
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CLEC16A-RMI2 (FusionGDB2 ID:17082) |
Fusion Gene Summary for CLEC16A-RMI2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CLEC16A-RMI2 | Fusion gene ID: 17082 | Hgene | Tgene | Gene symbol | CLEC16A | RMI2 | Gene ID | 23274 | 116028 |
| Gene name | C-type lectin domain containing 16A | RecQ mediated genome instability 2 | |
| Synonyms | Gop-1|KIAA0350 | BLAP18|C16orf75 | |
| Cytomap | 16p13.13 | 16p13.13 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein CLEC16AC-type lectin domain family 16 member A | recQ-mediated genome instability protein 2BLM-associated protein of 18 kDaRMI2, RecQ mediated genome instability 2, homolog | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q2KHT3 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000409552, ENST00000409790, ENST00000381822, ENST00000465491, | ENST00000312499, ENST00000381820, ENST00000572992, ENST00000576027, ENST00000572173, | |
| Fusion gene scores | * DoF score | 13 X 9 X 9=1053 | 6 X 2 X 4=48 |
| # samples | 16 | 6 | |
| ** MAII score | log2(16/1053*10)=-2.71836162613835 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(6/48*10)=0.321928094887362 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: CLEC16A [Title/Abstract] AND RMI2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | RMI2(11439623)-CLEC16A(11214472), # samples:3 CLEC16A(11220003)-RMI2(11444499), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across CLEC16A (5'-gene) Fusion gene breakpoints across CLEC16A (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
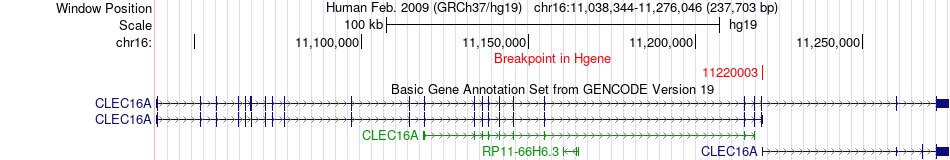 |
 Fusion gene breakpoints across RMI2 (3'-gene) Fusion gene breakpoints across RMI2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
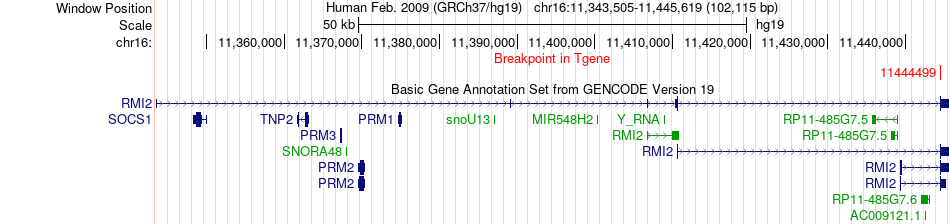 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | CESC | TCGA-FU-A40J-01A | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
Top |
