
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CTDSP2-LGR5 (FusionGDB2 ID:20175) |
Fusion Gene Summary for CTDSP2-LGR5 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CTDSP2-LGR5 | Fusion gene ID: 20175 | Hgene | Tgene | Gene symbol | CTDSP2 | LGR5 | Gene ID | 10106 | 8549 |
| Gene name | CTD small phosphatase 2 | leucine rich repeat containing G protein-coupled receptor 5 | |
| Synonyms | OS4|PSR2|SCP2 | FEX|GPR49|GPR67|GRP49|HG38 | |
| Cytomap | 12q14.1 | 12q21.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | carboxy-terminal domain RNA polymerase II polypeptide A small phosphatase 2CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 2NLI-interacting factor 2conserved gene amplified in osteosarcomanuclear LIM interactor-intera | leucine-rich repeat-containing G-protein coupled receptor 5G-protein coupled receptor 49G-protein coupled receptor 67G-protein coupled receptor HG38orphan G protein-coupled receptor HG38 | |
| Modification date | 20200313 | 20200329 | |
| UniProtAcc | O14595 | O75473 | |
| Ensembl transtripts involved in fusion gene | ENST00000398073, ENST00000547701, ENST00000548823, | ENST00000536515, ENST00000540815, ENST00000266674, | |
| Fusion gene scores | * DoF score | 32 X 8 X 10=2560 | 13 X 15 X 9=1755 |
| # samples | 37 | 20 | |
| ** MAII score | log2(37/2560*10)=-2.79054663437105 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(20/1755*10)=-3.1333991254172 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CTDSP2 [Title/Abstract] AND LGR5 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CTDSP2(58217687)-LGR5(71946853), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | CTDSP2-LGR5 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. CTDSP2-LGR5 seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | CTDSP2 | GO:0006470 | protein dephosphorylation | 12721286 |
| Tgene | LGR5 | GO:0090263 | positive regulation of canonical Wnt signaling pathway | 21693646|22815884 |
 Fusion gene breakpoints across CTDSP2 (5'-gene) Fusion gene breakpoints across CTDSP2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
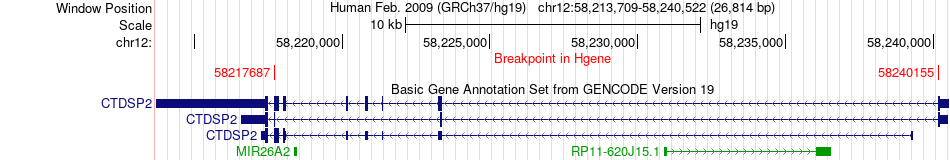 |
 Fusion gene breakpoints across LGR5 (3'-gene) Fusion gene breakpoints across LGR5 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
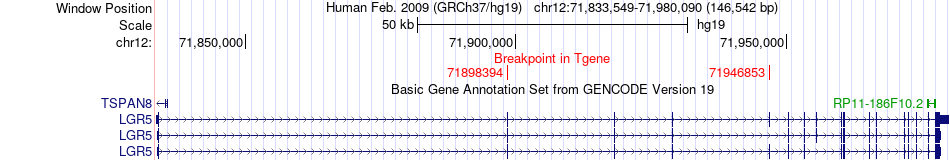 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SARC | TCGA-DX-A1KZ-01A | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| ChimerDB4 | SARC | TCGA-DX-AB3A-01A | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
| ChimerDB4 | SARC | TCGA-RN-AAAQ-01A | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
Top |
