
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CHRDL2-COPB1 (FusionGDB2 ID:16641) |
Fusion Gene Summary for CHRDL2-COPB1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CHRDL2-COPB1 | Fusion gene ID: 16641 | Hgene | Tgene | Gene symbol | CHRDL2 | COPB1 | Gene ID | 25884 | 1315 |
| Gene name | chordin like 2 | COPI coat complex subunit beta 1 | |
| Synonyms | BNF1|CHL2 | COPB | |
| Cytomap | 11q13.4 | 11p15.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | chordin-like protein 2breast tumor novel factor 1chordin-related protein 2 | coatomer subunit betabeta coat proteinbeta-copcoatomer protein complex subunit beta 1 | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | Q6WN34 | P53618 | |
| Ensembl transtripts involved in fusion gene | ENST00000263671, ENST00000376332, ENST00000534159, | ENST00000526191, ENST00000249923, ENST00000439561, | |
| Fusion gene scores | * DoF score | 2 X 2 X 2=8 | 7 X 8 X 2=112 |
| # samples | 2 | 8 | |
| ** MAII score | log2(2/8*10)=1.32192809488736 | log2(8/112*10)=-0.485426827170242 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CHRDL2 [Title/Abstract] AND COPB1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CHRDL2(74429765)-COPB1(14491109), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across CHRDL2 (5'-gene) Fusion gene breakpoints across CHRDL2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
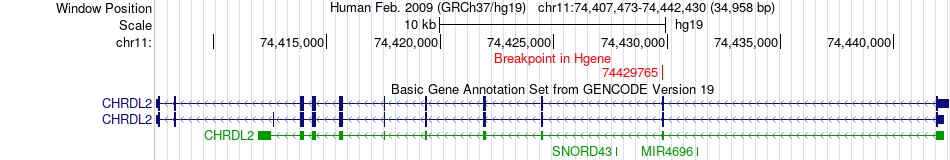 |
 Fusion gene breakpoints across COPB1 (3'-gene) Fusion gene breakpoints across COPB1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
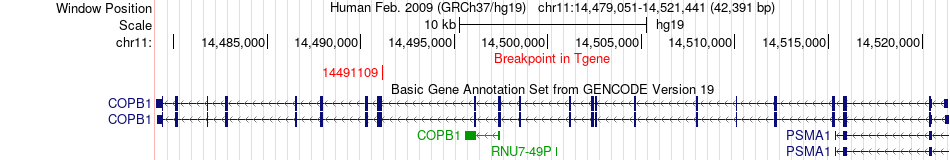 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-E2-A1IU-01A | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
Top |
Fusion Gene ORF analysis for CHRDL2-COPB1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000263671 | ENST00000526191 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
| 5CDS-intron | ENST00000376332 | ENST00000526191 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
| 5UTR-3CDS | ENST00000534159 | ENST00000249923 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
| 5UTR-3CDS | ENST00000534159 | ENST00000439561 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
| 5UTR-intron | ENST00000534159 | ENST00000526191 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
| In-frame | ENST00000263671 | ENST00000249923 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
| In-frame | ENST00000263671 | ENST00000439561 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
