
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CADPS-FHIT (FusionGDB2 ID:12483) |
Fusion Gene Summary for CADPS-FHIT |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CADPS-FHIT | Fusion gene ID: 12483 | Hgene | Tgene | Gene symbol | CADPS | FHIT | Gene ID | 8618 | 2272 |
| Gene name | calcium dependent secretion activator | fragile histidine triad diadenosine triphosphatase | |
| Synonyms | CADPS1|CAPS|CAPS1|UNC-31 | AP3Aase|FRA3B | |
| Cytomap | 3p14.2 | 3p14.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | calcium-dependent secretion activator 1CAPS-1Ca++-dependent secretion activatorCa2+ dependent secretion activatorCa2+-dependent activator protein for secretionCa2+-regulated cytoskeletal proteincalcium-dependent activator protein for secretion 1 | bis(5'-adenosyl)-triphosphataseAP3A hydrolasediadenosine 5',5'''-P1,P3-triphosphate hydrolasedinucleosidetriphosphatase | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q9ULU8 | P49789 | |
| Ensembl transtripts involved in fusion gene | ENST00000283269, ENST00000357948, ENST00000383710, ENST00000490353, ENST00000462768, | ENST00000341848, ENST00000466788, ENST00000468189, ENST00000476844, ENST00000492590, | |
| Fusion gene scores | * DoF score | 3 X 3 X 3=27 | 27 X 20 X 11=5940 |
| # samples | 3 | 32 | |
| ** MAII score | log2(3/27*10)=0.15200309344505 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(32/5940*10)=-4.21431912080077 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CADPS [Title/Abstract] AND FHIT [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CADPS(62860264)-FHIT(59908140), # samples:1 FHIT(61027719)-CADPS(62484943), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | FHIT | GO:0006163 | purine nucleotide metabolic process | 9323207 |
 Fusion gene breakpoints across CADPS (5'-gene) Fusion gene breakpoints across CADPS (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
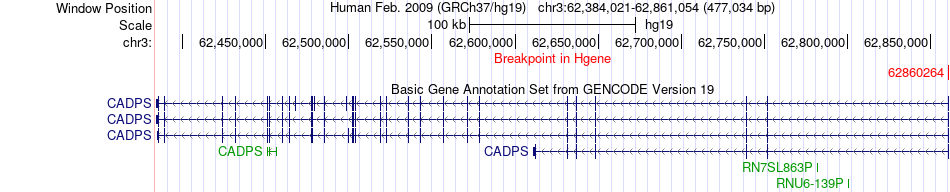 |
 Fusion gene breakpoints across FHIT (3'-gene) Fusion gene breakpoints across FHIT (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
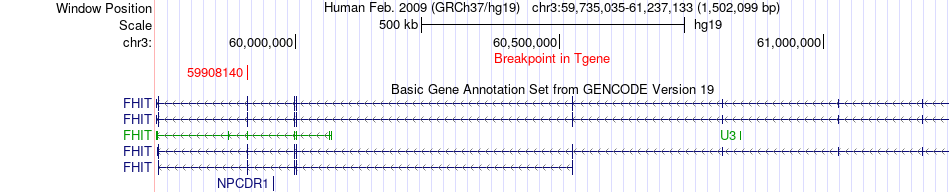 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-E9-A22E-01A | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
Top |
Fusion Gene ORF analysis for CADPS-FHIT |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000283269 | ENST00000341848 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000283269 | ENST00000466788 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000283269 | ENST00000468189 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000357948 | ENST00000341848 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000357948 | ENST00000466788 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000357948 | ENST00000468189 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000383710 | ENST00000341848 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000383710 | ENST00000466788 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000383710 | ENST00000468189 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000490353 | ENST00000341848 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000490353 | ENST00000466788 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| 5CDS-5UTR | ENST00000490353 | ENST00000468189 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| In-frame | ENST00000283269 | ENST00000476844 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| In-frame | ENST00000283269 | ENST00000492590 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| In-frame | ENST00000357948 | ENST00000476844 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| In-frame | ENST00000357948 | ENST00000492590 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| In-frame | ENST00000383710 | ENST00000476844 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| In-frame | ENST00000383710 | ENST00000492590 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| In-frame | ENST00000490353 | ENST00000476844 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| In-frame | ENST00000490353 | ENST00000492590 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| intron-3CDS | ENST00000462768 | ENST00000476844 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| intron-3CDS | ENST00000462768 | ENST00000492590 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| intron-5UTR | ENST00000462768 | ENST00000341848 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| intron-5UTR | ENST00000462768 | ENST00000466788 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
| intron-5UTR | ENST00000462768 | ENST00000468189 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000383710 | CADPS | chr3 | 62860264 | - | ENST00000476844 | FHIT | chr3 | 59908140 | - | 1220 | 791 | 871 | 41 | 276 |
| ENST00000383710 | CADPS | chr3 | 62860264 | - | ENST00000492590 | FHIT | chr3 | 59908140 | - | 1231 | 791 | 278 | 955 | 225 |
| ENST00000357948 | CADPS | chr3 | 62860264 | - | ENST00000476844 | FHIT | chr3 | 59908140 | - | 1179 | 750 | 830 | 0 | 276 |
| ENST00000357948 | CADPS | chr3 | 62860264 | - | ENST00000492590 | FHIT | chr3 | 59908140 | - | 1190 | 750 | 237 | 914 | 225 |
| ENST00000283269 | CADPS | chr3 | 62860264 | - | ENST00000476844 | FHIT | chr3 | 59908140 | - | 1082 | 653 | 733 | 2 | 244 |
| ENST00000283269 | CADPS | chr3 | 62860264 | - | ENST00000492590 | FHIT | chr3 | 59908140 | - | 1093 | 653 | 140 | 817 | 225 |
| ENST00000490353 | CADPS | chr3 | 62860264 | - | ENST00000476844 | FHIT | chr3 | 59908140 | - | 870 | 441 | 0 | 605 | 201 |
| ENST00000490353 | CADPS | chr3 | 62860264 | - | ENST00000492590 | FHIT | chr3 | 59908140 | - | 881 | 441 | 0 | 605 | 201 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000383710 | ENST00000476844 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - | 0.002684913 | 0.9973151 |
| ENST00000383710 | ENST00000492590 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - | 0.002579735 | 0.9974203 |
| ENST00000357948 | ENST00000476844 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - | 0.003773011 | 0.996227 |
| ENST00000357948 | ENST00000492590 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - | 0.003299749 | 0.9967002 |
| ENST00000283269 | ENST00000476844 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - | 0.003689748 | 0.9963103 |
| ENST00000283269 | ENST00000492590 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - | 0.003597081 | 0.996403 |
| ENST00000490353 | ENST00000476844 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - | 0.002611053 | 0.997389 |
| ENST00000490353 | ENST00000492590 | CADPS | chr3 | 62860264 | - | FHIT | chr3 | 59908140 | - | 0.002443738 | 0.9975562 |
Top |
Fusion Genomic Features for CADPS-FHIT |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
