
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CDC5L-HSP90AB1 (FusionGDB2 ID:14918) |
Fusion Gene Summary for CDC5L-HSP90AB1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CDC5L-HSP90AB1 | Fusion gene ID: 14918 | Hgene | Tgene | Gene symbol | CDC5L | HSP90AB1 | Gene ID | 988 | 3326 |
| Gene name | cell division cycle 5 like | heat shock protein 90 alpha family class B member 1 | |
| Synonyms | CDC5|CDC5-LIKE|CEF1|PCDC5RP|dJ319D22.1 | D6S182|HSP84|HSP90B|HSPC2|HSPCB | |
| Cytomap | 6p21.1 | 6p21.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | cell division cycle 5-like proteinCDC5 cell division cycle 5-likeCdc5-related proteindJ319D22.1 (CDC5-like protein)pombe cdc5-related protein | heat shock protein HSP 90-betaHSP90-betaheat shock 84 kDaheat shock 90kD protein 1, betaheat shock protein 90 kDaheat shock protein 90kDa alpha (cytosolic), class B member 1heat shock protein 90kDa alpha family class B member 1 | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | Q99459 | P08238 | |
| Ensembl transtripts involved in fusion gene | ENST00000371477, | ENST00000353801, ENST00000371554, ENST00000371646, | |
| Fusion gene scores | * DoF score | 10 X 4 X 7=280 | 17 X 17 X 9=2601 |
| # samples | 11 | 20 | |
| ** MAII score | log2(11/280*10)=-1.34792330342031 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(20/2601*10)=-3.70099449416827 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CDC5L [Title/Abstract] AND HSP90AB1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CDC5L(44376369)-HSP90AB1(44216367), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | CDC5L-HSP90AB1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. CDC5L-HSP90AB1 seems lost the major protein functional domain in Hgene partner, which is a transcription factor due to the frame-shifted ORF. CDC5L-HSP90AB1 seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. CDC5L-HSP90AB1 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | CDC5L | GO:0000398 | mRNA splicing, via spliceosome | 28076346 |
| Hgene | CDC5L | GO:0045944 | positive regulation of transcription by RNA polymerase II | 11082045 |
| Tgene | HSP90AB1 | GO:0007004 | telomere maintenance via telomerase | 10197982 |
| Tgene | HSP90AB1 | GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway | 24613385 |
| Tgene | HSP90AB1 | GO:0031396 | regulation of protein ubiquitination | 16809764 |
| Tgene | HSP90AB1 | GO:0032435 | negative regulation of proteasomal ubiquitin-dependent protein catabolic process | 24613385 |
| Tgene | HSP90AB1 | GO:0032516 | positive regulation of phosphoprotein phosphatase activity | 26593036 |
| Tgene | HSP90AB1 | GO:0051131 | chaperone-mediated protein complex assembly | 10811660 |
| Tgene | HSP90AB1 | GO:0051973 | positive regulation of telomerase activity | 10197982 |
| Tgene | HSP90AB1 | GO:1901389 | negative regulation of transforming growth factor beta activation | 20599762 |