| In-frame | ENST00000376332 | ENST00000249923 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
| In-frame | ENST00000376332 | ENST00000439561 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000263671 | CHRDL2 | chr11 | 74429765 | - | ENST00000249923 | COPB1 | chr11 | 14491109 | - | 1925 | 482 | 8 | 1606 | 532 |
| ENST00000263671 | CHRDL2 | chr11 | 74429765 | - | ENST00000439561 | COPB1 | chr11 | 14491109 | - | 1881 | 482 | 8 | 1606 | 532 |
| ENST00000376332 | CHRDL2 | chr11 | 74429765 | - | ENST00000249923 | COPB1 | chr11 | 14491109 | - | 2135 | 692 | 149 | 1816 | 555 |
| ENST00000376332 | CHRDL2 | chr11 | 74429765 | - | ENST00000439561 | COPB1 | chr11 | 14491109 | - | 2091 | 692 | 149 | 1816 | 555 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000263671 | ENST00000249923 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - | 0.000355508 | 0.9996445 |
| ENST00000263671 | ENST00000439561 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - | 0.00042853 | 0.9995715 |
| ENST00000376332 | ENST00000249923 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - | 0.000554317 | 0.9994456 |
| ENST00000376332 | ENST00000439561 | CHRDL2 | chr11 | 74429765 | - | COPB1 | chr11 | 14491109 | - | 0.000663581 | 0.9993364 |
Top |
Fusion Genomic Features for CHRDL2-COPB1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
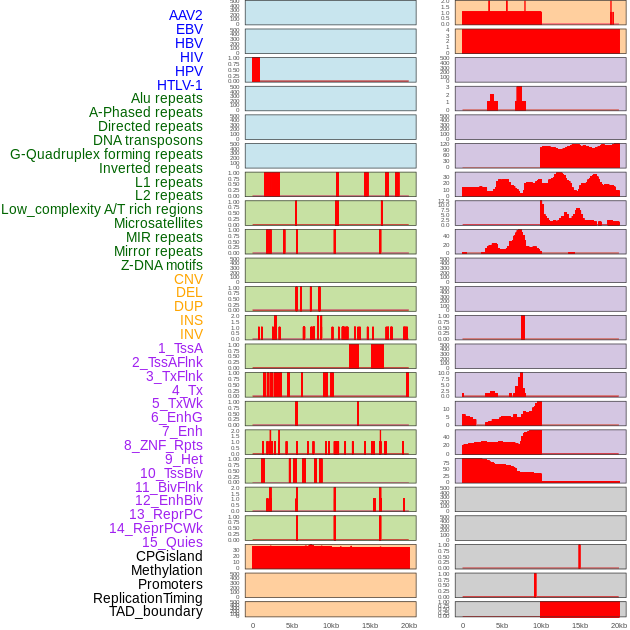 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for CHRDL2-COPB1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr11:74429765/chr11:14491109) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CHRDL2 | COPB1 |
| FUNCTION: May inhibit BMPs activity by blocking their interaction with their receptors. Has a negative regulator effect on the cartilage formation/regeneration from immature mesenchymal cells, by preventing or reducing the rate of matrix accumulation (By similarity). Implicated in tumor angiogenesis. May play a role during myoblast and osteoblast differentiation, and maturation. {ECO:0000250, ECO:0000269|PubMed:12853144, ECO:0000269|PubMed:15094188}. | FUNCTION: The coatomer is a cytosolic protein complex that binds to dilysine motifs and reversibly associates with Golgi non-clathrin-coated vesicles, which further mediate biosynthetic protein transport from the ER, via the Golgi up to the trans Golgi network. Coatomer complex is required for budding from Golgi membranes, and is essential for the retrograde Golgi-to-ER transport of dilysine-tagged proteins. In mammals, the coatomer can only be recruited