
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CDC34-MIER2 (FusionGDB2 ID:14815) |
Fusion Gene Summary for CDC34-MIER2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CDC34-MIER2 | Fusion gene ID: 14815 | Hgene | Tgene | Gene symbol | CDC34 | MIER2 | Gene ID | 997 | 54531 |
| Gene name | cell division cycle 34, ubiqiutin conjugating enzyme | MIER family member 2 | |
| Synonyms | E2-CDC34|UBC3|UBCH3|UBE2R1 | KIAA1193|Mi-er2 | |
| Cytomap | 19p13.3 | 19p13.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ubiquitin-conjugating enzyme E2 R1(E3-independent) E2 ubiquitin-conjugating enzyme R1E2 ubiquitin-conjugating enzyme R1cell division cycle 34 homologubiquitin carrier proteinubiquitin-conjugating enzyme E2-32 KDA complementingubiquitin-conjugating e | mesoderm induction early response protein 2mesoderm induction early response 1, family member 2 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | P49427 | Q8N344 | |
| Ensembl transtripts involved in fusion gene | ENST00000215574, | ENST00000592722, ENST00000264819, | |
| Fusion gene scores | * DoF score | 3 X 3 X 3=27 | 3 X 3 X 3=27 |
| # samples | 3 | 3 | |
| ** MAII score | log2(3/27*10)=0.15200309344505 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(3/27*10)=0.15200309344505 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: CDC34 [Title/Abstract] AND MIER2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CDC34(532108)-MIER2(336173), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | CDC34 | GO:0000209 | protein polyubiquitination | 20347421 |
| Hgene | CDC34 | GO:0016567 | protein ubiquitination | 10373550 |
| Hgene | CDC34 | GO:0043161 | proteasome-mediated ubiquitin-dependent protein catabolic process | 10373550|20347421 |
| Hgene | CDC34 | GO:0043951 | negative regulation of cAMP-mediated signaling | 10373550 |
| Hgene | CDC34 | GO:0070936 | protein K48-linked ubiquitination | 20061386 |
| Tgene | MIER2 | GO:0016575 | histone deacetylation | 28046085 |
 Fusion gene breakpoints across CDC34 (5'-gene) Fusion gene breakpoints across CDC34 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
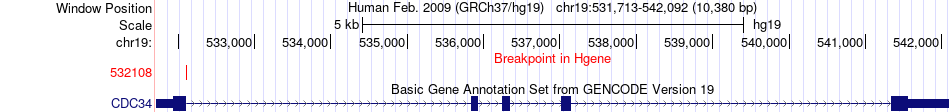 |
 Fusion gene breakpoints across MIER2 (3'-gene) Fusion gene breakpoints across MIER2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
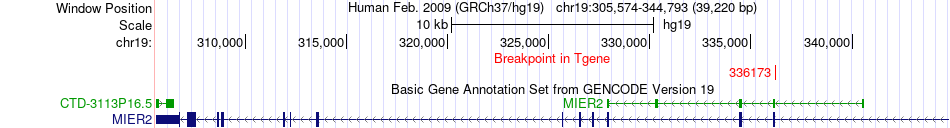 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SKCM | TCGA-GN-A265-06A | CDC34 | chr19 | 532108 | + | MIER2 | chr19 | 336173 | - |
Top |
Fusion Gene ORF analysis for CDC34-MIER2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000215574 | ENST00000592722 | CDC34 | chr19 | 532108 | + | MIER2 | chr19 | 336173 | - |