Fusion Gene ORF analysis for CTDSP2-LGR5 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000398073 | ENST00000536515 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| 5CDS-intron | ENST00000398073 | ENST00000540815 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| 5CDS-intron | ENST00000547701 | ENST00000536515 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| 5CDS-intron | ENST00000547701 | ENST00000540815 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| 5CDS-intron | ENST00000548823 | ENST00000536515 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| 5CDS-intron | ENST00000548823 | ENST00000540815 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| Frame-shift | ENST00000398073 | ENST00000266674 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| Frame-shift | ENST00000398073 | ENST00000266674 | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
| Frame-shift | ENST00000398073 | ENST00000536515 | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
| Frame-shift | ENST00000398073 | ENST00000540815 | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
| Frame-shift | ENST00000548823 | ENST00000266674 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| Frame-shift | ENST00000548823 | ENST00000266674 | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
| Frame-shift | ENST00000548823 | ENST00000536515 | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
| Frame-shift | ENST00000548823 | ENST00000540815 | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
| In-frame | ENST00000547701 | ENST00000266674 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + |
| intron-3CDS | ENST00000547701 | ENST00000266674 | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
| intron-3CDS | ENST00000547701 | ENST00000536515 | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
| intron-3CDS | ENST00000547701 | ENST00000540815 | CTDSP2 | chr12 | 58240155 | - | LGR5 | chr12 | 71898394 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000547701 | CTDSP2 | chr12 | 58217687 | - | ENST00000266674 | LGR5 | chr12 | 71946853 | + | 4593 | 721 | 689 | 3016 | 775 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000547701 | ENST00000266674 | CTDSP2 | chr12 | 58217687 | - | LGR5 | chr12 | 71946853 | + | 0.001305822 | 0.99869424 |
Top |
Fusion Genomic Features for CTDSP2-LGR5 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CTDSP2 | chr12 | 58217686 | - | LGR5 | chr12 | 71946852 | + | 0.009957893 | 0.9900421 |
| CTDSP2 | chr12 | 58217686 | - | LGR5 | chr12 | 71946852 | + | 0.009957893 | 0.9900421 |
| CTDSP2 | chr12 | 58240154 | - | LGR5 | chr12 | 71898393 | + | 8.29E-08 | 0.9999999 |
| CTDSP2 | chr12 | 58217686 | - | LGR5 | chr12 | 71946852 | + | 0.009957893 | 0.9900421 |
| CTDSP2 | chr12 | 58217686 | - | LGR5 | chr12 | 71946852 | + | 0.009957893 | 0.9900421 |
| CTDSP2 | chr12 | 58240154 | - | LGR5 | chr12 | 71898393 | + | 8.29E-08 | 0.9999999 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
