 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 345_367 | 0 | 517.0 | Zinc finger | C2H2-type 10 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 373_395 | 0 | 517.0 | Zinc finger | C2H2-type 11 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 401_423 | 0 | 517.0 | Zinc finger | C2H2-type 12 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 429_451 | 0 | 517.0 | Zinc finger | C2H2-type 13 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 457_479 | 0 | 517.0 | Zinc finger | C2H2-type 14 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 485_507 | 0 | 517.0 | Zinc finger | C2H2-type 15 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000545872 | - | 2 | 3 | 93_115 | 0 | 517.0 | Zinc finger | C2H2-type 1%3B degenerate |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 121_143 | 0 | 517.0 | Zinc finger | C2H2-type 2 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 149_171 | 0 | 517.0 | Zinc finger | C2H2-type 3 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 177_199 | 0 | 517.0 | Zinc finger | C2H2-type 4 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 205_227 | 0 | 517.0 | Zinc finger | C2H2-type 5 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 233_255 | 0 | 517.0 | Zinc finger | C2H2-type 6 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 261_283 | 0 | 517.0 | Zinc finger | C2H2-type 7 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 289_311 | 0 | 517.0 | Zinc finger | C2H2-type 8 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 317_339 | 0 | 517.0 | Zinc finger | C2H2-type 9 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 345_367 | 0 | 517.0 | Zinc finger | C2H2-type 10 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 373_395 | 0 | 517.0 | Zinc finger | C2H2-type 11 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 401_423 | 0 | 517.0 | Zinc finger | C2H2-type 12 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 429_451 | 0 | 517.0 | Zinc finger | C2H2-type 13 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 457_479 | 0 | 517.0 | Zinc finger | C2H2-type 14 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 485_507 | 0 | 517.0 | Zinc finger | C2H2-type 15 |