by membranes associated to ADP-ribosylation factors (ARFs), which are small GTP-binding proteins; the complex also influences the Golgi structural integrity, as well as the processing, activity, and endocytic recycling of LDL receptors. Plays a functional role in facilitating the transport of kappa-type opioid receptor mRNAs into axons and enhances translation of these proteins. Required for limiting lipid storage in lipid droplets. Involved in lipid homeostasis by regulating the presence of perilipin family members PLIN2 and PLIN3 at the lipid droplet surface and promoting the association of adipocyte surface triglyceride lipase (PNPLA2) with the lipid droplet to mediate lipolysis (By similarity). Involved in the Golgi disassembly and reassembly processes during cell cycle. Involved in autophagy by playing a role in early endosome function. Plays a role in organellar compartmentalization of secretory compartments including endoplasmic reticulum (ER)-Golgi intermediate compartment (ERGIC), Golgi, trans-Golgi network (TGN) and recycling endosomes, and in biosynthetic transport of CAV1. Promotes degradation of Nef cellular targets CD4 and MHC class I antigens by facilitating their trafficking to degradative compartments. {ECO:0000250, ECO:0000269|PubMed:18385291, ECO:0000269|PubMed:18725938, ECO:0000269|PubMed:19364919, ECO:0000269|PubMed:20056612}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CHRDL2 | chr11:74429765 | chr11:14491109 | ENST00000263671 | - | 2 | 12 | 109_175 | 65 | 452.0 | Domain | VWFC 2 |
| Hgene | CHRDL2 | chr11:74429765 | chr11:14491109 | ENST00000263671 | - | 2 | 12 | 250_315 | 65 | 452.0 | Domain | VWFC 3 |
| Hgene | CHRDL2 | chr11:74429765 | chr11:14491109 | ENST00000263671 | - | 2 | 12 | 31_96 | 65 | 452.0 | Domain | VWFC 1 |
| Hgene | CHRDL2 | chr11:74429765 | chr11:14491109 | ENST00000376332 | - | 2 | 11 | 109_175 | 65 | 430.0 | Domain | VWFC 2 |
| Hgene | CHRDL2 | chr11:74429765 | chr11:14491109 | ENST00000376332 | - | 2 | 11 | 250_315 | 65 | 430.0 | Domain | VWFC 3 |
| Hgene | CHRDL2 | chr11:74429765 | chr11:14491109 | ENST00000376332 | - | 2 | 11 | 31_96 | 65 | 430.0 | Domain | VWFC 1 |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000249923 | 13 | 22 | 132_168 | 579 | 954.0 | Repeat | Note=HEAT 2 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000249923 | 13 | 22 | 240_276 | 579 | 954.0 | Repeat | Note=HEAT 3 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000249923 | 13 | 22 | 277_314 | 579 | 954.0 | Repeat | Note=HEAT 4 