| Tgene | HSP90AB1 | GO:1905323 | telomerase holoenzyme complex assembly | 10197982 |
| Tgene | HSP90AB1 | GO:2000010 | positive regulation of protein localization to cell surface | 23431407 |
 Fusion gene breakpoints across CDC5L (5'-gene) Fusion gene breakpoints across CDC5L (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
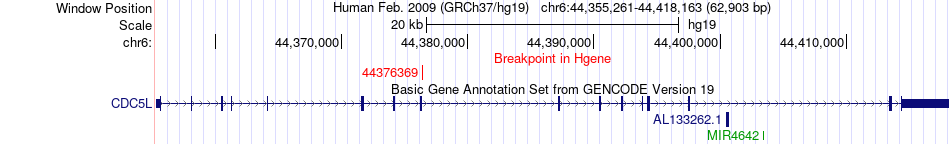 |
 Fusion gene breakpoints across HSP90AB1 (3'-gene) Fusion gene breakpoints across HSP90AB1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
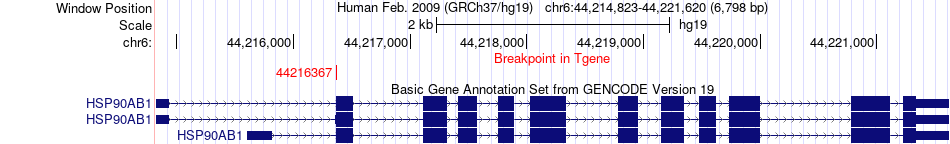 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-CD-8535-01A | CDC5L | chr6 | 44376369 | + | HSP90AB1 | chr6 | 44216367 | + |
Top |
Fusion Gene ORF analysis for CDC5L-HSP90AB1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000371477 | ENST00000353801 | CDC5L | chr6 | 44376369 | + | HSP90AB1 | chr6 | 44216367 | + |
| In-frame | ENST00000371477 | ENST00000371554 | CDC5L | chr6 | 44376369 | + | HSP90AB1 | chr6 | 44216367 | + |
| In-frame | ENST00000371477 | ENST00000371646 | CDC5L | chr6 | 44376369 | + | HSP90AB1 | chr6 | 44216367 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000371477 | CDC5L | chr6 | 44376369 | + | ENST00000371646 | HSP90AB1 | chr6 | 44216367 | + | 3851 | 1391 | 275 | 3565 | 1096 |
| ENST00000371477 | CDC5L | chr6 | 44376369 | + | ENST00000371554 | HSP90AB1 | chr6 | 44216367 | + | 3851 | 1391 | 275 | 3565 | 1096 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000371477 | ENST00000371646 | CDC5L | chr6 | 44376369 | + | HSP90AB1 | chr6 | 44216367 | + | 0.00254313 | 0.99745685 |
| ENST00000371477 | ENST00000371554 | CDC5L | chr6 | 44376369 | + | HSP90AB1 | chr6 | 44216367 | + | 0.00254313 | 0.99745685 |
Top |
Fusion Genomic Features for CDC5L-HSP90AB1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CDC5L | chr6 | 44376369 | + | HSP90AB1 | chr6 | 44216366 | + | 3.78E-09 | 1 |
| CDC5L | chr6 | 44376369 | + | HSP90AB1 | chr6 | 44216366 | + | 3.78E-09 | 1 |
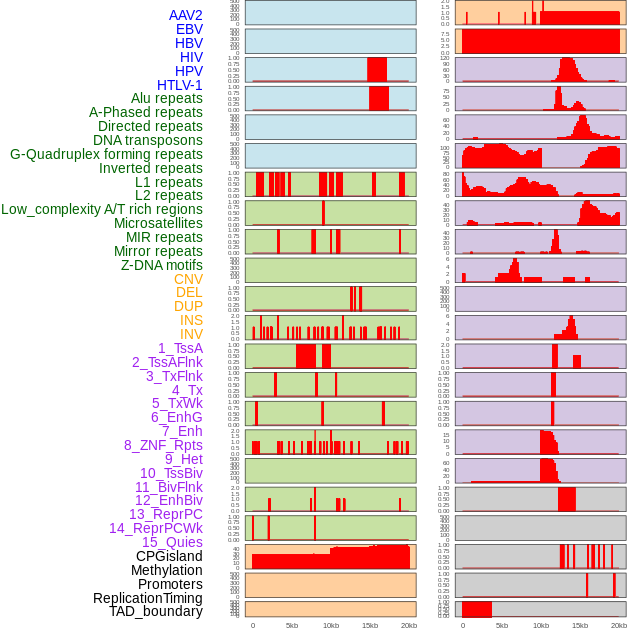
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
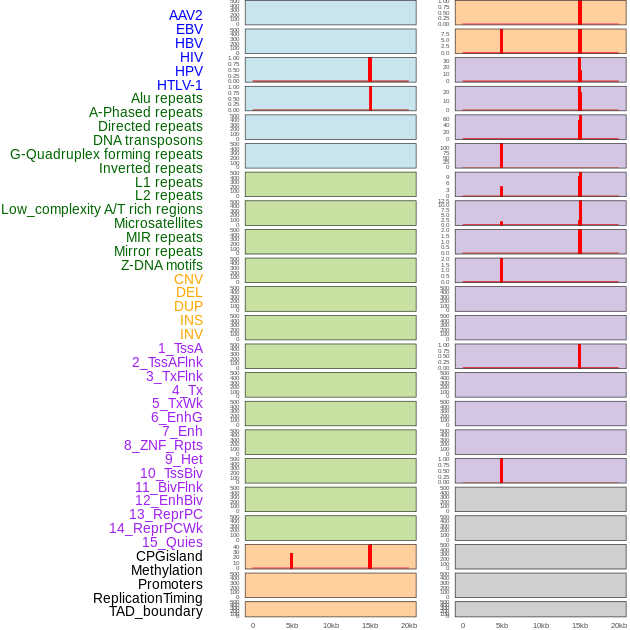
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for CDC5L-HSP90AB1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr6:44376369/chr6:44216367) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CDC5L | HSP90AB1 |
| FUNCTION: DNA-binding protein involved in cell cycle control. May act as a transcription activator. Plays role in pre-mRNA splicing as core component of precatalytic, catalytic and postcatalytic spliceosomal complexes (PubMed:11991638, PubMed:20176811, PubMed:28502770, PubMed:28076346, PubMed:29361316, PubMed:29360106, PubMed:29301961, PubMed:30728453, PubMed:30705154). Component of the PRP19-CDC5L complex that forms an integral part of the spliceosome and is required for activating pre-mRNA splicing. The PRP19-CDC5L complex may also play a role in the response to DNA