| Hgene | ZNF83 | chr19:53138318 | chr19:52705196 | ENST00000597597 | - | 1 | 2 | 93_115 | 0 | 517.0 | Zinc finger | C2H2-type 1%3B degenerate |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 8_399 | 26 | 590.0 | Region | Note=PP2A subunit B binding | |
| Tgene | PPP2R1A | chr19:53138318 | chr19:52705196 | ENST00000322088 | 0 | 15 | 8_46 | 26 | 590.0 | Repeat | Note=HEAT 1 |
Top |
Fusion Gene Sequence for ZNF83-PPP2R1A |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >103051_103051_1_ZNF83-PPP2R1A_ZNF83_chr19_53138318_ENST00000594682_PPP2R1A_chr19_52705196_ENST00000322088_length(transcript)=3488nt_BP=343nt GCAAATCTGGAAGCGGATCGCGTGGAGTGACGGTCACACCGCGGCGAATTAATTTCCAAAGACTCATGTTACATGAGAAAGCCACCAAGA AGACCAAAGAAAAGGAGACAAGGATGGCTCTTCCTCAGGGATGCTTGACTTTCAGGGATGTGGCTATAGAATTCTCTTTGGAGGAGTGGA AATGCCTGAACCCTGCACAGAGGGCTTTATACAGGGCCGTGATGTTGGAGAACTACAGGAACCTGGAGTCTGTGGGATTGACTTCTAAGG ACTCTTGGTACATGAGGAAGAAACCCGGAAGGGGAAGAGGAAAGCAAAGGCGTCAGGAATGGTTCTTCCTCAGCTTCGCCTCAACAGCAT CAAGAAGCTGTCCACCATCGCCTTGGCCCTTGGGGTTGAAAGGACCCGAAGTGAGCTTCTGCCTTTCCTTACAGATACCATCTATGATGA AGATGAGGTCCTCCTGGCCCTGGCAGAACAGCTGGGAACCTTCACTACCCTGGTGGGAGGCCCAGAGTACGTGCACTGCCTGCTGCCACC GCTGGAGTCGCTGGCCACAGTGGAGGAGACAGTGGTGCGGGACAAGGCAGTGGAGTCCTTACGGGCCATCTCACACGAGCACTCGCCCTC TGACCTGGAGGCGCACTTTGTGCCGCTAGTGAAGCGGCTGGCGGGCGGCGACTGGTTCACCTCCCGCACCTCGGCCTGCGGCCTCTTCTC CGTCTGCTACCCCCGAGTGTCCAGTGCTGTGAAGGCGGAACTTCGACAGTACTTCCGGAACCTGTGCTCAGATGACACCCCCATGGTGCG GCGGGCCGCAGCCTCCAAGCTGGGGGAGTTTGCCAAGGTGCTGGAGCTGGACAACGTCAAGAGTGAGATCATCCCCATGTTCTCCAACCT GGCCTCTGACGAGCAGGACTCGGTGCGGCTGCTGGCGGTGGAGGCGTGCGTGAACATCGCCCAGCTTCTGCCCCAGGAGGATCTGGAGGC CCTGGTGATGCCCACTCTGCGCCAGGCCGCTGAAGACAAGTCCTGGCGCGTCCGCTACATGGTGGCTGACAAGTTCACAGAGCTCCAGAA AGCAGTGGGGCCTGAGATCACCAAGACAGACCTGGTCCCTGCCTTCCAGAACCTGATGAAAGACTGTGAGGCCGAGGTGAGGGCCGCAGC