
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CABLES1-FTO (FusionGDB2 ID:12246) |
Fusion Gene Summary for CABLES1-FTO |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CABLES1-FTO | Fusion gene ID: 12246 | Hgene | Tgene | Gene symbol | CABLES1 | FTO | Gene ID | 91768 | 79068 |
| Gene name | Cdk5 and Abl enzyme substrate 1 | FTO alpha-ketoglutarate dependent dioxygenase | |
| Synonyms | CABL1|CABLES|HsT2563|IK3-1 | ALKBH9|BMIQ14|GDFD | |
| Cytomap | 18q11.2 | 16q12.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | CDK5 and ABL1 enzyme substrate 1interactor with CDK3 1 | alpha-ketoglutarate-dependent dioxygenase FTOAlkB homolog 9U6 small nuclear RNA (2'-O-methyladenosine-N(6)-)-demethylase FTOU6 small nuclear RNA N(6)-methyladenosine-demethylase FTOfat mass and obesity associatedfat mass and obesity-associated protei | |
| Modification date | 20200313 | 20200329 | |
| UniProtAcc | Q8TDN4 | Q9C0B1 | |
| Ensembl transtripts involved in fusion gene | ENST00000256925, ENST00000400473, ENST00000420687, ENST00000585061, | ENST00000472835, ENST00000471389, ENST00000394647, ENST00000431610, ENST00000460382, ENST00000463855, | |
| Fusion gene scores | * DoF score | 16 X 13 X 3=624 | 12 X 10 X 7=840 |
| # samples | 21 | 15 | |
| ** MAII score | log2(21/624*10)=-1.57115670119613 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(15/840*10)=-2.48542682717024 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CABLES1 [Title/Abstract] AND FTO [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CABLES1(20716571)-FTO(54145674), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | CABLES1-FTO seems lost the major protein functional domain in Hgene partner, which is a tumor suppressor due to the frame-shifted ORF. CABLES1-FTO seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | FTO | GO:0006307 | DNA dealkylation involved in DNA repair | 18775698|20376003 |
| Tgene | FTO | GO:0035552 | oxidative single-stranded DNA demethylation | 18775698|20376003 |
| Tgene | FTO | GO:0035553 | oxidative single-stranded RNA demethylation | 18775698|22002720|25452335|26457839|28002401|30197295 |
| Tgene | FTO | GO:0042245 | RNA repair | 18775698 |
| Tgene | FTO | GO:0061157 | mRNA destabilization | 28002401|30197295 |
| Tgene | FTO | GO:0070989 | oxidative demethylation | 18775698 |
| Tgene | FTO | GO:0080111 | DNA demethylation | 18775698 |
 Fusion gene breakpoints across CABLES1 (5'-gene) Fusion gene breakpoints across CABLES1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
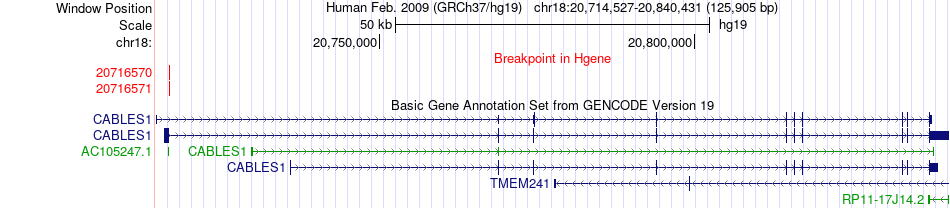 |
 Fusion gene breakpoints across FTO (3'-gene) Fusion gene breakpoints across FTO (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
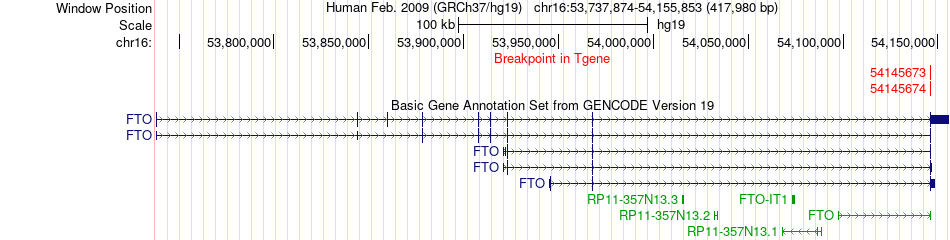 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-S3-AA12-01A | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| ChimerDB4 | BRCA | TCGA-S3-AA12-01A | CABLES1 | chr18 | 20716571 | - | FTO | chr16 | 54145674 | + |
| ChimerDB4 | BRCA | TCGA-S3-AA12-01A | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| ChimerDB4 | BRCA | TCGA-S3-AA12 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
Top |
Fusion Gene ORF analysis for CABLES1-FTO |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000256925 | ENST00000472835 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| 5CDS-3UTR | ENST00000256925 | ENST00000472835 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| 5CDS-3UTR | ENST00000256925 | ENST00000472835 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| Frame-shift | ENST00000256925 | ENST00000471389 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| Frame-shift | ENST00000256925 | ENST00000471389 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| Frame-shift | ENST00000256925 | ENST00000471389 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| In-frame | ENST00000256925 | ENST00000394647 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| In-frame | ENST00000256925 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| In-frame | ENST00000256925 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| In-frame | ENST00000256925 | ENST00000431610 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| In-frame | ENST00000256925 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| In-frame | ENST00000256925 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| In-frame | ENST00000256925 | ENST00000460382 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| In-frame | ENST00000256925 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| In-frame | ENST00000256925 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| In-frame | ENST00000256925 | ENST00000463855 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| In-frame | ENST00000256925 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| In-frame | ENST00000256925 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000394647 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000400473 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000431610 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000400473 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000460382 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000400473 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000463855 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000400473 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000471389 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000400473 | ENST00000471389 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000400473 | ENST00000471389 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000394647 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000420687 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000431610 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000420687 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000460382 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000420687 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000463855 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000420687 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000471389 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000420687 | ENST00000471389 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000420687 | ENST00000471389 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000394647 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000585061 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000431610 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000585061 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000460382 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000585061 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000463855 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000585061 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000471389 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3CDS | ENST00000585061 | ENST00000471389 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3CDS | ENST00000585061 | ENST00000471389 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3UTR | ENST00000400473 | ENST00000472835 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3UTR | ENST00000400473 | ENST00000472835 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3UTR | ENST00000400473 | ENST00000472835 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3UTR | ENST00000420687 | ENST00000472835 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3UTR | ENST00000420687 | ENST00000472835 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3UTR | ENST00000420687 | ENST00000472835 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
| intron-3UTR | ENST00000585061 | ENST00000472835 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + |
| intron-3UTR | ENST00000585061 | ENST00000472835 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + |
| intron-3UTR | ENST00000585061 | ENST00000472835 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000256925 | CABLES1 | chr18 | 20716571 | + | ENST00000394647 | FTO | chr16 | 54145674 | + | 1171 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716571 | + | ENST00000431610 | FTO | chr16 | 54145674 | + | 1222 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716571 | + | ENST00000460382 | FTO | chr16 | 54145674 | + | 2071 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716571 | + | ENST00000463855 | FTO | chr16 | 54145674 | + | 3547 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716571 | + | ENST00000394647 | FTO | chr16 | 54145673 | + | 1171 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716571 | + | ENST00000431610 | FTO | chr16 | 54145673 | + | 1222 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716571 | + | ENST00000460382 | FTO | chr16 | 54145673 | + | 2071 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716571 | + | ENST00000463855 | FTO | chr16 | 54145673 | + | 3547 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716570 | + | ENST00000394647 | FTO | chr16 | 54145673 | + | 1171 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716570 | + | ENST00000431610 | FTO | chr16 | 54145673 | + | 1222 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716570 | + | ENST00000460382 | FTO | chr16 | 54145673 | + | 2071 | 845 | 0 | 998 | 332 |
| ENST00000256925 | CABLES1 | chr18 | 20716570 | + | ENST00000463855 | FTO | chr16 | 54145673 | + | 3547 | 845 | 0 | 998 | 332 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000256925 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + | 0.017564785 | 0.9824353 |
| ENST00000256925 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + | 0.03375883 | 0.9662411 |