| In-frame | ENST00000215574 | ENST00000264819 | CDC34 | chr19 | 532108 | + | MIER2 | chr19 | 336173 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000215574 | CDC34 | chr19 | 532108 | + | ENST00000264819 | MIER2 | chr19 | 336173 | - | 3139 | 395 | 50 | 2023 | 657 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000215574 | ENST00000264819 | CDC34 | chr19 | 532108 | + | MIER2 | chr19 | 336173 | - | 0.004338872 | 0.9956611 |
Top |
Fusion Genomic Features for CDC34-MIER2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
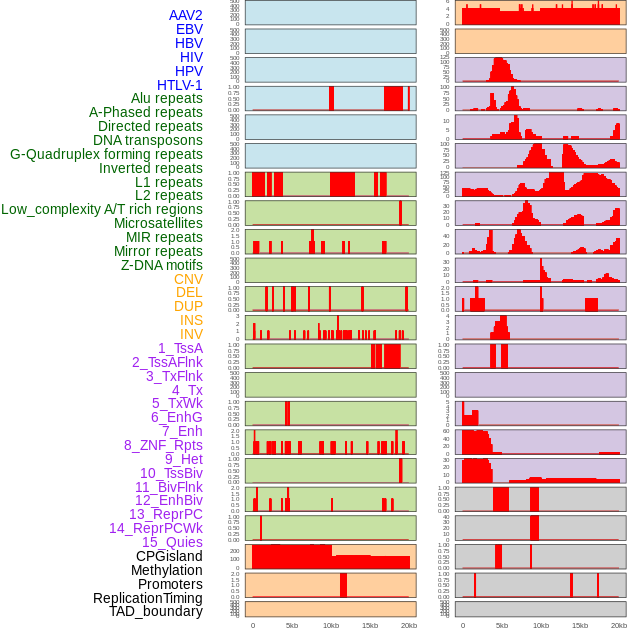 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for CDC34-MIER2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:532108/chr19:336173) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CDC34 | MIER2 |
| FUNCTION: Accepts ubiquitin from the E1 complex and catalyzes its covalent attachment to other proteins. In vitro catalyzes 'Lys-48'-linked polyubiquitination (PubMed:22496338). Cooperates with the E2 UBCH5C and the SCF(FBXW11) E3 ligase complex for the polyubiquitination of NFKBIA leading to its subsequent proteasomal degradation. Performs ubiquitin chain elongation building ubiquitin chains from the UBE2D3-primed NFKBIA-linked ubiquitin. UBE2D3 acts as an initiator E2, priming the phosphorylated NFKBIA target at positions 'Lys-21' and/or 'Lys-22' with a monoubiquitin. Cooperates with the SCF(SKP2) E3 ligase complex to regulate cell proliferation through ubiquitination and degradation of MYBL2 and KIP1. Involved in ubiquitin conjugation and degradation of CREM isoform ICERIIgamma and ATF15 resulting in abrogation of ICERIIgamma- and ATF5-mediated repression of cAMP-induced transcription during both meiotic and mitotic cell cycles. Involved in the regulation of the cell cycle G2/M phase through its targeting of the WEE1 kinase for ubiquitination and degradation. Also involved in the degradation of beta-catenin. Is target of human herpes virus 1 protein ICP0, leading to ICP0-dependent dynamic interaction with proteasomes (PubMed:10329681, PubMed:10373550, PubMed:10871850, PubMed:11675391, PubMed:12037680, PubMed:15652359, PubMed:17461777, PubMed:17698585, PubMed:19112177, PubMed:19126550, PubMed:19945379, PubMed:20061386, PubMed:20347421). {ECO:0000269|PubMed:10329681, ECO:0000269|PubMed:10373550, ECO:0000269|PubMed:10871850, ECO:0000269|PubMed:11675391, ECO:0000269|PubMed:12037680, ECO:0000269|PubMed:15652359, ECO:0000269|PubMed:17461777, ECO:0000269|PubMed:17698585, ECO:0000269|PubMed:19112177, ECO:0000269|PubMed:19126550, ECO:0000269|PubMed:19945379, ECO:0000269|PubMed:20061386, ECO:0000269|PubMed:20347421, ECO:0000269|PubMed:22496338}. | FUNCTION: Transcriptional repressor. {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | MIER2 | chr19:532108 | chr19:336173 | ENST00000264819 | 0 | 14 | 195_292 | 3 | 546.0 | Domain | ELM2 | |