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000249923 | 13 | 22 | 316_353 | 579 | 954.0 | Repeat | Note=HEAT 5 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000249923 | 13 | 22 | 396_433 | 579 | 954.0 | Repeat | Note=HEAT 6 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000249923 | 13 | 22 | 96_131 | 579 | 954.0 | Repeat | Note=HEAT 1 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000439561 | 13 | 22 | 132_168 | 579 | 954.0 | Repeat | Note=HEAT 2 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000439561 | 13 | 22 | 240_276 | 579 | 954.0 | Repeat | Note=HEAT 3 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000439561 | 13 | 22 | 277_314 | 579 | 954.0 | Repeat | Note=HEAT 4 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000439561 | 13 | 22 | 316_353 | 579 | 954.0 | Repeat | Note=HEAT 5 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000439561 | 13 | 22 | 396_433 | 579 | 954.0 | Repeat | Note=HEAT 6 | |
| Tgene | COPB1 | chr11:74429765 | chr11:14491109 | ENST00000439561 | 13 | 22 | 96_131 | 579 | 954.0 | Repeat | Note=HEAT 1 |
Top |
Fusion Gene Sequence for CHRDL2-COPB1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >16641_16641_1_CHRDL2-COPB1_CHRDL2_chr11_74429765_ENST00000263671_COPB1_chr11_14491109_ENST00000249923_length(transcript)=1925nt_BP=482nt CCCTTTCTTTGATCGCCTCTCCCTTCTGCTGGACCTTCCTTCGTCTCTCCATCTCTCCCTCCTTTCCCCGCGTTCTCTTTCCACCTTTCT CTTCTTCCCACCTTAGACCTCCCTTCCTGCCCTCCTTTCCTGCCCACCGCTGCTTCCTGGCCCTTCTCCGACCCCGCTCTAGCAGCAGAC CTCCTGGGGTCTGTGGGTTGATCTGTGGCCCCTGTGCCTCCGTGTCCTTTTCGTCTCCCTTCCTCCCGACTCCGCTCCCGGACCAGCGGC CTGACCCTGGGGAAAGGATGGTTCCCGAGGTGAGGGTCCTCTCCTCCTTGCTGGGACTCGCGCTGCTCTGGTTCCCCCTGGACTCCCACG CTCGAGCCCGCCCAGACATGTTCTGCCTTTTCCATGGGAAGAGATACTCCCCCGGCGAGAGCTGGCACCCCTACTTGGAGCCACAAGGCC TGATGTACTGCCTGCGCTGTACCTGCTCAGAGTCTTTTGTTGCTGAGGCTATGTTGCTCATGGCTACTATCCTGCATTTGGGAAAATCCT CTCTTCCTAAGAAGCCAATTACTGATGATGATGTGGATCGAATTTCCCTGTGCCTCAAGGTCTTGTCTGAATGTTCACCTTTAATGAATG ACATTTTCAATAAGGAATGCAGACAGTCCCTTTCTCACATGTTATCTGCTAAACTAGAAGAAGAGAAATTATCCCAAAAGAAAGAATCTG AAAAGAGGAATGTGACAGTACAGCCTGATGACCCCATTTCCTTCATGCAACTAACTGCTAAGAATGAAATGAACTGCAAGGAAGATCAGT TTCAGCTGAGTTTACTGGCAGCAATGGGTAACACACAGAGGAAAGAGGCAGCAGATCCCCTAGCATCTAAACTTAACAAGGTCACCCAAT TGACAGGTTTCTCAGATCCTGTATATGCAGAAGCTTACGTTCATGTCAACCAATATGATATTGTCCTGGATGTACTTGTTGTGAACCAAA CCAGTGATACTTTGCAGAATTGCACATTAGAACTAGCTACACTAGGGGATCTGAAACTTGTGGAAAAGCCGTCTCCTTTGACTCTTGCTC CTCATGACTTCGCAAATATTAAAGCTAACGTCAAAGTAGCATCAACAGAAAATGGAATAATTTTTGGTAATATAGTTTATGATGTCTCTG