Fusion Gene ORF analysis for CLEC16A-RMI2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000409552 | ENST00000312499 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5CDS-intron | ENST00000409552 | ENST00000381820 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5CDS-intron | ENST00000409552 | ENST00000572992 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5CDS-intron | ENST00000409552 | ENST00000576027 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5CDS-intron | ENST00000409790 | ENST00000312499 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5CDS-intron | ENST00000409790 | ENST00000381820 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5CDS-intron | ENST00000409790 | ENST00000572992 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5CDS-intron | ENST00000409790 | ENST00000576027 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5UTR-3CDS | ENST00000381822 | ENST00000572173 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5UTR-intron | ENST00000381822 | ENST00000312499 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5UTR-intron | ENST00000381822 | ENST00000381820 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5UTR-intron | ENST00000381822 | ENST00000572992 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| 5UTR-intron | ENST00000381822 | ENST00000576027 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| In-frame | ENST00000409552 | ENST00000572173 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| In-frame | ENST00000409790 | ENST00000572173 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| intron-3CDS | ENST00000465491 | ENST00000572173 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| intron-intron | ENST00000465491 | ENST00000312499 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| intron-intron | ENST00000465491 | ENST00000381820 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| intron-intron | ENST00000465491 | ENST00000572992 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
| intron-intron | ENST00000465491 | ENST00000576027 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000409790 | CLEC16A | chr16 | 11220003 | + | ENST00000572173 | RMI2 | chr16 | 11444499 | + | 3985 | 2871 | 230 | 3019 | 929 |
| ENST00000409552 | CLEC16A | chr16 | 11220003 | + | ENST00000572173 | RMI2 | chr16 | 11444499 | + | 3868 | 2754 | 137 | 2689 | 850 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000409790 | ENST00000572173 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + | 0.001653455 | 0.99834657 |
