
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ZNF880-ATOH8 (FusionGDB2 ID:103083) |
Fusion Gene Summary for ZNF880-ATOH8 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ZNF880-ATOH8 | Fusion gene ID: 103083 | Hgene | Tgene | Gene symbol | ZNF880 | ATOH8 | Gene ID | 400713 | 84913 |
| Gene name | zinc finger protein 880 | atonal bHLH transcription factor 8 | |
| Synonyms | - | HATH6|bHLHa21 | |
| Cytomap | 19q13.41 | 2p11.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | zinc finger protein 880zinc finger protein LOC400713 | protein atonal homolog 8atonal homolog 6atonal homolog 8atonal homolog bHLH transcription factor 8basic helix loop helix transcription factor 6class A basic helix-loop-helix protein 21helix-loop-helix protein hATH-6 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | . | Q96SQ7 | |
| Ensembl transtripts involved in fusion gene | ENST00000344085, ENST00000422689, ENST00000424032, ENST00000597976, ENST00000600321, ENST00000595099, | ENST00000463422, ENST00000306279, | |
| Fusion gene scores | * DoF score | 2 X 2 X 2=8 | 2 X 2 X 2=8 |
| # samples | 2 | 2 | |
| ** MAII score | log2(2/8*10)=1.32192809488736 | log2(2/8*10)=1.32192809488736 | |
| Context | PubMed: ZNF880 [Title/Abstract] AND ATOH8 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ZNF880(52873230)-ATOH8(86014008), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | ATOH8 | GO:0045603 | positive regulation of endothelial cell differentiation | 24463812 |
| Tgene | ATOH8 | GO:0045893 | positive regulation of transcription, DNA-templated | 24236640 |
| Tgene | ATOH8 | GO:0060395 | SMAD protein signal transduction | 24236640 |
 Fusion gene breakpoints across ZNF880 (5'-gene) Fusion gene breakpoints across ZNF880 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
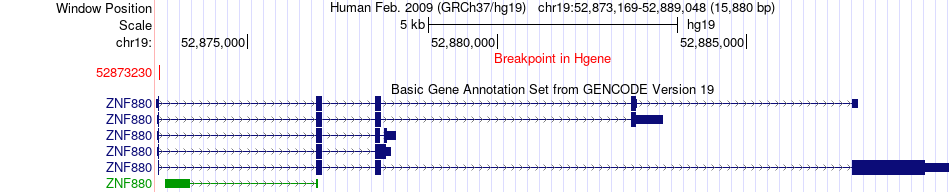 |
 Fusion gene breakpoints across ATOH8 (3'-gene) Fusion gene breakpoints across ATOH8 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
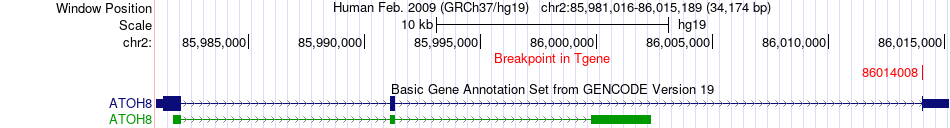 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-BH-A0E2-01A | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
Top |
Fusion Gene ORF analysis for ZNF880-ATOH8 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000344085 | ENST00000463422 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| 5CDS-intron | ENST00000422689 | ENST00000463422 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| 5CDS-intron | ENST00000424032 | ENST00000463422 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| 5CDS-intron | ENST00000597976 | ENST00000463422 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| 5CDS-intron | ENST00000600321 | ENST00000463422 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| In-frame | ENST00000344085 