CTCCCACAAGGTCAAAGAGTTCTGTGAAAACCTCTCAGCTGACTGTCGGGAGAATGTGATCATGTCCCAGATCTTGCCCTGCATCAAGGA GCTGGTGTCCGATGCCAACCAACATGTCAAGTCTGCCCTGGCCTCAGTCATCATGGGTCTCTCTCCCATCTTGGGCAAAGACAACACCAT CGAGCACCTCTTGCCCCTCTTCCTGGCTCAGCTGAAGGATGAGTGCCCTGAGGTACGGCTGAACATCATCTCTAACCTGGACTGTGTGAA CGAGGTGATTGGCATCCGGCAGCTGTCCCAGTCCCTGCTCCCTGCCATTGTGGAGCTGGCTGAGGACGCCAAGTGGCGGGTGCGGCTGGC CATCATTGAGTACATGCCCCTCCTGGCTGGACAGCTGGGAGTGGAGTTCTTTGATGAGAAACTTAACTCCTTGTGCATGGCCTGGCTTGT GGATCATGTATATGCCATCCGCGAGGCAGCCACCAGCAACCTGAAGAAGCTAGTGGAAAAGTTTGGGAAGGAGTGGGCCCATGCCACAAT CATCCCCAAGGTCTTGGCCATGTCCGGAGACCCCAACTACCTGCACCGCATGACTACGCTCTTCTGCATCAATGTGCTGTCTGAGGTCTG TGGGCAGGACATCACCACCAAGCACATGCTACCCACGGTTCTGCGCATGGCTGGGGACCCGGTTGCCAATGTCCGCTTCAATGTGGCCAA GTCTCTGCAGAAGATAGGGCCCATCCTGGACAACAGCACCTTGCAGAGTGAAGTCAAGCCCATCCTAGAGAAGCTGACCCAGGACCAGGA TGTGGACGTCAAATACTTTGCCCAGGAGGCTCTGACTGTTCTGTCTCTCGCCTGATGCTGGAAGAGGAGCAAACACTGGCCTCTGGTGTC CACCCTCCAACCCCCACAAGTCCCTCTTTGGGGAGACACTGGGGGGCCTTTGGCTGTCACTCCCTGTGCATGGTCTGACCCCAGGCCCCT TCCCCCAGCACGGTTCCTCCTCTCCCCAGCCTGGGAAGATGTCTCACTGTCCACCTCCCAACGGGCTAGGGGAGCACGGGGTTGGACAGG ACAGTGACCTTGGGAGGAAGGGGCTACTCCGCCCACGTCAGGGAGAGATGTGAGCATCCCGGGTCACTGGATCCTGCTGCTGTAATGGGA ACCCCTCCCCCATTTACTTCTCCACCTCCCGTCCTCCCCATCATTGGTTTTTTTTTGTGTGTCAACTGTGCCGTTTTTATTTTATTCCTT TTATTTTCCCCCTTTTCACAGAGAAATAAAGGTCTAGAAGTAGTTGGTCCTCTGGCCTTAGTCATCAGCTTGGAGGAGGGGGCACCACCC CAGAGCCACAACATCTGCGCTTTTCTCTGGAGAAGATCTTGTTACAGGACCCATTATACCCCTATGTCCCGAAAGAATACTCTGGGGATC CCCCCAGGAGTGCTGCTGGCCTTTGGGGTAGAGGGTCCATGAGGTGCTCTGGGTGGTGTCCTGTAGTGCAGTTCAGATTCATAGATGTCG GCCGAGTATTTGGGGGGTTAAGAGCAGGTGCCTTGGAACCAGACTGCCTGGGTCCAAATTGTGGCTCCTTCAGTTAATAGACGTGTGACT AGGAGTAGCTTGTTAAGTCTTTATGAGCTCAGTTTTGTATTGTGTAAAATGAGAGTTACAGTAATACCTAGATTGCAGGGTGGGTGGCGA GGACCAAGTGAATTAGTACCTGGAAAGTGTGTAACAGTTTCAACATGAGAAGTACACAACATGAGAAGCAGAGTTAGCTGTACATTGCCA GAGAACCCCCTGTGGGCCATTTCCTGTGTTCTTGAAGAAGACTTGGGCTTGGACCCTGTCCCCAGGAGGTCTCAGAGTAATCAGGACGGT ACTAGGGAGAGAACTGTGGCAAGATAGATCAGTTATCTCGGTATCTATTCTCCCCTTCCTAATGTAGAAGCTCTGATTTTTAACTGGGCA