
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CTNNB1-CHEK1 (FusionGDB2 ID:20315) |
Fusion Gene Summary for CTNNB1-CHEK1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CTNNB1-CHEK1 | Fusion gene ID: 20315 | Hgene | Tgene | Gene symbol | CTNNB1 | CHEK1 | Gene ID | 1499 | 1111 |
| Gene name | catenin beta 1 | checkpoint kinase 1 | |
| Synonyms | CTNNB|EVR7|MRD19|NEDSDV|armadillo | CHK1 | |
| Cytomap | 3p22.1 | 11q24.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | catenin beta-1catenin (cadherin-associated protein), beta 1, 88kDa | serine/threonine-protein kinase Chk1CHK1 checkpoint homologCheckpoint, S. pombe, homolog of, 1Chk1-Scell cycle checkpoint kinase | |
| Modification date | 20200327 | 20200313 | |
| UniProtAcc | P35222 | O14757 | |
| Ensembl transtripts involved in fusion gene | ENST00000349496, ENST00000396183, ENST00000396185, ENST00000453024, ENST00000405570, ENST00000471014, | ENST00000532449, ENST00000278916, ENST00000427383, ENST00000428830, ENST00000438015, ENST00000524737, ENST00000534070, ENST00000544373, | |
| Fusion gene scores | * DoF score | 19 X 14 X 13=3458 | 3 X 3 X 3=27 |
| # samples | 21 | 3 | |
| ** MAII score | log2(21/3458*10)=-4.04147663597616 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(3/27*10)=0.15200309344505 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: CTNNB1 [Title/Abstract] AND CHEK1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CTNNB1(41241161)-CHEK1(125496643), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | CTNNB1 | GO:0000209 | protein polyubiquitination | 29374064 |
| Hgene | CTNNB1 | GO:0008285 | negative regulation of cell proliferation | 12970740 |
| Hgene | CTNNB1 | GO:0030997 | regulation of centriole-centriole cohesion | 18086858 |
| Hgene | CTNNB1 | GO:0032355 | response to estradiol | 15304487 |
| Hgene | CTNNB1 | GO:0033234 | negative regulation of protein sumoylation | 22155184 |
| Hgene | CTNNB1 | GO:0043065 | positive regulation of apoptotic process | 12651860|12970740 |
| Hgene | CTNNB1 | GO:0043161 | proteasome-mediated ubiquitin-dependent protein catabolic process | 29374064 |
| Hgene | CTNNB1 | GO:0043525 | positive regulation of neuron apoptotic process | 19591802 |
| Hgene | CTNNB1 | GO:0045893 | positive regulation of transcription, DNA-templated | 12970740|18787224 |
| Hgene | CTNNB1 | GO:0045944 | positive regulation of transcription by RNA polymerase II | 9065402|11751639|12651860|14660579|18193033 |
| Hgene | CTNNB1 | GO:0060070 | canonical Wnt signaling pathway | 10644691|12937339|19187541 |
| Hgene | CTNNB1 | GO:0071681 | cellular response to indole-3-methanol | 10868478 |
| Hgene | CTNNB1 | GO:0090279 | regulation of calcium ion import | 19996314 |
| Hgene | CTNNB1 | GO:1904798 | positive regulation of core promoter binding | 22723415 |
| Hgene | CTNNB1 | GO:2000008 | regulation of protein localization to cell surface | 19996314 |
| Tgene | CHEK1 | GO:0000077 | DNA damage checkpoint | 16963448 |
| Tgene | CHEK1 | GO:0006915 | apoptotic process | 23028632 |
| Tgene | CHEK1 | GO:0006975 | DNA damage induced protein phosphorylation | 16963448 |
| Tgene | CHEK1 | GO:0010569 | regulation of double-strand break repair via homologous recombination | 15665856 |
| Tgene | CHEK1 | GO:0018107 | peptidyl-threonine phosphorylation | 15665856 |
| Tgene | CHEK1 | GO:0045787 | positive regulation of cell cycle | 26296656 |
| Tgene | CHEK1 | GO:0045839 | negative regulation of mitotic nuclear division | 15311285 |
| Tgene | CHEK1 | GO:0046602 | regulation of mitotic centrosome separation | 15311285 |
 Fusion gene breakpoints across CTNNB1 (5'-gene) Fusion gene breakpoints across CTNNB1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Fusion gene breakpoints across CHEK1 (3'-gene) Fusion gene breakpoints across CHEK1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | ESCA | TCGA-L5-A4OW | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