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| In-frame | ENST00000422689 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| In-frame | ENST00000424032 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| In-frame | ENST00000597976 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| In-frame | ENST00000600321 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| intron-3CDS | ENST00000595099 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
| intron-intron | ENST00000595099 | ENST00000463422 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000424032 | ZNF880 | chr19 | 52873230 | + | ENST00000306279 | ATOH8 | chr2 | 86014008 | + | 1243 | 61 | 576 | 1220 | 214 |
| ENST00000600321 | ZNF880 | chr19 | 52873230 | + | ENST00000306279 | ATOH8 | chr2 | 86014008 | + | 1224 | 42 | 557 | 1201 | 214 |
| ENST00000344085 | ZNF880 | chr19 | 52873230 | + | ENST00000306279 | ATOH8 | chr2 | 86014008 | + | 1221 | 39 | 554 | 1198 | 214 |
| ENST00000597976 | ZNF880 | chr19 | 52873230 | + | ENST00000306279 | ATOH8 | chr2 | 86014008 | + | 1214 | 32 | 547 | 1191 | 214 |
| ENST00000422689 | ZNF880 | chr19 | 52873230 | + | ENST00000306279 | ATOH8 | chr2 | 86014008 | + | 1209 | 27 | 542 | 1186 | 214 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000424032 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + | 0.86501074 | 0.13498929 |
| ENST00000600321 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + | 0.8418689 | 0.1581311 |
| ENST00000344085 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + | 0.80859137 | 0.1914086 |
| ENST00000597976 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + | 0.87159693 | 0.1284031 |
| ENST00000422689 | ENST00000306279 | ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014008 | + | 0.92500454 | 0.074995466 |
Top |
Fusion Genomic Features for ZNF880-ATOH8 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014007 | + | 0.026251424 | 0.97374856 |
| ZNF880 | chr19 | 52873230 | + | ATOH8 | chr2 | 86014007 | + | 0.026251424 | 0.97374856 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
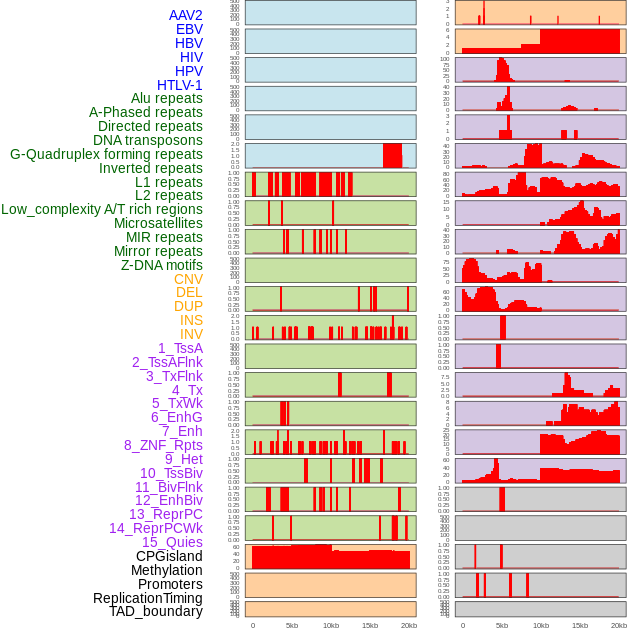 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
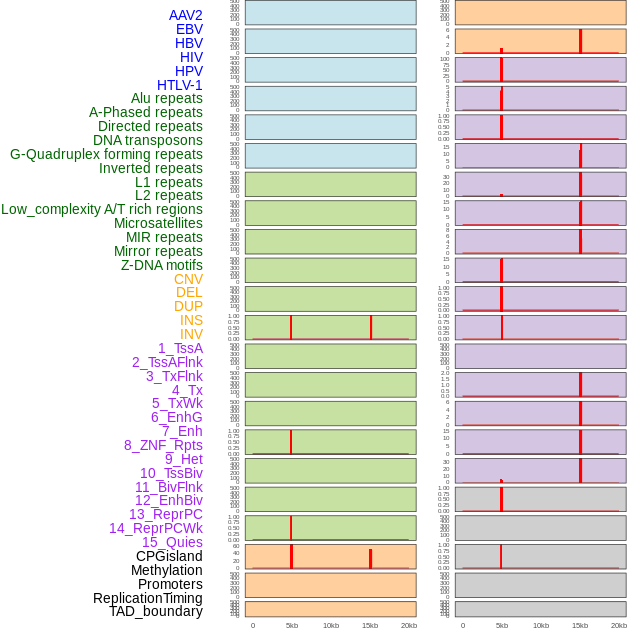 |
Top |
Fusion Protein Features for ZNF880-ATOH8 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:52873230/chr2:86014008) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | ATOH8 |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Transcription factor that binds a palindromic (canonical) core consensus DNA sequence 5'-CANNTG- 3' known as an E-box element, possibly as a heterodimer with other bHLH proteins (PubMed:24236640). Regulates endothelial cell proliferation, migration and tube-like structures formation (PubMed:24463812). Modulates