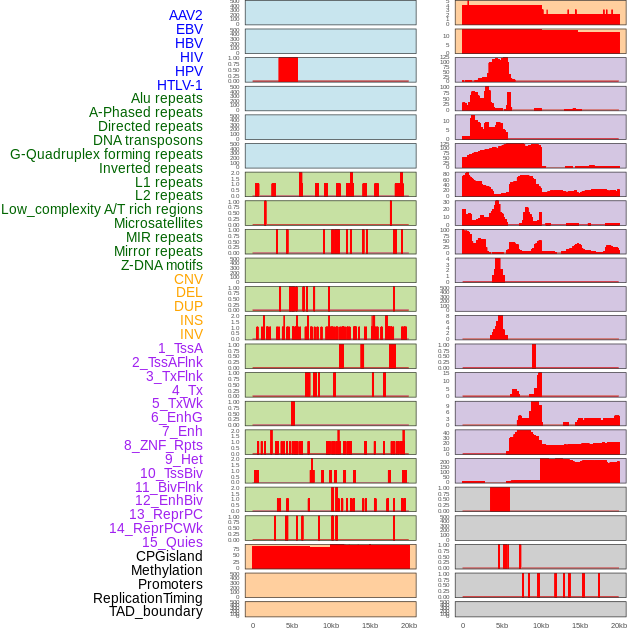 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
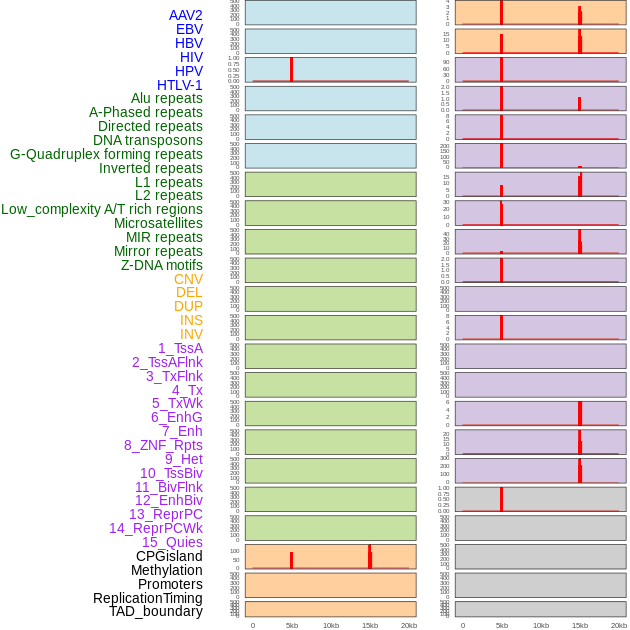 |
Top |
Fusion Protein Features for CTDSP2-LGR5 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:58217687/chr12:71946853) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CTDSP2 | LGR5 |
| FUNCTION: Preferentially catalyzes the dephosphorylation of 'Ser-5' within the tandem 7 residue repeats in the C-terminal domain (CTD) of the largest RNA polymerase II subunit POLR2A. Negatively regulates RNA polymerase II transcription, possibly by controlling the transition from initiation/capping to processive transcript elongation. Recruited by REST to neuronal genes that contain RE-1 elements, leading to neuronal gene silencing in non-neuronal cells. May contribute to the development of sarcomas. {ECO:0000269|PubMed:12721286, ECO:0000269|PubMed:15681389}. | FUNCTION: Receptor for R-spondins that potentiates the canonical Wnt signaling pathway and acts as a stem cell marker of the intestinal epithelium and the hair follicle. Upon binding to R-spondins (RSPO1, RSPO2, RSPO3 or RSPO4), associates with phosphorylated LRP6 and frizzled receptors that are activated by extracellular Wnt receptors, triggering the canonical Wnt signaling pathway to increase expression of target genes. In contrast to classical G-protein coupled receptors, does not activate heterotrimeric G-proteins to transduce the signal. Involved in the development and/or maintenance of the adult intestinal stem cells during postembryonic development. {ECO:0000269|PubMed:21693646, ECO:0000269|PubMed:21727895, ECO:0000269|PubMed:21909076, ECO:0000269|PubMed:22815884, ECO:0000269|PubMed:23809763}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 163_184 | 142 | 908.0 | Repeat | Note=LRR 5 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 187_208 | 142 | 908.0 | Repeat | Note=LRR 6 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 211_232 | 142 | 908.0 | Repeat | Note=LRR 7 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 235_256 | 142 | 908.0 | Repeat | Note=LRR 8 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 258_279 | 142 | 908.0 | Repeat | Note=LRR 9 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 282_303 | 142 | 908.0 | Repeat | Note=LRR 10 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 306_328 | 142 | 908.0 | Repeat | Note=LRR 11 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 329_350 | 142 | 908.0 | Repeat | Note=LRR 12 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 353_374 | 142 | 908.0 | Repeat | Note=LRR 13 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 375_396 | 142 | 908.0 | Repeat | Note=LRR 14 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 399_420 | 142 | 908.0 | Repeat | Note=LRR 15 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 423_446 | 142 | 