
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CCDC28A-CLEC16A (FusionGDB2 ID:13766) |
Fusion Gene Summary for CCDC28A-CLEC16A |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CCDC28A-CLEC16A | Fusion gene ID: 13766 | Hgene | Tgene | Gene symbol | CCDC28A | CLEC16A | Gene ID | 25901 | 23274 |
| Gene name | coiled-coil domain containing 28A | C-type lectin domain containing 16A | |
| Synonyms | C6orf80|CCRL1AP | Gop-1|KIAA0350 | |
| Cytomap | 6q24.1 | 16p13.13 | |
| Type of gene | protein-coding | protein-coding | |
| Description | coiled-coil domain-containing protein 28Achemokine C-C motif receptor-like 1 adjacent | protein CLEC16AC-type lectin domain family 16 member A | |
| Modification date | 20200321 | 20200313 | |
| UniProtAcc | Q8IWP9 | Q2KHT3 | |
| Ensembl transtripts involved in fusion gene | ENST00000332797, | ENST00000381822, ENST00000409552, ENST00000465491, ENST00000409790, | |
| Fusion gene scores | * DoF score | 4 X 5 X 4=80 | 14 X 13 X 10=1820 |
| # samples | 6 | 17 | |
| ** MAII score | log2(6/80*10)=-0.415037499278844 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(17/1820*10)=-3.42033179894836 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CCDC28A [Title/Abstract] AND CLEC16A [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CCDC28A(139101122)-CLEC16A(11260245), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across CCDC28A (5'-gene) Fusion gene breakpoints across CCDC28A (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
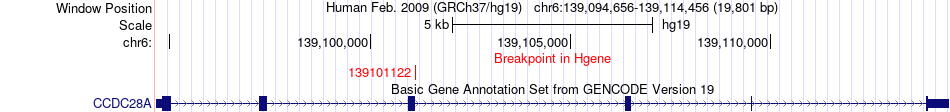 |
 Fusion gene breakpoints across CLEC16A (3'-gene) Fusion gene breakpoints across CLEC16A (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
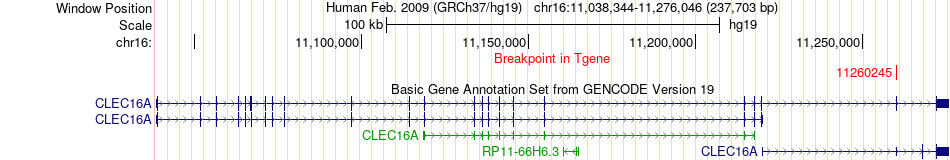 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LUAD | TCGA-50-5049-01A | CCDC28A | chr6 | 139101122 | + | CLEC16A | chr16 | 11260245 | + |
Top |
Fusion Gene ORF analysis for CCDC28A-CLEC16A |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000332797 | ENST00000381822 | CCDC28A | chr6 | 139101122 | + | CLEC16A | chr16 | 11260245 | + |
| 5CDS-intron | ENST00000332797 | ENST00000409552 | CCDC28A | chr6 | 139101122 | + | CLEC16A | chr16 | 11260245 | + |