| ENST00000256925 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + | 0.21201108 | 0.78798884 |
| ENST00000256925 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145674 | + | 0.16161251 | 0.83838755 |
| ENST00000256925 | ENST00000394647 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + | 0.017564785 | 0.9824353 |
| ENST00000256925 | ENST00000431610 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + | 0.03375883 | 0.9662411 |
| ENST00000256925 | ENST00000460382 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + | 0.21201108 | 0.78798884 |
| ENST00000256925 | ENST00000463855 | CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + | 0.16161251 | 0.83838755 |
| ENST00000256925 | ENST00000394647 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + | 0.017564785 | 0.9824353 |
| ENST00000256925 | ENST00000431610 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + | 0.03375883 | 0.9662411 |
| ENST00000256925 | ENST00000460382 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + | 0.21201108 | 0.78798884 |
| ENST00000256925 | ENST00000463855 | CABLES1 | chr18 | 20716570 | + | FTO | chr16 | 54145673 | + | 0.16161251 | 0.83838755 |
Top |
Fusion Genomic Features for CABLES1-FTO |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + | 1.27E-07 | 0.9999999 |
| CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + | 1.27E-07 | 0.9999999 |
| CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + | 1.27E-07 | 0.9999999 |
| CABLES1 | chr18 | 20716571 | + | FTO | chr16 | 54145673 | + | 1.27E-07 | 0.9999999 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
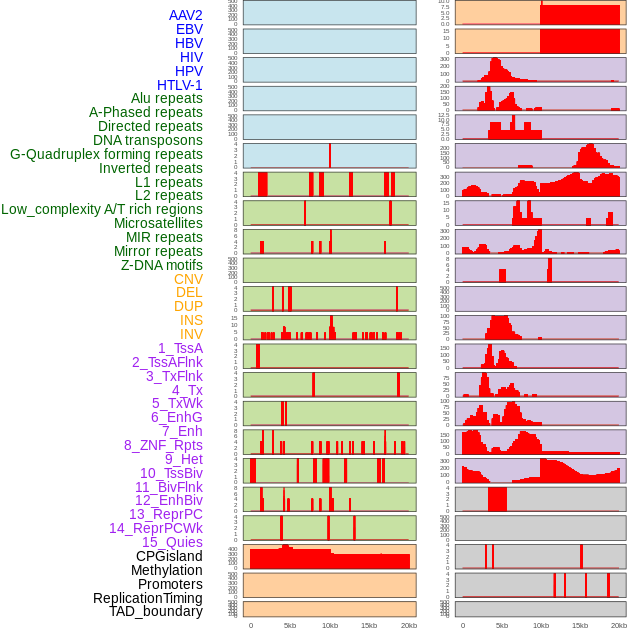 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
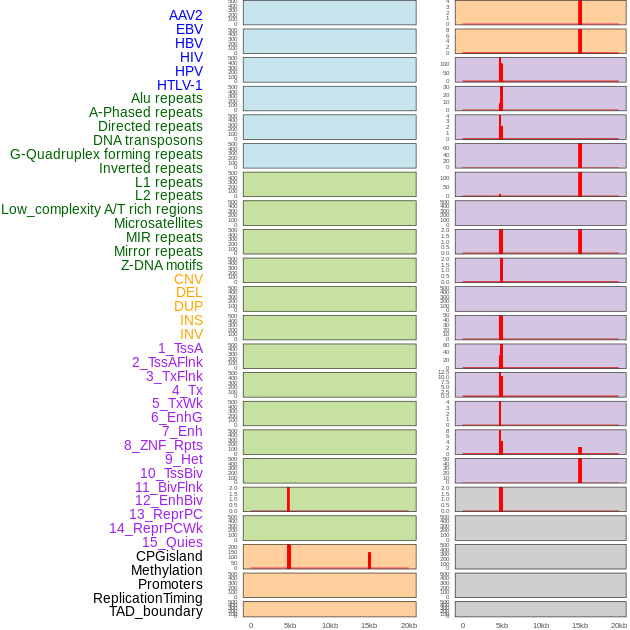 |
Top |
Fusion Protein Features for CABLES1-FTO |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr18:20716571/chr16:54145674) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CABLES1 | FTO |
| FUNCTION: Cyclin-dependent kinase binding protein. Enhances cyclin-dependent kinase tyrosine phosphorylation by nonreceptor tyrosine kinases, such as that of CDK5 by activated ABL1, which leads to increased CDK5 activity and is critical for neuronal development, and that of CDK2 by WEE1, which leads to decreased CDK2 activity and growth inhibition. Positively affects neuronal outgrowth. Plays a role as a regulator for p53/p73-induced cell death (By similarity). {ECO:0000250}. | FUNCTION: RNA demethylase that mediates oxidative demethylation of different RNA species, such as mRNAs, tRNAs and snRNAs, and acts as a regulator of fat mass, adipogenesis and energy homeostasis (PubMed:22002720, PubMed:26458103, PubMed:28002401, PubMed:30197295, PubMed:26457839, PubMed:25452335). Specifically demethylates N(6)-methyladenosine (m6A) RNA, the most prevalent internal modification of messenger RNA (mRNA) in higher eukaryotes (PubMed:22002720, PubMed:26458103, PubMed:30197295, PubMed:26457839, PubMed:25452335). M6A demethylation by FTO affects mRNA expression and stability (PubMed:30197295). Also able to demethylate m6A in U6 small nuclear RNA (snRNA) (PubMed:30197295). Mediates demethylation of N(6),2'-O-dimethyladenosine cap (m6A(m)), by demethylating the N(6)-methyladenosine at the second transcribed position of mRNAs and U6 snRNA (PubMed:28002401, PubMed:30197295). Demethylation of m6A(m) in the 5'-cap by FTO affects mRNA stability by promoting susceptibility to decapping (PubMed:28002401). Also acts as a tRNA demethylase by removing N(1)-methyladenine from various tRNAs (PubMed:30197295). Has no activity towards 1-methylguanine (PubMed:20376003). Has no detectable activity towards double-stranded DNA (PubMed:20376003). Also able to repair alkylated DNA and RNA by oxidative demethylation: demethylates single-stranded RNA containing 3-methyluracil, single-stranded DNA containing 3-methylthymine and has low demethylase activity towards single-stranded DNA containing 1-methyladenine or 3-methylcytosine (PubMed:18775698, PubMed:20376003). Ability to repair alkylated DNA and RNA is however unsure in vivo (PubMed:18775698, PubMed:20376003). Involved in the regulation of fat mass, adipogenesis and body weight, thereby contributing to the regulation of body size and body fat accumulation (PubMed:18775698, PubMed:20376003). Involved in the regulation of thermogenesis and the control of adipocyte differentiation into brown or white fat cells (PubMed:26287746). Regulates activity of the dopaminergic midbrain circuitry via its ability to demethylate m6A in mRNAs (By similarity). Plays an oncogenic role in a number of acute myeloid leukemias by enhancing leukemic oncogene-mediated cell transformation: acts by mediating m6A demethylation of target transcripts such as MYC, CEBPA, ASB2 and RARA, leading to promote their expression (PubMed:28017614, PubMed:29249359). {ECO:0000250|UniProtKB:Q8BGW1, ECO:0000269|PubMed:18775698, ECO:0000269|PubMed:20376003, ECO:0000269|PubMed:22002720, ECO:0000269|PubMed:25452335, ECO:0000269|PubMed:26287746, ECO:0000269|PubMed:26457839, ECO:0000269|PubMed:26458103, ECO:0000269|PubMed:28002401, ECO:0000269|PubMed:28017614, ECO:0000269|PubMed:29249359, ECO:0000269|PubMed:30197295}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CABLES1 | chr18:20716570 | chr16:54145673 | ENST00000256925 | + | 1 | 10 | 2_134 | 281 | 634.0 | Compositional bias | Note=Ala-rich |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145673 | ENST00000256925 | + | 1 | 10 | 2_134 | 281 | 634.0 | Compositional bias | Note=Ala-rich |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145674 | ENST00000256925 | + | 1 | 10 | 2_134 | 281 | 634.0 | Compositional bias | Note=Ala-rich |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000431610 | 3 | 5 | 213_224 | 55 | 107.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000431610 | 3 | 5 | 231_234 | 55 | 107.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000431610 | 3 | 5 | 316_318 | 55 | 107.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000460382 | 2 | 4 | 213_224 | 55 | 107.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000460382 | 2 | 4 | 231_234 | 55 | 107.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000460382 | 2 | 4 | 316_318 | 55 | 107.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000463855 | 1 | 3 | 213_224 | 76 | 128.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000463855 | 1 | 3 | 231_234 | 76 | 128.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000463855 | 1 | 3 | 316_318 | 76 | 128.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000431610 | 3 | 5 | 213_224 | 55 | 107.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000431610 | 3 | 5 | 231_234 | 55 | 107.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000431610 | 3 | 5 | 316_318 | 55 | 107.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000460382 | 2 | 4 | 213_224 | 55 | 107.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000460382 | 2 | 4 | 231_234 | 55 | 107.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000460382 | 2 | 4 | 316_318 | 55 | 107.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000463855 | 1 | 3 | 213_224 | 76 | 128.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000463855 | 1 | 3 | 231_234 | 76 | 128.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000463855 | 1 | 3 | 316_318 | 76 | 128.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000431610 | 3 | 5 | 213_224 | 55 | 107.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000431610 | 3 | 5 | 231_234 | 55 | 107.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000431610 | 3 | 5 | 316_318 | 55 | 107.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000460382 | 2 | 4 | 213_224 | 55 | 107.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000460382 | 2 | 4 | 231_234 | 55 | 107.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000460382 | 2 | 4 | 316_318 | 55 | 107.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000463855 | 1 | 3 | 213_224 | 76 | 128.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000463855 | 1 | 3 | 231_234 | 76 | 128.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000463855 | 1 | 3 | 316_318 | 76 | 128.0 | Region | Alpha-ketoglutarate binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CABLES1 | chr18:20716570 | chr16:54145673 | ENST00000400473 | + | 1 | 10 | 2_134 | 0 | 307.0 | Compositional bias | Note=Ala-rich |
| Hgene | CABLES1 | chr18:20716570 | chr16:54145673 | ENST00000420687 | + | 1 | 10 | 2_134 | 0 | 369.0 | Compositional bias | Note=Ala-rich |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145673 | ENST00000400473 | + | 1 | 10 | 2_134 | 0 | 307.0 | Compositional bias | Note=Ala-rich |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145673 | ENST00000420687 | + | 1 | 10 | 2_134 | 0 | 369.0 | Compositional bias | Note=Ala-rich |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145674 | ENST00000400473 | + | 1 | 10 | 2_134 | 0 | 307.0 | Compositional bias | Note=Ala-rich |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145674 | ENST00000420687 | + | 1 | 10 | 2_134 | 0 | 369.0 | Compositional bias | Note=Ala-rich |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000431610 | 3 | 5 | 32_327 | 55 | 107.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000460382 | 2 | 4 | 32_327 | 55 | 107.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000463855 | 1 | 3 | 32_327 | 76 | 128.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000471389 | 7 | 9 | 213_224 | 454 | 506.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000471389 | 7 | 9 | 231_234 | 454 | 506.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000471389 | 7 | 9 | 316_318 | 454 | 506.