| ENST00000409552 | ENST00000572173 | CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444499 | + | 0.002155987 | 0.997844 |
Top |
Fusion Genomic Features for CLEC16A-RMI2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444498 | + | 4.90E-05 | 0.999951 |
| CLEC16A | chr16 | 11220003 | + | RMI2 | chr16 | 11444498 | + | 4.90E-05 | 0.999951 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
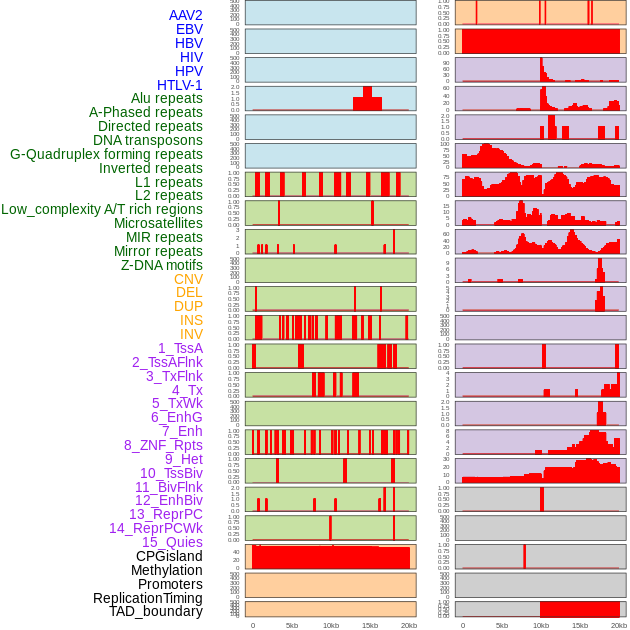 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
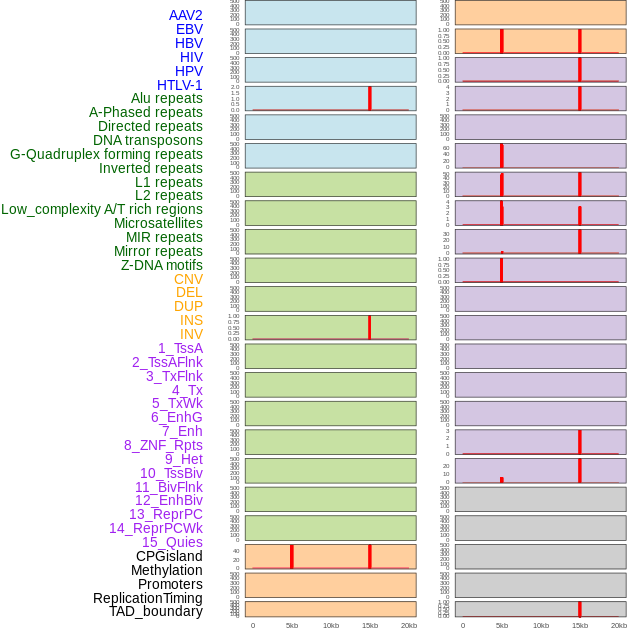 |
Top |
Fusion Protein Features for CLEC16A-RMI2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr16:11439623/chr16:11214472) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CLEC16A | . |
| FUNCTION: Regulator of mitophagy through the upstream regulation of the RNF41/NRDP1-PRKN pathway. Mitophagy is a selective form of autophagy necessary for mitochondrial quality control. The RNF41/NRDP1-PRKN pathway regulates autophagosome-lysosome fusion during late mitophagy. May protect RNF41/NRDP1 from proteosomal degradation, RNF41/NRDP1 which regulates proteosomal degradation of PRKN. Plays a key role in beta cells functions by regulating mitophagy/autophagy and mitochondrial health. {ECO:0000269|PubMed:24949970}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CLEC16A | chr16:11220003 | chr16:11444499 | ENST00000409790 | + | 22 | 24 | 51_198 | 880 | 1054.0 | Domain | FPL |
| Tgene | RMI2 | chr16:11220003 | chr16:11444499 | ENST00000381820 | 0 | 2 | 44_114 | 35 | 85.0 | DNA binding | Note=OB | |
| Tgene | RMI2 | chr16:11220003 | chr16:11444499 | ENST00000572173 | 3 | 5 | 44_114 | 35 | 85.0 | DNA binding | Note=OB |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CLEC16A | chr16:11220003 | chr16:11444499 | ENST00000381822 | + | 1 | 4 | 892_933 | 0 | 141.0 | Compositional bias | Note=Ser-rich |
| Hgene | CLEC16A | chr16:11220003 | chr16:11444499 | ENST00000409552 | + | 1 | 21 | 892_933 | 0 | 907.0 | Compositional bias | Note=Ser-rich |
| Hgene | CLEC16A | chr16:11220003 | chr16:11444499 | ENST00000409790 | + | 22 | 24 | 892_933 | 880 | 1054.0 | Compositional bias | Note=Ser-rich |