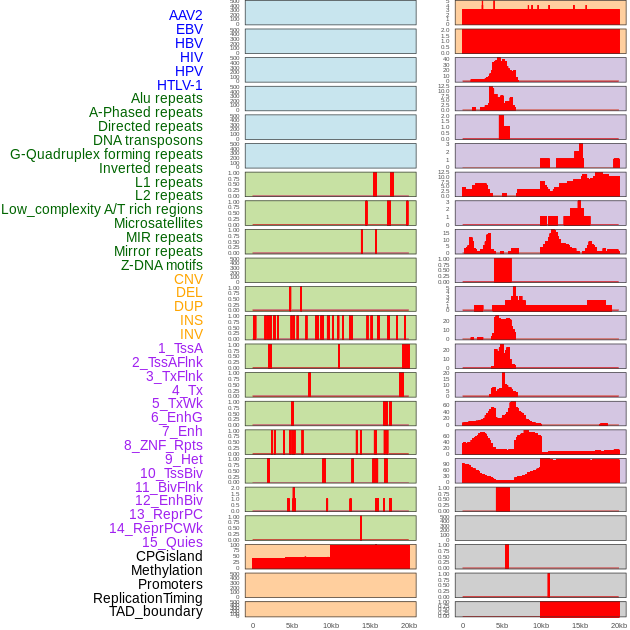 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for CADPS-FHIT |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr3:62860264/chr3:59908140) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CADPS | FHIT |
| FUNCTION: Calcium-binding protein involved in exocytosis of vesicles filled with neurotransmitters and neuropeptides. Probably acts upstream of fusion in the biogenesis or maintenance of mature secretory vesicles. Regulates catecholamine loading of DCVs. May specifically mediate the Ca(2+)-dependent exocytosis of large dense-core vesicles (DCVs) and other dense-core vesicles by acting as a PtdIns(4,5)P2-binding protein that acts at prefusion step following ATP-dependent priming and participates in DCVs-membrane fusion. However, it may also participate in small clear synaptic vesicles (SVs) exocytosis and it is unclear whether its function is related to Ca(2+) triggering (By similarity). {ECO:0000250}. | FUNCTION: Possesses dinucleoside triphosphate hydrolase activity (PubMed:12574506, PubMed:15182206, PubMed:8794732, PubMed:9323207, PubMed:9576908, PubMed:9543008). Cleaves P(1)-P(3)-bis(5'-adenosyl) triphosphate (Ap3A) to yield AMP and ADP (PubMed:12574506, PubMed:15182206, PubMed:8794732, PubMed:9323207, PubMed:9576908, PubMed:9543008). Can also hydrolyze P(1)-P(4)-bis(5'-adenosyl) tetraphosphate (Ap4A), but has extremely low activity with ATP (PubMed:8794732). Exhibits adenylylsulfatase activity, hydrolyzing adenosine 5'-phosphosulfate to yield AMP and sulfate (PubMed:18694747). Exhibits adenosine 5'-monophosphoramidase activity, hydrolyzing purine nucleotide phosphoramidates with a single phosphate group such as adenosine 5'monophosphoramidate (AMP-NH2) to yield AMP and NH2 (PubMed:18694747). Exhibits adenylylsulfate-ammonia adenylyltransferase, catalyzing the ammonolysis of adenosine 5'-phosphosulfate resulting in the formation of adenosine 5'-phosphoramidate (PubMed:26181368). Also catalyzes the ammonolysis of adenosine 5-phosphorofluoridate and diadenosine triphosphate (PubMed:26181368). Modulates transcriptional activation by CTNNB1 and thereby contributes to regulate the expression of genes essential for cell proliferation and survival, such as CCND1 and BIRC5 (PubMed:18077326). Plays a role in the induction of apoptosis via SRC and AKT1 signaling pathways (PubMed:16407838). Inhibits MDM2-mediated proteasomal degradation of p53/TP53 and thereby plays a role in p53/TP53-mediated apoptosis (PubMed:15313915). Induction of apoptosis depends on the ability of FHIT to bind P(1)-P(3)-bis(5'-adenosyl) triphosphate or related compounds, but does not require its catalytic activity, it may in part come from the mitochondrial form, which sensitizes the low-affinity Ca(2+) transporters, enhancing mitochondrial calcium uptake (PubMed:12574506, PubMed:19622739). Functions as tumor suppressor (By similarity). {ECO:0000250|UniProtKB:O89106, ECO:0000269|PubMed:12574506, ECO:0000269|PubMed:15313915, ECO:0000269|PubMed:16407838, ECO:0000269|PubMed:18077326, ECO:0000269|PubMed:18694747, ECO:0000269|PubMed:19622739, ECO:0000269|PubMed:26181368, ECO:0000269|PubMed:8794732, ECO:0000269|PubMed:9323207, ECO:0000269|PubMed:9543008}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000341848 | 2 | 5 | 94_98 | 93 | 148.0 | Motif | Histidine triad motif | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000468189 | 6 | 9 | 94_98 | 93 | 148.0 | Motif | Histidine triad motif | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000476844 | 6 | 10 | 94_98 | 93 | 214.0 | Motif | Histidine triad motif | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000492590 | 6 | 10 | 94_98 | 93 | 205.33333333333334 | Motif | Histidine triad motif |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000283269 | - | 1 | 28 | 380_495 | 147 | 1315.0 | Domain | C2 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000283269 | - | 1 | 28 | 521_624 | 147 | 1315.0 | Domain | PH |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000283269 | - | 1 | 28 | 931_1111 | 147 | 1315.0 | Domain | MHD1 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000357948 | - | 1 | 27 | 380_495 | 147 | 1275.0 | Domain | C2 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000357948 | - | 1 | 27 | 521_624 | 147 | 1275.0 | Domain | PH |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000357948 | - | 1 | 27 | 931_1111 | 147 | 1275.0 | Domain | MHD1 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000383710 | - | 1 | 30 | 380_495 | 147 | 1354.0 | Domain | C2 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000383710 | - | 1 | 30 | 521_624 | 147 | 1354.0 | Domain | PH |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000383710 | - | 1 | 30 | 931_1111 | 147 | 1354.0 | Domain | MHD1 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000490353 | - | 1 | 7 | 380_495 | 147 | 458.0 | Domain | C2 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000490353 | - | 1 | 7 | 521_624 | 147 | 458.0 | Domain | PH |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000490353 | - | 1 | 7 | 931_1111 | 147 | 458.0 | Domain | MHD1 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000283269 | - | 1 | 28 | 1177_1353 | 147 | 1315.0 | Region | Mediates targeting and association with DCVs |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000357948 | - | 1 | 27 | 1177_1353 | 147 | 1275.0 | Region | Mediates targeting and association with DCVs |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000383710 | - | 1 | 30 | 1177_1353 | 147 | 1354.0 | Region | Mediates targeting and association with DCVs |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000490353 | - | 1 | 7 | 1177_1353 | 147 | 458.0 | Region | Mediates targeting and association with DCVs |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000341848 | 2 | 5 | 2_109 | 93 | 148.0 | Domain | HIT | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000468189 | 6 | 9 | 2_109 | 93 | 148.0 | Domain | HIT | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000476844 | 6 | 10 | 2_109 | 93 | 214.0 | Domain | HIT | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000492590 | 6 | 10 | 2_109 | 93 | 205.33333333333334 | Domain | HIT | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000341848 | 2 | 5 | 89_92 | 93 | 148.0 | Nucleotide binding | Substrate | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000468189 | 6 | 9 | 89_92 | 93 | 148.0 | Nucleotide binding | Substrate | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000476844 | 6 | 10 | 89_92 | 93 | 214.0 | Nucleotide binding | Substrate | |
| Tgene | FHIT | chr3:62860264 | chr3:59908140 | ENST00000492590 | 6 | 10 | 89_92 | 93 | 205.33333333333334 | Nucleotide binding | Substrate |
Top |