damage (DDR) (PubMed:20176811). {ECO:0000269|PubMed:10570151, ECO:0000269|PubMed:11082045, ECO:0000269|PubMed:11101529, ECO:0000269|PubMed:11544257, ECO:0000269|PubMed:11991638, ECO:0000269|PubMed:12927788, ECO:0000269|PubMed:18583928, ECO:0000269|PubMed:20176811, ECO:0000269|PubMed:24332808, ECO:0000269|PubMed:28076346, ECO:0000269|PubMed:28502770, ECO:0000269|PubMed:29301961, ECO:0000269|PubMed:29360106, ECO:0000269|PubMed:29361316, ECO:0000269|PubMed:30705154, ECO:0000269|PubMed:30728453, ECO:0000269|PubMed:9038199, ECO:0000269|PubMed:9468527, ECO:0000269|PubMed:9632794}. | FUNCTION: Molecular chaperone that promotes the maturation, structural maintenance and proper regulation of specific target proteins involved for instance in cell cycle control and signal transduction. Undergoes a functional cycle linked to its ATPase activity. This cycle probably induces conformational changes in the client proteins, thereby causing their activation. Interacts dynamically with various co-chaperones that modulate its substrate recognition, ATPase cycle and chaperone function (PubMed:16478993, PubMed:19696785). Engages with a range of client protein classes via its interaction with various co-chaperone proteins or complexes, that act as adapters, simultaneously able to interact with the specific client and the central chaperone itself. Recruitment of ATP and co-chaperone followed by client protein forms a functional chaperone. After the completion of the chaperoning process, properly folded client protein and co-chaperone leave HSP90 in an ADP-bound partially open conformation and finally, ADP is released from HSP90 which acquires an open conformation for the next cycle (PubMed:27295069, PubMed:26991466). Apart from its chaperone activity, it also plays a role in the regulation of the transcription machinery. HSP90 and its co-chaperones modulate transcription at least at three different levels. They first alter the steady-state levels of certain transcription factors in response to various physiological cues. Second, they modulate the activity of certain epigenetic modifiers, such as histone deacetylases or DNA methyl transferases, and thereby respond to the change in the environment. Third, they participate in the eviction of histones from the promoter region of certain genes and thereby turn on gene expression (PubMed:25973397). Antagonizes STUB1-mediated inhibition of TGF-beta signaling via inhibition of STUB1-mediated SMAD3 ubiquitination and degradation (PubMed:24613385). Promotes cell differentiation by chaperoning BIRC2 and thereby protecting from auto-ubiquitination and degradation by the proteasomal machinery (PubMed:18239673). Main chaperone involved in the phosphorylation/activation of the STAT1 by chaperoning both JAK2 and PRKCE under heat shock and in turn, activates its own transcription (PubMed:20353823). Involved in the translocation into ERGIC (endoplasmic