| 5CDS-intron | ENST00000332797 | ENST00000465491 | CCDC28A | chr6 | 139101122 | + | CLEC16A | chr16 | 11260245 | + |
| In-frame | ENST00000332797 | ENST00000409790 | CCDC28A | chr6 | 139101122 | + | CLEC16A | chr16 | 11260245 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000332797 | CCDC28A | chr6 | 139101122 | + | ENST00000409790 | CLEC16A | chr16 | 11260245 | + | 4767 | 747 | 155 | 1267 | 370 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000332797 | ENST00000409790 | CCDC28A | chr6 | 139101122 | + | CLEC16A | chr16 | 11260245 | + | 0.037112612 | 0.96288735 |
Top |
Fusion Genomic Features for CCDC28A-CLEC16A |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CCDC28A | chr6 | 139101122 | + | CLEC16A | chr16 | 11260244 | + | 4.54E-07 | 0.9999995 |
| CCDC28A | chr6 | 139101122 | + | CLEC16A | chr16 | 11260244 | + | 4.54E-07 | 0.9999995 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
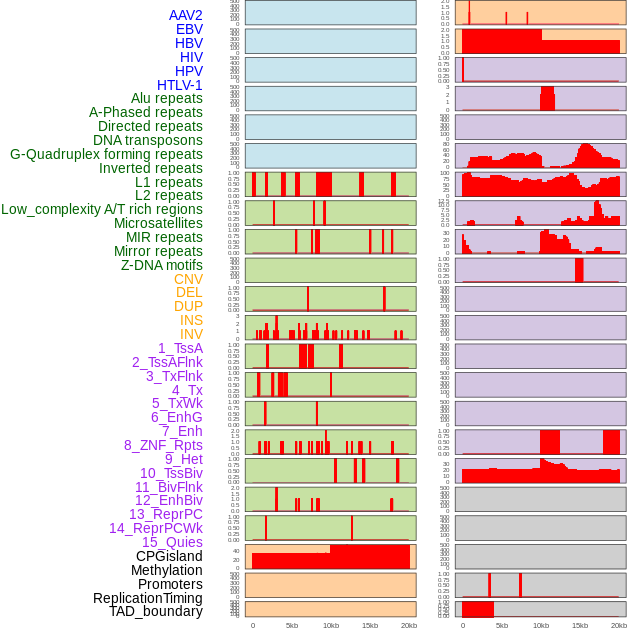 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
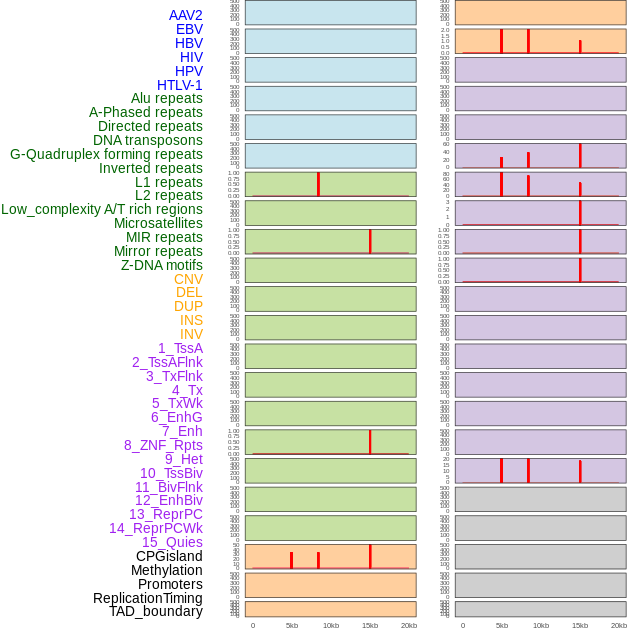 |
Top |
Fusion Protein Features for CCDC28A-CLEC16A |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr6:139101122/chr16:11260245) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CCDC28A | CLEC16A |
| FUNCTION: Regulator of mitophagy through the upstream regulation of the RNF41/NRDP1-PRKN pathway. Mitophagy is a selective form of autophagy necessary for mitochondrial quality control. The RNF41/NRDP1-PRKN pathway regulates autophagosome-lysosome fusion during late mitophagy. May protect RNF41/NRDP1 from proteosomal degradation, RNF41/NRDP1 which regulates proteosomal degradation of PRKN. Plays a key role in beta cells functions by regulating mitophagy/autophagy and mitochondrial health. {ECO:0000269|PubMed:24949970}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | CLEC16A | chr6:139101122 | chr16:11260245 | ENST00000381822 | 0 | 4 | 892_933 | 0 | 141.0 | Compositional bias | Note=Ser-rich | |
| Tgene | CLEC16A | chr6:139101122 | chr16:11260245 | ENST00000409552 | 0 | 21 | 892_933 | 0 | 907.0 | Compositional bias | Note=Ser-rich | |