908.0 | Repeat | Note=LRR 16 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 163_184 | 142 | 884.0 | Repeat | Note=LRR 5 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 187_208 | 142 | 884.0 | Repeat | Note=LRR 6 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 211_232 | 142 | 884.0 | Repeat | Note=LRR 7 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 235_256 | 142 | 884.0 | Repeat | Note=LRR 8 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 258_279 | 142 | 884.0 | Repeat | Note=LRR 9 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 282_303 | 142 | 884.0 | Repeat | Note=LRR 10 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 306_328 | 142 | 884.0 | Repeat | Note=LRR 11 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 329_350 | 142 | 884.0 | Repeat | Note=LRR 12 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 353_374 | 142 | 884.0 | Repeat | Note=LRR 13 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 375_396 | 142 | 884.0 | Repeat | Note=LRR 14 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 399_420 | 142 | 884.0 | Repeat | Note=LRR 15 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 423_446 | 142 | 884.0 | Repeat | Note=LRR 16 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 583_593 | 142 | 908.0 | Topological domain | Cytoplasmic | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 615_638 | 142 | 908.0 | Topological domain | Extracellular | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 660_682 | 142 | 908.0 | Topological domain | Cytoplasmic | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 704_722 | 142 | 908.0 | Topological domain | Extracellular | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 744_767 | 142 | 908.0 | Topological domain | Cytoplasmic | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 789_802 | 142 | 908.0 | Topological domain | Extracellular | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 824_907 | 142 | 908.0 | Topological domain | Cytoplasmic | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 583_593 | 142 | 884.0 | Topological domain | Cytoplasmic | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 615_638 | 142 | 884.0 | Topological domain | Extracellular | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 660_682 | 142 | 884.0 | Topological domain | Cytoplasmic | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 704_722 | 142 | 884.0 | Topological domain | Extracellular | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 744_767 | 142 | 884.0 | Topological domain | Cytoplasmic | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 789_802 | 142 | 884.0 | Topological domain | Extracellular | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 824_907 | 142 | 884.0 | Topological domain | Cytoplasmic | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 562_582 | 142 | 908.