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716570 | chr16:54145673 | ENST00000471389 | 7 | 9 | 32_327 | 454 | 506.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000431610 | 3 | 5 | 32_327 | 55 | 107.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000460382 | 2 | 4 | 32_327 | 55 | 107.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000463855 | 1 | 3 | 32_327 | 76 | 128.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000471389 | 7 | 9 | 213_224 | 454 | 506.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000471389 | 7 | 9 | 231_234 | 454 | 506.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000471389 | 7 | 9 | 316_318 | 454 | 506.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145673 | ENST00000471389 | 7 | 9 | 32_327 | 454 | 506.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000431610 | 3 | 5 | 32_327 | 55 | 107.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000460382 | 2 | 4 | 32_327 | 55 | 107.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000463855 | 1 | 3 | 32_327 | 76 | 128.0 | Region | Fe2OG dioxygenase domain | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000471389 | 7 | 9 | 213_224 | 454 | 506.0 | Region | Loop L1%3B predicted to block binding of double-stranded DNA or RNA | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000471389 | 7 | 9 | 231_234 | 454 | 506.0 | Region | Substrate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000471389 | 7 | 9 | 316_318 | 454 | 506.0 | Region | Alpha-ketoglutarate binding | |
| Tgene | FTO | chr18:20716571 | chr16:54145674 | ENST00000471389 | 7 | 9 | 32_327 | 454 | 506.0 | Region | Fe2OG dioxygenase domain |
Top |
Fusion Gene Sequence for CABLES1-FTO |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >12246_12246_1_CABLES1-FTO_CABLES1_chr18_20716570_ENST00000256925_FTO_chr16_54145673_ENST00000394647_length(transcript)=1171nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA >12246_12246_1_CABLES1-FTO_CABLES1_chr18_20716570_ENST00000256925_FTO_chr16_54145673_ENST00000394647_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_2_CABLES1-FTO_CABLES1_chr18_20716570_ENST00000256925_FTO_chr16_54145673_ENST00000431610_length(transcript)=1222nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA >12246_12246_2_CABLES1-FTO_CABLES1_chr18_20716570_ENST00000256925_FTO_chr16_54145673_ENST00000431610_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_3_CABLES1-FTO_CABLES1_chr18_20716570_ENST00000256925_FTO_chr16_54145673_ENST00000460382_length(transcript)=2071nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA AAATGACCCTGTGTTATAGTCTGATTTGGTGTTAAACAGGACCTTCTTCCCCCAAAATTGTTCAGATTATAAAATGTGAGCCATTCAGCC CCCAAGGTCCAGGGCAGGCGACAGGAACGAGCCCAGCGTGTGACAAAGCCTAACCTACTTTCCTCTTTCCCAAGCTTTTTCAGAGACTCT GGAGTGGACCCAGCCCTCTGGGGAAAGACAGAACTTAGAGACATCCCAGTTACTCACCACACCCATAGTGCTGTCCAATATGGTAGCCAC TAGCTAGCTGTGGCTACTTCAATTTAAATTCAGTTTTAATTTTAATTAAAAATGCAGCTCTTCAGTCGCCCTGGCCACATTTCAAGTGCT TAACAGCCTCATGTGGCTAGTGACTGCTGTATTGGACGGTACAGATATGGAACATTTTCATCATCGAAGAAAGTCCTATTGGACAACACT TCTATAAAAAGTTTGAGAGCAGGAATTCTCATTTCCATTCGTCTGTAGCTTCTATCCCCAAAGGCAAAGAAACTAAAAGAGAAATGACTC ATTGAAGATTGGCCTCTTTCCTTTCTCTAAGACAAACCTAAGTAAAAGCCTGAGCTTTGAGTCCTATGCTCAGCACACGGGAAGGAGATG TTAATAATTAAAATAAAGTTGATATCCTGTCTTTAGGGAGTTCCCTTGATCTCTTGAAAGAGACACAGCCCCATTTACATTATTTCGTGG ATTTCACCAGCATAGTATAGTTTTTTTCTGTAAGTCCCTCATTCTTATGTAATAACAGGTGGAACTGAGGTTTGAAGAACCTCAGTGGCC CATCCTGATGACATTGGAGACTCAAAGAGACAAGAGAGAGTAGGGTTTAAAACCTGAGCTTTAAGACTCCCACTAGCTTCGTGTCCTTTG >12246_12246_3_CABLES1-FTO_CABLES1_chr18_20716570_ENST00000256925_FTO_chr16_54145673_ENST00000460382_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_4_CABLES1-FTO_CABLES1_chr18_20716570_ENST00000256925_FTO_chr16_54145673_ENST00000463855_length(transcript)=3547nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA AAATGACCCTGTGTTATAGTCTGATTTGGTGTTAAACAGGACCTTCTTCCCCCAAAATTGTTCAGATTATAAAATGTGAGCCATTCAGCC CCCAAGGTCCAGGGCAGGCGACAGGAACGAGCCCAGCGTGTGACAAAGCCTAACCTACTTTCCTCTTTCCCAAGCTTTTTCAGAGACTCT GGAGTGGACCCAGCCCTCTGGGGAAAGACAGAACTTAGAGACATCCCAGTTACTCACCACACCCATAGTGCTGTCCAATATGGTAGCCAC TAGCTAGCTGTGGCTACTTCAATTTAAATTCAGTTTTAATTTTAATTAAAAATGCAGCTCTTCAGTCGCCCTGGCCACATTTCAAGTGCT TAACAGCCTCATGTGGCTAGTGACTGCTGTATTGGACGGTACAGATATGGAACATTTTCATCATCGAAGAAAGTCCTATTGGACAACACT TCTATAAAAAGTTTGAGAGCAGGAATTCTCATTTCCATTCGTCTGTAGCTTCTATCCCCAAAGGCAAAGAAACTAAAAGAGAAATGACTC ATTGAAGATTGGCCTCTTTCCTTTCTCTAAGACAAACCTAAGTAAAAGCCTGAGCTTTGAGTCCTATGCTCAGCACACGGGAAGGAGATG TTAATAATTAAAATAAAGTTGATATCCTGTCTTTAGGGAGTTCCCTTGATCTCTTGAAAGAGACACAGCCCCATTTACATTATTTCGTGG ATTTCACCAGCATAGTATAGTTTTTTTCTGTAAGTCCCTCATTCTTATGTAATAACAGGTGGAACTGAGGTTTGAAGAACCTCAGTGGCC CATCCTGATGACATTGGAGACTCAAAGAGACAAGAGAGAGTAGGGTTTAAAACCTGAGCTTTAAGACTCCCACTAGCTTCGTGTCCTTTG GCATGTTAACGTGCCTCAGTTTCCTCATCTGTATAATGGGGATATATGAAAGGCACCAGTCCTAAGGTGAACATTAAGTGAGATGATTCT