GATGACTACATGGATTCAAGGTATTTTCCTAGTGTCTCTTGAAATTAGGTGTGGTATTGTGACCAAATTCTAGCCAAAAAGATGTTGAAC AGAAGTGGTATATGCAGCTTCTAGGAAGTGTTCTTAAGGGGGAAGGCGTCATCCCCCACCTTTCCTCCTTCCTGCACTCTGGAATGTGGA CCAGATGGCTGGAGTAGGAGCTGCCACCTGGGGCCAGAAAATGATCTTAGGCGTGAAGGTCAAGTAAGGCAGGACAAGAGAGAAGGAGCC >103051_103051_1_ZNF83-PPP2R1A_ZNF83_chr19_53138318_ENST00000594682_PPP2R1A_chr19_52705196_ENST00000322088_length(amino acids)=586AA_BP=23 MVHEEETRKGKRKAKASGMVLPQLRLNSIKKLSTIALALGVERTRSELLPFLTDTIYDEDEVLLALAEQLGTFTTLVGGPEYVHCLLPPL ESLATVEETVVRDKAVESLRAISHEHSPSDLEAHFVPLVKRLAGGDWFTSRTSACGLFSVCYPRVSSAVKAELRQYFRNLCSDDTPMVRR AAASKLGEFAKVLELDNVKSEIIPMFSNLASDEQDSVRLLAVEACVNIAQLLPQEDLEALVMPTLRQAAEDKSWRVRYMVADKFTELQKA VGPEITKTDLVPAFQNLMKDCEAEVRAAASHKVKEFCENLSADCRENVIMSQILPCIKELVSDANQHVKSALASVIMGLSPILGKDNTIE HLLPLFLAQLKDECPEVRLNIISNLDCVNEVIGIRQLSQSLLPAIVELAEDAKWRVRLAIIEYMPLLAGQLGVEFFDEKLNSLCMAWLVD HVYAIREAATSNLKKLVEKFGKEWAHATIIPKVLAMSGDPNYLHRMTTLFCINVLSEVCGQDITTKHMLPTVLRMAGDPVANVRFNVAKS -------------------------------------------------------------- >103051_103051_2_ZNF83-PPP2R1A_ZNF83_chr19_53138318_ENST00000598536_PPP2R1A_chr19_52705196_ENST00000322088_length(transcript)=3558nt_BP=413nt AAATCTGGAAGCGGATCGCGTGGAGTGACGGTCACACCGCGGCGGGTTTCTATGATGACAGGATGGTCCTAAGTCTTCACACATCAGGGC CACAGCCCCAACCTCACAGGAAACAGAATTAATTTCCAAAGACTCATGTTACATGAGAAAGCCACCAAGAAGACCAAAGAAAAGGAGACA AGGATGGCTCTTCCTCAGGGATGCTTGACTTTCAGGGATGTGGCTATAGAATTCTCTTTGGAGGAGTGGAAATGCCTGAACCCTGCACAG AGGGCTTTATACAGGGCCGTGATGTTGGAGAACTACAGGAACCTGGAGTCTGTGGGATTGACTTCTAAGGACTCTTGGTACATGAGGAAG AAACCCGGAAGGGGAAGAGGAAAGCAAAGGCGTCAGGAATGGTTCTTCCTCAGCTTCGCCTCAACAGCATCAAGAAGCTGTCCACCATCG CCTTGGCCCTTGGGGTTGAAAGGACCCGAAGTGAGCTTCTGCCTTTCCTTACAGATACCATCTATGATGAAGATGAGGTCCTCCTGGCCC TGGCAGAACAGCTGGGAACCTTCACTACCCTGGTGGGAGGCCCAGAGTACGTGCACTGCCTGCTGCCACCGCTGGAGTCGCTGGCCACAG TGGAGGAGACAGTGGTGCGGGACAAGGCAGTGGAGTCCTTACGGGCCATCTCACACGAGCACTCGCCCTCTGACCTGGAGGCGCACTTTG TGCCGCTAGTGAAGCGGCTGGCGGGCGGCGACTGGTTCACCTCCCGCACCTCGGCCTGCGGCCTCTTCTCCGTCTGCTACCCCCGAGTGT CCAGTGCTGTGAAGGCGGAACTTCGACAGTACTTCCGGAACCTGTGCTCAGATGACACCCCCATGGTGCGGCGGGCCGCAGCCTCCAAGC