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 594_614 | 142 | 908.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 639_659 | 142 | 908.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 683_703 | 142 | 908.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 723_743 | 142 | 908.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 768_788 | 142 | 908.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 803_823 | 142 | 908.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 562_582 | 142 | 884.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 594_614 | 142 | 884.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 639_659 | 142 | 884.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 683_703 | 142 | 884.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 723_743 | 142 | 884.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 768_788 | 142 | 884.0 | Transmembrane | Helical%3B Name%3D6 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 803_823 | 142 | 884.0 | Transmembrane | Helical%3B Name%3D7 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CTDSP2 | chr12:58217687 | chr12:71946853 | ENST00000398073 | - | 7 | 8 | 97_255 | 230 | 272.0 | Domain | FCP1 homology |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 25_66 | 142 | 908.0 | Domain | Note=LRRNT | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 25_66 | 142 | 884.0 | Domain | Note=LRRNT | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 115_136 | 142 | 908.0 | Repeat | Note=LRR 3 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 139_160 | 142 | 908.0 | Repeat | Note=LRR 4 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 67_90 | 142 | 908.0 | Repeat | Note=LRR 1 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 91_112 | 142 | 908.0 | Repeat | Note=LRR 2 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 115_136 | 142 | 884.0 | Repeat | Note=LRR 3 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 139_160 | 142 | 884.0 | Repeat | Note=LRR 4 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 67_90 | 142 | 884.0 | Repeat | Note=LRR 1 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 91_112 | 142 | 884.0 | Repeat | Note=LRR 2 | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000266674 | 3 | 18 | 22_561 | 142 | 908.0 | Topological domain | Extracellular | |
| Tgene | LGR5 | chr12:58217687 | chr12:71946853 | ENST00000540815 | 3 | 17 | 22_561 | 142 | 884.0 | Topological domain | Extracellular |
Top |
