
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CALR-QSOX1 (FusionGDB2 ID:12613) |
Fusion Gene Summary for CALR-QSOX1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CALR-QSOX1 | Fusion gene ID: 12613 | Hgene | Tgene | Gene symbol | CALR | QSOX1 | Gene ID | 811 | 5768 |
| Gene name | calreticulin | quiescin sulfhydryl oxidase 1 | |
| Synonyms | CRT|HEL-S-99n|RO|SSA|cC1qR | Q6|QSCN6 | |
| Cytomap | 19p13.13 | 1q25.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | calreticulinCRP55ERp60HACBPSicca syndrome antigen A (autoantigen Ro; calreticulin)calregulinendoplasmic reticulum resident protein 60epididymis secretory sperm binding protein Li 99ngrp60 | sulfhydryl oxidase 1quiescin Q6 sulfhydryl oxidase 1testis tissue sperm-binding protein Li 62nthiol oxidase 1 | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | P27797 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000316448, | ENST00000367600, ENST00000367602, | |
| Fusion gene scores | * DoF score | 18 X 17 X 9=2754 | 8 X 9 X 3=216 |
| # samples | 24 | 9 | |
| ** MAII score | log2(24/2754*10)=-3.52042224852645 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(9/216*10)=-1.26303440583379 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CALR [Title/Abstract] AND QSOX1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CALR(13051468)-QSOX1(180165396), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | CALR-QSOX1 seems lost the major protein functional domain in Hgene partner, which is a cell metabolism gene due to the frame-shifted ORF. CALR-QSOX1 seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | CALR | GO:0000122 | negative regulation of transcription by RNA polymerase II | 8107809 |
| Hgene | CALR | GO:0006611 | protein export from nucleus | 11149926 |
| Hgene | CALR | GO:0017148 | negative regulation of translation | 14726956 |
| Hgene | CALR | GO:0033144 | negative regulation of intracellular steroid hormone receptor signaling pathway | 8107809 |
| Hgene | CALR | GO:0034504 | protein localization to nucleus | 15998798 |
| Hgene | CALR | GO:0045665 | negative regulation of neuron differentiation | 8107809 |