| Tgene | CLEC16A | chr6:139101122 | chr16:11260245 | ENST00000409790 | 21 | 24 | 892_933 | 880 | 1054.0 | Compositional bias | Note=Ser-rich | |
| Tgene | CLEC16A | chr6:139101122 | chr16:11260245 | ENST00000381822 | 0 | 4 | 51_198 | 0 | 141.0 | Domain | FPL | |
| Tgene | CLEC16A | chr6:139101122 | chr16:11260245 | ENST00000409552 | 0 | 21 | 51_198 | 0 | 907.0 | Domain | FPL |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CCDC28A | chr6:139101122 | chr16:11260245 | ENST00000332797 | + | 3 | 6 | 234_263 | 197 | 275.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Tgene | CLEC16A | chr6:139101122 | chr16:11260245 | ENST00000409790 | 21 | 24 | 51_198 | 880 | 1054.0 | Domain | FPL |
Top |
Fusion Gene Sequence for CCDC28A-CLEC16A |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >13766_13766_1_CCDC28A-CLEC16A_CCDC28A_chr6_139101122_ENST00000332797_CLEC16A_chr16_11260245_ENST00000409790_length(transcript)=4767nt_BP=747nt GGCGGAGTAGCTTCCGGAAAGGGTACTGCATTTCCCGTTTCTACCTCCACTGCACCCGCTTATTGCGTCTTGCTCCTGGGTCACAGAGCC TAAAACGACACACCCAACACGCCCGCCGGAGTTACAGCTAAAGGAAGGACAGGGGAAGCAATGAAATGCCGAGGGCGGAGCCAAGAGCGA CACTGGGGGAGCAGGAAAAGGCGGGGCTTCCGCTTGGGGCATGGAGGCTGTACCTCTTACGTCACTTCCGTAAACAAACGGAGCTGCGGA GGAGCGGGTCCCGGGATGTGACCGGGGCTCTGCTTGTGGCTGCGGCGGTGGCTTCTGAGGCTGTCGGGTCTTTGCGGGTTGCGGAAGGGG GCCCCAATACCCTTCTTCTTCAGGTCTTAAGAAGCTGGCCGTGGTGCAATAAGGAACTTAAAACAATGGAAGAGCGGAAAGTGAAGAGGA GGAGTCCTAAGTCTTTTAGTGCCCACTGTACTCAGGTTGTCAATGCCAAAAAAAATGCCATTCCAGTGAGTAAAAGCACAGGGTTTTCAA ATCCTGCATCACAGTCAACTTCACAGCGACCAAAGTTAAAAAGAGTGATGAAAGAAAAGACCAAACCTCAGGGTGGAGAGGGCAAAGGCG CTCAGTCAACTCCGATCCAGCACTCCTTCCTCACTGATGTCTCAGATGTTCAGGAGATGGAGAGAGGGCTGCTCAGTCTTTTGAATGATT TCCACTCTGGAAAACTTCAAGCATTTGGCTTCGCCGTGGCCCAGTGCATAAACCAGCACAGCTCCCCGTCCCTGTCCTCACAGTCGCCAC CCTCCGCCAGCGGGAGCCCCAGCGGCAGCGGGAGCACCAGCCACTGCGACTCTGGAGGCACCAGCTCGTCCTCCACCCCCTCCACAGCCC AGAGTCCAGCAGATGCCCCCATGAGTCCAGAACTGCCTAAGCCTCACCTTCCTGACCAGTTGGTAATCGTCAACGAAACGGAAGCAGACT CTAAGCCCAGCAAGAACGTGGCCAGGAGCGCAGCCGTGGAGACAGCCAGCCTGTCCCCCAGCCTCGTCCCTGCCCGGCAGCCCACCATTT CCCTGCTCTGCGAGGACACGGCTGACACGCTGAGCGTCGAATCGCTGACCCTTGTCCCCCCAGTTGACCCCCACAGCCTCCGCAGCCTCA