| Tgene | MIER2 | chr19:532108 | chr19:336173 | ENST00000264819 | 0 | 14 | 297_349 | 3 | 546.0 | Domain | SANT |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CDC34 | chr19:532108 | chr19:336173 | ENST00000215574 | + | 1 | 5 | 200_236 | 59 | 237.0 | Compositional bias | Note=Asp/Glu-rich (acidic) |
| Hgene | CDC34 | chr19:532108 | chr19:336173 | ENST00000215574 | + | 1 | 5 | 8_174 | 59 | 237.0 | Domain | UBC core |
| Hgene | CDC34 | chr19:532108 | chr19:336173 | ENST00000215574 | + | 1 | 5 | 190_236 | 59 | 237.0 | Region | Note=SCF-binding |
Top |
Fusion Gene Sequence for CDC34-MIER2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >14815_14815_1_CDC34-MIER2_CDC34_chr19_532108_ENST00000215574_MIER2_chr19_336173_ENST00000264819_length(transcript)=3139nt_BP=395nt GGCGCGCGGGCCCGGCCAAGGCAAGCGCCGGTGGGGCGGCGGCGCCAGAGCTGCTGGAGCGCTCGGGGTCCCCGGGCGGCGGCGGCGGCG CAGAGGAGGAGGCAGGCGGCGGCCCCGGTGGCTCCCCCCCGGACGGTGCGCGGCCCGGCCCGTCTCGCGAACTCGCGGTGGTCGCGCGGC CCCGCGCTGCTCCGACCCCGGGCCCCTCCGCCGCCGCCATGGCTCGGCCGCTAGTGCCCAGCTCGCAGAAGGCGCTGCTGCTGGAGCTCA AGGGGCTGCAGGAAGAGCCGGTCGAGGGATTCCGCGTGACACTGGTGGACGAGGGCGATCTATACAACTGGGAGGTGGCCATCTTCGGGC CCCCCAACACCTACTACGAGGGCGGCTACTTCAAGGCCTCCTCGCTGGGGAGGCAGAGTCCTCGCGTGGTCTCCTGCCTCGAGCACAGCC TGTGCCCAGGGGAGCCGGGCTTGCAGACAACAGCAGTGGTGTCCATGGGCTCTGGAGACCATCAGTTCAACCTCGCAGAGATCCTGTCAC AGAACTACAGTGTTAGGGGGGAGTGCGAGGAGGCCTCGAGGTGCCCAGACAAGCCCAAGGAGGAGCTGGAGAAGGACTTCATCTCCCAGA GCAACGACATGCCCTTTGATGAGCTGCTTGCGCTCTATGGCTACGAGGCGTCAGACCCCATTTCAGACCGGGAGAGTGAGGGTGGTGACG TGGCCCCGAACCTCCCAGACATGACCCTGGACAAAGAACAAATAGCGAAGGATTTGCTTTCAGGGGAAGAAGAGGAAGAGACGCAATCAT CTGCTGACGACCTCACCCCGTCCGTGACCTCCCACGAGGCCTCCGACCTCTTCCCTAACCGGAGTGGATCTCGTTTCCTGGCTGATGAAG ACAGAGAGCCTGGCTCTTCTGCCTCCTCCGACACCGAGGAGGACTCTCTTCCTGCCAACAAATGTAAGAAGGAGATCATGGTGGGACCTC AGTTCCAAGCTGACCTCAGCAACCTGCACTTGAACCGGCACTGTGAGAAGATCTACGAGAACGAAGACCAGCTGCTCTGGGACCCCAGCG TCCTCCCTGAGAGGGAGGTGGAGGAGTTCCTGTACAGGGCGGTGAAGCGGCGTTGGCACGAGATGGCCGGGCCTCAGCTCCCAGAGGGAG AAGCCGTGAAAGACAGTGAGCAGGCGCTGTACGAGTTGGTGAAATGCAACTTCAATGTGGAGGAGGCCCTGCGAAGGCTGCGGTTCAACG TGAAGGTGATCCGAGATGGGCTCTGTGCTTGGAGTGAAGAGGAGTGCAGGAACTTTGAGCACGGCTTCCGTGTGCATGGAAAGAACTTTC ACCTGATCCAGGCCAACAAGGTGCGCACACGGTCAGTGGGCGAGTGTGTCGAGTACTACTACCTGTGGAAGAAGTCGGAGCGCTACGACT ACTTCGCCCAGCAGACGCGGCTGGGCCGGAGGAAGTACGTCCCGTCCGGAACCACGGACGCAGACCAGGACCTGGATGGCAGCGACCCCG