| Hgene | CALR | GO:0045892 | negative regulation of transcription, DNA-templated | 8107809 |
| Hgene | CALR | GO:0048387 | negative regulation of retinoic acid receptor signaling pathway | 8107809 |
 Fusion gene breakpoints across CALR (5'-gene) Fusion gene breakpoints across CALR (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
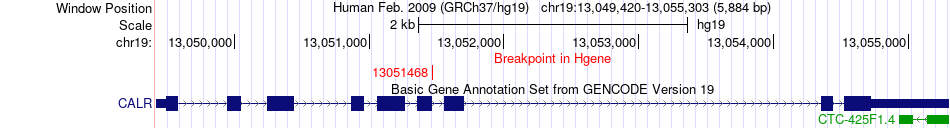 |
 Fusion gene breakpoints across QSOX1 (3'-gene) Fusion gene breakpoints across QSOX1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
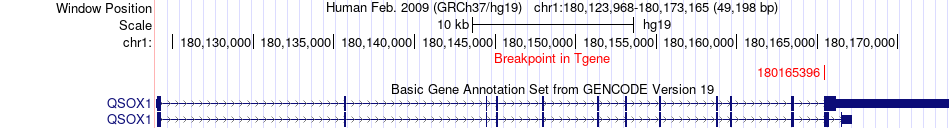 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | OV | TCGA-13-0919 | CALR | chr19 | 13051468 | + | QSOX1 | chr1 | 180165396 | + |
Top |
Fusion Gene ORF analysis for CALR-QSOX1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000316448 | ENST00000367600 | CALR | chr19 | 13051468 | + | QSOX1 | chr1 | 180165396 | + |
| In-frame | ENST00000316448 | ENST00000367602 | CALR | chr19 | 13051468 | + | QSOX1 | chr1 | 180165396 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000316448 | CALR | chr19 | 13051468 | + | ENST00000367602 | QSOX1 | chr1 | 180165396 | + | 8658 | 889 | 52 | 1176 | 374 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000316448 | ENST00000367602 | CALR | chr19 | 13051468 | + | QSOX1 | chr1 | 180165396 | + | 0.002033868 | 0.9979662 |
Top |
Fusion Genomic Features for CALR-QSOX1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CALR | chr19 | 13051468 | + | QSOX1 | chr1 | 180165396 | + | 3.54E-05 | 0.9999646 |
| CALR | chr19 | 13051468 | + | QSOX1 | chr1 | 180165396 | + | 3.54E-05 | 0.9999646 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
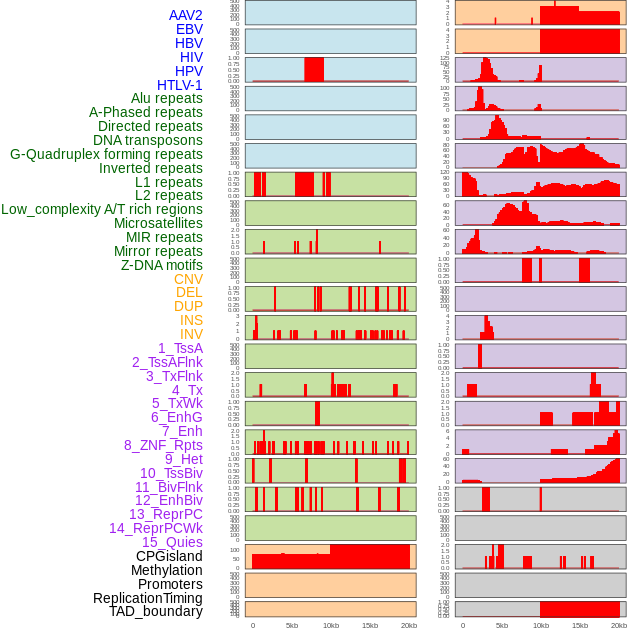 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
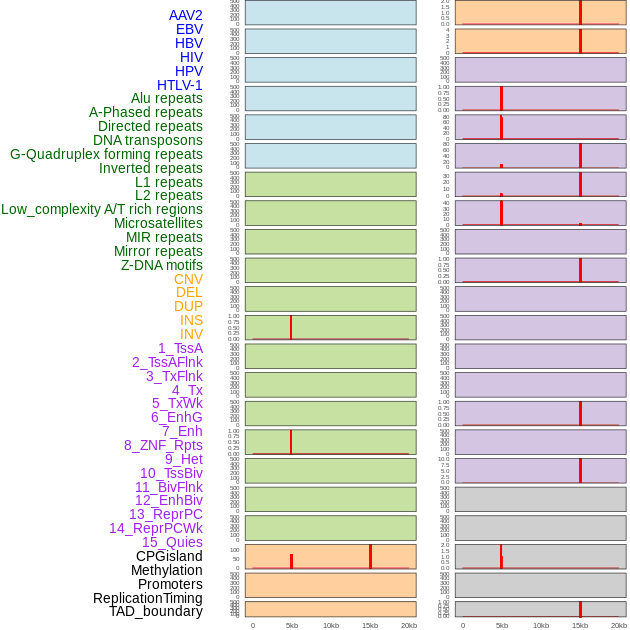 |
Top |
Fusion Protein Features for CALR-QSOX1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:13051468/chr1:180165396) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CALR | . |