Top |
Fusion Gene ORF analysis for CTNNB1-CHEK1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5UTR-3UTR | ENST00000349496 | ENST00000532449 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-3UTR | ENST00000396183 | ENST00000532449 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-3UTR | ENST00000396185 | ENST00000532449 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-3UTR | ENST00000453024 | ENST00000532449 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000349496 | ENST00000278916 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000349496 | ENST00000427383 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000349496 | ENST00000428830 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000349496 | ENST00000438015 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000349496 | ENST00000524737 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000349496 | ENST00000534070 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000349496 | ENST00000544373 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396183 | ENST00000278916 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396183 | ENST00000427383 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396183 | ENST00000428830 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396183 | ENST00000438015 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396183 | ENST00000524737 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396183 | ENST00000534070 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396183 | ENST00000544373 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396185 | ENST00000278916 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396185 | ENST00000427383 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396185 | ENST00000428830 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396185 | ENST00000438015 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396185 | ENST00000524737 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396185 | ENST00000534070 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000396185 | ENST00000544373 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000453024 | ENST00000278916 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000453024 | ENST00000427383 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000453024 | ENST00000428830 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000453024 | ENST00000438015 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000453024 | ENST00000524737 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000453024 | ENST00000534070 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| 5UTR-5UTR | ENST00000453024 | ENST00000544373 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-3UTR | ENST00000405570 | ENST00000532449 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-3UTR | ENST00000471014 | ENST00000532449 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000405570 | ENST00000278916 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000405570 | ENST00000427383 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000405570 | ENST00000428830 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000405570 | ENST00000438015 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000405570 | ENST00000524737 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000405570 | ENST00000534070 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000405570 | ENST00000544373 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000471014 | ENST00000278916 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000471014 | ENST00000427383 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000471014 | ENST00000428830 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000471014 | ENST00000438015 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000471014 | ENST00000524737 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000471014 | ENST00000534070 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