Fusion Gene Sequence for CADPS-FHIT |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >12483_12483_1_CADPS-FHIT_CADPS_chr3_62860264_ENST00000283269_FHIT_chr3_59908140_ENST00000476844_length(transcript)=1082nt_BP=653nt CTCCTCCCTTCGCGCTTCTGGGTCTGAGCCCAGCTCGCGACCGCCGGGCAGAGGATCAGTCGCGGCGGCCGAGGCTGAGCAGCAGCGCTC TCGCTCCCTGACCTGGGGAGAAGCGCCCACCCGGGAGAGCTGATCCCCGGCTGCCTCCAGCGCCCCCCACCTTTTGCACTCCAAGCCGGG GGCTCCAGAGACCCCGCTCCCCAGGCGCCACTATGCTGGACCCTTCGTCCAGCGAAGAAGAATCGGATGAGATCGTGGAGGAGGAGAGCG GCAAGGAGGTGCTCGGCTCGGCCCCGTCCGGCGCGCGCCTGTCTCCCAGCCGTACCAGCGAGGGCTCGGCCGGCAGCGCCGGGCTGGGGG GCGGCGGCGCCGGCGCCGGAGCCGGGGTGGGTGCAGGCGGCGGCGGGGGCAGCGGCGCGAGCAGCGGCGGCGGGGCCGGGGGGCTGCAAC CCAGCAGCCGCGCTGGCGGCGGCCGGCCCTCCAGCCCCAGCCCGTCGGTGGTGAGCGAGAAGGAGAAGGAAGAGTTGGAGCGGCTGCAGA AAGAGGAGGAGGAGAGGAAGAAGAGGCTGCAGCTGTATGTGTTCGTGATGCGCTGCATCGCCTACCCCTTTAATGCCAAGCAGCCCACCG ACATGGCTCGCCGGCAGCAGAAGCACGTTCACGTCCATGTTCTTCCCAGGAAGGCTGGAGACTTTCACAGGAATGACAGCATCTATGAGG AGCTCCAGAAACATGACAAGGAGGACTTTCCTGCCTCTTGGAGATCAGAGGAGGAAATGGCAGCAGAAGCCGCAGCTCTGCGGGTCTACT TTCAGTGACACAGATCCTGAATTCCAGCAAAAGAGCTATTGCCAACCAGTTTGAAGACCGCCCCCCGCCTCTCCCCAAGAGGAACTGAAT CAGCATGAAAATGCAGTTTCTTCATCTCACCATCCTGTATTCTTCAACCAGTGATCCCCCACCTCGGTCACTCCAACTCCCTTAAAATAC CTAGACCTAAACGGCTCAGACAGGCAGATTTGAGGTTTCCCCCTGTCTCCTTATTCGGCAGCCTTATGATTAAACTTCCTTCTCTGCTGC >12483_12483_1_CADPS-FHIT_CADPS_chr3_62860264_ENST00000283269_FHIT_chr3_59908140_ENST00000476844_length(amino acids)=244AA_BP=4 MFLELLIDAVIPVKVSSLPGKNMDVNVLLLPASHVGGLLGIKGVGDAAHHEHIQLQPLLPLLLLFLQPLQLFLLLLAHHRRAGAGGPAAA SAAAGLQPPGPAAAARAAAPAAACTHPGSGAGAAAPQPGAAGRALAGTAGRQARAGRGRAEHLLAALLLHDLIRFFFAGRRVQHSGAWGA -------------------------------------------------------------- >12483_12483_2_CADPS-FHIT_CADPS_chr3_62860264_ENST00000283269_FHIT_chr3_59908140_ENST00000492590_length(transcript)=1093nt_BP=653nt CTCCTCCCTTCGCGCTTCTGGGTCTGAGCCCAGCTCGCGACCGCCGGGCAGAGGATCAGTCGCGGCGGCCGAGGCTGAGCAGCAGCGCTC TCGCTCCCTGACCTGGGGAGAAGCGCCCACCCGGGAGAGCTGATCCCCGGCTGCCTCCAGCGCCCCCCACCTTTTGCACTCCAAGCCGGG GGCTCCAGAGACCCCGCTCCCCAGGCGCCACTATGCTGGACCCTTCGTCCAGCGAAGAAGAATCGGATGAGATCGTGGAGGAGGAGAGCG GCAAGGAGGTGCTCGGCTCGGCCCCGTCCGGCGCGCGCCTGTCTCCCAGCCGTACCAGCGAGGGCTCGGCCGGCAGCGCCGGGCTGGGGG GCGGCGGCGCCGGCGCCGGAGCCGGGGTGGGTGCAGGCGGCGGCGGGGGCAGCGGCGCGAGCAGCGGCGGCGGGGCCGGGGGGCTGCAAC CCAGCAGCCGCGCTGGCGGCGGCCGGCCCTCCAGCCCCAGCCCGTCGGTGGTGAGCGAGAAGGAGAAGGAAGAGTTGGAGCGGCTGCAGA AAGAGGAGGAGGAGAGGAAGAAGAGGCTGCAGCTGTATGTGTTCGTGATGCGCTGCATCGCCTACCCCTTTAATGCCAAGCAGCCCACCG ACATGGCTCGCCGGCAGCAGAAGCACGTTCACGTCCATGTTCTTCCCAGGAAGGCTGGAGACTTTCACAGGAATGACAGCATCTATGAGG AGCTCCAGAAACATGACAAGGAGGACTTTCCTGCCTCTTGGAGATCAGAGGAGGAAATGGCAGCAGAAGCCGCAGCTCTGCGGGTCTACT TTCAGTGACACAGATGTTTTTCAGATCCTGAATTCCAGCAAAAGAGCTATTGCCAACCAGTTTGAAGACCGCCCCCCGCCTCTCCCCAAG AGGAACTGAATCAGCATGAAAATGCAGTTTCTTCATCTCACCATCCTGTATTCTTCAACCAGTGATCCCCCACCTCGGTCACTCCAACTC CCTTAAAATACCTAGACCTAAACGGCTCAGACAGGCAGATTTGAGGTTTCCCCCTGTCTCCTTATTCGGCAGCCTTATGATTAAACTTCC >12483_12483_2_CADPS-FHIT_CADPS_chr3_62860264_ENST00000283269_FHIT_chr3_59908140_ENST00000492590_length(amino acids)=225AA_BP=171 MPPAPPTFCTPSRGLQRPRSPGATMLDPSSSEEESDEIVEEESGKEVLGSAPSGARLSPSRTSEGSAGSAGLGGGGAGAGAGVGAGGGGG SGASSGGGAGGLQPSSRAGGGRPSSPSPSVVSEKEKEELERLQKEEEERKKRLQLYVFVMRCIAYPFNAKQPTDMARRQQKHVHVHVLPR -------------------------------------------------------------- >12483_12483_3_CADPS-FHIT_CADPS_chr3_62860264_ENST00000357948_FHIT_chr3_59908140_ENST00000476844_length(transcript)=1179nt_BP=750nt