
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CLK1-HSPD1 (FusionGDB2 ID:17194) |
Fusion Gene Summary for CLK1-HSPD1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CLK1-HSPD1 | Fusion gene ID: 17194 | Hgene | Tgene | Gene symbol | CLK1 | HSPD1 | Gene ID | 78989 | 3329 |
| Gene name | collectin subfamily member 11 | heat shock protein family D (Hsp60) member 1 | |
| Synonyms | 3MC2|CL-K1-I|CL-K1-II|CL-K1-IIa|CL-K1-IIb|CLK1 | CPN60|GROEL|HLD4|HSP-60|HSP60|HSP65|HuCHA60|SPG13 | |
| Cytomap | 2p25.3 | 2q33.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | collectin-11Collectin K1collectin kidney protein 1collectin sub-family member 11 | 60 kDa heat shock protein, mitochondrial60 kDa chaperoninP60 lymphocyte proteinchaperonin 60epididymis secretory sperm binding proteinheat shock 60kDa protein 1 (chaperonin)heat shock protein 65mitochondrial matrix protein P1short heat shock prote | |
| Modification date | 20200313 | 20200315 | |
| UniProtAcc | P49759 | P10809 | |
| Ensembl transtripts involved in fusion gene | ENST00000321356, ENST00000434813, ENST00000409769, ENST00000492793, | ENST00000544407, ENST00000345042, ENST00000388968, | |
| Fusion gene scores | * DoF score | 7 X 7 X 5=245 | 14 X 10 X 6=840 |
| # samples | 9 | 16 | |
| ** MAII score | log2(9/245*10)=-1.4447848426729 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(16/840*10)=-2.39231742277876 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CLK1 [Title/Abstract] AND HSPD1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CLK1(201725960)-HSPD1(198353971), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | CLK1-HSPD1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. CLK1-HSPD1 seems lost the major protein functional domain in Hgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. CLK1-HSPD1 seems lost the major protein functional domain in Hgene partner, which is a kinase due to the frame-shifted ORF. CLK1-HSPD1 seems lost the major protein functional domain in Tgene partner, which is a cell metabolism gene due to the frame-shifted ORF. CLK1-HSPD1 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. CLK1-HSPD1 seems lost the major protein functional domain in Tgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | CLK1 | GO:0006956 | complement activation | 23954398 |
| Hgene | CLK1 | GO:0019730 | antimicrobial humoral response | 20956340 |
| Tgene | HSPD1 | GO:0002368 | B cell cytokine production | 16148103 |
| Tgene | HSPD1 | GO:0002755 | MyD88-dependent toll-like receptor signaling pathway | 16148103 |
| Tgene | HSPD1 | GO:0002842 | positive regulation of T cell mediated immune response to tumor cell | 10663613 |
| Tgene | HSPD1 | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process | 17823127 |
| Tgene | HSPD1 | GO:0006986 | response to unfolded protein | 11050098 |
| Tgene | HSPD1 | GO:0032727 | positive regulation of interferon-alpha production | 17164250 |