TGGGGGAGTTTGCCAAGGTGCTGGAGCTGGACAACGTCAAGAGTGAGATCATCCCCATGTTCTCCAACCTGGCCTCTGACGAGCAGGACT CGGTGCGGCTGCTGGCGGTGGAGGCGTGCGTGAACATCGCCCAGCTTCTGCCCCAGGAGGATCTGGAGGCCCTGGTGATGCCCACTCTGC GCCAGGCCGCTGAAGACAAGTCCTGGCGCGTCCGCTACATGGTGGCTGACAAGTTCACAGAGCTCCAGAAAGCAGTGGGGCCTGAGATCA CCAAGACAGACCTGGTCCCTGCCTTCCAGAACCTGATGAAAGACTGTGAGGCCGAGGTGAGGGCCGCAGCCTCCCACAAGGTCAAAGAGT TCTGTGAAAACCTCTCAGCTGACTGTCGGGAGAATGTGATCATGTCCCAGATCTTGCCCTGCATCAAGGAGCTGGTGTCCGATGCCAACC AACATGTCAAGTCTGCCCTGGCCTCAGTCATCATGGGTCTCTCTCCCATCTTGGGCAAAGACAACACCATCGAGCACCTCTTGCCCCTCT TCCTGGCTCAGCTGAAGGATGAGTGCCCTGAGGTACGGCTGAACATCATCTCTAACCTGGACTGTGTGAACGAGGTGATTGGCATCCGGC AGCTGTCCCAGTCCCTGCTCCCTGCCATTGTGGAGCTGGCTGAGGACGCCAAGTGGCGGGTGCGGCTGGCCATCATTGAGTACATGCCCC TCCTGGCTGGACAGCTGGGAGTGGAGTTCTTTGATGAGAAACTTAACTCCTTGTGCATGGCCTGGCTTGTGGATCATGTATATGCCATCC GCGAGGCAGCCACCAGCAACCTGAAGAAGCTAGTGGAAAAGTTTGGGAAGGAGTGGGCCCATGCCACAATCATCCCCAAGGTCTTGGCCA TGTCCGGAGACCCCAACTACCTGCACCGCATGACTACGCTCTTCTGCATCAATGTGCTGTCTGAGGTCTGTGGGCAGGACATCACCACCA AGCACATGCTACCCACGGTTCTGCGCATGGCTGGGGACCCGGTTGCCAATGTCCGCTTCAATGTGGCCAAGTCTCTGCAGAAGATAGGGC CCATCCTGGACAACAGCACCTTGCAGAGTGAAGTCAAGCCCATCCTAGAGAAGCTGACCCAGGACCAGGATGTGGACGTCAAATACTTTG CCCAGGAGGCTCTGACTGTTCTGTCTCTCGCCTGATGCTGGAAGAGGAGCAAACACTGGCCTCTGGTGTCCACCCTCCAACCCCCACAAG TCCCTCTTTGGGGAGACACTGGGGGGCCTTTGGCTGTCACTCCCTGTGCATGGTCTGACCCCAGGCCCCTTCCCCCAGCACGGTTCCTCC TCTCCCCAGCCTGGGAAGATGTCTCACTGTCCACCTCCCAACGGGCTAGGGGAGCACGGGGTTGGACAGGACAGTGACCTTGGGAGGAAG GGGCTACTCCGCCCACGTCAGGGAGAGATGTGAGCATCCCGGGTCACTGGATCCTGCTGCTGTAATGGGAACCCCTCCCCCATTTACTTC TCCACCTCCCGTCCTCCCCATCATTGGTTTTTTTTTGTGTGTCAACTGTGCCGTTTTTATTTTATTCCTTTTATTTTCCCCCTTTTCACA GAGAAATAAAGGTCTAGAAGTAGTTGGTCCTCTGGCCTTAGTCATCAGCTTGGAGGAGGGGGCACCACCCCAGAGCCACAACATCTGCGC TTTTCTCTGGAGAAGATCTTGTTACAGGACCCATTATACCCCTATGTCCCGAAAGAATACTCTGGGGATCCCCCCAGGAGTGCTGCTGGC CTTTGGGGTAGAGGGTCCATGAGGTGCTCTGGGTGGTGTCCTGTAGTGCAGTTCAGATTCATAGATGTCGGCCGAGTATTTGGGGGGTTA AGAGCAGGTGCCTTGGAACCAGACTGCCTGGGTCCAAATTGTGGCTCCTTCAGTTAATAGACGTGTGACTAGGAGTAGCTTGTTAAGTCT