AGTTACAGACTTAGAACAATTTCCAGCACATAGTTAAATATCCAGGAAATTCTGGTACTGTTATGTGTGGGTGAGCTGACCTGGATGTAG ATGTTTTCCTCTCTCTTGCTGACCCCTCCGCCAGTTTTGTCTTGTGATGCCATTAACACATCTCTCCCTTTCTGACCTGGCTCCTGCCCA TTGGTGTCCCAAGAAATCGTGAGAATAGTTAGCCCCCCGTCTCCCCAGCCTGTTGCTTTCTCGTGTAGTTGTTCACAGTAGTTGAGAAGT TGAAGAGCTTTTGCCTATTGAAGGTGCACTGAGAATAAACTCTTTCCTGCCACCAGAATTGCAGTGGTTCACGGCCTGCACTCATTCCCA TGAATGCAGTTAATAGCCACAGAAATGTCACATTAAGCAAAGCAGCCAGGGTCTCATCGTGTTGAGACTCGAGTCTCTCAGACCTTGGAT TCATTCCCTGGTGTCTTTGAGCCTCAGTTTCCTCATTGGTAAAAGAGAAGTGAAGCAGTGTCTCACAGGGTCATTACAGAGATTAAATGA AATAAATGAAATAACATAGACCAGGAGGGCGTGGTGTTTAAAAGTCACAGATGGGGCACCCTCGGGCCATCCAGCCCAGTGTTTTCTTTA GCCCCTATGATGTTCATTTTTTGTTATATCCCATTAGGTGCCCATATTTAAAAATTGGGAGATTTCACATAAAATTAAAAGGTCTGCATT TTCTTTTTTCTTTTCTTTTTTTTTTTTTTTGAGACACAGTCTCACTCTGTCACCAGGCTAGAGTGCAGTGGCACGATCTCAGCTCACTGC AACCTCTGCCTCCCAGGTTCAAGTAATTCTCCTGCCTCAGCCTCCCAAGTAGCTGGGACTACAGGCACGTGCCACCACGCCCAGCTAATT TTTGTATTTTTAGCAGAGATGGGGTTTCACCACATTGGCCAGGATGGTCTCGATCTCAACCTCGTGATCCACCCACCTCGGTCTCCCAAA GCGCTGGGATTACAGGCGTGAGCCACCGCGCCAAGCCAAGGTCTGCATTTTTCTTTAGAACTCAGAACACCCAATAGTCCTAGGCCCCCA TCCTCGCATGGCAGCAAGCTAAATAAGCATCTTCCCACTGCGAGTTGGGGCATGACCCAGCCTATGGTTTGCCATACTCCCTCTTTTTCT CCGTTTTTTCATTAATTGTGAACCTGACCTGCATCACCCTTTCATGTCAGTGCTCTCCAAACCTGCTTGCTTGCACCCCTCTAGTCGAAA TATTTTGTGCTTACCCCAATATATGTGTGTGACTATTGAACTCTATTCGTAGACTGCTTGTACTAATGTCATTTGCATCATAAAATATTC >12246_12246_4_CABLES1-FTO_CABLES1_chr18_20716570_ENST00000256925_FTO_chr16_54145673_ENST00000463855_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_5_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145673_ENST00000394647_length(transcript)=1171nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA >12246_12246_5_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145673_ENST00000394647_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_6_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145673_ENST00000431610_length(transcript)=1222nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA >12246_12246_6_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145673_ENST00000431610_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_7_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145673_ENST00000460382_length(transcript)=2071nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA AAATGACCCTGTGTTATAGTCTGATTTGGTGTTAAACAGGACCTTCTTCCCCCAAAATTGTTCAGATTATAAAATGTGAGCCATTCAGCC CCCAAGGTCCAGGGCAGGCGACAGGAACGAGCCCAGCGTGTGACAAAGCCTAACCTACTTTCCTCTTTCCCAAGCTTTTTCAGAGACTCT GGAGTGGACCCAGCCCTCTGGGGAAAGACAGAACTTAGAGACATCCCAGTTACTCACCACACCCATAGTGCTGTCCAATATGGTAGCCAC TAGCTAGCTGTGGCTACTTCAATTTAAATTCAGTTTTAATTTTAATTAAAAATGCAGCTCTTCAGTCGCCCTGGCCACATTTCAAGTGCT TAACAGCCTCATGTGGCTAGTGACTGCTGTATTGGACGGTACAGATATGGAACATTTTCATCATCGAAGAAAGTCCTATTGGACAACACT TCTATAAAAAGTTTGAGAGCAGGAATTCTCATTTCCATTCGTCTGTAGCTTCTATCCCCAAAGGCAAAGAAACTAAAAGAGAAATGACTC ATTGAAGATTGGCCTCTTTCCTTTCTCTAAGACAAACCTAAGTAAAAGCCTGAGCTTTGAGTCCTATGCTCAGCACACGGGAAGGAGATG TTAATAATTAAAATAAAGTTGATATCCTGTCTTTAGGGAGTTCCCTTGATCTCTTGAAAGAGACACAGCCCCATTTACATTATTTCGTGG ATTTCACCAGCATAGTATAGTTTTTTTCTGTAAGTCCCTCATTCTTATGTAATAACAGGTGGAACTGAGGTTTGAAGAACCTCAGTGGCC CATCCTGATGACATTGGAGACTCAAAGAGACAAGAGAGAGTAGGGTTTAAAACCTGAGCTTTAAGACTCCCACTAGCTTCGTGTCCTTTG >12246_12246_7_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145673_ENST00000460382_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_8_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145673_ENST00000463855_length(transcript)=3547nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA AAATGACCCTGTGTTATAGTCTGATTTGGTGTTAAACAGGACCTTCTTCCCCCAAAATTGTTCAGATTATAAAATGTGAGCCATTCAGCC CCCAAGGTCCAGGGCAGGCGACAGGAACGAGCCCAGCGTGTGACAAAGCCTAACCTACTTTCCTCTTTCCCAAGCTTTTTCAGAGACTCT GGAGTGGACCCAGCCCTCTGGGGAAAGACAGAACTTAGAGACATCCCAGTTACTCACCACACCCATAGTGCTGTCCAATATGGTAGCCAC TAGCTAGCTGTGGCTACTTCAATTTAAATTCAGTTTTAATTTTAATTAAAAATGCAGCTCTTCAGTCGCCCTGGCCACATTTCAAGTGCT TAACAGCCTCATGTGGCTAGTGACTGCTGTATTGGACGGTACAGATATGGAACATTTTCATCATCGAAGAAAGTCCTATTGGACAACACT TCTATAAAAAGTTTGAGAGCAGGAATTCTCATTTCCATTCGTCTGTAGCTTCTATCCCCAAAGGCAAAGAAACTAAAAGAGAAATGACTC ATTGAAGATTGGCCTCTTTCCTTTCTCTAAGACAAACCTAAGTAAAAGCCTGAGCTTTGAGTCCTATGCTCAGCACACGGGAAGGAGATG TTAATAATTAAAATAAAGTTGATATCCTGTCTTTAGGGAGTTCCCTTGATCTCTTGAAAGAGACACAGCCCCATTTACATTATTTCGTGG ATTTCACCAGCATAGTATAGTTTTTTTCTGTAAGTCCCTCATTCTTATGTAATAACAGGTGGAACTGAGGTTTGAAGAACCTCAGTGGCC CATCCTGATGACATTGGAGACTCAAAGAGACAAGAGAGAGTAGGGTTTAAAACCTGAGCTTTAAGACTCCCACTAGCTTCGTGTCCTTTG GCATGTTAACGTGCCTCAGTTTCCTCATCTGTATAATGGGGATATATGAAAGGCACCAGTCCTAAGGTGAACATTAAGTGAGATGATTCT AGTTACAGACTTAGAACAATTTCCAGCACATAGTTAAATATCCAGGAAATTCTGGTACTGTTATGTGTGGGTGAGCTGACCTGGATGTAG ATGTTTTCCTCTCTCTTGCTGACCCCTCCGCCAGTTTTGTCTTGTGATGCCATTAACACATCTCTCCCTTTCTGACCTGGCTCCTGCCCA TTGGTGTCCCAAGAAATCGTGAGAATAGTTAGCCCCCCGTCTCCCCAGCCTGTTGCTTTCTCGTGTAGTTGTTCACAGTAGTTGAGAAGT