GAGCAGCAAGTGACAGAAATTGTGTGGTTCTCAGTGATATTCACATCGACATCATGGACTATATCCAGCCTGCAACTTGCACTGATGCAG AATTCCGTCAGATGTGGGCCGAATTTGAATGGGAAAACAAAGTGACAGTTAACACCAACATGGTTGATTTAAATGACTACTTACAGCACA TATTAAAGTCAACCAATATGAAATGCCTGACTCCAGAAAAGGCCCTTTCTGGTTACTGTGGCTTTATGGCAGCCAACCTTTATGCTCGTT CCATATTTGGTGAAGATGCACTTGCAAATGTCAGCATTGAGAAGCCAATTCACCAGGGACCAGATGCTGCTGTTACCGGCCATATAAGAA TTCGTGCAAAGAGCCAGGGAATGGCCTTAAGTCTTGGAGATAAAATCAACTTGTCACAGAAGAAAACTAGTATATAAAAATAAACAAAAA GTCCTTGAAGCTTTACAGTTAATTTAGGTATGGGCTTACTGGACTCCAACATCTTTTGTACTCTTTCATGCTTATATAGAATCTGAGTTC ATGCTGAATACTTTTCAGCCAATAATTTATAGCCTTTCCCTTAAATCAAGATTGAGTTTAAAATTATAGTTTGTCTTTTGTCTTAACAGT TCTGAATGCTGTCCTCAAAGTATATAATGTTTCATGTACCAAGACCCTTTTCACAGTACAATAAACAGATCTATTCATAAATTTTTGTTA >16641_16641_1_CHRDL2-COPB1_CHRDL2_chr11_74429765_ENST00000263671_COPB1_chr11_14491109_ENST00000249923_length(amino acids)=532AA_BP=156 MIASPFCWTFLRLSISPSFPRVLFPPFSSSHLRPPFLPSFPAHRCFLALLRPRSSSRPPGVCGLICGPCASVSFSSPFLPTPLPDQRPDP GERMVPEVRVLSSLLGLALLWFPLDSHARARPDMFCLFHGKRYSPGESWHPYLEPQGLMYCLRCTCSESFVAEAMLLMATILHLGKSSLP KKPITDDDVDRISLCLKVLSECSPLMNDIFNKECRQSLSHMLSAKLEEEKLSQKKESEKRNVTVQPDDPISFMQLTAKNEMNCKEDQFQL SLLAAMGNTQRKEAADPLASKLNKVTQLTGFSDPVYAEAYVHVNQYDIVLDVLVVNQTSDTLQNCTLELATLGDLKLVEKPSPLTLAPHD FANIKANVKVASTENGIIFGNIVYDVSGAASDRNCVVLSDIHIDIMDYIQPATCTDAEFRQMWAEFEWENKVTVNTNMVDLNDYLQHILK -------------------------------------------------------------- >16641_16641_2_CHRDL2-COPB1_CHRDL2_chr11_74429765_ENST00000263671_COPB1_chr11_14491109_ENST00000439561_length(transcript)=1881nt_BP=482nt CCCTTTCTTTGATCGCCTCTCCCTTCTGCTGGACCTTCCTTCGTCTCTCCATCTCTCCCTCCTTTCCCCGCGTTCTCTTTCCACCTTTCT CTTCTTCCCACCTTAGACCTCCCTTCCTGCCCTCCTTTCCTGCCCACCGCTGCTTCCTGGCCCTTCTCCGACCCCGCTCTAGCAGCAGAC CTCCTGGGGTCTGTGGGTTGATCTGTGGCCCCTGTGCCTCCGTGTCCTTTTCGTCTCCCTTCCTCCCGACTCCGCTCCCGGACCAGCGGC CTGACCCTGGGGAAAGGATGGTTCCCGAGGTGAGGGTCCTCTCCTCCTTGCTGGGACTCGCGCTGCTCTGGTTCCCCCTGGACTCCCACG CTCGAGCCCGCCCAGACATGTTCTGCCTTTTCCATGGGAAGAGATACTCCCCCGGCGAGAGCTGGCACCCCTACTTGGAGCCACAAGGCC TGATGTACTGCCTGCGCTGTACCTGCTCAGAGTCTTTTGTTGCTGAGGCTATGTTGCTCATGGCTACTATCCTGCATTTGGGAAAATCCT CTCTTCCTAAGAAGCCAATTACTGATGATGATGTGGATCGAATTTCCCTGTGCCTCAAGGTCTTGTCTGAATGTTCACCTTTAATGAATG ACATTTTCAATAAGGAATGCAGACAGTCCCTTTCTCACATGTTATCTGCTAAACTAGAAGAAGAGAAATTATCCCAAAAGAAAGAATCTG AAAAGAGGAATGTGACAGTACAGCCTGATGACCCCATTTCCTTCATGCAACTAACTGCTAAGAATGAAATGAACTGCAAGGAAGATCAGT TTCAGCTGAGTTTACTGGCAGCAATGGGTAACACACAGAGGAAAGAGGCAGCAGATCCCCTAGCATCTAAACTTAACAAGGTCACCCAAT TGACAGGTTTCTCAGATCCTGTATATGCAGAAGCTTACGTTCATGTCAACCAATATGATATTGTCCTGGATGTACTTGTTGTGAACCAAA