TTATGAGCTCAGTTTTGTATTGTGTAAAATGAGAGTTACAGTAATACCTAGATTGCAGGGTGGGTGGCGAGGACCAAGTGAATTAGTACC TGGAAAGTGTGTAACAGTTTCAACATGAGAAGTACACAACATGAGAAGCAGAGTTAGCTGTACATTGCCAGAGAACCCCCTGTGGGCCAT TTCCTGTGTTCTTGAAGAAGACTTGGGCTTGGACCCTGTCCCCAGGAGGTCTCAGAGTAATCAGGACGGTACTAGGGAGAGAACTGTGGC AAGATAGATCAGTTATCTCGGTATCTATTCTCCCCTTCCTAATGTAGAAGCTCTGATTTTTAACTGGGCAGATGACTACATGGATTCAAG GTATTTTCCTAGTGTCTCTTGAAATTAGGTGTGGTATTGTGACCAAATTCTAGCCAAAAAGATGTTGAACAGAAGTGGTATATGCAGCTT CTAGGAAGTGTTCTTAAGGGGGAAGGCGTCATCCCCCACCTTTCCTCCTTCCTGCACTCTGGAATGTGGACCAGATGGCTGGAGTAGGAG CTGCCACCTGGGGCCAGAAAATGATCTTAGGCGTGAAGGTCAAGTAAGGCAGGACAAGAGAGAAGGAGCCTGCCTCCCCGAGGCTGTGAA >103051_103051_2_ZNF83-PPP2R1A_ZNF83_chr19_53138318_ENST00000598536_PPP2R1A_chr19_52705196_ENST00000322088_length(amino acids)=586AA_BP=23 MVHEEETRKGKRKAKASGMVLPQLRLNSIKKLSTIALALGVERTRSELLPFLTDTIYDEDEVLLALAEQLGTFTTLVGGPEYVHCLLPPL ESLATVEETVVRDKAVESLRAISHEHSPSDLEAHFVPLVKRLAGGDWFTSRTSACGLFSVCYPRVSSAVKAELRQYFRNLCSDDTPMVRR AAASKLGEFAKVLELDNVKSEIIPMFSNLASDEQDSVRLLAVEACVNIAQLLPQEDLEALVMPTLRQAAEDKSWRVRYMVADKFTELQKA VGPEITKTDLVPAFQNLMKDCEAEVRAAASHKVKEFCENLSADCRENVIMSQILPCIKELVSDANQHVKSALASVIMGLSPILGKDNTIE HLLPLFLAQLKDECPEVRLNIISNLDCVNEVIGIRQLSQSLLPAIVELAEDAKWRVRLAIIEYMPLLAGQLGVEFFDEKLNSLCMAWLVD HVYAIREAATSNLKKLVEKFGKEWAHATIIPKVLAMSGDPNYLHRMTTLFCINVLSEVCGQDITTKHMLPTVLRMAGDPVANVRFNVAKS -------------------------------------------------------------- >103051_103051_3_ZNF83-PPP2R1A_ZNF83_chr19_53138318_ENST00000601257_PPP2R1A_chr19_52705196_ENST00000322088_length(transcript)=3568nt_BP=423nt AGATTCTCGCAAATCTGGAAGCGGATCGCGTGGAGTGACGGTCACACCGCGGCGGGTTTCTATGATGACAGGATGGTCCTAAGTCTTCAC ACATCAGGGCCACAGCCCCAACCTCACAGGAAACAGAATTAATTTCCAAAGACTCATGTTACATGAGAAAGCCACCAAGAAGACCAAAGA AAAGGAGACAAGGATGGCTCTTCCTCAGGGATGCTTGACTTTCAGGGATGTGGCTATAGAATTCTCTTTGGAGGAGTGGAAATGCCTGAA CCCTGCACAGAGGGCTTTATACAGGGCCGTGATGTTGGAGAACTACAGGAACCTGGAGTCTGTGGGATTGACTTCTAAGGACTCTTGGTA CATGAGGAAGAAACCCGGAAGGGGAAGAGGAAAGCAAAGGCGTCAGGAATGGTTCTTCCTCAGCTTCGCCTCAACAGCATCAAGAAGCTG