| FUNCTION: Calcium-binding chaperone that promotes folding, oligomeric assembly and quality control in the endoplasmic reticulum (ER) via the calreticulin/calnexin cycle. This lectin interacts transiently with almost all of the monoglucosylated glycoproteins that are synthesized in the ER (PubMed:7876246). Interacts with the DNA-binding domain of NR3C1 and mediates its nuclear export (PubMed:11149926). Involved in maternal gene expression regulation. May participate in oocyte maturation via the regulation of calcium homeostasis (By similarity). Present in the cortical granules of non-activated oocytes, is exocytosed during the cortical reaction in response to oocyte activation and might participate in the block to polyspermy (By similarity). {ECO:0000250|UniProtKB:P28491, ECO:0000250|UniProtKB:Q8K3H7, ECO:0000269|PubMed:11149926, ECO:0000269|PubMed:7876246}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 18_197 | 272 | 418.0 | Region | Note=N-domain |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 191_255 | 272 | 418.0 | Region | Note=4 X approximate repeats |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 191_202 | 272 | 418.0 | Repeat | Note=1-1 |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 210_221 | 272 | 418.0 | Repeat | Note=1-2 |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 227_238 | 272 | 418.0 | Repeat | Note=1-3 |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 244_255 | 272 | 418.0 | Repeat | Note=1-4 |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 259_269 | 272 | 418.0 | Repeat | Note=2-1 |
| Tgene | QSOX1 | chr19:13051468 | chr1:180165396 | ENST00000367600 | 10 | 13 | 710_730 | 489 | 605.0 | Transmembrane | Helical | |
| Tgene | QSOX1 | chr19:13051468 | chr1:180165396 | ENST00000367602 | 10 | 12 | 710_730 | 489 | 748.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 351_408 | 272 | 418.0 | Compositional bias | Note=Asp/Glu/Lys-rich |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 414_417 | 272 | 418.0 | Motif | Note=Prevents secretion from ER |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 198_308 | 272 | 418.0 | Region | Note=P-domain |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 259_297 | 272 | 418.0 | Region | Note=3 X approximate repeats |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 309_417 | 272 | 418.0 | Region | Note=C-domain |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 273_283 | 272 | 418.0 | Repeat | Note=2-2 |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 287_297 | 272 | 418.0 | Repeat | Note=2-3 |