| Hgene | CLEC16A | chr16:11220003 | chr16:11444499 | ENST00000381822 | + | 1 | 4 | 51_198 | 0 | 141.0 | Domain | FPL |
| Hgene | CLEC16A | chr16:11220003 | chr16:11444499 | ENST00000409552 | + | 1 | 21 | 51_198 | 0 | 907.0 | Domain | FPL |
| Tgene | RMI2 | chr16:11220003 | chr16:11444499 | ENST00000312499 | 0 | 2 | 44_114 | 98 | 148.0 | DNA binding | Note=OB |
Top |
Fusion Gene Sequence for CLEC16A-RMI2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >17082_17082_1_CLEC16A-RMI2_CLEC16A_chr16_11220003_ENST00000409552_RMI2_chr16_11444499_ENST00000572173_length(transcript)=3868nt_BP=2754nt CGGGGAGGAAGGCGGCTCGCGGTTCCTCCACCGCCTCCGCCGCCGCATCCTCCGCTTGTGCTACCGCCGCGGGCGCTGGGCCGCTCTGCT GGTCCGGCATGAGACCGTGAGACGAGAGACGGGTCGGGGCCGCCGACATGTTTGGCCGCTCGCGGAGCTGGGTGGGCGGGGGCCATGGCA AGACTTCCCGCAACATCCACTCCTTGGACCACCTCAAGTATCTGTACCACGTTTTGACCAAAAACACCACAGTCACAGAACAGAACCGGA ACCTGCTAGTGGAGACCATCCGTTCCATCACTGAGATCCTGATCTGGGGAGATCAAAATGACAGCTCTGTATTTGACTTCTTCCTGGAGA AGAATATGTTTGTTTTCTTCTTGAACATCTTGCGGCAAAAGTCGGGCCGTTACGTGTGCGTTCAGCTGCTGCAGACCTTGAACATCCTCT TTGAGAACATCAGTCACGAGACCTCACTTTATTATTTGCTCTCAAATAACTACGTAAATTCTATCATCGTTCATAAATTTGACTTTTCTG ATGAGGAGATTATGGCCTATTATATATCGTTCCTGAAAACACTTTCGTTAAAACTCAACAACCACACTGTCCATTTCTTTTATAATGAGC ACACCAATGACTTTGCCCTGTACACAGAAGCCATCAAGTTTTTCAACCACCCTGAAAGCATGGTTAGAATTGCTGTAAGAACCATAACTT TGAATGTCTATAAAGTGGATAACCAGGCCATGCTGCACTACATCCGAGATAAAACTGCTGTTCCTTACTTCTCCAATTTGGTCTGGTTCA TTGGGAGCCATGTGATCGAACTCGATGACTGCGTGCAGACTGATGAGGAGCATCGGAATCGGGGTAAACTGAGTGATCTGGTGGCAGAGC ACCTAGACCACCTGCACTATCTCAATGACATCCTGATCATCAACTGTGAGTTCCTCAACGATGTGCTCACTGACCACCTGCTCAACAGGC TCTTCCTGCCCCTCTACGTGTACTCACTGGAGAACCAGGACAAGGGAGGAGAACGGCCGAAAATTAGCCTGCCGGTGTCTCTTTATCTTC TGTCACAGGTCTTCTTAATTATACATCATGCACCGCTGGTGAACTCGTTAGCTGAAGTCATTCTGAATGGTGATCTGTCTGAGATGTACG CTAAGACTGAACAGGATATTCAGAGAAGTTCTGCCAAGCCCAGCATTCGGTGCTTCATTAAACCCACCGAGACACTCGAGCGGTCCCTTG AGATGAACAAGCACAAGGGCAAGAGGCGGGTGCAAAAGAGACCCAACTACAAAAACGTTGGGGAAGAAGAAGATGAGGAGAAAGGGCCCA CCGAGGATGCCCAAGAAGACGCCGAGAAGGCTAAAGAGATCGAGATGGTGATCATGGAGCGTAGCAAGCTCTCAGAGCTGGCCGCCAGCA CCTCCGTGCAGGAGCAGAACACCACGGACGAGGAGAAAAGCGCCGCCGCCACCTGCTCTGAGAGCACGCAATGGAGCAGACCCTTCCTGG