TGAAGAGCTTTTGCCTATTGAAGGTGCACTGAGAATAAACTCTTTCCTGCCACCAGAATTGCAGTGGTTCACGGCCTGCACTCATTCCCA TGAATGCAGTTAATAGCCACAGAAATGTCACATTAAGCAAAGCAGCCAGGGTCTCATCGTGTTGAGACTCGAGTCTCTCAGACCTTGGAT TCATTCCCTGGTGTCTTTGAGCCTCAGTTTCCTCATTGGTAAAAGAGAAGTGAAGCAGTGTCTCACAGGGTCATTACAGAGATTAAATGA AATAAATGAAATAACATAGACCAGGAGGGCGTGGTGTTTAAAAGTCACAGATGGGGCACCCTCGGGCCATCCAGCCCAGTGTTTTCTTTA GCCCCTATGATGTTCATTTTTTGTTATATCCCATTAGGTGCCCATATTTAAAAATTGGGAGATTTCACATAAAATTAAAAGGTCTGCATT TTCTTTTTTCTTTTCTTTTTTTTTTTTTTTGAGACACAGTCTCACTCTGTCACCAGGCTAGAGTGCAGTGGCACGATCTCAGCTCACTGC AACCTCTGCCTCCCAGGTTCAAGTAATTCTCCTGCCTCAGCCTCCCAAGTAGCTGGGACTACAGGCACGTGCCACCACGCCCAGCTAATT TTTGTATTTTTAGCAGAGATGGGGTTTCACCACATTGGCCAGGATGGTCTCGATCTCAACCTCGTGATCCACCCACCTCGGTCTCCCAAA GCGCTGGGATTACAGGCGTGAGCCACCGCGCCAAGCCAAGGTCTGCATTTTTCTTTAGAACTCAGAACACCCAATAGTCCTAGGCCCCCA TCCTCGCATGGCAGCAAGCTAAATAAGCATCTTCCCACTGCGAGTTGGGGCATGACCCAGCCTATGGTTTGCCATACTCCCTCTTTTTCT CCGTTTTTTCATTAATTGTGAACCTGACCTGCATCACCCTTTCATGTCAGTGCTCTCCAAACCTGCTTGCTTGCACCCCTCTAGTCGAAA TATTTTGTGCTTACCCCAATATATGTGTGTGACTATTGAACTCTATTCGTAGACTGCTTGTACTAATGTCATTTGCATCATAAAATATTC >12246_12246_8_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145673_ENST00000463855_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_9_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145674_ENST00000394647_length(transcript)=1171nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA >12246_12246_9_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145674_ENST00000394647_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_10_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145674_ENST00000431610_length(transcript)=1222nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA >12246_12246_10_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145674_ENST00000431610_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_11_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145674_ENST00000460382_length(transcript)=2071nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA AAATGACCCTGTGTTATAGTCTGATTTGGTGTTAAACAGGACCTTCTTCCCCCAAAATTGTTCAGATTATAAAATGTGAGCCATTCAGCC CCCAAGGTCCAGGGCAGGCGACAGGAACGAGCCCAGCGTGTGACAAAGCCTAACCTACTTTCCTCTTTCCCAAGCTTTTTCAGAGACTCT GGAGTGGACCCAGCCCTCTGGGGAAAGACAGAACTTAGAGACATCCCAGTTACTCACCACACCCATAGTGCTGTCCAATATGGTAGCCAC TAGCTAGCTGTGGCTACTTCAATTTAAATTCAGTTTTAATTTTAATTAAAAATGCAGCTCTTCAGTCGCCCTGGCCACATTTCAAGTGCT TAACAGCCTCATGTGGCTAGTGACTGCTGTATTGGACGGTACAGATATGGAACATTTTCATCATCGAAGAAAGTCCTATTGGACAACACT TCTATAAAAAGTTTGAGAGCAGGAATTCTCATTTCCATTCGTCTGTAGCTTCTATCCCCAAAGGCAAAGAAACTAAAAGAGAAATGACTC ATTGAAGATTGGCCTCTTTCCTTTCTCTAAGACAAACCTAAGTAAAAGCCTGAGCTTTGAGTCCTATGCTCAGCACACGGGAAGGAGATG TTAATAATTAAAATAAAGTTGATATCCTGTCTTTAGGGAGTTCCCTTGATCTCTTGAAAGAGACACAGCCCCATTTACATTATTTCGTGG ATTTCACCAGCATAGTATAGTTTTTTTCTGTAAGTCCCTCATTCTTATGTAATAACAGGTGGAACTGAGGTTTGAAGAACCTCAGTGGCC CATCCTGATGACATTGGAGACTCAAAGAGACAAGAGAGAGTAGGGTTTAAAACCTGAGCTTTAAGACTCCCACTAGCTTCGTGTCCTTTG >12246_12246_11_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145674_ENST00000460382_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- >12246_12246_12_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145674_ENST00000463855_length(transcript)=3547nt_BP=845nt ATGGCGGCGGCGGCGGCGGCCGCCACCACGGCCGCCTGCAGCAGCGGCAGCGCCGGCACCGACGCCGCGGGCGCCAGCGGATTGCAGCAG CCGCCGCCGCAGCCCCAGCCTCAGCCCGCGGCCGCCGCGCCGGCCCAGCCGCCGCCCGAACCCCCCCGGAAGCCGCGCATGGACCCGCGG CGCCGCCAGGCTGCCCTCTCCTTCCTCACCAACATCTCGCTGGACGGCCGGCTGCCGCCGCAGGACGCGGAGTGGGGCGGTGGCGAGGAG GGCGGCGCGGCCAAGCCGGGCGCCGGCGGCGCCTGCGGCGCGAGGACTCGGTTCAGCTTGCTCGCCGCTGCCGAGCGGGGCGGCTGCATC GCGCTCGCCGCGCCGGGCACGCCGGCTGCGGGGTTAGCCGCTGGGTCCGGCCCCTGCCTCCCACAGCCCTCGTCGCTGCCGCCCTTGATT CCTGGCGGCCATGCGACCGTGTCCGGCCCCGGGGTGGCGCGGGGGTTCGCGAGTCCCCTGGGCGCCGGCCGGGCGTCGGGGGAGCAGTGG CAGCCGCCCCGGCCGGCGCCTCTCGCCGCCTGTGCCCAACTGCAGCTGCTCGACGGGTCCGGGGCCGCCGGGCAGGAGGAGTTGGAGGAG GACGATGCCTTTATCAGCGTGCAGGTGCCGGCGGCCGCCTTTTTGGGCTCCGGGACCCCCGGGAGTGGGAGCGGCAGTCGGGGACGCCTC AACTCGTTCACTCAGGGAATCCTGCCCATCGCCTTCTCCAGGCCGACTTCGCAGAACTACTGCTCCCTGGAGCAGCCAGGCCAGGGCGGC AGCACCAGCGCCTTCGAGCAGCTGCAGAGGTCCCGGTGCCAGTCACGAATTGCCCGAACATTACCTGCTGATCAGAAGCCAGAATGTCGG CCATACTGGGAAAAGGATGATGCTTCGATGCCTCTGCCGTTTGACCTCACAGACATCGTTTCAGAACTCAGAGGTCAGCTTCTGGAAGCA AAACCCTAGAAGGAGCACAAGTCTCAGGCGGAGGAGAAAAAGAGATCGGCTTTTCTCCTCCAACGTTGTCATGGGCTTAAGCAAGAGCAG TGGAGACTTCTCTTGGCCCCTAGATTGTAGCACCCGGGTCCCAATCCAAAACAGCTAGGAAATGGTGCCCATGAAGTTTTAAATGTTTTA AAATGACCCTGTGTTATAGTCTGATTTGGTGTTAAACAGGACCTTCTTCCCCCAAAATTGTTCAGATTATAAAATGTGAGCCATTCAGCC CCCAAGGTCCAGGGCAGGCGACAGGAACGAGCCCAGCGTGTGACAAAGCCTAACCTACTTTCCTCTTTCCCAAGCTTTTTCAGAGACTCT GGAGTGGACCCAGCCCTCTGGGGAAAGACAGAACTTAGAGACATCCCAGTTACTCACCACACCCATAGTGCTGTCCAATATGGTAGCCAC TAGCTAGCTGTGGCTACTTCAATTTAAATTCAGTTTTAATTTTAATTAAAAATGCAGCTCTTCAGTCGCCCTGGCCACATTTCAAGTGCT TAACAGCCTCATGTGGCTAGTGACTGCTGTATTGGACGGTACAGATATGGAACATTTTCATCATCGAAGAAAGTCCTATTGGACAACACT TCTATAAAAAGTTTGAGAGCAGGAATTCTCATTTCCATTCGTCTGTAGCTTCTATCCCCAAAGGCAAAGAAACTAAAAGAGAAATGACTC