reticulum-Golgi intermediate compartment) of leaderless cargos (lacking the secretion signal sequence) such as the interleukin 1/IL-1; the translocation process is mediated by the cargo receptor TMED10 (PubMed:32272059). {ECO:0000269|PubMed:16478993, ECO:0000269|PubMed:18239673, ECO:0000269|PubMed:19696785, ECO:0000269|PubMed:20353823, ECO:0000269|PubMed:24613385, ECO:0000269|PubMed:32272059, ECO:0000303|PubMed:25973397, ECO:0000303|PubMed:26991466, ECO:0000303|PubMed:27295069}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 142_245 | 364 | 803.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 31_54 | 364 | 803.0 | DNA binding | H-T-H motif |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 82_104 | 364 | 803.0 | DNA binding | H-T-H motif |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 1_56 | 364 | 803.0 | Domain | HTH myb-type 1 |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 57_108 | 364 | 803.0 | Domain | HTH myb-type 2 |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 165_271 | 364 | 803.0 | Motif | Nuclear localization signal |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000353801 | 0 | 12 | 720_724 | 0 | 725.0 | Motif | Note=TPR repeat-binding | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371554 | 0 | 12 | 720_724 | 0 | 725.0 | Motif | Note=TPR repeat-binding | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371646 | 0 | 12 | 720_724 | 0 | 725.0 | Motif | Note=TPR repeat-binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 676_701 | 364 | 803.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 764_802 | 364 | 803.0 | Coiled coil | Ontology_term=ECO:0000255 |
Top |
Fusion Gene Sequence for CDC5L-HSP90AB1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >14918_14918_1_CDC5L-HSP90AB1_CDC5L_chr6_44376369_ENST00000371477_HSP90AB1_chr6_44216367_ENST00000371554_length(transcript)=3851nt_BP=1391nt TTTCTTCCATTCTGTTTTGGACATGCGCGGGGCGTAAGGCGCTTTGGCCAGAGTGGTTTGATGGTTTTGTGTTAGTCTTCCTGCAGCTGA GCTTCCACAAGACACTTTTTCCCTTTCAGCCACCCAATATCTCGCGAGACTTGCTATTCTGCGCCCCCTCCCACCTTGCGCTTGCGCAGA TCTTCAAAGCAGAAGGTCGCGCTTGGAGGAAGTGGCGGCTTTGAGTCCGGTGGCCCAATCGCTGTTACTACTTCTCTGAAGCTCCTCTCG GCTGCTTGCCGAGACACCCTGCCGCCAAGATGCCTCGAATTATGATCAAGGGGGGCGTATGGAGGAATACCGAGGATGAAATTCTGAAAG CAGCGGTAATGAAATATGGGAAAAATCAGTGGTCTAGGATTGCCTCATTGCTGCATAGAAAATCAGCAAAGCAGTGCAAAGCCAGATGGT ATGAATGGCTGGATCCAAGCATTAAGAAGACAGAATGGTCCAGAGAAGAAGAGGAAAAACTCTTGCACTTGGCCAAGTTGATGCCAACTC AGTGGAGGACCATTGCTCCAATCATTGGAAGAACAGCGGCCCAGTGCTTAGAACACTATGAATTTCTTCTGGATAAAGCTGCCCAAAGAG ACAATGAAGAGGAAACAACAGATGATCCACGAAAACTTAAACCTGGAGAAATAGATCCAAATCCAGAAACAAAACCAGCGCGGCCTGATC CAATTGATATGGATGAGGATGAACTTGAGATGCTTTCTGAAGCCAGAGCCCGCTTGGCTAATACTCAGGGAAAGAAGGCCAAGAGGAAAG CAAGAGAGAAACAATTGGAAGAAGCAAGACGTCTTGCTGCCCTCCAAAAAAGAAGAGAACTTCGAGCAGCTGGCATAGAAATTCAGAAGA AAAGAAAAAGGAAGAGAGGAGTTGATTATAATGCCGAAATCCCATTTGAAAAAAAGCCTGCCCTTGGTTTTTATGATACTTCTGAGGAAA