| Tgene | QSOX1 | chr19:13051468 | chr1:180165396 | ENST00000367600 | 10 | 13 | 36_156 | 489 | 605.0 | Domain | Thioredoxin | |
| Tgene | QSOX1 | chr19:13051468 | chr1:180165396 | ENST00000367600 | 10 | 13 | 396_503 | 489 | 605.0 | Domain | ERV/ALR sulfhydryl oxidase | |
| Tgene | QSOX1 | chr19:13051468 | chr1:180165396 | ENST00000367602 | 10 | 12 | 36_156 | 489 | 748.0 | Domain | Thioredoxin | |
| Tgene | QSOX1 | chr19:13051468 | chr1:180165396 | ENST00000367602 | 10 | 12 | 396_503 | 489 | 748.0 | Domain | ERV/ALR sulfhydryl oxidase | |
| Tgene | QSOX1 | chr19:13051468 | chr1:180165396 | ENST00000367600 | 10 | 13 | 478_485 | 489 | 605.0 | Nucleotide binding | FAD | |
| Tgene | QSOX1 | chr19:13051468 | chr1:180165396 | ENST00000367602 | 10 | 12 | 478_485 | 489 | 748.0 | Nucleotide binding | FAD |
Top |
Fusion Gene Sequence for CALR-QSOX1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >12613_12613_1_CALR-QSOX1_CALR_chr19_13051468_ENST00000316448_QSOX1_chr1_180165396_ENST00000367602_length(transcript)=8658nt_BP=889nt CCGTCCGTACTGCAGAGCCGCTGCCGGAGGGTCGTTTTAAAGGGCCCGCGCGTTGCCGCCCCCTCGGCCCGCCATGCTGCTATCCGTGCC GCTGCTGCTCGGCCTCCTCGGCCTGGCCGTCGCCGAGCCTGCCGTCTACTTCAAGGAGCAGTTTCTGGACGGAGACGGGTGGACTTCCCG CTGGATCGAATCCAAACACAAGTCAGATTTTGGCAAATTCGTTCTCAGTTCCGGCAAGTTCTACGGTGACGAGGAGAAAGATAAAGGTTT GCAGACAAGCCAGGATGCACGCTTTTATGCTCTGTCGGCCAGTTTCGAGCCTTTCAGCAACAAAGGCCAGACGCTGGTGGTGCAGTTCAC GGTGAAACATGAGCAGAACATCGACTGTGGGGGCGGCTATGTGAAGCTGTTTCCTAATAGTTTGGACCAGACAGACATGCACGGAGACTC AGAATACAACATCATGTTTGGTCCCGACATCTGTGGCCCTGGCACCAAGAAGGTTCATGTCATCTTCAACTACAAGGGCAAGAACGTGCT GATCAACAAGGACATCCGTTGCAAGGATGATGAGTTTACACACCTGTACACACTGATTGTGCGGCCAGACAACACCTATGAGGTGAAGAT TGACAACAGCCAGGTGGAGTCCGGCTCCTTGGAAGACGATTGGGACTTCCTGCCACCCAAGAAGATAAAGGATCCTGATGCTTCAAAACC GGAAGACTGGGATGAGCGGGCCAAGATCGATGATCCCACAGACTCCAAGCCTGAGGACTGGGACAAGCCCGAGCATATCCCTGACCCTGA TGCTAAGAAGCCCGAGGACTGGGATGAAGAGATGGACGGAGAGTGGGAACCCCCAGTGATTCAGAACCCTGAGTACAAGGTGCCCCCAGC GAGGACCCCCAGTTCCCCAAGGTGCAGTGGCCACCCCGTGAACTTTGTTCTGCCTGCCACAATGAACGCCTGGATGTGCCCGTGTGGGAC