ATATGGTGTACCACGCGCTGGACAGCCCGGATGATGATTACCATGCCCTGTTCGTGCTCTGCCTCCTCTATGCCATGTCTCATAATAAAG GCATGGATCCTGAAAAATTAGAGCGAATCCAGCTCCCCGTGCCAAATGCGGCCGAGAAGACCACCTACAACCACCCGCTAGCTGAAAGAC TCATCAGGATCATGAACAACGCTGCCCAGCCAGATGGGAAGATCCGGCTGGCGACGCTGGAGCTGAGCTGCCTGCTTCTGAAGCAGCAAG TCCTGATGAGTGCTGGCTGCATCATGAAGGACGTGCACCTGGCCTGCCTGGAGGGTGCGAGAGAAGAAAGTGTTCACCTTGTACGACATT TTTATAAGGGAGAAGACATTTTTTTGGACATGTTTGAAGATGAGTATAGGAGCATGACAATGAAGCCCATGAACGTGGAATATCTCATGA TGGACGCCTCCATCCTGCTGCCCCCAACAGGCACGCCACTGACGGGCATTGACTTCGTGAAGCGGCTGCCGTGTGGCGATGTGGAGAAGA CCCGGCGGGCCATCCGGGTGTTCTTCATGCTGCGTTCCCTGTCACTGCAATTGCGAGGGGAGCCTGAGACACAGTTGCCGCTGACTCGGG AGGAGGACCTGATCAAGACTGATGATGTCCTGGATCTGAATAACAGCGACTTGATTGCATGTACAGTGATCACCAAGGATGGCGGCATGG TCCAGCGATTCCTGGCTGTGGATATTTACCAGATGAGTTTGGTGGAGCCTGATGTGTCCAGGCTTGGCTGGGGAGTGGTCAAGTTTGCAG GCCTATTGCAGGACATGCAGGTGACTGGCGTGGAGGACGACAGCCGTGCCCTGAACATCACCATCCACAAGCCTGCGTCCAGCCCCCATT CCAAGCCCTTCCCCATCCTCCAGGCCACCTTCATCTTCTCAGACCACATCCGCTGCATCATCGCCAAGCAGCGCCTGGCCAAAGGCCGCA TCCAGGCAAGGCGCATGAAGATGCAGAGAATAGCTGGTGAGCCAGCCCCCCGCCCTGCGCCACAGCTCGTTCATCATGGTGGGCGTTCAA GGTCTTTCTCTCTGTGGAGCCTGTGTGAGCTGCCCTTCCTCTCTCAGAAGCCCCGTCGGCTGGCAGCACCAGCTTCTTAGAATTTGTCAA AGCACAGCGCAAGATTAAACACAGCCCCCAAAATAAATTTTGGCTTTCCCTTTAGAAAGTATGTGATGGTGATGGGAGTGGTTCAGGCCT GCAGCCCTGAGCCCTGCCTGCAGGCTGTGAAGATGACAGACCTTTCTGATAATCCCATCCATGAAAGTATGTGGGAACTGGAGGTAGAAG ATTTACACAGGAATATTCCTTAGAGTATGTTGGAACTGTCGTTAAAAACAACCAAAATCCCGAAACTATTTAGAAGCTTATAATGATGTG GATTTCATGGACACTTTTCAATGCGTATTTTTCAAATGCTTCTCAGAGAGCCTTGCTTTGGTTGACCAAGGAGTCCGGATGTAGGAATGT TTAAATCCTCGGATACTTCAGTGACACAGCCTCTGCTGCCCCTTGCTTTGCCTGTGTTTGCTGATGAAAAGCAGATGCTTGTGTTTCATT TTCCTTCCTGGTTTGTGTGTGTTAATTCTCTCTCTCTCTCTCAGACACAGAAGTCTCATGTTGCATTTTCCAAATTTTATGAGTGATGAT ACTTTTTCCATTACTGCTGCGTCCCTGTTTTACAATGCAAAATTTAAGTACGGTCGTTGCCCATGGTGATTAAAGTGTGGTTATGGGCAG GAAGACAGACTGTGTAAAAAAGGAATGACATCCTGGCTCCTCATCTTCTTCATCAGCAACTACCATAACCAGTTTGCGAGTCAAATGGCA TTTCCTAACGGCAGGCATGGCGGCCCCTGAAAGACAACAGCTCCCTTTCTGCTTCGGACACCACTCAAACATTTAGACGCAGCTCTATCC CTTTTCCTAGCTAGAGAAGGTGATGCCTTCTTCCATTACTCAGAGATGTTGAGACGTTTTCAGAATTTCTTGTTGAAATGAAAAACATCA AGATAAAGGACGCCTTTCAGGCATTAGCTAAACTTCCACTTCATAACTTTCGGCGAGACATGGTGAGCCTCCTGGTGTAGAGTTCTTTTG