TCAGCAGCCGCAGATGGCATCCGGCTGCGGGCTCGGGGCTCGCAATTGATTCTCCCCCTTGCCCACCTCGAGTCCACGGACGCACCTCTC CCTTCCCCTCCTCCCTTCGCGCTTCTGGGTCTGAGCCCAGCTCGCGACCGCCGGGCAGAGGATCAGTCGCGGCGGCCGAGGCTGAGCAGC AGCGCTCTCGCTCCCTGACCTGGGGAGAAGCGCCCACCCGGGAGAGCTGATCCCCGGCTGCCTCCAGCGCCCCCCACCTTTTGCACTCCA AGCCGGGGGCTCCAGAGACCCCGCTCCCCAGGCGCCACTATGCTGGACCCTTCGTCCAGCGAAGAAGAATCGGATGAGATCGTGGAGGAG GAGAGCGGCAAGGAGGTGCTCGGCTCGGCCCCGTCCGGCGCGCGCCTGTCTCCCAGCCGTACCAGCGAGGGCTCGGCCGGCAGCGCCGGG CTGGGGGGCGGCGGCGCCGGCGCCGGAGCCGGGGTGGGTGCAGGCGGCGGCGGGGGCAGCGGCGCGAGCAGCGGCGGCGGGGCCGGGGGG CTGCAACCCAGCAGCCGCGCTGGCGGCGGCCGGCCCTCCAGCCCCAGCCCGTCGGTGGTGAGCGAGAAGGAGAAGGAAGAGTTGGAGCGG CTGCAGAAAGAGGAGGAGGAGAGGAAGAAGAGGCTGCAGCTGTATGTGTTCGTGATGCGCTGCATCGCCTACCCCTTTAATGCCAAGCAG CCCACCGACATGGCTCGCCGGCAGCAGAAGCACGTTCACGTCCATGTTCTTCCCAGGAAGGCTGGAGACTTTCACAGGAATGACAGCATC TATGAGGAGCTCCAGAAACATGACAAGGAGGACTTTCCTGCCTCTTGGAGATCAGAGGAGGAAATGGCAGCAGAAGCCGCAGCTCTGCGG GTCTACTTTCAGTGACACAGATCCTGAATTCCAGCAAAAGAGCTATTGCCAACCAGTTTGAAGACCGCCCCCCGCCTCTCCCCAAGAGGA ACTGAATCAGCATGAAAATGCAGTTTCTTCATCTCACCATCCTGTATTCTTCAACCAGTGATCCCCCACCTCGGTCACTCCAACTCCCTT AAAATACCTAGACCTAAACGGCTCAGACAGGCAGATTTGAGGTTTCCCCCTGTCTCCTTATTCGGCAGCCTTATGATTAAACTTCCTTCT >12483_12483_3_CADPS-FHIT_CADPS_chr3_62860264_ENST00000357948_FHIT_chr3_59908140_ENST00000476844_length(amino acids)=276AA_BP=4 MFLELLIDAVIPVKVSSLPGKNMDVNVLLLPASHVGGLLGIKGVGDAAHHEHIQLQPLLPLLLLFLQPLQLFLLLLAHHRRAGAGGPAAA SAAAGLQPPGPAAAARAAAPAAACTHPGSGAGAAAPQPGAAGRALAGTAGRQARAGRGRAEHLLAALLLHDLIRFFFAGRRVQHSGAWGA GSLEPPAWSAKGGGRWRQPGISSPGWALLPRSGSESAAAQPRPPRLILCPAVASWAQTQKREGRRGRERCVRGLEVGKGENQLRAPSPQP -------------------------------------------------------------- >12483_12483_4_CADPS-FHIT_CADPS_chr3_62860264_ENST00000357948_FHIT_chr3_59908140_ENST00000492590_length(transcript)=1190nt_BP=750nt TCAGCAGCCGCAGATGGCATCCGGCTGCGGGCTCGGGGCTCGCAATTGATTCTCCCCCTTGCCCACCTCGAGTCCACGGACGCACCTCTC CCTTCCCCTCCTCCCTTCGCGCTTCTGGGTCTGAGCCCAGCTCGCGACCGCCGGGCAGAGGATCAGTCGCGGCGGCCGAGGCTGAGCAGC AGCGCTCTCGCTCCCTGACCTGGGGAGAAGCGCCCACCCGGGAGAGCTGATCCCCGGCTGCCTCCAGCGCCCCCCACCTTTTGCACTCCA AGCCGGGGGCTCCAGAGACCCCGCTCCCCAGGCGCCACTATGCTGGACCCTTCGTCCAGCGAAGAAGAATCGGATGAGATCGTGGAGGAG GAGAGCGGCAAGGAGGTGCTCGGCTCGGCCCCGTCCGGCGCGCGCCTGTCTCCCAGCCGTACCAGCGAGGGCTCGGCCGGCAGCGCCGGG CTGGGGGGCGGCGGCGCCGGCGCCGGAGCCGGGGTGGGTGCAGGCGGCGGCGGGGGCAGCGGCGCGAGCAGCGGCGGCGGGGCCGGGGGG CTGCAACCCAGCAGCCGCGCTGGCGGCGGCCGGCCCTCCAGCCCCAGCCCGTCGGTGGTGAGCGAGAAGGAGAAGGAAGAGTTGGAGCGG CTGCAGAAAGAGGAGGAGGAGAGGAAGAAGAGGCTGCAGCTGTATGTGTTCGTGATGCGCTGCATCGCCTACCCCTTTAATGCCAAGCAG CCCACCGACATGGCTCGCCGGCAGCAGAAGCACGTTCACGTCCATGTTCTTCCCAGGAAGGCTGGAGACTTTCACAGGAATGACAGCATC TATGAGGAGCTCCAGAAACATGACAAGGAGGACTTTCCTGCCTCTTGGAGATCAGAGGAGGAAATGGCAGCAGAAGCCGCAGCTCTGCGG GTCTACTTTCAGTGACACAGATGTTTTTCAGATCCTGAATTCCAGCAAAAGAGCTATTGCCAACCAGTTTGAAGACCGCCCCCCGCCTCT CCCCAAGAGGAACTGAATCAGCATGAAAATGCAGTTTCTTCATCTCACCATCCTGTATTCTTCAACCAGTGATCCCCCACCTCGGTCACT CCAACTCCCTTAAAATACCTAGACCTAAACGGCTCAGACAGGCAGATTTGAGGTTTCCCCCTGTCTCCTTATTCGGCAGCCTTATGATTA >12483_12483_4_CADPS-FHIT_CADPS_chr3_62860264_ENST00000357948_FHIT_chr3_59908140_ENST00000492590_length(amino acids)=225AA_BP=171 MPPAPPTFCTPSRGLQRPRSPGATMLDPSSSEEESDEIVEEESGKEVLGSAPSGARLSPSRTSEGSAGSAGLGGGGAGAGAGVGAGGGGG SGASSGGGAGGLQPSSRAGGGRPSSPSPSVVSEKEKEELERLQKEEEERKKRLQLYVFVMRCIAYPFNAKQPTDMARRQQKHVHVHVLPR -------------------------------------------------------------- >12483_12483_5_CADPS-FHIT_CADPS_chr3_62860264_ENST00000383710_FHIT_chr3_59908140_ENST00000476844_length(transcript)=1220nt_BP=791nt ACAGCTTGCAACACTCGGCATCTTTTCTGGAGGCGCCTCCTTCAGCAGCCGCAGATGGCATCCGGCTGCGGGCTCGGGGCTCGCAATTGA TTCTCCCCCTTGCCCACCTCGAGTCCACGGACGCACCTCTCCCTTCCCCTCCTCCCTTCGCGCTTCTGGGTCTGAGCCCAGCTCGCGACC GCCGGGCAGAGGATCAGTCGCGGCGGCCGAGGCTGAGCAGCAGCGCTCTCGCTCCCTGACCTGGGGAGAAGCGCCCACCCGGGAGAGCTG ATCCCCGGCTGCCTCCAGCGCCCCCCACCTTTTGCACTCCAAGCCGGGGGCTCCAGAGACCCCGCTCCCCAGGCGCCACTATGCTGGACC CTTCGTCCAGCGAAGAAGAATCGGATGAGATCGTGGAGGAGGAGAGCGGCAAGGAGGTGCTCGGCTCGGCCCCGTCCGGCGCGCGCCTGT CTCCCAGCCGTACCAGCGAGGGCTCGGCCGGCAGCGCCGGGCTGGGGGGCGGCGGCGCCGGCGCCGGAGCCGGGGTGGGTGCAGGCGGCG GCGGGGGCAGCGGCGCGAGCAGCGGCGGCGGGGCCGGGGGGCTGCAACCCAGCAGCCGCGCTGGCGGCGGCCGGCCCTCCAGCCCCAGCC CGTCGGTGGTGAGCGAGAAGGAGAAGGAAGAGTTGGAGCGGCTGCAGAAAGAGGAGGAGGAGAGGAAGAAGAGGCTGCAGCTGTATGTGT TCGTGATGCGCTGCATCGCCTACCCCTTTAATGCCAAGCAGCCCACCGACATGGCTCGCCGGCAGCAGAAGCACGTTCACGTCCATGTTC TTCCCAGGAAGGCTGGAGACTTTCACAGGAATGACAGCATCTATGAGGAGCTCCAGAAACATGACAAGGAGGACTTTCCTGCCTCTTGGA GATCAGAGGAGGAAATGGCAGCAGAAGCCGCAGCTCTGCGGGTCTACTTTCAGTGACACAGATCCTGAATTCCAGCAAAAGAGCTATTGC CAACCAGTTTGAAGACCGCCCCCCGCCTCTCCCCAAGAGGAACTGAATCAGCATGAAAATGCAGTTTCTTCATCTCACCATCCTGTATTC TTCAACCAGTGATCCCCCACCTCGGTCACTCCAACTCCCTTAAAATACCTAGACCTAAACGGCTCAGACAGGCAGATTTGAGGTTTCCCC >12483_12483_5_CADPS-FHIT_CADPS_chr3_62860264_ENST00000383710_FHIT_chr3_59908140_ENST00000476844_length(amino acids)=276AA_BP=4 MFLELLIDAVIPVKVSSLPGKNMDVNVLLLPASHVGGLLGIKGVGDAAHHEHIQLQPLLPLLLLFLQPLQLFLLLLAHHRRAGAGGPAAA SAAAGLQPPGPAAAARAAAPAAACTHPGSGAGAAAPQPGAAGRALAGTAGRQARAGRGRAEHLLAALLLHDLIRFFFAGRRVQHSGAWGA GSLEPPAWSAKGGGRWRQPGISSPGWALLPRSGSESAAAQPRPPRLILCPAVASWAQTQKREGRRGRERCVRGLEVGKGENQLRAPSPQP -------------------------------------------------------------- >12483_12483_6_CADPS-FHIT_CADPS_chr3_62860264_ENST00000383710_FHIT_chr3_59908140_ENST00000492590_length(transcript)=1231nt_BP=791nt ACAGCTTGCAACACTCGGCATCTTTTCTGGAGGCGCCTCCTTCAGCAGCCGCAGATGGCATCCGGCTGCGGGCTCGGGGCTCGCAATTGA TTCTCCCCCTTGCCCACCTCGAGTCCACGGACGCACCTCTCCCTTCCCCTCCTCCCTTCGCGCTTCTGGGTCTGAGCCCAGCTCGCGACC GCCGGGCAGAGGATCAGTCGCGGCGGCCGAGGCTGAGCAGCAGCGCTCTCGCTCCCTGACCTGGGGAGAAGCGCCCACCCGGGAGAGCTG ATCCCCGGCTGCCTCCAGCGCCCCCCACCTTTTGCACTCCAAGCCGGGGGCTCCAGAGACCCCGCTCCCCAGGCGCCACTATGCTGGACC CTTCGTCCAGCGAAGAAGAATCGGATGAGATCGTGGAGGAGGAGAGCGGCAAGGAGGTGCTCGGCTCGGCCCCGTCCGGCGCGCGCCTGT CTCCCAGCCGTACCAGCGAGGGCTCGGCCGGCAGCGCCGGGCTGGGGGGCGGCGGCGCCGGCGCCGGAGCCGGGGTGGGTGCAGGCGGCG GCGGGGGCAGCGGCGCGAGCAGCGGCGGCGGGGCCGGGGGGCTGCAACCCAGCAGCCGCGCTGGCGGCGGCCGGCCCTCCAGCCCCAGCC CGTCGGTGGTGAGCGAGAAGGAGAAGGAAGAGTTGGAGCGGCTGCAGAAAGAGGAGGAGGAGAGGAAGAAGAGGCTGCAGCTGTATGTGT TCGTGATGCGCTGCATCGCCTACCCCTTTAATGCCAAGCAGCCCACCGACATGGCTCGCCGGCAGCAGAAGCACGTTCACGTCCATGTTC TTCCCAGGAAGGCTGGAGACTTTCACAGGAATGACAGCATCTATGAGGAGCTCCAGAAACATGACAAGGAGGACTTTCCTGCCTCTTGGA GATCAGAGGAGGAAATGGCAGCAGAAGCCGCAGCTCTGCGGGTCTACTTTCAGTGACACAGATGTTTTTCAGATCCTGAATTCCAGCAAA AGAGCTATTGCCAACCAGTTTGAAGACCGCCCCCCGCCTCTCCCCAAGAGGAACTGAATCAGCATGAAAATGCAGTTTCTTCATCTCACC ATCCTGTATTCTTCAACCAGTGATCCCCCACCTCGGTCACTCCAACTCCCTTAAAATACCTAGACCTAAACGGCTCAGACAGGCAGATTT >12483_12483_6_CADPS-FHIT_CADPS_chr3_62860264_ENST00000383710_FHIT_chr3_59908140_ENST00000492590_length(amino acids)=225AA_BP=171 MPPAPPTFCTPSRGLQRPRSPGATMLDPSSSEEESDEIVEEESGKEVLGSAPSGARLSPSRTSEGSAGSAGLGGGGAGAGAGVGAGGGGG SGASSGGGAGGLQPSSRAGGGRPSSPSPSVVSEKEKEELERLQKEEEERKKRLQLYVFVMRCIAYPFNAKQPTDMARRQQKHVHVHVLPR -------------------------------------------------------------- >12483_12483_7_CADPS-FHIT_CADPS_chr3_62860264_ENST00000490353_FHIT_chr3_59908140_ENST00000476844_length(transcript)=870nt_BP=441nt ATGCTGGACCCTTCGTCCAGCGAAGAAGAATCGGATGAGATCGTGGAGGAGGAGAGCGGCAAGGAGGTGCTCGGCTCGGCCCCGTCCGGC GCGCGCCTGTCTCCCAGCCGTACCAGCGAGGGCTCGGCCGGCAGCGCCGGGCTGGGGGGCGGCGGCGCCGGCGCCGGAGCCGGGGTGGGT GCAGGCGGCGGCGGGGGCAGCGGCGCGAGCAGCGGCGGCGGGGCCGGGGGGCTGCAACCCAGCAGCCGCGCTGGCGGCGGCCGGCCCTCC AGCCCCAGCCCGTCGGTGGTGAGCGAGAAGGAGAAGGAAGAGTTGGAGCGGCTGCAGAAAGAGGAGGAGGAGAGGAAGAAGAGGCTGCAG CTGTATGTGTTCGTGATGCGCTGCATCGCCTACCCCTTTAATGCCAAGCAGCCCACCGACATGGCTCGCCGGCAGCAGAAGCACGTTCAC GTCCATGTTCTTCCCAGGAAGGCTGGAGACTTTCACAGGAATGACAGCATCTATGAGGAGCTCCAGAAACATGACAAGGAGGACTTTCCT GCCTCTTGGAGATCAGAGGAGGAAATGGCAGCAGAAGCCGCAGCTCTGCGGGTCTACTTTCAGTGACACAGATCCTGAATTCCAGCAAAA GAGCTATTGCCAACCAGTTTGAAGACCGCCCCCCGCCTCTCCCCAAGAGGAACTGAATCAGCATGAAAATGCAGTTTCTTCATCTCACCA TCCTGTATTCTTCAACCAGTGATCCCCCACCTCGGTCACTCCAACTCCCTTAAAATACCTAGACCTAAACGGCTCAGACAGGCAGATTTG >12483_12483_7_CADPS-FHIT_CADPS_chr3_62860264_ENST00000490353_FHIT_chr3_59908140_ENST00000476844_length(amino acids)=201AA_BP=147 MLDPSSSEEESDEIVEEESGKEVLGSAPSGARLSPSRTSEGSAGSAGLGGGGAGAGAGVGAGGGGGSGASSGGGAGGLQPSSRAGGGRPS SPSPSVVSEKEKEELERLQKEEEERKKRLQLYVFVMRCIAYPFNAKQPTDMARRQQKHVHVHVLPRKAGDFHRNDSIYEELQKHDKEDFP -------------------------------------------------------------- >12483_12483_8_CADPS-FHIT_CADPS_chr3_62860264_ENST00000490353_FHIT_chr3_59908140_ENST00000492590_length(transcript)=881nt_BP=441nt ATGCTGGACCCTTCGTCCAGCGAAGAAGAATCGGATGAGATCGTGGAGGAGGAGAGCGGCAAGGAGGTGCTCGGCTCGGCCCCGTCCGGC GCGCGCCTGTCTCCCAGCCGTACCAGCGAGGGCTCGGCCGGCAGCGCCGGGCTGGGGGGCGGCGGCGCCGGCGCCGGAGCCGGGGTGGGT GCAGGCGGCGGCGGGGGCAGCGGCGCGAGCAGCGGCGGCGGGGCCGGGGGGCTGCAACCCAGCAGCCGCGCTGGCGGCGGCCGGCCCTCC AGCCCCAGCCCGTCGGTGGTGAGCGAGAAGGAGAAGGAAGAGTTGGAGCGGCTGCAGAAAGAGGAGGAGGAGAGGAAGAAGAGGCTGCAG CTGTATGTGTTCGTGATGCGCTGCATCGCCTACCCCTTTAATGCCAAGCAGCCCACCGACATGGCTCGCCGGCAGCAGAAGCACGTTCAC GTCCATGTTCTTCCCAGGAAGGCTGGAGACTTTCACAGGAATGACAGCATCTATGAGGAGCTCCAGAAACATGACAAGGAGGACTTTCCT GCCTCTTGGAGATCAGAGGAGGAAATGGCAGCAGAAGCCGCAGCTCTGCGGGTCTACTTTCAGTGACACAGATGTTTTTCAGATCCTGAA TTCCAGCAAAAGAGCTATTGCCAACCAGTTTGAAGACCGCCCCCCGCCTCTCCCCAAGAGGAACTGAATCAGCATGAAAATGCAGTTTCT TCATCTCACCATCCTGTATTCTTCAACCAGTGATCCCCCACCTCGGTCACTCCAACTCCCTTAAAATACCTAGACCTAAACGGCTCAGAC >12483_12483_8_CADPS-FHIT_CADPS_chr3_62860264_ENST00000490353_FHIT_chr3_59908140_ENST00000492590_length(amino acids)=201AA_BP=147 MLDPSSSEEESDEIVEEESGKEVLGSAPSGARLSPSRTSEGSAGSAGLGGGGAGAGAGVGAGGGGGSGASSGGGAGGLQPSSRAGGGRPS SPSPSVVSEKEKEELERLQKEEEERKKRLQLYVFVMRCIAYPFNAKQPTDMARRQQKHVHVHVLPRKAGDFHRNDSIYEELQKHDKEDFP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CADPS-FHIT |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000283269 | - | 1 | 28 | 790_1129 | 147.0 | 1315.0 | DRD2 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000357948 | - | 1 | 27 | 790_1129 | 147.0 | 1275.0 | DRD2 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000383710 | - | 1 | 30 | 790_1129 | 147.0 | 1354.0 | DRD2 |
| Hgene | CADPS | chr3:62860264 | chr3:59908140 | ENST00000490353 | - | 1 | 7 | 790_1129 | 147.0 | 458.0 | DRD2 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CADPS-FHIT |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CADPS-FHIT |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