GTGGAAGCCACCCTCAACTTCCTCAAGGCCCACTTCTCCCCAAGCAACATCATCCTGGACTTCCCTGCAGCTGGGTCAGCTGCCCGGAGG GATGTGCAGAATGTGGCAGCCGCCCCAGAGCTGGCGATGGGAGCCCTGGAGCTGGAAAGCCGGAATTCAACTCTGGACCCTGGGAAGCCT GAGATGATGAAGTCCCCCACAAACACCACCCCACATGTGCCGGCTGAGGGACCTGAGGCAAGTCGACCCCCGAAGCTGCACCCTGGCCTC AGAGCTGCACCAGGCCAGGAGCCTCCTGAGCACATGGCAGAGCTTCAGAGGAATGAGCAGGAGCAGCCGCTTGGGCAGTGGCACTTGAGC AAGCGAGACACAGGGGCTGCATTGCTGGCTGAGTCCAGGGCTGAGAAGAACCGCCTCTGGGGCCCTTTGGAGGTCAGGCGCGTGGGCCGC AGCTCCAAGCAGCTGGTCGACATCCCTGAGGGCCAGCTGGAGGCCCGAGCTGGACGGGGCCGAGGCCAGTGGCTGCAGGTGCTGGGAGGG GGCTTCTCTTACCTGGACATCAGCCTCTGTGTGGGGCTCTATTCCCTGTCCTTCATGGGCCTGCTGGCCATGTACACCTACTTCCAGGCC AAGATAAGGGCCCTGAAGGGCCATGCTGGCCACCCTGCAGCCTGAACCACCTGGGGAGGAGGCGGGAGAGGGAGCTGCCATCTCTAGGCA CCTCAAGCCCCCTGACCCCATTCCCTCCCCTCCCACCCCTTGCTCCTTGTCTGGCCTAGAAGTGTGGGAAATTCAGGAAAACGAGTTGCT CCAGTGAAGCTTCTTGGGGTTGCTAGGACAGAGAGCTCCTTTGACACAAAAGACAGGAGCAGGGTCCAGGTTCCCCTGCTGTGCAGGGAG GGCAGCCCCGGGCAGTGGGCATAGGGCAGCTCAGTCCCTGGCCTCTTAGCACCACATTCCTGTTTTTCAGCTTATTTGAAGTCCTGCCTC ATTCTCACTGGAGCCTCAGTCTCTCCTGCTTGGTCTTGGCCCTCAACTGGGGCAAGTGAAGCCAGAGGAGGGTCCCCCAGCTGGGTGGGC TGGAATGGAACTCCTCACTAGCTGCTGGGGCTCCGCCCACCCTGCTCCCTTCCGGACAATGAAGAAGCCTTTGCACCCTGGGAGGAAGGA CCACCCCGGGCCCTCTATGCCTGGCCAGCCTCCAGCTCCTCAGACCTCCTGGGTGGGGTTTGGCTTCAGGGTGGGGTTTGGAAGCTTCTG GAAGTCGTGCTGGTCTCCCAGGTGAGGCAAGCCATGGTTGCTGGGCTGTAGGGTGAGTGGCTTGCTTGGTGGGACCTGACGAGTTGGTGG CATGGGAAGGATGTGGGTCTCTAGTGCCTTGCCCTGGCTTAGCTGCAGGAGAAGATGGCTGCTTTCACTTCCCCCCATTGAGCTCTGCTC CCTCTGAGCCTGGTCTTTTGTCCTTTTTTATTTTGGTCTCCAAGATGAATGCTCATCTTTGGAGGGTGCCAGGTAGAAGCTAGGGAGGGG AGTGTCTTCTCTCTCCAGGTTTCACCTTCCAGTGTGCAGAAGTTAGAAGGGTCTGGCGGGGGCAGTGCCTTACACATGCTTGATTCCCAC GCTACCCCCTGCCTTGGGAGGTGTGTGGAATAAATTATTTTTGTTAAGGCAAGCATTGGTGTGTTCTTTAACTTGCTACTTGGAGACCTA GTGTCCAGTCTGGACAGAATGCACAGAGACCAGCCTCACCTGGAATGCAGCCAGACTCGATGTCCCTCAGCACACACAGCTCCTGGCCAC AGACTGCCCATAGAACTGTCTGCACCCAAACCCCATGGCCTTTTCATGCACGGAGACAGGCCTCTGGATGTGCAGCCTTGCCACCCCCTG CCCCAATCCTCCCTGAGAGCTCCTGCCTCAGTGCCCTGGGCTGGTGAGGGAGAAGCCTGTCTGCACCTGCCTAATTCCAGCTCCTCCAGG AAGCAAGGGCTTGTTTGTCAGAGCTGCAGGGTTGGCCACTCGGTGGGGAGGAGTCAGCCAGGATTAGAGAGCCTGCCCCTAATCCGGCCT