ACTACCAAGCTCTTGACGCAGATTTCAGGAAATTAAGACAACAGGATCTTGATGGGGAGCTAAGATCTGAAAAAGAAGGAAGAGATAGAA AAAAAGACAAACAGCATTTGAAAAGGAAAAAAGAATCTGATTTACCATCAGCTATTCTTCAAACTAGTGGTGTTTCTGAATTTACTAAAA AGAGAAGCAAACTAGTACTTCCTGCCCCTCAGATTTCAGATGCAGAACTCCAGGAAGTTGTAAAAGTAGGCCAAGCGAGTGAAATTGCAC GTCAAACTGCCGAGGAATCTGGCATAACAAATTCTGCTTCCAGTACACTTTTGTCTGAGTACAATGTCACCAACAACAGCGTTGCTCTTA GAACACCACGAACACCAGCTTCCCAGGACAGAATTCTGCAGATGCCTGAGGAAGTGCACCATGGAGAGGAGGAGGTGGAGACTTTTGCCT TTCAGGCAGAAATTGCCCAACTCATGTCCCTCATCATCAATACCTTCTATTCCAACAAGGAGATTTTCCTTCGGGAGTTGATCTCTAATG CTTCTGATGCCTTGGACAAGATTCGCTATGAGAGCCTGACAGACCCTTCGAAGTTGGACAGTGGTAAAGAGCTGAAAATTGACATCATCC CCAACCCTCAGGAACGTACCCTGACTTTGGTAGACACAGGCATTGGCATGACCAAAGCTGATCTCATAAATAATTTGGGAACCATTGCCA AGTCTGGTACTAAAGCATTCATGGAGGCTCTTCAGGCTGGTGCAGACATCTCCATGATTGGGCAGTTTGGTGTTGGCTTTTATTCTGCCT ACTTGGTGGCAGAGAAAGTGGTTGTGATCACAAAGCACAACGATGATGAACAGTATGCTTGGGAGTCTTCTGCTGGAGGTTCCTTCACTG TGCGTGCTGACCATGGTGAGCCCATTGGCAGGGGTACCAAAGTGATCCTCCATCTTAAAGAAGATCAGACAGAGTACCTAGAAGAGAGGC GGGTCAAAGAAGTAGTGAAGAAGCATTCTCAGTTCATAGGCTATCCCATCACCCTTTATTTGGAGAAGGAACGAGAGAAGGAAATTAGTG ATGATGAGGCAGAGGAAGAGAAAGGTGAGAAAGAAGAGGAAGATAAAGATGATGAAGAAAAACCCAAGATCGAAGATGTGGGTTCAGATG AGGAGGATGACAGCGGTAAGGATAAGAAGAAGAAAACTAAGAAGATCAAAGAGAAATACATTGATCAGGAAGAACTAAACAAGACCAAGC CTATTTGGACCAGAAACCCTGATGACATCACCCAAGAGGAGTATGGAGAATTCTACAAGAGCCTCACTAATGACTGGGAAGACCACTTGG CAGTCAAGCACTTTTCTGTAGAAGGTCAGTTGGAATTCAGGGCATTGCTATTTATTCCTCGTCGGGCTCCCTTTGACCTTTTTGAGAACA AGAAGAAAAAGAACAACATCAAACTCTATGTCCGCCGTGTGTTCATCATGGACAGCTGTGATGAGTTGATACCAGAGTATCTCAATTTTA TCCGTGGTGTGGTTGACTCTGAGGATCTGCCCCTGAACATCTCCCGAGAAATGCTCCAGCAGAGCAAAATCTTGAAAGTCATTCGCAAAA ACATTGTTAAGAAGTGCCTTGAGCTCTTCTCTGAGCTGGCAGAAGACAAGGAGAATTACAAGAAATTCTATGAGGCATTCTCTAAAAATC TCAAGCTTGGAATCCACGAAGACTCCACTAACCGCCGCCGCCTGTCTGAGCTGCTGCGCTATCATACCTCCCAGTCTGGAGATGAGATGA CATCTCTGTCAGAGTATGTTTCTCGCATGAAGGAGACACAGAAGTCCATCTATTACATCACTGGTGAGAGCAAAGAGCAGGTGGCCAACT CAGCTTTTGTGGAGCGAGTGCGGAAACGGGGCTTCGAGGTGGTATATATGACCGAGCCCATTGACGAGTACTGTGTGCAGCAGCTCAAGG AATTTGATGGGAAGAGCCTGGTCTCAGTTACCAAGGAGGGTCTGGAGCTGCCTGAGGATGAGGAGGAGAAGAAGAAGATGGAAGAGAGCA AGGCAAAGTTTGAGAACCTCTGCAAGCTCATGAAAGAAATCTTAGATAAGAAGGTTGAGAAGGTGACAATCTCCAATAGACTTGTGTCTT CACCTTGCTGCATTGTGACCAGCACCTACGGCTGGACAGCCAATATGGAGCGGATCATGAAAGCCCAGGCACTTCGGGACAACTCCACCA TGGGCTATATGATGGCCAAAAAGCACCTGGAGATCAACCCTGACCACCCCATTGTGGAGACGCTGCGGCAGAAGGCTGAGGCCGACAAGA ATGATAAGGCAGTTAAGGACCTGGTGGTGCTGCTGTTTGAAACCGCCCTGCTATCTTCTGGCTTTTCCCTTGAGGATCCCCAGACCCACT CCAACCGCATCTATCGCATGATCAAGCTAGGTCTAGGTATTGATGAAGATGAAGTGGCAGCAGAGGAACCCAATGCTGCAGTTCCTGATG AGATCCCCCCTCTCGAGGGCGATGAGGATGCGTCTCGCATGGAAGAAGTCGATTAGGTTAGGAGTTCATAGTTGGAAAACTTGTGCCCTT GTATAGTGTCCCCATGGGCTCCCACTGCAGCCTCGAGTGCCCCTGTCCCACCTGGCTCCCCCTGCTGGTGTCTAGTGTTTTTTTCCCTCT CCTGTCCTTGTGTTGAAGGCAGTAAACTAAGGGTGTCAAGCCCCATTCCCTCTCTACTCTTGACAGCAGGATTGGATGTTGTGTATTGTG >14918_14918_1_CDC5L-HSP90AB1_CDC5L_chr6_44376369_ENST00000371477_HSP90AB1_chr6_44216367_ENST00000371554_length(amino