endothelial cell differentiation through NOS3 (PubMed:24463812). May be implicated in specification and differentiation of neuronal cell lineages in the brain (By similarity). May participate in kidney development and may be involved in podocyte differentiation (By similarity). During early embryonic development is involved in tissue-specific differentiation processes that are dependent on class II bHLH factors and namely modulates the differentiation program initiated by the pro-endocrine factor NEUROG3 (By similarity). During myogenesis, may play a role during the transition of myoblasts from the proliferative phase to the differentiation phase (By similarity). Positively regulates HAMP transcription in two ways, firstly by acting directly on the HAMP promoter via E-boxes binding and indirectly through increased phosphorylation of SMAD protein complex (PubMed:24236640). Repress NEUROG3-dependent gene activation in a gene-specific manner through at least two mechanisms; requires only either the sequestering of a general partner such as TCF3 through heterodimerization, either also requires binding of the bHLH domain to DNA via a basic motif (By similarity). {ECO:0000250|UniProtKB:Q99NA2, ECO:0000269|PubMed:24236640, ECO:0000269|PubMed:24463812}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 7_78 | 4 | 99.0 | Domain | KRAB |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 7_78 | 4 | 578.0 | Domain | KRAB |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 184_206 | 4 | 99.0 | Zinc finger | C2H2-type 1%3B degenerate |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 212_234 | 4 | 99.0 | Zinc finger | C2H2-type 2 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 240_262 | 4 | 99.0 | Zinc finger | C2H2-type 3 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 268_290 | 4 | 99.0 | Zinc finger | C2H2-type 4 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 296_318 | 4 | 99.0 | Zinc finger | C2H2-type 5 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 324_346 | 4 | 99.0 | Zinc finger | C2H2-type 6 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 352_374 | 4 | 99.0 | Zinc finger | C2H2-type 7 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 380_402 | 4 | 99.0 | Zinc finger | C2H2-type 8 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 408_430 | 4 | 99.0 | Zinc finger | C2H2-type 9 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 436_458 | 4 | 99.0 | Zinc finger | C2H2-type 10 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 464_486 | 4 | 99.0 | Zinc finger | C2H2-type 11 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 492_514 | 4 | 99.0 | Zinc finger | C2H2-type 12 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 520_542 | 4 | 99.0 | Zinc finger | C2H2-type 13 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000344085 | + | 1 | 4 | 548_570 | 4 | 99.0 | Zinc finger | C2H2-type 14 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 184_206 | 4 | 578.0 | Zinc finger | C2H2-type 1%3B degenerate |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 212_234 | 4 | 578.0 | Zinc finger | C2H2-type 2 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 240_262 | 4 | 578.0 | Zinc finger | C2H2-type 3 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 268_290 | 4 | 578.0 | Zinc finger | C2H2-type 4 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 296_318 | 4 | 578.0 | Zinc finger | C2H2-type 5 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 324_346 | 4 | 578.0 | Zinc finger | C2H2-type 6 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 352_374 | 4 | 578.0 | Zinc finger | C2H2-type 7 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 380_402 | 4 | 578.0 | Zinc finger | C2H2-type 8 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 408_430 | 4 | 578.0 | Zinc finger | C2H2-type 9 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 436_458 | 4 | 578.0 | Zinc finger | C2H2-type 10 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 464_486 | 4 | 578.0 | Zinc finger | C2H2-type 11 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 492_514 | 4 | 578.0 | Zinc finger | C2H2-type 12 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 520_542 | 4 | 578.0 | Zinc finger | C2H2-type 13 |
| Hgene | ZNF880 | chr19:52873230 | chr2:86014008 | ENST00000422689 | + | 1 | 4 | 548_570 | 4 | 578.0 | Zinc finger | C2H2-type 14 |
| Tgene | ATOH8 | chr19:52873230 | chr2:86014008 | ENST00000306279 | 1 | 3 | 69_187 | 320 | 322.0 | Compositional bias | Pro-rich | |
| Tgene | ATOH8 | chr19:52873230 | chr2:86014008 | ENST00000306279 | 1 | 3 | 230_282 | 320 | 322.0 | Domain | bHLH | |
| Tgene | ATOH8 | chr19:52873230 | chr2:86014008 | ENST00000306279 | 1 | 3 | 230_243 | 320 | 322.0 | Region | Basic motif%3B degenerate | |
| Tgene | ATOH8 | chr19:52873230 | chr2:86014008 | ENST00000306279 | 1 | 3 | 244_282 | 320 | 322.0 | Region | Helix-loop-helix motif |
Top |
Fusion Gene Sequence for ZNF880-ATOH8 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >103083_103083_1_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000344085_ATOH8_chr2_86014008_ENST00000306279_length(transcript)=1221nt_BP=39nt GGAAGCAGATTACGTGGAGTGACGGTCATGCTGCGGCGTGAGTGACTGGCTGCAGGCAAGACCAAGGCCACCACTGTGGGCCCTCCTTCC AGTCAGGCCTGAGGACAAGGTGAGCTCGCTGAGTCCAGCCTCGTGGTCTTCTCCAAGATGCCGCCAGATGCCCAGCCTACAGCCTCTCAG GGTCGGATCGGAGCACGCCTGCCTCCCTCTCCCCTCCGCCCTCACCCAGCCAATCCGAGGCTGCTTCGCACTTTGCCCTCTGCCTGGTGG GGAGGGGAGAGCTCAGCCCCCGACTCACTCAGACCCCAAGGCCCACTGTCCAGCTGCAGAAATTCGTTGCCAAAGATTGGACAGAGACAC