ATTGAAGATTGGCCTCTTTCCTTTCTCTAAGACAAACCTAAGTAAAAGCCTGAGCTTTGAGTCCTATGCTCAGCACACGGGAAGGAGATG TTAATAATTAAAATAAAGTTGATATCCTGTCTTTAGGGAGTTCCCTTGATCTCTTGAAAGAGACACAGCCCCATTTACATTATTTCGTGG ATTTCACCAGCATAGTATAGTTTTTTTCTGTAAGTCCCTCATTCTTATGTAATAACAGGTGGAACTGAGGTTTGAAGAACCTCAGTGGCC CATCCTGATGACATTGGAGACTCAAAGAGACAAGAGAGAGTAGGGTTTAAAACCTGAGCTTTAAGACTCCCACTAGCTTCGTGTCCTTTG GCATGTTAACGTGCCTCAGTTTCCTCATCTGTATAATGGGGATATATGAAAGGCACCAGTCCTAAGGTGAACATTAAGTGAGATGATTCT AGTTACAGACTTAGAACAATTTCCAGCACATAGTTAAATATCCAGGAAATTCTGGTACTGTTATGTGTGGGTGAGCTGACCTGGATGTAG ATGTTTTCCTCTCTCTTGCTGACCCCTCCGCCAGTTTTGTCTTGTGATGCCATTAACACATCTCTCCCTTTCTGACCTGGCTCCTGCCCA TTGGTGTCCCAAGAAATCGTGAGAATAGTTAGCCCCCCGTCTCCCCAGCCTGTTGCTTTCTCGTGTAGTTGTTCACAGTAGTTGAGAAGT TGAAGAGCTTTTGCCTATTGAAGGTGCACTGAGAATAAACTCTTTCCTGCCACCAGAATTGCAGTGGTTCACGGCCTGCACTCATTCCCA TGAATGCAGTTAATAGCCACAGAAATGTCACATTAAGCAAAGCAGCCAGGGTCTCATCGTGTTGAGACTCGAGTCTCTCAGACCTTGGAT TCATTCCCTGGTGTCTTTGAGCCTCAGTTTCCTCATTGGTAAAAGAGAAGTGAAGCAGTGTCTCACAGGGTCATTACAGAGATTAAATGA AATAAATGAAATAACATAGACCAGGAGGGCGTGGTGTTTAAAAGTCACAGATGGGGCACCCTCGGGCCATCCAGCCCAGTGTTTTCTTTA GCCCCTATGATGTTCATTTTTTGTTATATCCCATTAGGTGCCCATATTTAAAAATTGGGAGATTTCACATAAAATTAAAAGGTCTGCATT TTCTTTTTTCTTTTCTTTTTTTTTTTTTTTGAGACACAGTCTCACTCTGTCACCAGGCTAGAGTGCAGTGGCACGATCTCAGCTCACTGC AACCTCTGCCTCCCAGGTTCAAGTAATTCTCCTGCCTCAGCCTCCCAAGTAGCTGGGACTACAGGCACGTGCCACCACGCCCAGCTAATT TTTGTATTTTTAGCAGAGATGGGGTTTCACCACATTGGCCAGGATGGTCTCGATCTCAACCTCGTGATCCACCCACCTCGGTCTCCCAAA GCGCTGGGATTACAGGCGTGAGCCACCGCGCCAAGCCAAGGTCTGCATTTTTCTTTAGAACTCAGAACACCCAATAGTCCTAGGCCCCCA TCCTCGCATGGCAGCAAGCTAAATAAGCATCTTCCCACTGCGAGTTGGGGCATGACCCAGCCTATGGTTTGCCATACTCCCTCTTTTTCT CCGTTTTTTCATTAATTGTGAACCTGACCTGCATCACCCTTTCATGTCAGTGCTCTCCAAACCTGCTTGCTTGCACCCCTCTAGTCGAAA TATTTTGTGCTTACCCCAATATATGTGTGTGACTATTGAACTCTATTCGTAGACTGCTTGTACTAATGTCATTTGCATCATAAAATATTC >12246_12246_12_CABLES1-FTO_CABLES1_chr18_20716571_ENST00000256925_FTO_chr16_54145674_ENST00000463855_length(amino acids)=332AA_BP=280 MAAAAAAATTAACSSGSAGTDAAGASGLQQPPPQPQPQPAAAAPAQPPPEPPRKPRMDPRRRQAALSFLTNISLDGRLPPQDAEWGGGEE GGAAKPGAGGACGARTRFSLLAAAERGGCIALAAPGTPAAGLAAGSGPCLPQPSSLPPLIPGGHATVSGPGVARGFASPLGAGRASGEQW QPPRPAPLAACAQLQLLDGSGAAGQEELEEDDAFISVQVPAAAFLGSGTPGSGSGSRGRLNSFTQGILPIAFSRPTSQNYCSLEQPGQGG -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CABLES1-FTO |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Hgene | CABLES1 | chr18:20716570 | chr16:54145673 | ENST00000256925 | + | 1 | 10 | 1_109 | 281.6666666666667 | 634.0 | TDRD7 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145673 | ENST00000256925 | + | 1 | 10 | 1_109 | 281.6666666666667 | 634.0 | TDRD7 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145674 | ENST00000256925 | + | 1 | 10 | 1_109 | 281.6666666666667 | 634.0 | TDRD7 |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | CABLES1 | chr18:20716570 | chr16:54145673 | ENST00000256925 | + | 1 | 10 | 179_492 | 281.6666666666667 | 634.0 | CDK3 |
| Hgene | CABLES1 | chr18:20716570 | chr16:54145673 | ENST00000400473 | + | 1 | 10 | 179_492 | 0 | 307.0 | CDK3 |
| Hgene | CABLES1 | chr18:20716570 | chr16:54145673 | ENST00000420687 | + | 1 | 10 | 179_492 | 0 | 369.0 | CDK3 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145673 | ENST00000256925 | + | 1 | 10 | 179_492 | 281.6666666666667 | 634.0 | CDK3 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145673 | ENST00000400473 | + | 1 | 10 | 179_492 | 0 | 307.0 | CDK3 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145673 | ENST00000420687 | + | 1 | 10 | 179_492 | 0 | 369.0 | CDK3 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145674 | ENST00000256925 | + | 1 | 10 | 179_492 | 281.6666666666667 | 634.0 | CDK3 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145674 | ENST00000400473 | + | 1 | 10 | 179_492 | 0 | 307.0 | CDK3 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145674 | ENST00000420687 | + | 1 | 10 | 179_492 | 0 | 369.0 | CDK3 |
| Hgene | CABLES1 | chr18:20716570 | chr16:54145673 | ENST00000400473 | + | 1 | 10 | 1_109 | 0 | 307.0 | TDRD7 |
| Hgene | CABLES1 | chr18:20716570 | chr16:54145673 | ENST00000420687 | + | 1 | 10 | 1_109 | 0 | 369.0 | TDRD7 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145673 | ENST00000400473 | + | 1 | 10 | 1_109 | 0 | 307.0 | TDRD7 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145673 | ENST00000420687 | + | 1 | 10 | 1_109 | 0 | 369.0 | TDRD7 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145674 | ENST00000400473 | + | 1 | 10 | 1_109 | 0 | 307.0 | TDRD7 |
| Hgene | CABLES1 | chr18:20716571 | chr16:54145674 | ENST00000420687 | + | 1 | 10 | 1_109 | 0 | 369.0 | TDRD7 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CABLES1-FTO |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CABLES1-FTO |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