acids)=1096AA_BP=372 MPRHPAAKMPRIMIKGGVWRNTEDEILKAAVMKYGKNQWSRIASLLHRKSAKQCKARWYEWLDPSIKKTEWSREEEEKLLHLAKLMPTQW RTIAPIIGRTAAQCLEHYEFLLDKAAQRDNEEETTDDPRKLKPGEIDPNPETKPARPDPIDMDEDELEMLSEARARLANTQGKKAKRKAR EKQLEEARRLAALQKRRELRAAGIEIQKKRKRKRGVDYNAEIPFEKKPALGFYDTSEENYQALDADFRKLRQQDLDGELRSEKEGRDRKK DKQHLKRKKESDLPSAILQTSGVSEFTKKRSKLVLPAPQISDAELQEVVKVGQASEIARQTAEESGITNSASSTLLSEYNVTNNSVALRT PRTPASQDRILQMPEEVHHGEEEVETFAFQAEIAQLMSLIINTFYSNKEIFLRELISNASDALDKIRYESLTDPSKLDSGKELKIDIIPN PQERTLTLVDTGIGMTKADLINNLGTIAKSGTKAFMEALQAGADISMIGQFGVGFYSAYLVAEKVVVITKHNDDEQYAWESSAGGSFTVR ADHGEPIGRGTKVILHLKEDQTEYLEERRVKEVVKKHSQFIGYPITLYLEKEREKEISDDEAEEEKGEKEEEDKDDEEKPKIEDVGSDEE DDSGKDKKKKTKKIKEKYIDQEELNKTKPIWTRNPDDITQEEYGEFYKSLTNDWEDHLAVKHFSVEGQLEFRALLFIPRRAPFDLFENKK KKNNIKLYVRRVFIMDSCDELIPEYLNFIRGVVDSEDLPLNISREMLQQSKILKVIRKNIVKKCLELFSELAEDKENYKKFYEAFSKNLK LGIHEDSTNRRRLSELLRYHTSQSGDEMTSLSEYVSRMKETQKSIYYITGESKEQVANSAFVERVRKRGFEVVYMTEPIDEYCVQQLKEF DGKSLVSVTKEGLELPEDEEEKKKMEESKAKFENLCKLMKEILDKKVEKVTISNRLVSSPCCIVTSTYGWTANMERIMKAQALRDNSTMG YMMAKKHLEINPDHPIVETLRQKAEADKNDKAVKDLVVLLFETALLSSGFSLEDPQTHSNRIYRMIKLGLGIDEDEVAAEEPNAAVPDEI -------------------------------------------------------------- >14918_14918_2_CDC5L-HSP90AB1_CDC5L_chr6_44376369_ENST00000371477_HSP90AB1_chr6_44216367_ENST00000371646_length(transcript)=3851nt_BP=1391nt TTTCTTCCATTCTGTTTTGGACATGCGCGGGGCGTAAGGCGCTTTGGCCAGAGTGGTTTGATGGTTTTGTGTTAGTCTTCCTGCAGCTGA GCTTCCACAAGACACTTTTTCCCTTTCAGCCACCCAATATCTCGCGAGACTTGCTATTCTGCGCCCCCTCCCACCTTGCGCTTGCGCAGA TCTTCAAAGCAGAAGGTCGCGCTTGGAGGAAGTGGCGGCTTTGAGTCCGGTGGCCCAATCGCTGTTACTACTTCTCTGAAGCTCCTCTCG GCTGCTTGCCGAGACACCCTGCCGCCAAGATGCCTCGAATTATGATCAAGGGGGGCGTATGGAGGAATACCGAGGATGAAATTCTGAAAG CAGCGGTAATGAAATATGGGAAAAATCAGTGGTCTAGGATTGCCTCATTGCTGCATAGAAAATCAGCAAAGCAGTGCAAAGCCAGATGGT ATGAATGGCTGGATCCAAGCATTAAGAAGACAGAATGGTCCAGAGAAGAAGAGGAAAAACTCTTGCACTTGGCCAAGTTGATGCCAACTC AGTGGAGGACCATTGCTCCAATCATTGGAAGAACAGCGGCCCAGTGCTTAGAACACTATGAATTTCTTCTGGATAAAGCTGCCCAAAGAG ACAATGAAGAGGAAACAACAGATGATCCACGAAAACTTAAACCTGGAGAAATAGATCCAAATCCAGAAACAAAACCAGCGCGGCCTGATC CAATTGATATGGATGAGGATGAACTTGAGATGCTTTCTGAAGCCAGAGCCCGCTTGGCTAATACTCAGGGAAAGAAGGCCAAGAGGAAAG CAAGAGAGAAACAATTGGAAGAAGCAAGACGTCTTGCTGCCCTCCAAAAAAGAAGAGAACTTCGAGCAGCTGGCATAGAAATTCAGAAGA AAAGAAAAAGGAAGAGAGGAGTTGATTATAATGCCGAAATCCCATTTGAAAAAAAGCCTGCCCTTGGTTTTTATGATACTTCTGAGGAAA ACTACCAAGCTCTTGACGCAGATTTCAGGAAATTAAGACAACAGGATCTTGATGGGGAGCTAAGATCTGAAAAAGAAGGAAGAGATAGAA AAAAAGACAAACAGCATTTGAAAAGGAAAAAAGAATCTGATTTACCATCAGCTATTCTTCAAACTAGTGGTGTTTCTGAATTTACTAAAA AGAGAAGCAAACTAGTACTTCCTGCCCCTCAGATTTCAGATGCAGAACTCCAGGAAGTTGTAAAAGTAGGCCAAGCGAGTGAAATTGCAC GTCAAACTGCCGAGGAATCTGGCATAACAAATTCTGCTTCCAGTACACTTTTGTCTGAGTACAATGTCACCAACAACAGCGTTGCTCTTA GAACACCACGAACACCAGCTTCCCAGGACAGAATTCTGCAGATGCCTGAGGAAGTGCACCATGGAGAGGAGGAGGTGGAGACTTTTGCCT TTCAGGCAGAAATTGCCCAACTCATGTCCCTCATCATCAATACCTTCTATTCCAACAAGGAGATTTTCCTTCGGGAGTTGATCTCTAATG