Fusion Gene Sequence for CTDSP2-LGR5 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >20175_20175_1_CTDSP2-LGR5_CTDSP2_chr12_58217687_ENST00000547701_LGR5_chr12_71946853_ENST00000266674_length(transcript)=4593nt_BP=721nt GAAAAAGTGCGAGTTCTCTGTGGGCAGCGGCTGGCGGCCCCCGCCTTCCAGAGCGTCAGGTGGAGGCGCGAGGGGGCCCGCAGGCCCTCG CGGGGGCCTGGTCTCCAAGTCCTCTCCTAAGAAGCCTCGTGGACGTAACATCTTCAAGGCCCTTTTCTGCTGTTTTCGCGCCCAGCATGT TGGCCAGTCAAGTTCCTCCACTGAGCTCGCTGCGTATAAGGAGGAAGCAAACACCATTGCTAAGTCGGATCTGCTCCAGTGTCTCCAGTA CCAGTTCTACCAGATCCCAGGGACCTGCCTGCTCCCAGAGGTGACAGAGGAAGATCAAGGAAGGATCTGTGTGGTCATTGACCTCGATGA AACCCTTGTGCATAGCTCCTTTAAGCCAATCAACAATGCTGACTTCATAGTGCCTATAGAGATTGAGGGGACCACTCACCAGGTGTATGT GCTCAAGAGGCCTTATGTGGATGAGTTCCTGAGACGCATGGGGGAACTCTTTGAATGTGTTCTCTTCACTGCCAGCCTGGCCAAGTATGC CGACCCTGTGACAGACCTGCTGGACCGGTGTGGGGTGTTCCGGGCCCGCCTATTCCGTGAGTCTTGCGTGTTCCACCAGGGCTGCTACGT CAAGGACCTCAGCCGCCTGGGGAGGGACCTGAGAAAGACCCTCATCCTGGACAACTCGCCTGCTTCTTACATATTCCACCCCGAGAATGC AGCGTCTGGATGCTAACCACATCAGCTATGTGCCCCCAAGCTGTTTCAGTGGCCTGCATTCCCTGAGGCACCTGTGGCTGGATGACAATG CGTTAACAGAAATCCCCGTCCAGGCTTTTAGAAGTTTATCGGCATTGCAAGCCATGACCTTGGCCCTGAACAAAATACACCACATACCAG ACTATGCCTTTGGAAACCTCTCCAGCTTGGTAGTTCTACATCTCCATAACAATAGAATCCACTCCCTGGGAAAGAAATGCTTTGATGGGC TCCACAGCCTAGAGACTTTAGATTTAAATTACAATAACCTTGATGAATTCCCCACTGCAATTAGGACACTCTCCAACCTTAAAGAACTAG GATTTCATAGCAACAATATCAGGTCGATACCTGAGAAAGCATTTGTAGGCAACCCTTCTCTTATTACAATACATTTCTATGACAATCCCA TCCAGTTTGTTGGGAGATCTGCTTTTCAACATTTACCTGAACTAAGAACACTGACTCTGAATGGTGCCTCACAAATAACTGAATTTCCTG ATTTAACTGGAACTGCAAACCTGGAGAGTCTGACTTTAACTGGAGCACAGATCTCATCTCTTCCTCAAACCGTCTGCAATCAGTTACCTA ATCTCCAAGTGCTAGATCTGTCTTACAACCTATTAGAAGATTTACCCAGTTTTTCAGTCTGCCAAAAGCTTCAGAAAATTGACCTAAGAC ATAATGAAATCTACGAAATTAAAGTTGACACTTTCCAGCAGTTGCTTAGCCTCCGATCGCTGAATTTGGCTTGGAACAAAATTGCTATTA TTCACCCCAATGCATTTTCCACTTTGCCATCCCTAATAAAGCTGGACCTATCGTCCAACCTCCTGTCGTCTTTTCCTATAACTGGGTTAC ATGGTTTAACTCACTTAAAATTAACAGGAAATCATGCCTTACAGAGCTTGATATCATCTGAAAACTTTCCAGAACTCAAGGTTATAGAAA TGCCTTATGCTTACCAGTGCTGTGCATTTGGAGTGTGTGAGAATGCCTATAAGATTTCTAATCAATGGAATAAAGGTGACAACAGCAGTA TGGACGACCTTCATAAGAAAGATGCTGGAATGTTTCAGGCTCAAGATGAACGTGACCTTGAAGATTTCCTGCTTGACTTTGAGGAAGACC TGAAAGCCCTTCATTCAGTGCAGTGTTCACCTTCCCCAGGCCCCTTCAAACCCTGTGAACACCTGCTTGATGGCTGGCTGATCAGAATTG GAGTGTGGACCATAGCAGTTCTGGCACTTACTTGTAATGCTTTGGTGACTTCAACAGTTTTCAGATCCCCTCTGTACATTTCCCCCATTA AACTGTTAATTGGGGTCATCGCAGCAGTGAACATGCTCACGGGAGTCTCCAGTGCCGTGCTGGCTGGTGTGGATGCGTTCACTTTTGGCA GCTTTGCACGACATGGTGCCTGGTGGGAGAATGGGGTTGGTTGCCATGTCATTGGTTTTTTGTCCATTTTTGCTTCAGAATCATCTGTTT TCCTGCTTACTCTGGCAGCCCTGGAGCGTGGGTTCTCTGTGAAATATTCTGCAAAATTTGAAACGAAAGCTCCATTTTCTAGCCTGAAAG TAATCATTTTGCTCTGTGCCCTGCTGGCCTTGACCATGGCCGCAGTTCCCCTGCTGGGTGGCAGCAAGTATGGCGCCTCCCCTCTCTGCC TGCCTTTGCCTTTTGGGGAGCCCAGCACCATGGGCTACATGGTCGCTCTCATCTTGCTCAATTCCCTTTGCTTCCTCATGATGACCATTG CCTACACCAAGCTCTACTGCAATTTGGACAAGGGAGACCTGGAGAATATTTGGGACTGCTCTATGGTAAAACACATTGCCCTGTTGCTCT TCACCAACTGCATCCTAAACTGCCCTGTGGCTTTCTTGTCCTTCTCCTCTTTAATAAACCTTACATTTATCAGTCCTGAAGTAATTAAGT TTATCCTTCTGGTGGTAGTCCCACTTCCTGCATGTCTCAATCCCCTTCTCTACATCTTGTTCAATCCTCACTTTAAGGAGGATCTGGTGA GCCTGAGAAAGCAAACCTACGTCTGGACAAGATCAAAACACCCAAGCTTGATGTCAATTAACTCTGATGATGTCGAAAAACAGTCCTGTG