ATGGCCCCGGCCGTCCGCGCCCGGAGCAAGACACCCTGACTGGGATGCGCACAGATCCACTGAGCGTGGATGGCACGGCCGGTGGTCTCG ATGAGCCCGGAGTGGCCTCTGATGGACTCCCGTCCTCGGAGCCAGGGCCGTGTTCCTTCCAGCAGCTGGATGAGTCCCCCGCTGTACCCC TGTCCCATCGGCCCCCAGCCCTGGCCGACCCAGCCTCATACCAGCCAGCTGTCACTGCTCCGGAGCCAGACGCCAGCCCAAGGCTGGCCG TGGACTTCGCCCTGCCCAAGGAGCTGCCCCTCATCTCCAGCCATGTGGACCTCAGCGGGGATCCGGAGGAGACTGTGGCCCCAGCACAGG TGGCTTTGTCGGTCACCGAGTTTGGACTCATCGGCATTGGGGACGTGAACCCCTTCCTGGCCGCCCACCCCACGTGCCCGGCCCCCGGGC TACACTCGGAGCCCCTGTCACACTGTAACGTGATGACCTGCTGACTCCTGGCCGCGGGCGGCGTATGCGGCCCAGACTGGACTTAGCGCT GCCGCTGGGCCCGCCTCTGTCAGTCTTCCTGACCCCTTCCCCACCCCCCGGGCCTTGGGGTAGCACCTCCTTCTGCTTCAGAACACGTCA GGACTGGGGTGAGGTGGCTGGGCCGTGAGCCCTTGCCCCTGTCCACACAGAATGGACCCACGGCCCCACCCAGCGCCGTCAGCGCCCGGC ACTGCCACCCGGGTCCGGGCCGCTGCCTGCACGTGGGATCCGTCGGGCAGCCGGGGACAGAAGAGACCCCGCCGTTGGGACGCAGGGCAG AGCCGGCCACCTAGTCCCTTCCAGCCAGCAGAGGCGAGGGAAGGCGTCACTGCCCCGGCGGGGAGACGGGCAGGACGCCCTGCCCCGCAC CAGCAGCCTCCGCCGGGGCGCCCTCAGCTCCCTGCTTGGCTCTGTCTCTCCACACCCGGCAGGGCCGCGGGCTGCCCCAGCCCTGGGGGT CGTGGGCAGCTGCTACTCAGTGCCAACCCCGTGGGGCACAGAGCCATATACCTCGCTGTCCGGCCCCCACCCCAGCCTCGCCTTCCCACC CCATCGTCTCCACTTCAGGAAAAGCCGCACTTTACACCCCCACCTGCCTCTTCCCCCTCCATCCCTGCTCCCCGATCCTGAGCGGTTGGG GTGGGGTCCCTCAGCAACCCCAGGCGTGGGTTTGAGGAGACAGGTGATTTACATCCCCTTTGCTGTCCTCCCCCGGTACCAAGGCAGGGA GCCTCCGGAGGAGCCGGCCCTGCTGGCCACGCAGGGGCCAGACTCCAGCCTGTTTCCCCAGCCCTGCAGGTCTTCCTTCTGTGGGAAGCT TCCTAGCAAGATGGCTTGGAGTCCTGGTCCCCCTCCTCCCTGGCCCTCTCGTTCGTTTCTGTTTCTGTTTACACGTTGGAGTGGGGTCCT CCGTGGGCGGCGGCGCGCCCTGCCCCGGGTGTCGTCCGGCCTCTTGTGCTCGAGCCCCTTTCCGAGTTGGACTCGACCATCCCTCACCCC >14815_14815_1_CDC34-MIER2_CDC34_chr19_532108_ENST00000215574_MIER2_chr19_336173_ENST00000264819_length(amino acids)=657AA_BP=9 MLERSGSPGGGGGAEEEAGGGPGGSPPDGARPGPSRELAVVARPRAAPTPGPSAAAMARPLVPSSQKALLLELKGLQEEPVEGFRVTLVD EGDLYNWEVAIFGPPNTYYEGGYFKASSLGRQSPRVVSCLEHSLCPGEPGLQTTAVVSMGSGDHQFNLAEILSQNYSVRGECEEASRCPD KPKEELEKDFISQSNDMPFDELLALYGYEASDPISDRESEGGDVAPNLPDMTLDKEQIAKDLLSGEEEEETQSSADDLTPSVTSHEASDL FPNRSGSRFLADEDREPGSSASSDTEEDSLPANKCKKEIMVGPQFQADLSNLHLNRHCEKIYENEDQLLWDPSVLPEREVEEFLYRAVKR RWHEMAGPQLPEGEAVKDSEQALYELVKCNFNVEEALRRLRFNVKVIRDGLCAWSEEECRNFEHGFRVHGKNFHLIQANKVRTRSVGECV EYYYLWKKSERYDYFAQQTRLGRRKYVPSGTTDADQDLDGSDPDGPGRPRPEQDTLTGMRTDPLSVDGTAGGLDEPGVASDGLPSSEPGP CSFQQLDESPAVPLSHRPPALADPASYQPAVTAPEPDASPRLAVDFALPKELPLISSHVDLSGDPEETVAPAQVALSVTEFGLIGIGDVN -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CDC34-MIER2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CDC34-MIER2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CDC34-MIER2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