CCGGCATGCCCCCGCTGTCCACGCCGGCTGCCGCCTGCACAGAGCCCGTGGGCGAAGAGGCTGCATGTGCTGAGCCTGTGGGCACCGCTG AGGACTGAGTCAGTGCCGGGGCCTCCCTTTGTGTGTGTGGCCCCGCTGGTAGGGACCCCAGTGCCGCTGACTGGCAAGACACACTGGGAG CACCCACCATTCTGTGCGGCCCCCAGCAGCCATCTCAACCACCTATCCCTGCGCTCCCTTGAATGGGAAGAAGCCCCACGTTGTCCTTGA ATTCCTTTTTCACTTTGCATCTCTTCACGTGCAGGCTGGGACCAGCGGAGACACCGCGGCGAATGCAGATGACTGCACCGGCCACTCAGG GAGCTGCCTGGGCTCCGTGTCTCTGAGCCCCGGGTGGCAGGACCCACCGGCACCTCTTTCTTCCTCTGTCATATGGCTCCTCTGTCACCA GCCCCAGTGTGCACAGAAGAATTGGACCAGGTCACTGTACGTAGAAATTTGTAGAAAAGCAGACTTAGATAAACATCTCCTTTGGATATT TATTTCCGCTTTTGGCAGCAGGTGAACATTTATTTTTAAAACTTCTATTTAAAAGAAGTCCAAAAACATCAACACTAAGGTTTGATGTCA TGTGAAAAGTGTAATAATAACAGTTAAGATTTCATGATCATTTTCACTGGACCTTTCCTGATATTTTGTTTCAGAGTTCTTAGTGTGGCT TTTTCCATTTATTTAAGTGATTCTTTGTTACTCACTAACTCTGCAAGCCTGTGGAATAATGAAGTACCTTCCTGGAAAGTTTGGATTATT TTTTAAACAAAAACAAGGGAGATACATGTATTCTCAGGTACACACAGAGCTGAGAGGGCTGAATGGTTTTCTGCTATAGCAGCCGAGAGG CCTCCCATCATGGAAAGATTTCTCCAGGAAAAGGAGGAATGTAGCCAGCTCCCCACTCAGGACGCTTCCTCATTTCTCTTCACCAAAACC AAACAGAGACAGCTTCCAGCACCTTCTTCAGTGTTACCATCTCTAAGAAGGAACCAGTTGGGACCGTGAAGACTCCCGACCCTGTGGCCA TGATGGAAATCAAAGGAAGACACCCTCTACGTCACCTGCCCTCGACTGTGTGTGCCCACATGTGCCGAGAGATGGCCCAGAGCCAGTTCC CCTCCAGCTGCAAGGGCATGGTGTCCCCAGAGCTCTGAGTCTGTCACTCTCCCTCTGCTACTGCTGCTGATCTGAATATGGAAACCCCAT GGTTCCCTTCCCCATTCGGACTGGGTGTGTACAAGCAAGGACCCAGATGCATCAGACACAGCCCCCAAGATGTTCCTTTCTACTCGGCCA GCTCGGGAGCCAGACACAGCACTCACAGCCCAGGCCGTGATCCACCCTCCCCAAGTCCACCAGGGCCAGCGGCCCCTCACCTCTCTGGTC ACTGGTGAGACCTTCCACAACTTTCCTCCAGACCTGCCAGCAGATGTGCCCACCAGGGGCATTAGGTATCCGCCGGAGCCTGGCCATAGG GTAGTCTCGGGAGCCGCGCTGAGATCTTTTGCCACCTGCATTTTAGAAGAACATGGTCTCTGTCTCCTCGGCCCAGCCAGCTGTCCCGGC AAGGCCTGCCGAGGGCAGTTTTCAACCTCATGAAGGAAACACAGTCCTGCCAAGGAGGGGGAGTGGCGCCCATGGGGACAGGCCTCAGTC CTTAGAAGCCCTCTGGGTAGCTGTGCCCACCCAGCCTTCATGGCTGCAGGTACAAGGACCTTTGCTTCCATAGAGAAAACGCACAGCTCA GAAAGGGGGCCACATGGGCAGAAACCCAAAGGAAGGACAAACCACGACCACCGTGGCCATCTGCAGAATCCCTGGAAGAGAAGGAAGGCA GGGTGGAGCGGGGGGAAGACCATCATGGAGAGAAGGACCACAGCATCAGGAGACGGGACACGCCACACCCAGCAGGCAGCCTGTGTGTTG CTTAATTTTTTAAGAGCAAGAGGGGTAGAGAGGATCAAGCTGGCCCTGGCTGGAGATGGCTAGCCCCTGAGACATGCACTTCTGGTTTTG AAATGACTCTGTCTGTGGGGCAGCAGAAACTAGAGAAGGCAAGTGGCTGCCCCACCCCAAGGCGTGACCAGGAGGAACAGCCTGCAGCTC