TCTTTGTATGGAATGACTTTTTGCTGTGATGGTTTTGAATGTTGGGTTTCTGCTGTCTGCTTAGTACCCATGCCTGAATTTTTTGAGATT >17082_17082_1_CLEC16A-RMI2_CLEC16A_chr16_11220003_ENST00000409552_RMI2_chr16_11444499_ENST00000572173_length(amino acids)=850AA_BP= MFGRSRSWVGGGHGKTSRNIHSLDHLKYLYHVLTKNTTVTEQNRNLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFVFFLNILRQKSG RYVCVQLLQTLNILFENISHETSLYYLLSNNYVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFYNEHTNDFALYTEAIKFFN HPESMVRIAVRTITLNVYKVDNQAMLHYIRDKTAVPYFSNLVWFIGSHVIELDDCVQTDEEHRNRGKLSDLVAEHLDHLHYLNDILIINC EFLNDVLTDHLLNRLFLPLYVYSLENQDKGGERPKISLPVSLYLLSQVFLIIHHAPLVNSLAEVILNGDLSEMYAKTEQDIQRSSAKPSI RCFIKPTETLERSLEMNKHKGKRRVQKRPNYKNVGEEEDEEKGPTEDAQEDAEKAKEIEMVIMERSKLSELAASTSVQEQNTTDEEKSAA ATCSESTQWSRPFLDMVYHALDSPDDDYHALFVLCLLYAMSHNKGMDPEKLERIQLPVPNAAEKTTYNHPLAERLIRIMNNAAQPDGKIR LATLELSCLLLKQQVLMSAGCIMKDVHLACLEGAREESVHLVRHFYKGEDIFLDMFEDEYRSMTMKPMNVEYLMMDASILLPPTGTPLTG IDFVKRLPCGDVEKTRRAIRVFFMLRSLSLQLRGEPETQLPLTREEDLIKTDDVLDLNNSDLIACTVITKDGGMVQRFLAVDIYQMSLVE PDVSRLGWGVVKFAGLLQDMQVTGVEDDSRALNITIHKPASSPHSKPFPILQATFIFSDHIRCIIAKQRLAKGRIQARRMKMQRIAGEPA -------------------------------------------------------------- >17082_17082_2_CLEC16A-RMI2_CLEC16A_chr16_11220003_ENST00000409790_RMI2_chr16_11444499_ENST00000572173_length(transcript)=3985nt_BP=2871nt AACTGCATTTCCCAGCGCCCCACGCGGCGGCGGCCGTAAAGCGCGGCGGTCGAACGGCCGGTTCCGGCTGAATGTCAGTGCTGGGCTGTG GGCCGGGGAGGAAGGCGGCTCGCGGTTCCTCCACCGCCTCCGCCGCCGCATCCTCCGCTTGTGCTACCGCCGCGGGCGCTGGGCCGCTCT GCTGGTCCGGCATGAGACCGTGAGACGAGAGACGGGTCGGGGCCGCCGACATGTTTGGCCGCTCGCGGAGCTGGGTGGGCGGGGGCCATG GCAAGACTTCCCGCAACATCCACTCCTTGGACCACCTCAAGTATCTGTACCACGTTTTGACCAAAAACACCACAGTCACAGAACAGAACC GGAACCTGCTAGTGGAGACCATCCGTTCCATCACTGAGATCCTGATCTGGGGAGATCAAAATGACAGCTCTGTATTTGACTTCTTCCTGG AGAAGAATATGTTTGTTTTCTTCTTGAACATCTTGCGGCAAAAGTCGGGCCGTTACGTGTGCGTTCAGCTGCTGCAGACCTTGAACATCC TCTTTGAGAACATCAGTCACGAGACCTCACTTTATTATTTGCTCTCAAATAACTACGTAAATTCTATCATCGTTCATAAATTTGACTTTT CTGATGAGGAGATTATGGCCTATTATATATCGTTCCTGAAAACACTTTCGTTAAAACTCAACAACCACACTGTCCATTTCTTTTATAATG AGCACACCAATGACTTTGCCCTGTACACAGAAGCCATCAAGTTTTTCAACCACCCTGAAAGCATGGTTAGAATTGCTGTAAGAACCATAA CTTTGAATGTCTATAAAGTGTCATTGGATAACCAGGCCATGCTGCACTACATCCGAGATAAAACTGCTGTTCCTTACTTCTCCAATTTGG