CTTCTGATGCCTTGGACAAGATTCGCTATGAGAGCCTGACAGACCCTTCGAAGTTGGACAGTGGTAAAGAGCTGAAAATTGACATCATCC CCAACCCTCAGGAACGTACCCTGACTTTGGTAGACACAGGCATTGGCATGACCAAAGCTGATCTCATAAATAATTTGGGAACCATTGCCA AGTCTGGTACTAAAGCATTCATGGAGGCTCTTCAGGCTGGTGCAGACATCTCCATGATTGGGCAGTTTGGTGTTGGCTTTTATTCTGCCT ACTTGGTGGCAGAGAAAGTGGTTGTGATCACAAAGCACAACGATGATGAACAGTATGCTTGGGAGTCTTCTGCTGGAGGTTCCTTCACTG TGCGTGCTGACCATGGTGAGCCCATTGGCAGGGGTACCAAAGTGATCCTCCATCTTAAAGAAGATCAGACAGAGTACCTAGAAGAGAGGC GGGTCAAAGAAGTAGTGAAGAAGCATTCTCAGTTCATAGGCTATCCCATCACCCTTTATTTGGAGAAGGAACGAGAGAAGGAAATTAGTG ATGATGAGGCAGAGGAAGAGAAAGGTGAGAAAGAAGAGGAAGATAAAGATGATGAAGAAAAACCCAAGATCGAAGATGTGGGTTCAGATG AGGAGGATGACAGCGGTAAGGATAAGAAGAAGAAAACTAAGAAGATCAAAGAGAAATACATTGATCAGGAAGAACTAAACAAGACCAAGC CTATTTGGACCAGAAACCCTGATGACATCACCCAAGAGGAGTATGGAGAATTCTACAAGAGCCTCACTAATGACTGGGAAGACCACTTGG CAGTCAAGCACTTTTCTGTAGAAGGTCAGTTGGAATTCAGGGCATTGCTATTTATTCCTCGTCGGGCTCCCTTTGACCTTTTTGAGAACA AGAAGAAAAAGAACAACATCAAACTCTATGTCCGCCGTGTGTTCATCATGGACAGCTGTGATGAGTTGATACCAGAGTATCTCAATTTTA TCCGTGGTGTGGTTGACTCTGAGGATCTGCCCCTGAACATCTCCCGAGAAATGCTCCAGCAGAGCAAAATCTTGAAAGTCATTCGCAAAA ACATTGTTAAGAAGTGCCTTGAGCTCTTCTCTGAGCTGGCAGAAGACAAGGAGAATTACAAGAAATTCTATGAGGCATTCTCTAAAAATC TCAAGCTTGGAATCCACGAAGACTCCACTAACCGCCGCCGCCTGTCTGAGCTGCTGCGCTATCATACCTCCCAGTCTGGAGATGAGATGA CATCTCTGTCAGAGTATGTTTCTCGCATGAAGGAGACACAGAAGTCCATCTATTACATCACTGGTGAGAGCAAAGAGCAGGTGGCCAACT CAGCTTTTGTGGAGCGAGTGCGGAAACGGGGCTTCGAGGTGGTATATATGACCGAGCCCATTGACGAGTACTGTGTGCAGCAGCTCAAGG AATTTGATGGGAAGAGCCTGGTCTCAGTTACCAAGGAGGGTCTGGAGCTGCCTGAGGATGAGGAGGAGAAGAAGAAGATGGAAGAGAGCA AGGCAAAGTTTGAGAACCTCTGCAAGCTCATGAAAGAAATCTTAGATAAGAAGGTTGAGAAGGTGACAATCTCCAATAGACTTGTGTCTT CACCTTGCTGCATTGTGACCAGCACCTACGGCTGGACAGCCAATATGGAGCGGATCATGAAAGCCCAGGCACTTCGGGACAACTCCACCA TGGGCTATATGATGGCCAAAAAGCACCTGGAGATCAACCCTGACCACCCCATTGTGGAGACGCTGCGGCAGAAGGCTGAGGCCGACAAGA ATGATAAGGCAGTTAAGGACCTGGTGGTGCTGCTGTTTGAAACCGCCCTGCTATCTTCTGGCTTTTCCCTTGAGGATCCCCAGACCCACT CCAACCGCATCTATCGCATGATCAAGCTAGGTCTAGGTATTGATGAAGATGAAGTGGCAGCAGAGGAACCCAATGCTGCAGTTCCTGATG AGATCCCCCCTCTCGAGGGCGATGAGGATGCGTCTCGCATGGAAGAAGTCGATTAGGTTAGGAGTTCATAGTTGGAAAACTTGTGCCCTT GTATAGTGTCCCCATGGGCTCCCACTGCAGCCTCGAGTGCCCCTGTCCCACCTGGCTCCCCCTGCTGGTGTCTAGTGTTTTTTTCCCTCT CCTGTCCTTGTGTTGAAGGCAGTAAACTAAGGGTGTCAAGCCCCATTCCCTCTCTACTCTTGACAGCAGGATTGGATGTTGTGTATTGTG >14918_14918_2_CDC5L-HSP90AB1_CDC5L_chr6_44376369_ENST00000371477_HSP90AB1_chr6_44216367_ENST00000371646_length(amino acids)=1096AA_BP=372 MPRHPAAKMPRIMIKGGVWRNTEDEILKAAVMKYGKNQWSRIASLLHRKSAKQCKARWYEWLDPSIKKTEWSREEEEKLLHLAKLMPTQW RTIAPIIGRTAAQCLEHYEFLLDKAAQRDNEEETTDDPRKLKPGEIDPNPETKPARPDPIDMDEDELEMLSEARARLANTQGKKAKRKAR EKQLEEARRLAALQKRRELRAAGIEIQKKRKRKRGVDYNAEIPFEKKPALGFYDTSEENYQALDADFRKLRQQDLDGELRSEKEGRDRKK DKQHLKRKKESDLPSAILQTSGVSEFTKKRSKLVLPAPQISDAELQEVVKVGQASEIARQTAEESGITNSASSTLLSEYNVTNNSVALRT PRTPASQDRILQMPEEVHHGEEEVETFAFQAEIAQLMSLIINTFYSNKEIFLRELISNASDALDKIRYESLTDPSKLDSGKELKIDIIPN PQERTLTLVDTGIGMTKADLINNLGTIAKSGTKAFMEALQAGADISMIGQFGVGFYSAYLVAEKVVVITKHNDDEQYAWESSAGGSFTVR