GCTGGGTTTTACAAGGATCAGAGCTGCTGATAATGAACCTCATTAAGGGGGAGCAGGAGCCTCAATCCGATTTGGTTTTCTCTTTGACAT CTTCACTCTGCTCAGATGGCCTGGGTGCTATGTGGAGCAGGTGGGATGCCAAGGCCACTCCTGCTATGGGGCAGCTGGGGCTGGGGAGGG ATGGCAGTCTCCCTGCATGTTTCCCTCGACCTCTTTAGCTGCAGCGCCTTGCTGGGCTCCTGGGTTGGACTCCCTCTCTGTGCCCCTGCT CCAGGCACCCATTGGCTCCATCCTCCTGGTTGTGCTCTGCACCCCCTGCTCCCTTGGGCTGGCCCTGGCTGGGGGCCTGAGAGACAGACA GGAACCCACAATCAGGAGGCAACCCTGGCCTGCAAGAGGAAGACAGAGGCTCCCAGGGCCGGTGCCCTGTGTGCCCACTGCACCAAGGCC GCTGAATAAGCCTGCCCTTCACCCCCTAAGGGCTCCTTGCCCAATGCCAAGTGCTGGGGATTTCTGTCAGCAAGCCCTGTGGCTCCAGTG ACGGTATTTCTAAAGCCAAACTTAGTTACCTAGAATTAGCGCCATGTTGGAAACACTGTCGCAGCAGCCCGGGCTGCACAGTGTGTAGCC CAGCCTCCAGGTCCACGGAGTGGTGTGGACCTCCCACCTCACAGCTGCCTCTGGCAGCCAAGCCTCTTTTCGCCCGGCCCCAGCCCCTCT GGTTGATAAACGGGTGGGCCTCCTCAGCAGCGTGGCTGCCTTTCACCTTGATTTCCCCAGGGCTCTCGGCAACATCGATAAACCAGCCTC GCCCACCAGCTGGGCCCTCCCCCACCCAGTCTGCCAGGCTGGGAGCTGGAGCTTGCTGAGTCTTGAATGCCCTTCTAGATGGCTTCTCTA GAGGCTCTCCTGGCAAGAGAGGGTCCCAAGGGGAGCCCTGCAAAGCAAAGGCTCCTTGTCTGGGGCGGGATAGAGAATCTCGCCTCTGTC TGGTGTGTTACCTACTGGGGGCACAGGAACAATTTCCTCAAGGAGACAGTGGCATGGAGCTTTGAAAGACGAGTAGGTGTTAGCAAGGAA ATAAGGAGGAACGGGGGTTACGGGCAGAGGAGAAAGCACATGCCAAGTCAGCAAAGAAAAGTAGAATTCGAAAACTTTTTAAAAATATTA CTAAGGATTTTCACAATGCTGCACTGGGCTAGAAACTGAAGCTAAAACAGATACGTGGTCCCTGCTGCTATGGGGCTTCCGTTCTAGAGG CAAGGACAGGTTGTGATGAGGGTTCTGAAGGATAGAGACCAAGCAGGGAGGGTGTTGAGGAGGCTTCTGCGAGACCTGAAGGATGGGAAG CCAGGAAGTGGGAGGGGTGGGGGTCCAGGCTGGAGGGGCCCAATGTAGGTGTAGAGGGACTACAGCCCTGAGGGGCTGCTCCATGCGGCA TTCTTGGAGGTCCAAGAGGGGCAGCGCCACCTTGGGCCAGGCTCCTCTCCAGCAGCCTTGGTATGGGGTGGGGGTGGGAAGACCCCTGAT GCAGCCTGTCCTCTGGGTGTGGGGGTGGAGGAGAGGGCTCCAGGGGACCCAGCCAGCCTTGACCCTGGAGGAAAGTCAGGTGGGCTCAGA GAGAGCCCTCAAATCTGGGCCCTGGGTCAGGGTGGGGTCAAGTCCAGCCTTGAAGAGAATGGACTCTGGAATCTGACGTCTGGGTTTGAA TCCAGCTTCCAACACTCACTGGCTGTGTGACTTTGGGCAAGATAACTTATCCCCTTGTCCCCATGTGTCATTGGAGATAAACTACCCACC TACCAGATTGTTGTACCATGCTGGGCAAACACTAAGTGCTTAATAATGGTAGCCCACTGCTCATGTCGTGTCGTCTCCTGTCTTGATACC TGAAAACTCAAGGGGCCCTGTTGTGCTTGGAGGGGGGTGCCGTTCCCACAGGATATTCTGTGAAGGGTGTCCCTGGAGCTGGGCCTCCAG GTGGCAAGCCCCAAGCCACCCCGGCAGGTGCCACTGGCCAGAGTTGAACTGAGGTGTTGAGTGGAAAGTGCCTCCCCACCCCCATCTGCC ACAGTAGTATGGATTTCCTTCATGAGATGGGGCTCCTAAAATGCCTGAAATGTGTATGCTGTTGGAGAATTGATGTGTATGATTTATTGA