TCCACCATCGCCTTGGCCCTTGGGGTTGAAAGGACCCGAAGTGAGCTTCTGCCTTTCCTTACAGATACCATCTATGATGAAGATGAGGTC CTCCTGGCCCTGGCAGAACAGCTGGGAACCTTCACTACCCTGGTGGGAGGCCCAGAGTACGTGCACTGCCTGCTGCCACCGCTGGAGTCG CTGGCCACAGTGGAGGAGACAGTGGTGCGGGACAAGGCAGTGGAGTCCTTACGGGCCATCTCACACGAGCACTCGCCCTCTGACCTGGAG GCGCACTTTGTGCCGCTAGTGAAGCGGCTGGCGGGCGGCGACTGGTTCACCTCCCGCACCTCGGCCTGCGGCCTCTTCTCCGTCTGCTAC CCCCGAGTGTCCAGTGCTGTGAAGGCGGAACTTCGACAGTACTTCCGGAACCTGTGCTCAGATGACACCCCCATGGTGCGGCGGGCCGCA GCCTCCAAGCTGGGGGAGTTTGCCAAGGTGCTGGAGCTGGACAACGTCAAGAGTGAGATCATCCCCATGTTCTCCAACCTGGCCTCTGAC GAGCAGGACTCGGTGCGGCTGCTGGCGGTGGAGGCGTGCGTGAACATCGCCCAGCTTCTGCCCCAGGAGGATCTGGAGGCCCTGGTGATG CCCACTCTGCGCCAGGCCGCTGAAGACAAGTCCTGGCGCGTCCGCTACATGGTGGCTGACAAGTTCACAGAGCTCCAGAAAGCAGTGGGG CCTGAGATCACCAAGACAGACCTGGTCCCTGCCTTCCAGAACCTGATGAAAGACTGTGAGGCCGAGGTGAGGGCCGCAGCCTCCCACAAG GTCAAAGAGTTCTGTGAAAACCTCTCAGCTGACTGTCGGGAGAATGTGATCATGTCCCAGATCTTGCCCTGCATCAAGGAGCTGGTGTCC GATGCCAACCAACATGTCAAGTCTGCCCTGGCCTCAGTCATCATGGGTCTCTCTCCCATCTTGGGCAAAGACAACACCATCGAGCACCTC TTGCCCCTCTTCCTGGCTCAGCTGAAGGATGAGTGCCCTGAGGTACGGCTGAACATCATCTCTAACCTGGACTGTGTGAACGAGGTGATT GGCATCCGGCAGCTGTCCCAGTCCCTGCTCCCTGCCATTGTGGAGCTGGCTGAGGACGCCAAGTGGCGGGTGCGGCTGGCCATCATTGAG TACATGCCCCTCCTGGCTGGACAGCTGGGAGTGGAGTTCTTTGATGAGAAACTTAACTCCTTGTGCATGGCCTGGCTTGTGGATCATGTA TATGCCATCCGCGAGGCAGCCACCAGCAACCTGAAGAAGCTAGTGGAAAAGTTTGGGAAGGAGTGGGCCCATGCCACAATCATCCCCAAG GTCTTGGCCATGTCCGGAGACCCCAACTACCTGCACCGCATGACTACGCTCTTCTGCATCAATGTGCTGTCTGAGGTCTGTGGGCAGGAC ATCACCACCAAGCACATGCTACCCACGGTTCTGCGCATGGCTGGGGACCCGGTTGCCAATGTCCGCTTCAATGTGGCCAAGTCTCTGCAG AAGATAGGGCCCATCCTGGACAACAGCACCTTGCAGAGTGAAGTCAAGCCCATCCTAGAGAAGCTGACCCAGGACCAGGATGTGGACGTC AAATACTTTGCCCAGGAGGCTCTGACTGTTCTGTCTCTCGCCTGATGCTGGAAGAGGAGCAAACACTGGCCTCTGGTGTCCACCCTCCAA CCCCCACAAGTCCCTCTTTGGGGAGACACTGGGGGGCCTTTGGCTGTCACTCCCTGTGCATGGTCTGACCCCAGGCCCCTTCCCCCAGCA CGGTTCCTCCTCTCCCCAGCCTGGGAAGATGTCTCACTGTCCACCTCCCAACGGGCTAGGGGAGCACGGGGTTGGACAGGACAGTGACCT TGGGAGGAAGGGGCTACTCCGCCCACGTCAGGGAGAGATGTGAGCATCCCGGGTCACTGGATCCTGCTGCTGTAATGGGAACCCCTCCCC