CGAAGGAAATGGGGTGGTGAAACCCCACAGCGAAAAGCCACACCGTTGCTCTGTGACTTTTGCTCCTCCTGTTGCCTGAGCCCCATCTCA AGCCAAAGATGAGTCAGTGGTTCTGCTAGGAACTCATGGAATGGATGGGCATTTGATGACCCCTGGGGGTCATCTTGGCCCTCTGACCTG GTGCTCTCTCTCCACTGGGCCTTGTGCTGGCTGAGTGCAAGACAAGCCTTAGGGGCTGTGAGAGGGAGGCTGGGGTGCCTGGGCGGGGCT GGGAGTGGGACCTGAGATCCCTGCCCACTCTCTCCCCTTCATTGGCTGCCCAGGCCACTGGCCCCAGTTCTCAGTGTCCCTTGGGTCCAG GCTCCTTGGGCCCTAAGCATCACCAGAAGGGAGTAAGCAGGGAGAGAAGCAATATTACTCCCTCCCCTACACCAGGGACTTGCCCCAGGG CAGCTACCTATGGGTCTTTGCTTCCCCAGCCAGCCTCTCCTCACTGTGACCCACCCCCATGGGCCCCCGTCCCAGGCAGCCAGCACCATG GGCAGGCCCTGCCATGGACAGAAAAAGAGTTTTTCTCTTGTTCAGCCTGCACGTGGCCTGAGGAAGGAGTAGAGGCTGGGTTGGCTGGAG CCGTCCTACTGGGCAAGATGGCGCCCCACTTGGAGGGCGGTGGTCTGTTACAGGGTGTGCAGGGGCAGAGAAGGAAGGGACCAGGGGACT GGGCCAGTATGTGGAGGATGGGGCCTGCGTGTTCAAAGCCAAGGCCCGCCCCTTCCTTGTGCTCAAATGGCCAAAGCTGTTCACGTCTGT >103083_103083_1_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000344085_ATOH8_chr2_86014008_ENST00000306279_length(amino acids)=214AA_BP= MGLVLAECKTSLRGCEREAGVPGRGWEWDLRSLPTLSPSLAAQATGPSSQCPLGPGSLGPKHHQKGVSRERSNITPSPTPGTCPRAATYG SLLPQPASPHCDPPPWAPVPGSQHHGQALPWTEKEFFSCSACTWPEEGVEAGLAGAVLLGKMAPHLEGGGLLQGVQGQRRKGPGDWASMW -------------------------------------------------------------- >103083_103083_2_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000422689_ATOH8_chr2_86014008_ENST00000306279_length(transcript)=1209nt_BP=27nt CGTGGAGTGACGGTCATGCTGCGGCGTGAGTGACTGGCTGCAGGCAAGACCAAGGCCACCACTGTGGGCCCTCCTTCCAGTCAGGCCTGA GGACAAGGTGAGCTCGCTGAGTCCAGCCTCGTGGTCTTCTCCAAGATGCCGCCAGATGCCCAGCCTACAGCCTCTCAGGGTCGGATCGGA GCACGCCTGCCTCCCTCTCCCCTCCGCCCTCACCCAGCCAATCCGAGGCTGCTTCGCACTTTGCCCTCTGCCTGGTGGGGAGGGGAGAGC TCAGCCCCCGACTCACTCAGACCCCAAGGCCCACTGTCCAGCTGCAGAAATTCGTTGCCAAAGATTGGACAGAGACACCGAAGGAAATGG GGTGGTGAAACCCCACAGCGAAAAGCCACACCGTTGCTCTGTGACTTTTGCTCCTCCTGTTGCCTGAGCCCCATCTCAAGCCAAAGATGA GTCAGTGGTTCTGCTAGGAACTCATGGAATGGATGGGCATTTGATGACCCCTGGGGGTCATCTTGGCCCTCTGACCTGGTGCTCTCTCTC CACTGGGCCTTGTGCTGGCTGAGTGCAAGACAAGCCTTAGGGGCTGTGAGAGGGAGGCTGGGGTGCCTGGGCGGGGCTGGGAGTGGGACC TGAGATCCCTGCCCACTCTCTCCCCTTCATTGGCTGCCCAGGCCACTGGCCCCAGTTCTCAGTGTCCCTTGGGTCCAGGCTCCTTGGGCC CTAAGCATCACCAGAAGGGAGTAAGCAGGGAGAGAAGCAATATTACTCCCTCCCCTACACCAGGGACTTGCCCCAGGGCAGCTACCTATG GGTCTTTGCTTCCCCAGCCAGCCTCTCCTCACTGTGACCCACCCCCATGGGCCCCCGTCCCAGGCAGCCAGCACCATGGGCAGGCCCTGC CATGGACAGAAAAAGAGTTTTTCTCTTGTTCAGCCTGCACGTGGCCTGAGGAAGGAGTAGAGGCTGGGTTGGCTGGAGCCGTCCTACTGG GCAAGATGGCGCCCCACTTGGAGGGCGGTGGTCTGTTACAGGGTGTGCAGGGGCAGAGAAGGAAGGGACCAGGGGACTGGGCCAGTATGT GGAGGATGGGGCCTGCGTGTTCAAAGCCAAGGCCCGCCCCTTCCTTGTGCTCAAATGGCCAAAGCTGTTCACGTCTGTGCTCAACCATCT >103083_103083_2_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000422689_ATOH8_chr2_86014008_ENST00000306279_length(amino acids)=214AA_BP= MGLVLAECKTSLRGCEREAGVPGRGWEWDLRSLPTLSPSLAAQATGPSSQCPLGPGSLGPKHHQKGVSRERSNITPSPTPGTCPRAATYG