CCAGTGATACTTTGCAGAATTGCACATTAGAACTAGCTACACTAGGGGATCTGAAACTTGTGGAAAAGCCGTCTCCTTTGACTCTTGCTC CTCATGACTTCGCAAATATTAAAGCTAACGTCAAAGTAGCATCAACAGAAAATGGAATAATTTTTGGTAATATAGTTTATGATGTCTCTG GAGCAGCAAGTGACAGAAATTGTGTGGTTCTCAGTGATATTCACATCGACATCATGGACTATATCCAGCCTGCAACTTGCACTGATGCAG AATTCCGTCAGATGTGGGCCGAATTTGAATGGGAAAACAAAGTGACAGTTAACACCAACATGGTTGATTTAAATGACTACTTACAGCACA TATTAAAGTCAACCAATATGAAATGCCTGACTCCAGAAAAGGCCCTTTCTGGTTACTGTGGCTTTATGGCAGCCAACCTTTATGCTCGTT CCATATTTGGTGAAGATGCACTTGCAAATGTCAGCATTGAGAAGCCAATTCACCAGGGACCAGATGCTGCTGTTACCGGCCATATAAGAA TTCGTGCAAAGAGCCAGGGAATGGCCTTAAGTCTTGGAGATAAAATCAACTTGTCACAGAAGAAAACTAGTATATAAAAATAAACAAAAA GTCCTTGAAGCTTTACAGTTAATTTAGGTATGGGCTTACTGGACTCCAACATCTTTTGTACTCTTTCATGCTTATATAGAATCTGAGTTC ATGCTGAATACTTTTCAGCCAATAATTTATAGCCTTTCCCTTAAATCAAGATTGAGTTTAAAATTATAGTTTGTCTTTTGTCTTAACAGT >16641_16641_2_CHRDL2-COPB1_CHRDL2_chr11_74429765_ENST00000263671_COPB1_chr11_14491109_ENST00000439561_length(amino acids)=532AA_BP=156 MIASPFCWTFLRLSISPSFPRVLFPPFSSSHLRPPFLPSFPAHRCFLALLRPRSSSRPPGVCGLICGPCASVSFSSPFLPTPLPDQRPDP GERMVPEVRVLSSLLGLALLWFPLDSHARARPDMFCLFHGKRYSPGESWHPYLEPQGLMYCLRCTCSESFVAEAMLLMATILHLGKSSLP KKPITDDDVDRISLCLKVLSECSPLMNDIFNKECRQSLSHMLSAKLEEEKLSQKKESEKRNVTVQPDDPISFMQLTAKNEMNCKEDQFQL SLLAAMGNTQRKEAADPLASKLNKVTQLTGFSDPVYAEAYVHVNQYDIVLDVLVVNQTSDTLQNCTLELATLGDLKLVEKPSPLTLAPHD FANIKANVKVASTENGIIFGNIVYDVSGAASDRNCVVLSDIHIDIMDYIQPATCTDAEFRQMWAEFEWENKVTVNTNMVDLNDYLQHILK -------------------------------------------------------------- >16641_16641_3_CHRDL2-COPB1_CHRDL2_chr11_74429765_ENST00000376332_COPB1_chr11_14491109_ENST00000249923_length(transcript)=2135nt_BP=692nt GCCGCCGCGCCGCCGCCGCTGCTGGGACCGCCGCCCCGCCGGGCCCCAGGCCCCCGCGGCCCGACCCCGAGCCCACCGAGCAGGGGGCGC CGGCCGCCTCTAGCCCGCGCGCCCGCAACACCCGCTGGTCCCGCGCGCCTCGCCCGGCGCTGCGCGCTCGGCAGACGCCGGTCCTCTCCC TGCTTTCTCCTCCTCTCCCTCTCACCTTCCCCCTTTCTTTGATCGCCTCTCCCTTCTGCTGGACCTTCCTTCGTCTCTCCATCTCTCCCT CCTTTCCCCGCGTTCTCTTTCCACCTTTCTCTTCTTCCCACCTTAGACCTCCCTTCCTGCCCTCCTTTCCTGCCCACCGCTGCTTCCTGG CCCTTCTCCGACCCCGCTCTAGCAGCAGACCTCCTGGGGTCTGTGGGTTGATCTGTGGCCCCTGTGCCTCCGTGTCCTTTTCGTCTCCCT TCCTCCCGACTCCGCTCCCGGACCAGCGGCCTGACCCTGGGGAAAGGATGGTTCCCGAGGTGAGGGTCCTCTCCTCCTTGCTGGGACTCG CGCTGCTCTGGTTCCCCCTGGACTCCCACGCTCGAGCCCGCCCAGACATGTTCTGCCTTTTCCATGGGAAGAGATACTCCCCCGGCGAGA GCTGGCACCCCTACTTGGAGCCACAAGGCCTGATGTACTGCCTGCGCTGTACCTGCTCAGAGTCTTTTGTTGCTGAGGCTATGTTGCTCA TGGCTACTATCCTGCATTTGGGAAAATCCTCTCTTCCTAAGAAGCCAATTACTGATGATGATGTGGATCGAATTTCCCTGTGCCTCAAGG TCTTGTCTGAATGTTCACCTTTAATGAATGACATTTTCAATAAGGAATGCAGACAGTCCCTTTCTCACATGTTATCTGCTAAACTAGAAG