CATTTACTTCTCCACCTCCCGTCCTCCCCATCATTGGTTTTTTTTTGTGTGTCAACTGTGCCGTTTTTATTTTATTCCTTTTATTTTCCC CCTTTTCACAGAGAAATAAAGGTCTAGAAGTAGTTGGTCCTCTGGCCTTAGTCATCAGCTTGGAGGAGGGGGCACCACCCCAGAGCCACA ACATCTGCGCTTTTCTCTGGAGAAGATCTTGTTACAGGACCCATTATACCCCTATGTCCCGAAAGAATACTCTGGGGATCCCCCCAGGAG TGCTGCTGGCCTTTGGGGTAGAGGGTCCATGAGGTGCTCTGGGTGGTGTCCTGTAGTGCAGTTCAGATTCATAGATGTCGGCCGAGTATT TGGGGGGTTAAGAGCAGGTGCCTTGGAACCAGACTGCCTGGGTCCAAATTGTGGCTCCTTCAGTTAATAGACGTGTGACTAGGAGTAGCT TGTTAAGTCTTTATGAGCTCAGTTTTGTATTGTGTAAAATGAGAGTTACAGTAATACCTAGATTGCAGGGTGGGTGGCGAGGACCAAGTG AATTAGTACCTGGAAAGTGTGTAACAGTTTCAACATGAGAAGTACACAACATGAGAAGCAGAGTTAGCTGTACATTGCCAGAGAACCCCC TGTGGGCCATTTCCTGTGTTCTTGAAGAAGACTTGGGCTTGGACCCTGTCCCCAGGAGGTCTCAGAGTAATCAGGACGGTACTAGGGAGA GAACTGTGGCAAGATAGATCAGTTATCTCGGTATCTATTCTCCCCTTCCTAATGTAGAAGCTCTGATTTTTAACTGGGCAGATGACTACA TGGATTCAAGGTATTTTCCTAGTGTCTCTTGAAATTAGGTGTGGTATTGTGACCAAATTCTAGCCAAAAAGATGTTGAACAGAAGTGGTA TATGCAGCTTCTAGGAAGTGTTCTTAAGGGGGAAGGCGTCATCCCCCACCTTTCCTCCTTCCTGCACTCTGGAATGTGGACCAGATGGCT GGAGTAGGAGCTGCCACCTGGGGCCAGAAAATGATCTTAGGCGTGAAGGTCAAGTAAGGCAGGACAAGAGAGAAGGAGCCTGCCTCCCCG >103051_103051_3_ZNF83-PPP2R1A_ZNF83_chr19_53138318_ENST00000601257_PPP2R1A_chr19_52705196_ENST00000322088_length(amino acids)=586AA_BP=23 MVHEEETRKGKRKAKASGMVLPQLRLNSIKKLSTIALALGVERTRSELLPFLTDTIYDEDEVLLALAEQLGTFTTLVGGPEYVHCLLPPL ESLATVEETVVRDKAVESLRAISHEHSPSDLEAHFVPLVKRLAGGDWFTSRTSACGLFSVCYPRVSSAVKAELRQYFRNLCSDDTPMVRR AAASKLGEFAKVLELDNVKSEIIPMFSNLASDEQDSVRLLAVEACVNIAQLLPQEDLEALVMPTLRQAAEDKSWRVRYMVADKFTELQKA VGPEITKTDLVPAFQNLMKDCEAEVRAAASHKVKEFCENLSADCRENVIMSQILPCIKELVSDANQHVKSALASVIMGLSPILGKDNTIE HLLPLFLAQLKDECPEVRLNIISNLDCVNEVIGIRQLSQSLLPAIVELAEDAKWRVRLAIIEYMPLLAGQLGVEFFDEKLNSLCMAWLVD HVYAIREAATSNLKKLVEKFGKEWAHATIIPKVLAMSGDPNYLHRMTTLFCINVLSEVCGQDITTKHMLPTVLRMAGDPVANVRFNVAKS -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for ZNF83-PPP2R1A |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ZNF83-PPP2R1A |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ZNF83-PPP2R1A |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