ACTTTGCACTCATTTAAATCTCATTTAATCCTCCAACAACGCAAGATGAAGAAGCTGAGGCTTAAAGAGATTAAGAAACATCCTATCCCC TTTACTCCCACCAAAATAAATACATTTGCCCTATTCTCCCAAATGGTGAGAGGTAGATCCAGGACTTGACCCCAGATCCCACCCACTCAA CCCTGGAGCTGCAGTCTCTACCCCTTCTAACACATGTGATTCTGTTTCATGTCCTGTTCCATCGGTAGAGTTACACACTTTCCTGAGACT TTATATGGTGGTGCTGCTGGGTGCCAGCCCTTCTTCCCTCCAACAGTGGCAGTAACCACTGTTGTGTGTTGGTGTCAAGGAGTGGGGACA CCCTGAAGGAATCCGCCTTTTCTAGAGCTCTCGCTCCATTCCCTACATCGAGGCTGAAGTGCCCTGTCTCGGGTCCTTGTTTTTAGGCCT CGTTTGAAGGAGGGGAAGAAACCAGGTTCGTTGGGCGTGACCTTGGCTCAGCTCGTCTGTGAAATTGGGGCCGCAGACTTAGTTTGCGGG TCCCTGCTAAATTCCTCTCCCTCTTCCTGGGATTCCCATCCATGTATTGCCAGCTACATGTAGAGTGGAGAGGAGAGGTTTTTCCTGCAG GGCCTTCCAGAGAAAGGATTCTTGGCCAGAGGAAATCACAGCCTATAAGATCAATATCCTGATAGAATCCCGGGCCAGGTGTGAGAGGGG AGAGCTCACTCACACCAGCCTGACAGTCTTGGGGCAGAGCGCCCCATAGGCCTCTGCAGCCCCATCACTGCCTCATTCTCCCTCAGGTGC CAAGGAAGCTCCAGGGTGGGCCCAGTATCCTTCCACAGCATCCCTCCTCTTCCCCTTCAACACCCCTTTCACCCCTCCCTCCACCACCTC TGAAAGGTGCTCCTTGGGCAGGGCGATAGGATTCCCACCCCTAGCTTTAAGAACTAAACCAAACTTCAAGAGGAGCAATTTAATCTTCAT ACCTGACTCCTTGTCCCTCCTCAGGCATGCTGTTGTGCAGGACAGAGCCCTGGCTTCTCCTGGATGTCACACCTGGCCTGTCTCTGACCC TTGCTTTCCCTTTCTCTGCTGCCCTCAAGGGTCATTCACACCCAAGAGGGTAAGAAGCTGCCCTCTCATCCCTTCCTCTCCCACCCTGGG CCTCTGCCCTCTGCCTGGCCCCAGCAGCACAGTCTTCTCCATAGAGAGGGGCTGGGCCATGGGACCCCAGTTCCTGGGGTCTCGGTGACA CAGTTCACTGTTTGCTGCAGAGGAGGCCCCACTCGCAGCCTTGGCCTCTGCCCTGCAGAAGAATTCCCTGTAACTCACTCACCAATGGCC GTGGGAGCCAGTGGCCTCGGCGAGGGTCAGTTGGACTGCATAGCTGCCGGCTGAGTCTCTCTGGCCACCCTCACTGGCCCCATCACTCCA GGTTGCACTGCTGCCCAGGGGGTTTGTGCCAGGCCTGGGTCACAGTGGAAGAGAGTCTGGAAATCACAAGGCAGAGGAGGAGACCTTTGC CAATGTGAGAGAACAAAGTTAAGGGGCAGTCCATGGGGGCATTGACCTTACTTAGGTGAGGAGGACTCAGGGAACATCTACAGACTCTAG GGCTAGGGTGAGATCCTCCTGATAGTGCTGGTAGCTCCAGCTGCTGCTTTGGCAAGACTTGGTCGTCTCTATACTGAGAGAGCAAAAGCT CTTTTCCTGGGGCTGCATGTTCTTCCATAGGAAAGTGGGCTTCAGGTGCTGCCTATTAAACCTCCTGCCTCCCCTGCTGCAGTGCTTTGA TGCCTTTCACCTGTGTCTACCTGCGAACCTCTGCCCACCACGGAAGAGCAGCTCACATTTGGAAGGGGCTAAAATTTTTACCTAGTTTGA ATCTGCTCTGCAATGTCCCCATCAAGTTCTTAACCATCCCGGGTTCCAATGGTTACAGAGTTCTGCCCTGGGGAGGGAGGAATTGCTTAG CAAATGATGTTGAGAATAGTGATTATTTGGGGGAAAATTCAGGTAAGATCTCACCTTCATACCATATACCAAAATACATCTCAGTAGATT AGTAAGCTAATGCAAAAAAAAAAAAAAAAGGGGAAATATAATTGAATACTTCACTGTTTTTGAAAGCATGAAAACAATGGAAAATGCAAC