| Tgene | HSPD1 | GO:0032729 | positive regulation of interferon-gamma production | 17164250 |
| Tgene | HSPD1 | GO:0032733 | positive regulation of interleukin-10 production | 16148103 |
| Tgene | HSPD1 | GO:0032735 | positive regulation of interleukin-12 production | 17164250 |
| Tgene | HSPD1 | GO:0032755 | positive regulation of interleukin-6 production | 16148103 |
| Tgene | HSPD1 | GO:0042026 | protein refolding | 11050098 |
| Tgene | HSPD1 | GO:0042100 | B cell proliferation | 16148103 |
| Tgene | HSPD1 | GO:0042110 | T cell activation | 15371451|17164250|18256040 |
| Tgene | HSPD1 | GO:0042113 | B cell activation | 16148103 |
| Tgene | HSPD1 | GO:0043032 | positive regulation of macrophage activation | 17164250 |
| Tgene | HSPD1 | GO:0044406 | adhesion of symbiont to host | 20633027 |
| Tgene | HSPD1 | GO:0048291 | isotype switching to IgG isotypes | 16148103 |
| Tgene | HSPD1 | GO:0050870 | positive regulation of T cell activation | 16148103|17164250 |
 Fusion gene breakpoints across CLK1 (5'-gene) Fusion gene breakpoints across CLK1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
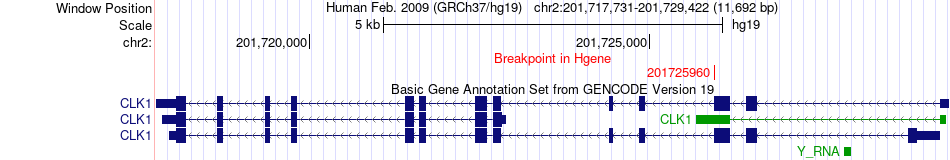 |
 Fusion gene breakpoints across HSPD1 (3'-gene) Fusion gene breakpoints across HSPD1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
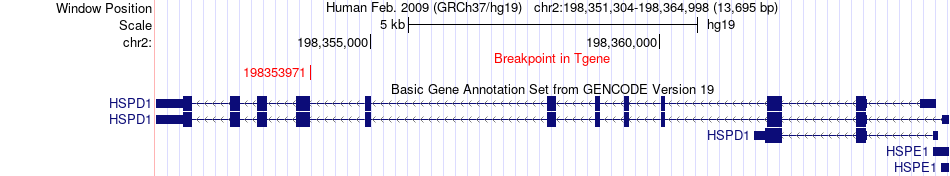 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-B6-A0I2 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
Top |
Fusion Gene ORF analysis for CLK1-HSPD1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000321356 | ENST00000544407 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| 5CDS-intron | ENST00000434813 | ENST00000544407 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| Frame-shift | ENST00000321356 | ENST00000345042 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| Frame-shift | ENST00000321356 | ENST00000388968 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| In-frame | ENST00000434813 | ENST00000345042 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| In-frame | ENST00000434813 | ENST00000388968 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| intron-3CDS | ENST00000409769 | ENST00000345042 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| intron-3CDS | ENST00000409769 | ENST00000388968 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| intron-3CDS | ENST00000492793 | ENST00000345042 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| intron-3CDS | ENST00000492793 | ENST00000388968 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| intron-intron | ENST00000409769 | ENST00000544407 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
| intron-intron | ENST00000492793 | ENST00000544407 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000434813 | CLK1 | chr2 | 201725960 | - | ENST00000388968 | HSPD1 | chr2 | 198353971 | - | 2069 | 851 | 335 | 1603 | 422 |