| intron-5UTR | ENST00000471014 | ENST00000544373 | CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496643 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
Top |
Fusion Genomic Features for CTNNB1-CHEK1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496642 | + | 3.66E-06 | 0.9999963 |
| CTNNB1 | chr3 | 41241161 | + | CHEK1 | chr11 | 125496642 | + | 3.66E-06 | 0.9999963 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
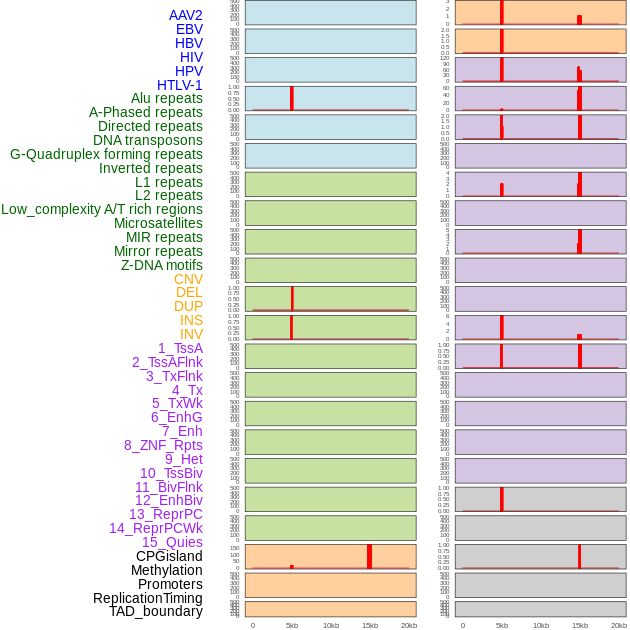 |
Top |
Fusion Protein Features for CTNNB1-CHEK1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/:41241161/:125496643) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CTNNB1 | CHEK1 |
| FUNCTION: Key downstream component of the canonical Wnt signaling pathway (PubMed:17524503, PubMed:18077326, PubMed:18086858, PubMed:18957423, PubMed:21262353, PubMed:22155184, PubMed:22647378, PubMed:22699938). In the absence of Wnt, forms a complex with AXIN1, AXIN2, APC, CSNK1A1 and GSK3B that promotes phosphorylation on N-terminal Ser and Thr residues and ubiquitination of CTNNB1 via BTRC and its subsequent degradation by the proteasome (PubMed:17524503, PubMed:18077326, PubMed:18086858, PubMed:18957423, PubMed:21262353, PubMed:22155184, PubMed:22647378, PubMed:22699938). In the presence of Wnt ligand, CTNNB1 is not ubiquitinated and accumulates in the nucleus, where it acts as a coactivator for transcription factors of the TCF/LEF family, leading to activate Wnt responsive genes (PubMed:17524503, PubMed:18077326, PubMed:18086858, PubMed:18957423, PubMed:21262353, PubMed:22155184, PubMed:22647378, PubMed:22699938). Involved in the regulation of cell adhesion, as component of an E-cadherin:catenin adhesion complex (By similarity). Acts as a negative regulator of centrosome cohesion (PubMed:18086858). Involved in the CDK2/PTPN6/CTNNB1/CEACAM1 pathway of insulin internalization (PubMed:21262353). Blocks anoikis of malignant kidney and intestinal epithelial cells and promotes their anchorage-independent growth by down-regulating DAPK2 (PubMed:18957423). Disrupts PML function and PML-NB formation by inhibiting RANBP2-mediated sumoylation of PML (PubMed:22155184). Promotes neurogenesis by maintaining sympathetic neuroblasts within the cell cycle (By similarity). Involved in chondrocyte differentiation via interaction with SOX9: SOX9-binding competes with the binding sites of TCF/LEF within CTNNB1, thereby inhibiting the Wnt signaling (By similarity). {ECO:0000250|UniProtKB:Q02248, ECO:0000269|PubMed:17524503, ECO:0000269|PubMed:18077326, ECO:0000269|PubMed:18086858, ECO:0000269|PubMed:18957423, ECO:0000269|PubMed:21262353, ECO:0000269|PubMed:22155184, ECO:0000269|PubMed:22647378, ECO:0000269|PubMed:22699938}. | FUNCTION: Serine/threonine-protein kinase which is required for checkpoint-mediated cell cycle arrest and activation of DNA repair in response to the presence of DNA damage or unreplicated DNA (PubMed:11535615, PubMed:12446774, PubMed:12399544, PubMed:14559997, PubMed:14988723, PubMed:15311285, PubMed:15665856, PubMed:15650047). May also negatively regulate cell cycle progression during unperturbed cell cycles (PubMed:11535615, PubMed:12446774, PubMed:12399544, PubMed:14559997, PubMed:14988723, PubMed:15311285, PubMed:15665856, PubMed:15650047). This regulation is achieved by a number of mechanisms that together help to preserve the integrity of the genome (PubMed:11535615, PubMed:12446774, PubMed:12399544, PubMed:14559997, PubMed:14988723, PubMed:15311285, PubMed:15665856, PubMed:15650047). Recognizes the substrate consensus sequence [R-X-X-S/T] (PubMed:11535615, PubMed:12446774, PubMed:12399544, PubMed:14559997, PubMed:14988723, PubMed:15311285, PubMed:15665856, PubMed:15650047). Binds to and phosphorylates CDC25A, CDC25B and CDC25C (PubMed:9278511, PubMed:12676583, PubMed:14681206, PubMed:12676925, PubMed:12759351, PubMed:19734889, PubMed:14559997). Phosphorylation