AAGAGAAATTATCCCAAAAGAAAGAATCTGAAAAGAGGAATGTGACAGTACAGCCTGATGACCCCATTTCCTTCATGCAACTAACTGCTA AGAATGAAATGAACTGCAAGGAAGATCAGTTTCAGCTGAGTTTACTGGCAGCAATGGGTAACACACAGAGGAAAGAGGCAGCAGATCCCC TAGCATCTAAACTTAACAAGGTCACCCAATTGACAGGTTTCTCAGATCCTGTATATGCAGAAGCTTACGTTCATGTCAACCAATATGATA TTGTCCTGGATGTACTTGTTGTGAACCAAACCAGTGATACTTTGCAGAATTGCACATTAGAACTAGCTACACTAGGGGATCTGAAACTTG TGGAAAAGCCGTCTCCTTTGACTCTTGCTCCTCATGACTTCGCAAATATTAAAGCTAACGTCAAAGTAGCATCAACAGAAAATGGAATAA TTTTTGGTAATATAGTTTATGATGTCTCTGGAGCAGCAAGTGACAGAAATTGTGTGGTTCTCAGTGATATTCACATCGACATCATGGACT ATATCCAGCCTGCAACTTGCACTGATGCAGAATTCCGTCAGATGTGGGCCGAATTTGAATGGGAAAACAAAGTGACAGTTAACACCAACA TGGTTGATTTAAATGACTACTTACAGCACATATTAAAGTCAACCAATATGAAATGCCTGACTCCAGAAAAGGCCCTTTCTGGTTACTGTG GCTTTATGGCAGCCAACCTTTATGCTCGTTCCATATTTGGTGAAGATGCACTTGCAAATGTCAGCATTGAGAAGCCAATTCACCAGGGAC CAGATGCTGCTGTTACCGGCCATATAAGAATTCGTGCAAAGAGCCAGGGAATGGCCTTAAGTCTTGGAGATAAAATCAACTTGTCACAGA AGAAAACTAGTATATAAAAATAAACAAAAAGTCCTTGAAGCTTTACAGTTAATTTAGGTATGGGCTTACTGGACTCCAACATCTTTTGTA CTCTTTCATGCTTATATAGAATCTGAGTTCATGCTGAATACTTTTCAGCCAATAATTTATAGCCTTTCCCTTAAATCAAGATTGAGTTTA AAATTATAGTTTGTCTTTTGTCTTAACAGTTCTGAATGCTGTCCTCAAAGTATATAATGTTTCATGTACCAAGACCCTTTTCACAGTACA >16641_16641_3_CHRDL2-COPB1_CHRDL2_chr11_74429765_ENST00000376332_COPB1_chr11_14491109_ENST00000249923_length(amino acids)=555AA_BP=179 MRARQTPVLSLLSPPLPLTFPLSLIASPFCWTFLRLSISPSFPRVLFPPFSSSHLRPPFLPSFPAHRCFLALLRPRSSSRPPGVCGLICG PCASVSFSSPFLPTPLPDQRPDPGERMVPEVRVLSSLLGLALLWFPLDSHARARPDMFCLFHGKRYSPGESWHPYLEPQGLMYCLRCTCS ESFVAEAMLLMATILHLGKSSLPKKPITDDDVDRISLCLKVLSECSPLMNDIFNKECRQSLSHMLSAKLEEEKLSQKKESEKRNVTVQPD DPISFMQLTAKNEMNCKEDQFQLSLLAAMGNTQRKEAADPLASKLNKVTQLTGFSDPVYAEAYVHVNQYDIVLDVLVVNQTSDTLQNCTL ELATLGDLKLVEKPSPLTLAPHDFANIKANVKVASTENGIIFGNIVYDVSGAASDRNCVVLSDIHIDIMDYIQPATCTDAEFRQMWAEFE WENKVTVNTNMVDLNDYLQHILKSTNMKCLTPEKALSGYCGFMAANLYARSIFGEDALANVSIEKPIHQGPDAAVTGHIRIRAKSQGMAL -------------------------------------------------------------- >16641_16641_4_CHRDL2-COPB1_CHRDL2_chr11_74429765_ENST00000376332_COPB1_chr11_14491109_ENST00000439561_length(transcript)=2091nt_BP=692nt GCCGCCGCGCCGCCGCCGCTGCTGGGACCGCCGCCCCGCCGGGCCCCAGGCCCCCGCGGCCCGACCCCGAGCCCACCGAGCAGGGGGCGC CGGCCGCCTCTAGCCCGCGCGCCCGCAACACCCGCTGGTCCCGCGCGCCTCGCCCGGCGCTGCGCGCTCGGCAGACGCCGGTCCTCTCCC TGCTTTCTCCTCCTCTCCCTCTCACCTTCCCCCTTTCTTTGATCGCCTCTCCCTTCTGCTGGACCTTCCTTCGTCTCTCCATCTCTCCCT CCTTTCCCCGCGTTCTCTTTCCACCTTTCTCTTCTTCCCACCTTAGACCTCCCTTCCTGCCCTCCTTTCCTGCCCACCGCTGCTTCCTGG CCCTTCTCCGACCCCGCTCTAGCAGCAGACCTCCTGGGGTCTGTGGGTTGATCTGTGGCCCCTGTGCCTCCGTGTCCTTTTCGTCTCCCT