| ENST00000434813 | CLK1 | chr2 | 201725960 | - | ENST00000345042 | HSPD1 | chr2 | 198353971 | - | 2064 | 851 | 335 | 1603 | 422 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000434813 | ENST00000388968 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - | 0.000370214 | 0.99962974 |
| ENST00000434813 | ENST00000345042 | CLK1 | chr2 | 201725960 | - | HSPD1 | chr2 | 198353971 | - | 0.000390713 | 0.9996093 |
Top |
Fusion Genomic Features for CLK1-HSPD1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
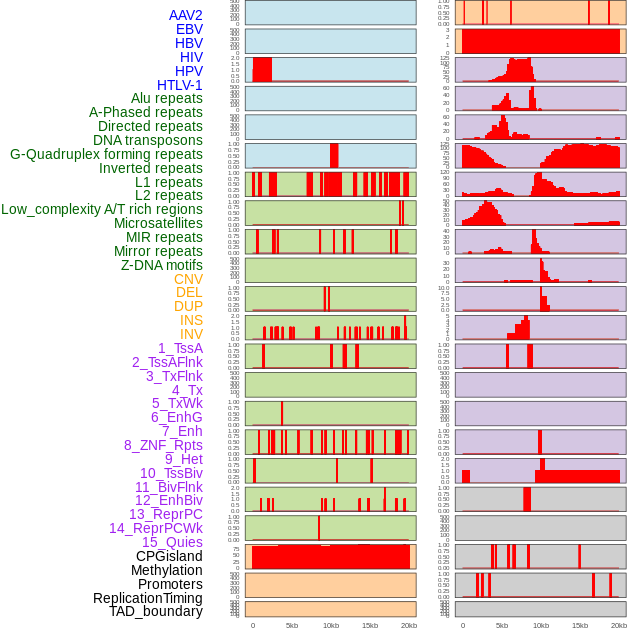 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for CLK1-HSPD1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr2:201725960/chr2:198353971) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CLK1 | HSPD1 |
| FUNCTION: Dual specificity kinase acting on both serine/threonine and tyrosine-containing substrates. Phosphorylates serine- and arginine-rich (SR) proteins of the spliceosomal complex and may be a constituent of a network of regulatory mechanisms that enable SR proteins to control RNA splicing. Phosphorylates: SRSF1, SRSF3 and PTPN1. Regulates the alternative splicing of tissue factor (F3) pre-mRNA in endothelial cells and adenovirus E1A pre-mRNA. {ECO:0000269|PubMed:10480872, ECO:0000269|PubMed:19168442}. | FUNCTION: Chaperonin implicated in mitochondrial protein import and macromolecular assembly. Together with Hsp10, facilitates the correct folding of imported proteins. May also prevent misfolding and promote the refolding and proper assembly of unfolded polypeptides generated under stress conditions in the mitochondrial matrix (PubMed:1346131, PubMed:11422376). The functional units of these chaperonins consist of heptameric rings of the large subunit Hsp60, which function as a back-to-back double ring. In a cyclic reaction, Hsp60 ring complexes bind one unfolded substrate protein per ring, followed by the binding of ATP and association with 2 heptameric rings of the co-chaperonin Hsp10. This leads to sequestration of the substrate protein in the inner cavity of Hsp60 where, for a certain period of time, it can fold undisturbed by other cell components. Synchronous hydrolysis of ATP in all Hsp60 subunits results in the dissociation of the chaperonin rings and the release of ADP and the folded substrate protein (Probable). {ECO:0000269|PubMed:11422376, ECO:0000269|PubMed:1346131, ECO:0000305|PubMed:25918392}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CLK1 | chr2:201725960 | chr2:198353971 | ENST00000434813 | - | 3 | 13 | 167_175 | 172 | 527.0 | Nucleotide binding | ATP |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CLK1 | chr2:201725960 | chr2:198353971 | ENST00000321356 | - | 3 | 13 | 161_477 | 130 | 485.0 | Domain | Protein kinase |
| Hgene | CLK1 | chr2:201725960 | chr2:198353971 | ENST00000434813 | - | 3 | 13 | 161_477 | 172 | 527.0 | Domain | Protein kinase |
| Hgene | CLK1 | chr2:201725960 | chr2:198353971 | ENST00000321356 | - | 3 | 13 | 167_175 | 130 | 485.0 | Nucleotide binding | ATP |