of CDC25A at 'Ser-178' and 'Thr-507' and phosphorylation of CDC25C at 'Ser-216' creates binding sites for 14-3-3 proteins which inhibit CDC25A and CDC25C (PubMed:9278511). Phosphorylation of CDC25A at 'Ser-76', 'Ser-124', 'Ser-178', 'Ser-279' and 'Ser-293' promotes proteolysis of CDC25A (PubMed:9278511, PubMed:12676583, PubMed:14681206, PubMed:12676925, PubMed:12759351, PubMed:19734889). Phosphorylation of CDC25A at 'Ser-76' primes the protein for subsequent phosphorylation at 'Ser-79', 'Ser-82' and 'Ser-88' by NEK11, which is required for polyubiquitination and degradation of CDCD25A (PubMed:9278511, PubMed:19734889, PubMed:20090422). Inhibition of CDC25 leads to increased inhibitory tyrosine phosphorylation of CDK-cyclin complexes and blocks cell cycle progression (PubMed:9278511). Also phosphorylates NEK6 (PubMed:18728393). Binds to and phosphorylates RAD51 at 'Thr-309', which promotes the release of RAD51 from BRCA2 and enhances the association of RAD51 with chromatin, thereby promoting DNA repair by homologous recombination (PubMed:15665856). Phosphorylates multiple sites within the C-terminus of TP53, which promotes activation of TP53 by acetylation and promotes cell cycle arrest and suppression of cellular proliferation (PubMed:10673501, PubMed:15659650, PubMed:16511572). Also promotes repair of DNA cross-links through phosphorylation of FANCE (PubMed:17296736). Binds to and phosphorylates TLK1 at 'Ser-743', which prevents the TLK1-dependent phosphorylation of the chromatin assembly factor ASF1A (PubMed:12660173, PubMed:12955071). This may enhance chromatin assembly both in the presence or absence of DNA damage (PubMed:12660173, PubMed:12955071). May also play a role in replication fork maintenance through regulation of PCNA (PubMed:18451105). May regulate the transcription of genes that regulate cell-cycle progression through the phosphorylation of histones (By similarity). Phosphorylates histone H3.1 (to form H3T11ph), which leads to epigenetic inhibition of a subset of genes (By similarity). May also phosphorylate RB1 to promote its interaction with the E2F family of transcription factors and subsequent cell cycle arrest (PubMed:17380128). Phosphorylates SPRTN, promoting SPRTN recruitment to chromatin (PubMed:31316063). Reduces replication stress and activates the G2/M checkpoint, by phosphorylating and inactivating PABIR1/FAM122A and promoting the serine/threonine-protein phosphatase 2A-mediated dephosphorylation and stabilization of WEE1 levels and activity (PubMed:33108758). {ECO:0000250|UniProtKB:O35280, ECO:0000269|PubMed:10673501, ECO:0000269|PubMed:11535615, ECO:0000269|PubMed:12399544, ECO:0000269|PubMed:12446774, ECO:0000269|PubMed:12660173, ECO:0000269|PubMed:12676583, ECO:0000269|PubMed:12676925, ECO:0000269|PubMed:12759351, ECO:0000269|PubMed:12955071, ECO:0000269|PubMed:14559997, ECO:0000269|PubMed:14681206, ECO:0000269|PubMed:14988723, ECO:0000269|PubMed:15311285, ECO:0000269|PubMed:15650047, ECO:0000269|PubMed:15659650, ECO:0000269|PubMed:15665856, ECO:0000269|PubMed:16511572, ECO:0000269|PubMed:17296736, ECO:0000269|PubMed:17380128, ECO:0000269|PubMed:18451105, ECO:0000269|PubMed:18728393, ECO:0000269|PubMed:19734889, ECO:0000269|PubMed:20090422, ECO:0000269|PubMed:31316063, ECO:0000269|PubMed:33108758, ECO:0000269|PubMed:9278511}.; FUNCTION: [Isoform 2]: Endogenous repressor of isoform 1, interacts with, and antagonizes CHK1 to promote the S to G2/M phase transition. {ECO:0000269|PubMed:22184239}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Gene Sequence for CTNNB1-CHEK1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
Top |
Fusion Gene PPI Analysis for CTNNB1-CHEK1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CTNNB1-CHEK1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CTNNB1-CHEK1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