TCCTCCCGACTCCGCTCCCGGACCAGCGGCCTGACCCTGGGGAAAGGATGGTTCCCGAGGTGAGGGTCCTCTCCTCCTTGCTGGGACTCG CGCTGCTCTGGTTCCCCCTGGACTCCCACGCTCGAGCCCGCCCAGACATGTTCTGCCTTTTCCATGGGAAGAGATACTCCCCCGGCGAGA GCTGGCACCCCTACTTGGAGCCACAAGGCCTGATGTACTGCCTGCGCTGTACCTGCTCAGAGTCTTTTGTTGCTGAGGCTATGTTGCTCA TGGCTACTATCCTGCATTTGGGAAAATCCTCTCTTCCTAAGAAGCCAATTACTGATGATGATGTGGATCGAATTTCCCTGTGCCTCAAGG TCTTGTCTGAATGTTCACCTTTAATGAATGACATTTTCAATAAGGAATGCAGACAGTCCCTTTCTCACATGTTATCTGCTAAACTAGAAG AAGAGAAATTATCCCAAAAGAAAGAATCTGAAAAGAGGAATGTGACAGTACAGCCTGATGACCCCATTTCCTTCATGCAACTAACTGCTA AGAATGAAATGAACTGCAAGGAAGATCAGTTTCAGCTGAGTTTACTGGCAGCAATGGGTAACACACAGAGGAAAGAGGCAGCAGATCCCC TAGCATCTAAACTTAACAAGGTCACCCAATTGACAGGTTTCTCAGATCCTGTATATGCAGAAGCTTACGTTCATGTCAACCAATATGATA TTGTCCTGGATGTACTTGTTGTGAACCAAACCAGTGATACTTTGCAGAATTGCACATTAGAACTAGCTACACTAGGGGATCTGAAACTTG TGGAAAAGCCGTCTCCTTTGACTCTTGCTCCTCATGACTTCGCAAATATTAAAGCTAACGTCAAAGTAGCATCAACAGAAAATGGAATAA TTTTTGGTAATATAGTTTATGATGTCTCTGGAGCAGCAAGTGACAGAAATTGTGTGGTTCTCAGTGATATTCACATCGACATCATGGACT ATATCCAGCCTGCAACTTGCACTGATGCAGAATTCCGTCAGATGTGGGCCGAATTTGAATGGGAAAACAAAGTGACAGTTAACACCAACA TGGTTGATTTAAATGACTACTTACAGCACATATTAAAGTCAACCAATATGAAATGCCTGACTCCAGAAAAGGCCCTTTCTGGTTACTGTG GCTTTATGGCAGCCAACCTTTATGCTCGTTCCATATTTGGTGAAGATGCACTTGCAAATGTCAGCATTGAGAAGCCAATTCACCAGGGAC CAGATGCTGCTGTTACCGGCCATATAAGAATTCGTGCAAAGAGCCAGGGAATGGCCTTAAGTCTTGGAGATAAAATCAACTTGTCACAGA AGAAAACTAGTATATAAAAATAAACAAAAAGTCCTTGAAGCTTTACAGTTAATTTAGGTATGGGCTTACTGGACTCCAACATCTTTTGTA CTCTTTCATGCTTATATAGAATCTGAGTTCATGCTGAATACTTTTCAGCCAATAATTTATAGCCTTTCCCTTAAATCAAGATTGAGTTTA AAATTATAGTTTGTCTTTTGTCTTAACAGTTCTGAATGCTGTCCTCAAAGTATATAATGTTTCATGTACCAAGACCCTTTTCACAGTACA >16641_16641_4_CHRDL2-COPB1_CHRDL2_chr11_74429765_ENST00000376332_COPB1_chr11_14491109_ENST00000439561_length(amino acids)=555AA_BP=179 MRARQTPVLSLLSPPLPLTFPLSLIASPFCWTFLRLSISPSFPRVLFPPFSSSHLRPPFLPSFPAHRCFLALLRPRSSSRPPGVCGLICG PCASVSFSSPFLPTPLPDQRPDPGERMVPEVRVLSSLLGLALLWFPLDSHARARPDMFCLFHGKRYSPGESWHPYLEPQGLMYCLRCTCS ESFVAEAMLLMATILHLGKSSLPKKPITDDDVDRISLCLKVLSECSPLMNDIFNKECRQSLSHMLSAKLEEEKLSQKKESEKRNVTVQPD DPISFMQLTAKNEMNCKEDQFQLSLLAAMGNTQRKEAADPLASKLNKVTQLTGFSDPVYAEAYVHVNQYDIVLDVLVVNQTSDTLQNCTL ELATLGDLKLVEKPSPLTLAPHDFANIKANVKVASTENGIIFGNIVYDVSGAASDRNCVVLSDIHIDIMDYIQPATCTDAEFRQMWAEFE WENKVTVNTNMVDLNDYLQHILKSTNMKCLTPEKALSGYCGFMAANLYARSIFGEDALANVSIEKPIHQGPDAAVTGHIRIRAKSQGMAL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CHRDL2-COPB1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CHRDL2-COPB1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CHRDL2-COPB1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