| Tgene | HSPD1 | chr2:201725960 | chr2:198353971 | ENST00000345042 | 7 | 12 | 111_115 | 323 | 574.0 | Nucleotide binding | ATP | |
| Tgene | HSPD1 | chr2:201725960 | chr2:198353971 | ENST00000388968 | 7 | 12 | 111_115 | 323 | 574.0 | Nucleotide binding | ATP |
Top |
Fusion Gene Sequence for CLK1-HSPD1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >17194_17194_1_CLK1-HSPD1_CLK1_chr2_201725960_ENST00000434813_HSPD1_chr2_198353971_ENST00000345042_length(transcript)=2064nt_BP=851nt AAGACGCAGGAGGGAGCGGACTAGGTGACAGGGCCGTTCCTGTGAGCCTCGCGGGCGCCTGGCGATGCCCCCTTTTCCTGCTTGTTTGCT GCCCGCCGTCCCCGGCGCGACGACTGCCTGCTCCCTTCACTCCCAGGCTGCACAGTGGCGGCGCGCCCCTCTTTCCTGCGCGGCCGAGCC TGTCGCCGCCGGATCCGGCCTGCGGGGGTAGTTACGGTGTTTGCTAGGCCGGCCGCCCTCTTGGAGCTTCTGCCCTCCGCTGAGGAAGCG GCGCCGCCTGACGCGGGACGGTCGGGCGGGCGCCATGTTGTGAACCGCCTCGGCGGAGCTGTAAGATGGCGGCTGGGCGGAGGCCGGCTT CGGCCCTGTGGCCGGAAAGGCGAGGCTCCCCGTTGAGGGGGGATTTGCTGGGGTTCCAGAATGTGCGTGAGCCAAGCAGCTGTGGGGAAA CGTTGTCTGGAATGAGACACTCAAAGAGAACTTACTGTCCTGATTGGGATGACAAGGATTGGGATTATGGAAAATGGAGGAGCAGCAGCA GTCATAAAAGAAGGAAGAGATCACATAGCAGTGCCCAGGAGAACAAGCGCTGCAAATACAATCACTCTAAAATGTGTGATAGCCATTATT TGGAAAGCAGGTCTATAAATGAGAAAGATTATCATAGTCGACGCTACATTGATGAGTACAGAAATGACTACACTCAAGGATGTGAACCTG GACATCGCCAAAGAGACCATGAAAGCCGGTATCAGAACCATAGTAGCAAGTCTTCTGGTAGAAGTGGAAGAAGTAGTTATAAAAGCAAAC ACAGGATTCACCACAGTACTTCACATCGTCGTTCACATGGGGTGTTTGGAGAAGAGGGATTGACCCTGAATCTTGAAGACGTTCAGCCTC ATGACTTAGGAAAAGTTGGAGAGGTCATTGTGACCAAAGACGATGCCATGCTCTTAAAAGGAAAAGGTGACAAGGCTCAAATTGAAAAAC GTATTCAAGAAATCATTGAGCAGTTAGATGTCACAACTAGTGAATATGAAAAGGAAAAACTGAATGAACGGCTTGCAAAACTTTCAGATG GAGTGGCTGTGCTGAAGGTTGGTGGGACAAGTGATGTTGAAGTGAATGAAAAGAAAGACAGAGTTACAGATGCCCTTAATGCTACAAGAG CTGCTGTTGAAGAAGGCATTGTTTTGGGAGGGGGTTGTGCCCTCCTTCGATGCATTCCAGCCTTGGACTCATTGACTCCAGCTAATGAAG ATCAAAAAATTGGTATAGAAATTATTAAAAGAACACTCAAAATTCCAGCAATGACCATTGCTAAGAATGCAGGTGTTGAAGGATCTTTGA TAGTTGAGAAAATTATGCAAAGTTCCTCAGAAGTTGGTTATGATGCTATGGCTGGAGATTTTGTGAATATGGTGGAAAAAGGAATCATTG ACCCAACAAAGGTTGTGAGAACTGCTTTATTGGATGCTGCTGGTGTGGCCTCTCTGTTAACTACAGCAGAAGTTGTAGTCACAGAAATTC CTAAAGAAGAGAAGGACCCTGGAATGGGTGCAATGGGTGGAATGGGAGGTGGTATGGGAGGTGGCATGTTCTAACTCCTAGACTAGTGCT TTACCTTTATTAATGAACTGTGACAGGAAGCCCAAGGCAGTGTTCCTCACCAATAACTTCAGAGAAGTCAGTTGGAGAAAATGAAGAAAA AGGCTGGCTGAAAATCACTATAACCATCAGTTACTGGTTTCAGTTGACAAAATATATAATGGTTTACTGCTGTCATTGTCCATGCCTACA GATAATTTATTTTGTATTTTTGAATAAAAAACATTTGTACATTCCTGATACTGGGTACAAGAGCCATGTACCAGTGTACTGCTTTCAACT TAAATCACTGAGGCATTTTTACTACTATTCTGTTAAAATCAGGATTTTAGTGCTTGCCACCACCAGATGAGAAGTTAAGCAGCCTTTCTG >17194_17194_1_CLK1-HSPD1_CLK1_chr2_201725960_ENST00000434813_HSPD1_chr2_198353971_ENST00000345042_length(amino acids)=422AA_BP=172 MAAGRRPASALWPERRGSPLRGDLLGFQNVREPSSCGETLSGMRHSKRTYCPDWDDKDWDYGKWRSSSSHKRRKRSHSSAQENKRCKYNH SKMCDSHYLESRSINEKDYHSRRYIDEYRNDYTQGCEPGHRQRDHESRYQNHSSKSSGRSGRSSYKSKHRIHHSTSHRRSHGVFGEEGLT LNLEDVQPHDLGKVGEVIVTKDDAMLLKGKGDKAQIEKRIQEIIEQLDVTTSEYEKEKLNERLAKLSDGVAVLKVGGTSDVEVNEKKDRV TDALNATRAAVEEGIVLGGGCALLRCIPALDSLTPANEDQKIGIEIIKRTLKIPAMTIAKNAGVEGSLIVEKIMQSSSEVGYDAMAGDFV -------------------------------------------------------------- >17194_17194_2_CLK1-HSPD1_CLK1_chr2_201725960_ENST00000434813_HSPD1_chr2_198353971_ENST00000388968_length(transcript)=2069nt_BP=851nt AAGACGCAGGAGGGAGCGGACTAGGTGACAGGGCCGTTCCTGTGAGCCTCGCGGGCGCCTGGCGATGCCCCCTTTTCCTGCTTGTTTGCT GCCCGCCGTCCCCGGCGCGACGACTGCCTGCTCCCTTCACTCCCAGGCTGCACAGTGGCGGCGCGCCCCTCTTTCCTGCGCGGCCGAGCC TGTCGCCGCCGGATCCGGCCTGCGGGGGTAGTTACGGTGTTTGCTAGGCCGGCCGCCCTCTTGGAGCTTCTGCCCTCCGCTGAGGAAGCG GCGCCGCCTGACGCGGGACGGTCGGGCGGGCGCCATGTTGTGAACCGCCTCGGCGGAGCTGTAAGATGGCGGCTGGGCGGAGGCCGGCTT CGGCCCTGTGGCCGGAAAGGCGAGGCTCCCCGTTGAGGGGGGATTTGCTGGGGTTCCAGAATGTGCGTGAGCCAAGCAGCTGTGGGGAAA CGTTGTCTGGAATGAGACACTCAAAGAGAACTTACTGTCCTGATTGGGATGACAAGGATTGGGATTATGGAAAATGGAGGAGCAGCAGCA GTCATAAAAGAAGGAAGAGATCACATAGCAGTGCCCAGGAGAACAAGCGCTGCAAATACAATCACTCTAAAATGTGTGATAGCCATTATT TGGAAAGCAGGTCTATAAATGAGAAAGATTATCATAGTCGACGCTACATTGATGAGTACAGAAATGACTACACTCAAGGATGTGAACCTG GACATCGCCAAAGAGACCATGAAAGCCGGTATCAGAACCATAGTAGCAAGTCTTCTGGTAGAAGTGGAAGAAGTAGTTATAAAAGCAAAC ACAGGATTCACCACAGTACTTCACATCGTCGTTCACATGGGGTGTTTGGAGAAGAGGGATTGACCCTGAATCTTGAAGACGTTCAGCCTC ATGACTTAGGAAAAGTTGGAGAGGTCATTGTGACCAAAGACGATGCCATGCTCTTAAAAGGAAAAGGTGACAAGGCTCAAATTGAAAAAC GTATTCAAGAAATCATTGAGCAGTTAGATGTCACAACTAGTGAATATGAAAAGGAAAAACTGAATGAACGGCTTGCAAAACTTTCAGATG GAGTGGCTGTGCTGAAGGTTGGTGGGACAAGTGATGTTGAAGTGAATGAAAAGAAAGACAGAGTTACAGATGCCCTTAATGCTACAAGAG CTGCTGTTGAAGAAGGCATTGTTTTGGGAGGGGGTTGTGCCCTCCTTCGATGCATTCCAGCCTTGGACTCATTGACTCCAGCTAATGAAG ATCAAAAAATTGGTATAGAAATTATTAAAAGAACACTCAAAATTCCAGCAATGACCATTGCTAAGAATGCAGGTGTTGAAGGATCTTTGA TAGTTGAGAAAATTATGCAAAGTTCCTCAGAAGTTGGTTATGATGCTATGGCTGGAGATTTTGTGAATATGGTGGAAAAAGGAATCATTG ACCCAACAAAGGTTGTGAGAACTGCTTTATTGGATGCTGCTGGTGTGGCCTCTCTGTTAACTACAGCAGAAGTTGTAGTCACAGAAATTC CTAAAGAAGAGAAGGACCCTGGAATGGGTGCAATGGGTGGAATGGGAGGTGGTATGGGAGGTGGCATGTTCTAACTCCTAGACTAGTGCT TTACCTTTATTAATGAACTGTGACAGGAAGCCCAAGGCAGTGTTCCTCACCAATAACTTCAGAGAAGTCAGTTGGAGAAAATGAAGAAAA AGGCTGGCTGAAAATCACTATAACCATCAGTTACTGGTTTCAGTTGACAAAATATATAATGGTTTACTGCTGTCATTGTCCATGCCTACA GATAATTTATTTTGTATTTTTGAATAAAAAACATTTGTACATTCCTGATACTGGGTACAAGAGCCATGTACCAGTGTACTGCTTTCAACT TAAATCACTGAGGCATTTTTACTACTATTCTGTTAAAATCAGGATTTTAGTGCTTGCCACCACCAGATGAGAAGTTAAGCAGCCTTTCTG >17194_17194_2_CLK1-HSPD1_CLK1_chr2_201725960_ENST00000434813_HSPD1_chr2_198353971_ENST00000388968_length(amino acids)=422AA_BP=172 MAAGRRPASALWPERRGSPLRGDLLGFQNVREPSSCGETLSGMRHSKRTYCPDWDDKDWDYGKWRSSSSHKRRKRSHSSAQENKRCKYNH SKMCDSHYLESRSINEKDYHSRRYIDEYRNDYTQGCEPGHRQRDHESRYQNHSSKSSGRSGRSSYKSKHRIHHSTSHRRSHGVFGEEGLT LNLEDVQPHDLGKVGEVIVTKDDAMLLKGKGDKAQIEKRIQEIIEQLDVTTSEYEKEKLNERLAKLSDGVAVLKVGGTSDVEVNEKKDRV TDALNATRAAVEEGIVLGGGCALLRCIPALDSLTPANEDQKIGIEIIKRTLKIPAMTIAKNAGVEGSLIVEKIMQSSSEVGYDAMAGDFV -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CLK1-HSPD1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CLK1-HSPD1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CLK1-HSPD1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