SLLPQPASPHCDPPPWAPVPGSQHHGQALPWTEKEFFSCSACTWPEEGVEAGLAGAVLLGKMAPHLEGGGLLQGVQGQRRKGPGDWASMW -------------------------------------------------------------- >103083_103083_3_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000424032_ATOH8_chr2_86014008_ENST00000306279_length(transcript)=1243nt_BP=61nt GCGCGCAGTTTCCTGGAGACCCGGAAGCAGATTACGTGGAGTGACGGTCATGCTGCGGCGTGAGTGACTGGCTGCAGGCAAGACCAAGGC CACCACTGTGGGCCCTCCTTCCAGTCAGGCCTGAGGACAAGGTGAGCTCGCTGAGTCCAGCCTCGTGGTCTTCTCCAAGATGCCGCCAGA TGCCCAGCCTACAGCCTCTCAGGGTCGGATCGGAGCACGCCTGCCTCCCTCTCCCCTCCGCCCTCACCCAGCCAATCCGAGGCTGCTTCG CACTTTGCCCTCTGCCTGGTGGGGAGGGGAGAGCTCAGCCCCCGACTCACTCAGACCCCAAGGCCCACTGTCCAGCTGCAGAAATTCGTT GCCAAAGATTGGACAGAGACACCGAAGGAAATGGGGTGGTGAAACCCCACAGCGAAAAGCCACACCGTTGCTCTGTGACTTTTGCTCCTC CTGTTGCCTGAGCCCCATCTCAAGCCAAAGATGAGTCAGTGGTTCTGCTAGGAACTCATGGAATGGATGGGCATTTGATGACCCCTGGGG GTCATCTTGGCCCTCTGACCTGGTGCTCTCTCTCCACTGGGCCTTGTGCTGGCTGAGTGCAAGACAAGCCTTAGGGGCTGTGAGAGGGAG GCTGGGGTGCCTGGGCGGGGCTGGGAGTGGGACCTGAGATCCCTGCCCACTCTCTCCCCTTCATTGGCTGCCCAGGCCACTGGCCCCAGT TCTCAGTGTCCCTTGGGTCCAGGCTCCTTGGGCCCTAAGCATCACCAGAAGGGAGTAAGCAGGGAGAGAAGCAATATTACTCCCTCCCCT ACACCAGGGACTTGCCCCAGGGCAGCTACCTATGGGTCTTTGCTTCCCCAGCCAGCCTCTCCTCACTGTGACCCACCCCCATGGGCCCCC GTCCCAGGCAGCCAGCACCATGGGCAGGCCCTGCCATGGACAGAAAAAGAGTTTTTCTCTTGTTCAGCCTGCACGTGGCCTGAGGAAGGA GTAGAGGCTGGGTTGGCTGGAGCCGTCCTACTGGGCAAGATGGCGCCCCACTTGGAGGGCGGTGGTCTGTTACAGGGTGTGCAGGGGCAG AGAAGGAAGGGACCAGGGGACTGGGCCAGTATGTGGAGGATGGGGCCTGCGTGTTCAAAGCCAAGGCCCGCCCCTTCCTTGTGCTCAAAT >103083_103083_3_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000424032_ATOH8_chr2_86014008_ENST00000306279_length(amino acids)=214AA_BP= MGLVLAECKTSLRGCEREAGVPGRGWEWDLRSLPTLSPSLAAQATGPSSQCPLGPGSLGPKHHQKGVSRERSNITPSPTPGTCPRAATYG SLLPQPASPHCDPPPWAPVPGSQHHGQALPWTEKEFFSCSACTWPEEGVEAGLAGAVLLGKMAPHLEGGGLLQGVQGQRRKGPGDWASMW -------------------------------------------------------------- >103083_103083_4_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000597976_ATOH8_chr2_86014008_ENST00000306279_length(transcript)=1214nt_BP=32nt GATTACGTGGAGTGACGGTCATGCTGCGGCGTGAGTGACTGGCTGCAGGCAAGACCAAGGCCACCACTGTGGGCCCTCCTTCCAGTCAGG CCTGAGGACAAGGTGAGCTCGCTGAGTCCAGCCTCGTGGTCTTCTCCAAGATGCCGCCAGATGCCCAGCCTACAGCCTCTCAGGGTCGGA TCGGAGCACGCCTGCCTCCCTCTCCCCTCCGCCCTCACCCAGCCAATCCGAGGCTGCTTCGCACTTTGCCCTCTGCCTGGTGGGGAGGGG AGAGCTCAGCCCCCGACTCACTCAGACCCCAAGGCCCACTGTCCAGCTGCAGAAATTCGTTGCCAAAGATTGGACAGAGACACCGAAGGA AATGGGGTGGTGAAACCCCACAGCGAAAAGCCACACCGTTGCTCTGTGACTTTTGCTCCTCCTGTTGCCTGAGCCCCATCTCAAGCCAAA GATGAGTCAGTGGTTCTGCTAGGAACTCATGGAATGGATGGGCATTTGATGACCCCTGGGGGTCATCTTGGCCCTCTGACCTGGTGCTCT CTCTCCACTGGGCCTTGTGCTGGCTGAGTGCAAGACAAGCCTTAGGGGCTGTGAGAGGGAGGCTGGGGTGCCTGGGCGGGGCTGGGAGTG GGACCTGAGATCCCTGCCCACTCTCTCCCCTTCATTGGCTGCCCAGGCCACTGGCCCCAGTTCTCAGTGTCCCTTGGGTCCAGGCTCCTT