TGATGTGAGTAATCCATCTACTTGACGTCACTTTTAAGAAATAAGGCCAGGCACGGTGGCTCACGCTTGTAATCCTAGCACTTTGGGAGG CCGAGGTGGGTGGATCACGAGGTCAGGAGTTTGAGACCAGCTTGGCCAACATGGTGAAACCCCCATCTCTACTAAAATATTACACAGGCA TTGTGGCACACATCTGTAATCCCAGCTACTCAGGAGGCTGAGGCAGGAGAATCACTTGAACCTGGGAGGTGGAGGTTGCAGTGAGCTGAG ATCGCGCCATTGCACTCCAGCCTGGGGGACAGAGCGAGACTCTGTCTCAAAAAAATAAAAAAATAAAAATAAAAAAATAAAGTAAAGCTA CATATTTAAAAGCTGGGGAAATATATGCTGTAAATATGACAAAGGGCTGTGACTCTTCTGCAACACACCATTTTAAGAATGAGCTAGGTC TGCATTTATTGACCTGATAAAATGTCCCAAGAAGGACACATTTTTATAAACATAAGTTACTATTGTTATATATATGCTAACGTTTGTCTA GAAAAAGTCTAGAAAAATACACACTGAACTGTGAATAGCAGTTATATTTAGAAAGCAACATTAGGATGAACTTAAGTCAGGGCTTGGACT AGAGCAAGGCAAGTGAGGTGCCAGGATGCAAAAGTTAAGGAGACACCTGACCTCAGGGGCCCACAACTGCAGAGCCCCTGACAGTGAGTG TCTCAGTAGCCTCGCCCTAGTCCTGGCTTTCACTTAAATATGTTTTCGAATGTTTGTAATAAAATATTTCTTTTGGAAGGCAAAAATAGC AAGGAAAATAAATAGGATTAAAGGATACTAAAGCCCATTCAATAAAATTTGTAGGAGGCCACTGATTTGGCCTAGGCTCCTATGATGTAA CCAACAGACCAAACCAATATGGAGTCATTCATGCTAAATGAAACTAGTTAGGAGTATATTCATATAACAAGTGGCTAAGTGTCAGTTACA GCCTCTGAGCTTTAGTCAATTGTAGGTGGTCAATAGATTCAGATAAGGCAATCCAGTTAAACTGTATATCCCACTTGCTGTTTTCTGTCC CTAAATTCTCTCTGAGTACACTGCAGCCCTGAAGTTCTTTGAACCTGTTCTGGGTTCTGAGGGCTGTCCAATTACTGAATCATACATAAA AGCCAAGTAAGATTTTAAAATTTGTTATAATTTTGGTTTTTAACAACTTCTGTTGTAAATAATAAAAGCACTAAATCCCTACTGAAAATA >12613_12613_1_CALR-QSOX1_CALR_chr19_13051468_ENST00000316448_QSOX1_chr1_180165396_ENST00000367602_length(amino acids)=374AA_BP=279 MPPPRPAMLLSVPLLLGLLGLAVAEPAVYFKEQFLDGDGWTSRWIESKHKSDFGKFVLSSGKFYGDEEKDKGLQTSQDARFYALSASFEP FSNKGQTLVVQFTVKHEQNIDCGGGYVKLFPNSLDQTDMHGDSEYNIMFGPDICGPGTKKVHVIFNYKGKNVLINKDIRCKDDEFTHLYT LIVRPDNTYEVKIDNSQVESGSLEDDWDFLPPKKIKDPDASKPEDWDERAKIDDPTDSKPEDWDKPEHIPDPDAKKPEDWDEEMDGEWEP PVIQNPEYKVPPARTPSSPRCSGHPVNFVLPATMNAWMCPCGTWKPPSTSSRPTSPQATSSWTSLQLGQLPGGMCRMWQPPQSWRWEPWS -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CALR-QSOX1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Hgene | CALR | chr19:13051468 | chr1:180165396 | ENST00000316448 | + | 6 | 9 | 237_270 | 272.0 | 418.0 | PPIB |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CALR-QSOX1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CALR-QSOX1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