TCTGGTTCATTGGGAGCCATGTGATCGAACTCGATGACTGCGTGCAGACTGATGAGGAGCATCGGAATCGGGGTAAACTGAGTGATCTGG TGGCAGAGCACCTAGACCACCTGCACTATCTCAATGACATCCTGATCATCAACTGTGAGTTCCTCAACGATGTGCTCACTGACCACCTGC TCAACAGGCTCTTCCTGCCCCTCTACGTGTACTCACTGGAGAACCAGGACAAGGGAGGAGAACGGCCGAAAATTAGCCTGCCGGTGTCTC TTTATCTTCTGTCACAGGTCTTCTTAATTATACATCATGCACCGCTGGTGAACTCGTTAGCTGAAGTCATTCTGAATGGTGATCTGTCTG AGATGTACGCTAAGACTGAACAGGATATTCAGAGAAGTTCTGCCAAGCCCAGCATTCGGTGCTTCATTAAACCCACCGAGACACTCGAGC GGTCCCTTGAGATGAACAAGCACAAGGGCAAGAGGCGGGTGCAAAAGAGACCCAACTACAAAAACGTTGGGGAAGAAGAAGATGAGGAGA AAGGGCCCACCGAGGATGCCCAAGAAGACGCCGAGAAGGCTAAAGGTACAGAGGGTGGTTCAAAAGGCATCAAGACGAGTGGGGAGAGTG AAGAGATCGAGATGGTGATCATGGAGCGTAGCAAGCTCTCAGAGCTGGCCGCCAGCACCTCCGTGCAGGAGCAGAACACCACGGACGAGG AGAAAAGCGCCGCCGCCACCTGCTCTGAGAGCACGCAATGGAGCAGACCCTTCCTGGATATGGTGTACCACGCGCTGGACAGCCCGGATG ATGATTACCATGCCCTGTTCGTGCTCTGCCTCCTCTATGCCATGTCTCATAATAAAGGCATGGATCCTGAAAAATTAGAGCGAATCCAGC TCCCCGTGCCAAATGCGGCCGAGAAGACCACCTACAACCACCCGCTAGCTGAAAGACTCATCAGGATCATGAACAACGCTGCCCAGCCAG ATGGGAAGATCCGGCTGGCGACGCTGGAGCTGAGCTGCCTGCTTCTGAAGCAGCAAGTCCTGATGAGTGCTGGCTGCATCATGAAGGACG TGCACCTGGCCTGCCTGGAGGGTGCGAGAGAAGAAAGTGTTCACCTTGTACGACATTTTTATAAGGGAGAAGACATTTTTTTGGACATGT TTGAAGATGAGTATAGGAGCATGACAATGAAGCCCATGAACGTGGAATATCTCATGATGGACGCCTCCATCCTGCTGCCCCCAACAGGCA CGCCACTGACGGGCATTGACTTCGTGAAGCGGCTGCCGTGTGGCGATGTGGAGAAGACCCGGCGGGCCATCCGGGTGTTCTTCATGCTGC GTTCCCTGTCACTGCAATTGCGAGGGGAGCCTGAGACACAGTTGCCGCTGACTCGGGAGGAGGACCTGATCAAGACTGATGATGTCCTGG ATCTGAATAACAGCGACTTGATTGCATGTACAGTGATCACCAAGGATGGCGGCATGGTCCAGCGATTCCTGGCTGTGGATATTTACCAGA TGAGTTTGGTGGAGCCTGATGTGTCCAGGCTTGGCTGGGGAGTGGTCAAGTTTGCAGGCCTATTGCAGGACATGCAGGTGACTGGCGTGG AGGACGACAGCCGTGCCCTGAACATCACCATCCACAAGCCTGCGTCCAGCCCCCATTCCAAGCCCTTCCCCATCCTCCAGGCCACCTTCA TCTTCTCAGACCACATCCGCTGCATCATCGCCAAGCAGCGCCTGGCCAAAGGCCGCATCCAGGCAAGGCGCATGAAGATGCAGAGAATAG CTGCCCTCCTGGACCTCCCAATCCAGCCCACCACTGAAGTCCTGGGGTTTGGACTCGGCTCCTCCACCTCCACTCAGCACCTGCCTTTCC GCTTCTACGACCAGGGGCGCCGGGGCAGCAGCGACCCCACAGTGCAGCGCTCCGTGTTTGCATCGGTGGACAAGGTGCCAGGAAAGTATG TGATGGTGATGGGAGTGGTTCAGGCCTGCAGCCCTGAGCCCTGCCTGCAGGCTGTGAAGATGACAGACCTTTCTGATAATCCCATCCATG AAAGTATGTGGGAACTGGAGGTAGAAGATTTACACAGGAATATTCCTTAGAGTATGTTGGAACTGTCGTTAAAAACAACCAAAATCCCGA