ACTCCATGCCACACGGGTGGGCCACCAGCCTGCTGTCAGAAGTCTCTGGGCTCCAACTGGTCTTGTAACCACTGAGCACTGAAGGAGAGA GGTCTTGGTCAGGGCTGGACAGCATGCCCGGGAGGACCAGCAGAGGATTAAAGGTGACTGGGAGGACCAGCGGAGGATAAAAGACACTGC TCAGGGCAGGGCTTCTACCCTGCATCCCTGGCCAAGAAAAGGGCAGTCCCCATGTGGGCTTGCAGGGTCACTCTCAGGGGCCTCTTTCAG CTGGGGCTGGCAACTTGCGTCTGGGGGACACCTCCAGGTGTGTGGGGTGAGGATTTCCTATAACCAGGGCTCCCAGAAGCTTTGCTTATG TAAGGAGGTCTGGGAGCCAGCCCATTGGAGGCCACCAGCCATTTTGGCTTCAAAGGACCCCACCTCACCCAGGTCTCAGCGGCAGTGGGC ACAGCTATGTCTTCAGGAGCTCCCGTCAAACCTCATAGCTGGGGCGCTCCCAGACAGGCCAGTCCAGACAGGACACGCTGGGCCCCTGGC ATCCAGAGGAAGAGCCAGGAGTGTGGGAAGGCCCACAGTGGGGGCTGTGGCTTCTGACACTCAGGTCATAGCCTCAGAGGTCTGAGGTCA GCCCCCACAGACCCATCCGGCCCGCCCCCCAAGTCCCTGCAGAGAGCACTTAGAGTTATGGCCCAGGCCCTGGTCCACCCTTCCCCTGTG CACCTCCGGCTGGGTTTGCCAAGTCAGGGAGCAGGGCTGGCCGCAGGAACTCCCAAACCTTGGCTTTGAATATTGTTGTGGAGGTGTGCT CGTCCCTTTCTGGACGTGCAAGGTACCTGTCCCAGCAGGTCAGATGGGGCCAGCTGAGGCGCTCCCCCAGGCAGGAAGGGCCAGCCTTCA CCATCGCGTGGGATTGGGAGGAGGGGCCTCCGTGAGCAGCCCCTCCTCTGCCGCTGTCCCAGCCCAGTCCCTCTCCCGGAGCCTTGGCAG CCTCCCACAACCCAGACACTTGCGTTCACAAGCAACCTAAGGGGCAGGTGAAGAAGCGCAGCCCTGCCAGACGCGCTAGATTCCTCTAAG GTCTCTGAGATGCACCGTTTTTTAAAAAGGCGTGGGGTGAACTGATTTTGATCTTCTTGTCTAGATGCAATAAATAAATCTGAAGCATTT AATGTAGTCATCTTGACATTGGGCCTACACTGTACGAGTTCCTTATGTTTCCTTGAGCTAAAAATATGTAAATAATTTTTGTCCCAGTGA GAACCGAGGGTTAGAAAACCTCGATGCCTCTGAGCCTCGGGACCGCTCTAGGGAAGTACCTGCTTTCGCCAGCATGACTCATGCTTCGTG >13766_13766_1_CCDC28A-CLEC16A_CCDC28A_chr6_139101122_ENST00000332797_CLEC16A_chr16_11260245_ENST00000409790_length(amino acids)=370AA_BP=197 MPRAEPRATLGEQEKAGLPLGAWRLYLLRHFRKQTELRRSGSRDVTGALLVAAAVASEAVGSLRVAEGGPNTLLLQVLRSWPWCNKELKT MEERKVKRRSPKSFSAHCTQVVNAKKNAIPVSKSTGFSNPASQSTSQRPKLKRVMKEKTKPQGGEGKGAQSTPIQHSFLTDVSDVQEMER GLLSLLNDFHSGKLQAFGFAVAQCINQHSSPSLSSQSPPSASGSPSGSGSTSHCDSGGTSSSSTPSTAQSPADAPMSPELPKPHLPDQLV IVNETEADSKPSKNVARSAAVETASLSPSLVPARQPTISLLCEDTADTLSVESLTLVPPVDPHSLRSLTGMPPLSTPAAACTEPVGEEAA -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CCDC28A-CLEC16A |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CCDC28A-CLEC16A |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CCDC28A-CLEC16A |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