ACTCAACTCAAGCCTTGGTAACCTTTACCAGCTCCAGCATCACTTATGACCTGCCTCCCAGTTCCGTGCCATCACCAGCTTATCCAGTGA CTGAGAGCTGCCATCTTTCCTCTGTGGCATTTGTCCCATGTCTCTAATTAATATGTGAAGGAAAATGTTTTCAAAGGTTGAGAACCTGAA AATGTGAGATTGAGTATATCAGAGCAGTAATTAATAAGAAGAGCTGAGGTGAAACTCGGTTTAAAAACCAAAAAAGAATCTCTCAGTTAG TAAGAAAAGGCTGAAAACCTCTTGATACTTGAGAGTGAATATAAGTCTAAATGCTGCTTTGTATAATTTGTTCAGCTAAGGGATAGATCG ATCACACTATTTAAGTGAGCCCAGATCAAAAAAGCAGATTGAAATTTTCTTTAGAAAAGATTCTCCATGATTTGAATTGCATTCTCTTTA AACTCACCAATGTAATCATTTTGGGAGGAGGGAGAACCCACTTGCTTTCCAAATGGGTTTATTTAAACCCACAAACTCAAGAGGTTGTTG GGGGAATTAGGAAAATAAGGGTTTTCAATGACCTACATTGCTAGGTAGAGGCTGTGATCCATGGGATTTCATTCTAATGACCATGTGAAG ATGTTTGAGTCCTCCTTTGCCTTTCCTCAGAAAGAATCCTTCTAAGGCACAAATCCCTTAGATGGATAATGTAAGGTATTGTTAACTCAC TCATATTGAGATCATTTTTAGAGATACCAGGTTTTATGTATCAGCACTAGATGGTTCCACCCTCATGGGATAAAACTGCTTACAAGTATT TTGAAAGAAAAACTGACCAAAATTCTTAAATTGTTACTAAGGCAATCATGCACAGGTGACGTATGTCTTATCTGATTTGTTTTTAACTCC TTGGTGCCCAAAGCTCAGAAGGGAATTCCACTGCCAGCAATGAACATACCTGGAAAAGAAAGTAAGCAATCTGGGATTTTTTTTCTGGGT TAGTAAAGAATTTTTGCAATAAGTTTTATCAGTTGATTCAAACTGATGTGCATCTTAATGATCAAATGTGCACATTACATAAATTAAGTC CACTGATACAACTTCTTACACATGTATCTCTAGTAGCTCTGGCAAACCCAATATCTGACACCACTTTGGACTCAAGAGACTCAGTAACGT ATTATCCTGTTTATTTAGCTTGGTTTTAGCTGTGTTCTCTCTGGATAACCCACTTGATGTTAGGAACATTACTTCTCTGCTTATTCCATA TTAATACTGTGTTAGGTATTTTAAGAAGCAAGTTATTAAATAAGAAAAGTCAAAGTATTAATTCTTACCTTCTATTATCCTATATTAGCT TCAATACATCCAAACCAAATGGCTGTTAGGTAGATTTATTTTTATATAAGCATGTTTATTTTGATCAGATGTTTTAACTTGGATTTGAAA AAATACATTTATGAGATGTTTTATAAGATGTGTAAATATAGAACTGTATTTATTACTATAGTAAAGGTTCAGTAACATTAAGGACCATGA TAATGATAATAAACCTTGTACAGTGGCATATTCTTTGATTTATATTGTGTTTCTCTGCCCATTTTCTTTAAATTCATTAACTGTATATAT GTAAATATATAGTACTTGTAAATAGATTCCAAATTTGCTTTTCTATTGGGTAAAAAATAAATTTGTAATAAAATGTGTGACTATGAAACA >20175_20175_1_CTDSP2-LGR5_CTDSP2_chr12_58217687_ENST00000547701_LGR5_chr12_71946853_ENST00000266674_length(amino acids)=775AA_BP=11 MLLTYSTPRMQRLDANHISYVPPSCFSGLHSLRHLWLDDNALTEIPVQAFRSLSALQAMTLALNKIHHIPDYAFGNLSSLVVLHLHNNRI HSLGKKCFDGLHSLETLDLNYNNLDEFPTAIRTLSNLKELGFHSNNIRSIPEKAFVGNPSLITIHFYDNPIQFVGRSAFQHLPELRTLTL NGASQITEFPDLTGTANLESLTLTGAQISSLPQTVCNQLPNLQVLDLSYNLLEDLPSFSVCQKLQKIDLRHNEIYEIKVDTFQQLLSLRS LNLAWNKIAIIHPNAFSTLPSLIKLDLSSNLLSSFPITGLHGLTHLKLTGNHALQSLISSENFPELKVIEMPYAYQCCAFGVCENAYKIS NQWNKGDNSSMDDLHKKDAGMFQAQDERDLEDFLLDFEEDLKALHSVQCSPSPGPFKPCEHLLDGWLIRIGVWTIAVLALTCNALVTSTV FRSPLYISPIKLLIGVIAAVNMLTGVSSAVLAGVDAFTFGSFARHGAWWENGVGCHVIGFLSIFASESSVFLLTLAALERGFSVKYSAKF ETKAPFSSLKVIILLCALLALTMAAVPLLGGSKYGASPLCLPLPFGEPSTMGYMVALILLNSLCFLMMTIAYTKLYCNLDKGDLENIWDC SMVKHIALLLFTNCILNCPVAFLSFSSLINLTFISPEVIKFILLVVVPLPACLNPLLYILFNPHFKEDLVSLRKQTYVWTRSKHPSLMSI -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CTDSP2-LGR5 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CTDSP2-LGR5 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CTDSP2-LGR5 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