ADHGEPIGRGTKVILHLKEDQTEYLEERRVKEVVKKHSQFIGYPITLYLEKEREKEISDDEAEEEKGEKEEEDKDDEEKPKIEDVGSDEE DDSGKDKKKKTKKIKEKYIDQEELNKTKPIWTRNPDDITQEEYGEFYKSLTNDWEDHLAVKHFSVEGQLEFRALLFIPRRAPFDLFENKK KKNNIKLYVRRVFIMDSCDELIPEYLNFIRGVVDSEDLPLNISREMLQQSKILKVIRKNIVKKCLELFSELAEDKENYKKFYEAFSKNLK LGIHEDSTNRRRLSELLRYHTSQSGDEMTSLSEYVSRMKETQKSIYYITGESKEQVANSAFVERVRKRGFEVVYMTEPIDEYCVQQLKEF DGKSLVSVTKEGLELPEDEEEKKKMEESKAKFENLCKLMKEILDKKVEKVTISNRLVSSPCCIVTSTYGWTANMERIMKAQALRDNSTMG YMMAKKHLEINPDHPIVETLRQKAEADKNDKAVKDLVVLLFETALLSSGFSLEDPQTHSNRIYRMIKLGLGIDEDEVAAEEPNAAVPDEI -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CDC5L-HSP90AB1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000353801 | 0 | 12 | 215_552 | 0 | 725.0 | AHSA1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371554 | 0 | 12 | 215_552 | 0.0 | 725.0 | AHSA1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371646 | 0 | 12 | 215_552 | 0.0 | 725.0 | AHSA1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000353801 | 0 | 12 | 2_214 | 0 | 725.0 | BIRC2 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371554 | 0 | 12 | 2_214 | 0.0 | 725.0 | BIRC2 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371646 | 0 | 12 | 2_214 | 0.0 | 725.0 | BIRC2 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000353801 | 0 | 12 | 620_723 | 0 | 725.0 | NR1D1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371554 | 0 | 12 | 620_723 | 0.0 | 725.0 | NR1D1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371646 | 0 | 12 | 620_723 | 0.0 | 725.0 | NR1D1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000353801 | 0 | 12 | 264_608 | 0 | 725.0 | NR3C1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000353801 | 0 | 12 | 9_231 | 0 | 725.0 | NR3C1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371554 | 0 | 12 | 264_608 | 0.0 | 725.0 | NR3C1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371554 | 0 | 12 | 9_231 | 0.0 | 725.0 | NR3C1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371646 | 0 | 12 | 264_608 | 0.0 | 725.0 | NR3C1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371646 | 0 | 12 | 9_231 | 0.0 | 725.0 | NR3C1 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000353801 | 0 | 12 | 2_527 | 0 | 725.0 | TP53 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371554 | 0 | 12 | 2_527 | 0.0 | 725.0 | TP53 | |
| Tgene | HSP90AB1 | chr6:44376369 | chr6:44216367 | ENST00000371646 | 0 | 12 | 2_527 | 0.0 | 725.0 | TP53 |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 501_659 | 364.0 | 803.0 | DAPK3 |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 706_800 | 364.0 | 803.0 | PLRG1 |
| Hgene | CDC5L | chr6:44376369 | chr6:44216367 | ENST00000371477 | + | 8 | 16 | 260_606 | 364.0 | 803.0 | PPP1R8 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CDC5L-HSP90AB1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CDC5L-HSP90AB1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