AACTATTTAGAAGCTTATAATGATGTGGATTTCATGGACACTTTTCAATGCGTATTTTTCAAATGCTTCTCAGAGAGCCTTGCTTTGGTT GACCAAGGAGTCCGGATGTAGGAATGTTTAAATCCTCGGATACTTCAGTGACACAGCCTCTGCTGCCCCTTGCTTTGCCTGTGTTTGCTG ATGAAAAGCAGATGCTTGTGTTTCATTTTCCTTCCTGGTTTGTGTGTGTTAATTCTCTCTCTCTCTCTCAGACACAGAAGTCTCATGTTG CATTTTCCAAATTTTATGAGTGATGATACTTTTTCCATTACTGCTGCGTCCCTGTTTTACAATGCAAAATTTAAGTACGGTCGTTGCCCA TGGTGATTAAAGTGTGGTTATGGGCAGGAAGACAGACTGTGTAAAAAAGGAATGACATCCTGGCTCCTCATCTTCTTCATCAGCAACTAC CATAACCAGTTTGCGAGTCAAATGGCATTTCCTAACGGCAGGCATGGCGGCCCCTGAAAGACAACAGCTCCCTTTCTGCTTCGGACACCA CTCAAACATTTAGACGCAGCTCTATCCCTTTTCCTAGCTAGAGAAGGTGATGCCTTCTTCCATTACTCAGAGATGTTGAGACGTTTTCAG AATTTCTTGTTGAAATGAAAAACATCAAGATAAAGGACGCCTTTCAGGCATTAGCTAAACTTCCACTTCATAACTTTCGGCGAGACATGG TGAGCCTCCTGGTGTAGAGTTCTTTTGTCTTTGTATGGAATGACTTTTTGCTGTGATGGTTTTGAATGTTGGGTTTCTGCTGTCTGCTTA GTACCCATGCCTGAATTTTTTGAGATTGTAAATATCAAAGGAGTTAGATTGTGTTCTGACATGGTTGTAGACTTTCACCTGGATTATTGA >17082_17082_2_CLEC16A-RMI2_CLEC16A_chr16_11220003_ENST00000409790_RMI2_chr16_11444499_ENST00000572173_length(amino acids)=929AA_BP=0 MFGRSRSWVGGGHGKTSRNIHSLDHLKYLYHVLTKNTTVTEQNRNLLVETIRSITEILIWGDQNDSSVFDFFLEKNMFVFFLNILRQKSG RYVCVQLLQTLNILFENISHETSLYYLLSNNYVNSIIVHKFDFSDEEIMAYYISFLKTLSLKLNNHTVHFFYNEHTNDFALYTEAIKFFN HPESMVRIAVRTITLNVYKVSLDNQAMLHYIRDKTAVPYFSNLVWFIGSHVIELDDCVQTDEEHRNRGKLSDLVAEHLDHLHYLNDILII NCEFLNDVLTDHLLNRLFLPLYVYSLENQDKGGERPKISLPVSLYLLSQVFLIIHHAPLVNSLAEVILNGDLSEMYAKTEQDIQRSSAKP SIRCFIKPTETLERSLEMNKHKGKRRVQKRPNYKNVGEEEDEEKGPTEDAQEDAEKAKGTEGGSKGIKTSGESEEIEMVIMERSKLSELA ASTSVQEQNTTDEEKSAAATCSESTQWSRPFLDMVYHALDSPDDDYHALFVLCLLYAMSHNKGMDPEKLERIQLPVPNAAEKTTYNHPLA ERLIRIMNNAAQPDGKIRLATLELSCLLLKQQVLMSAGCIMKDVHLACLEGAREESVHLVRHFYKGEDIFLDMFEDEYRSMTMKPMNVEY LMMDASILLPPTGTPLTGIDFVKRLPCGDVEKTRRAIRVFFMLRSLSLQLRGEPETQLPLTREEDLIKTDDVLDLNNSDLIACTVITKDG GMVQRFLAVDIYQMSLVEPDVSRLGWGVVKFAGLLQDMQVTGVEDDSRALNITIHKPASSPHSKPFPILQATFIFSDHIRCIIAKQRLAK GRIQARRMKMQRIAALLDLPIQPTTEVLGFGLGSSTSTQHLPFRFYDQGRRGSSDPTVQRSVFASVDKVPGKYVMVMGVVQACSPEPCLQ -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CLEC16A-RMI2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CLEC16A-RMI2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CLEC16A-RMI2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