GGGCCCTAAGCATCACCAGAAGGGAGTAAGCAGGGAGAGAAGCAATATTACTCCCTCCCCTACACCAGGGACTTGCCCCAGGGCAGCTAC CTATGGGTCTTTGCTTCCCCAGCCAGCCTCTCCTCACTGTGACCCACCCCCATGGGCCCCCGTCCCAGGCAGCCAGCACCATGGGCAGGC CCTGCCATGGACAGAAAAAGAGTTTTTCTCTTGTTCAGCCTGCACGTGGCCTGAGGAAGGAGTAGAGGCTGGGTTGGCTGGAGCCGTCCT ACTGGGCAAGATGGCGCCCCACTTGGAGGGCGGTGGTCTGTTACAGGGTGTGCAGGGGCAGAGAAGGAAGGGACCAGGGGACTGGGCCAG TATGTGGAGGATGGGGCCTGCGTGTTCAAAGCCAAGGCCCGCCCCTTCCTTGTGCTCAAATGGCCAAAGCTGTTCACGTCTGTGCTCAAC >103083_103083_4_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000597976_ATOH8_chr2_86014008_ENST00000306279_length(amino acids)=214AA_BP= MGLVLAECKTSLRGCEREAGVPGRGWEWDLRSLPTLSPSLAAQATGPSSQCPLGPGSLGPKHHQKGVSRERSNITPSPTPGTCPRAATYG SLLPQPASPHCDPPPWAPVPGSQHHGQALPWTEKEFFSCSACTWPEEGVEAGLAGAVLLGKMAPHLEGGGLLQGVQGQRRKGPGDWASMW -------------------------------------------------------------- >103083_103083_5_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000600321_ATOH8_chr2_86014008_ENST00000306279_length(transcript)=1224nt_BP=42nt CCCGGAAGCAGATTACGTGGAGTGACGGTCATGCTGCGGCGTGAGTGACTGGCTGCAGGCAAGACCAAGGCCACCACTGTGGGCCCTCCT TCCAGTCAGGCCTGAGGACAAGGTGAGCTCGCTGAGTCCAGCCTCGTGGTCTTCTCCAAGATGCCGCCAGATGCCCAGCCTACAGCCTCT CAGGGTCGGATCGGAGCACGCCTGCCTCCCTCTCCCCTCCGCCCTCACCCAGCCAATCCGAGGCTGCTTCGCACTTTGCCCTCTGCCTGG TGGGGAGGGGAGAGCTCAGCCCCCGACTCACTCAGACCCCAAGGCCCACTGTCCAGCTGCAGAAATTCGTTGCCAAAGATTGGACAGAGA CACCGAAGGAAATGGGGTGGTGAAACCCCACAGCGAAAAGCCACACCGTTGCTCTGTGACTTTTGCTCCTCCTGTTGCCTGAGCCCCATC TCAAGCCAAAGATGAGTCAGTGGTTCTGCTAGGAACTCATGGAATGGATGGGCATTTGATGACCCCTGGGGGTCATCTTGGCCCTCTGAC CTGGTGCTCTCTCTCCACTGGGCCTTGTGCTGGCTGAGTGCAAGACAAGCCTTAGGGGCTGTGAGAGGGAGGCTGGGGTGCCTGGGCGGG GCTGGGAGTGGGACCTGAGATCCCTGCCCACTCTCTCCCCTTCATTGGCTGCCCAGGCCACTGGCCCCAGTTCTCAGTGTCCCTTGGGTC CAGGCTCCTTGGGCCCTAAGCATCACCAGAAGGGAGTAAGCAGGGAGAGAAGCAATATTACTCCCTCCCCTACACCAGGGACTTGCCCCA GGGCAGCTACCTATGGGTCTTTGCTTCCCCAGCCAGCCTCTCCTCACTGTGACCCACCCCCATGGGCCCCCGTCCCAGGCAGCCAGCACC ATGGGCAGGCCCTGCCATGGACAGAAAAAGAGTTTTTCTCTTGTTCAGCCTGCACGTGGCCTGAGGAAGGAGTAGAGGCTGGGTTGGCTG GAGCCGTCCTACTGGGCAAGATGGCGCCCCACTTGGAGGGCGGTGGTCTGTTACAGGGTGTGCAGGGGCAGAGAAGGAAGGGACCAGGGG ACTGGGCCAGTATGTGGAGGATGGGGCCTGCGTGTTCAAAGCCAAGGCCCGCCCCTTCCTTGTGCTCAAATGGCCAAAGCTGTTCACGTC >103083_103083_5_ZNF880-ATOH8_ZNF880_chr19_52873230_ENST00000600321_ATOH8_chr2_86014008_ENST00000306279_length(amino acids)=214AA_BP= MGLVLAECKTSLRGCEREAGVPGRGWEWDLRSLPTLSPSLAAQATGPSSQCPLGPGSLGPKHHQKGVSRERSNITPSPTPGTCPRAATYG SLLPQPASPHCDPPPWAPVPGSQHHGQALPWTEKEFFSCSACTWPEEGVEAGLAGAVLLGKMAPHLEGGGLLQGVQGQRRKGPGDWASMW -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for ZNF880-ATOH8 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ZNF880-ATOH8 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ZNF880-ATOH8 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
