
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CYLD-HAGH (FusionGDB2 ID:20992) |
Fusion Gene Summary for CYLD-HAGH |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CYLD-HAGH | Fusion gene ID: 20992 | Hgene | Tgene | Gene symbol | CYLD | HAGH | Gene ID | 1540 | 3029 |
| Gene name | CYLD lysine 63 deubiquitinase | hydroxyacylglutathione hydrolase | |
| Synonyms | BRSS|CDMT|CYLD1|CYLDI|EAC|MFT|MFT1|SBS|TEM|USPL2 | GLO2|GLX2|GLXII|HAGH1 | |
| Cytomap | 16q12.1 | 16p13.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ubiquitin carboxyl-terminal hydrolase CYLDcylindromatosis (turban tumor syndrome)deubiquitinating enzyme CYLDprobable ubiquitin carboxyl-terminal hydrolase CYLDubiquitin specific peptidase like 2ubiquitin thioesterase CYLDubiquitin thiolesterase CYL | hydroxyacylglutathione hydrolase, mitochondrialglyoxalase IIhydroxyacylglutathione hydroxylase | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | Q9NQC7 | Q6PII5 | |
| Ensembl transtripts involved in fusion gene | ENST00000311559, ENST00000398568, ENST00000427738, ENST00000540145, ENST00000564326, ENST00000566206, ENST00000568704, ENST00000569418, | ENST00000566709, ENST00000567398, ENST00000397353, ENST00000397356, ENST00000455446, | |
| Fusion gene scores | * DoF score | 6 X 4 X 4=96 | 4 X 2 X 4=32 |
| # samples | 6 | 4 | |
| ** MAII score | log2(6/96*10)=-0.678071905112638 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(4/32*10)=0.321928094887362 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: CYLD [Title/Abstract] AND HAGH [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CYLD(50788335)-HAGH(1859834), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | CYLD-HAGH seems lost the major protein functional domain in Hgene partner, which is a CGC due to the frame-shifted ORF. CYLD-HAGH seems lost the major protein functional domain in Hgene partner, which is a tumor suppressor due to the frame-shifted ORF. CYLD-HAGH seems lost the major protein functional domain in Tgene partner, which is a cell metabolism gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | CYLD | GO:0010803 | regulation of tumor necrosis factor-mediated signaling pathway | 26997266|27458237|27591049 |
| Hgene | CYLD | GO:0016579 | protein deubiquitination | 29291351 |
| Hgene | CYLD | GO:0032088 | negative regulation of NF-kappaB transcription factor activity | 18313383 |
| Hgene | CYLD | GO:0045087 | innate immune response | 26997266 |
| Hgene | CYLD | GO:0046329 | negative regulation of JNK cascade | 29291351 |
| Hgene | CYLD | GO:0050727 | regulation of inflammatory response | 27591049 |
| Hgene | CYLD | GO:0060544 | regulation of necroptotic process | 27458237 |
| Hgene | CYLD | GO:0070536 | protein K63-linked deubiquitination | 18313383|18636086|26997266|27458237|27591049|29291351 |
| Hgene | CYLD | GO:1901223 | negative regulation of NIK/NF-kappaB signaling | 18313383 |
| Hgene | CYLD | GO:1903753 | negative regulation of p38MAPK cascade | 29291351 |
| Hgene | CYLD | GO:1990108 | protein linear deubiquitination | 26997266|27458237|27591049 |
| Tgene | HAGH | GO:0006750 | glutathione biosynthetic process | 8550579 |
 Fusion gene breakpoints across CYLD (5'-gene) Fusion gene breakpoints across CYLD (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
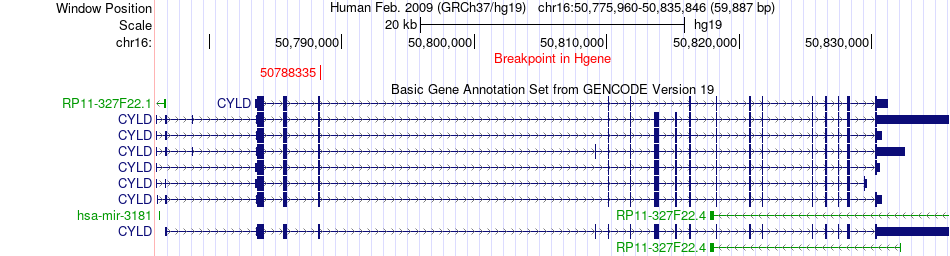 |
 Fusion gene breakpoints across HAGH (3'-gene) Fusion gene breakpoints across HAGH (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
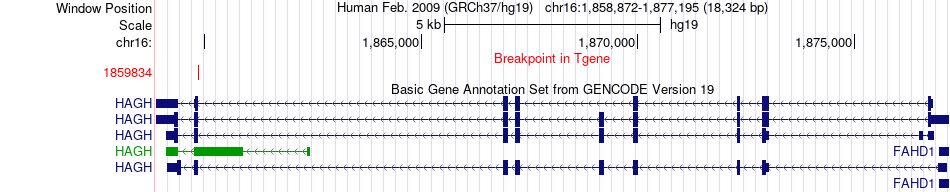 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | GBM | TCGA-14-0736-02A | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
Top |
Fusion Gene ORF analysis for CYLD-HAGH |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000311559 | ENST00000566709 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000311559 | ENST00000567398 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000398568 | ENST00000566709 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000398568 | ENST00000567398 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000427738 | ENST00000566709 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000427738 | ENST00000567398 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000540145 | ENST00000566709 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000540145 | ENST00000567398 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000564326 | ENST00000566709 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000564326 | ENST00000567398 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000566206 | ENST00000566709 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000566206 | ENST00000567398 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000568704 | ENST00000566709 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000568704 | ENST00000567398 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000569418 | ENST00000566709 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| 5CDS-5UTR | ENST00000569418 | ENST00000567398 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000311559 | ENST00000397353 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000311559 | ENST00000397356 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000398568 | ENST00000397353 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000398568 | ENST00000397356 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000427738 | ENST00000397353 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000427738 | ENST00000397356 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000540145 | ENST00000397353 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000540145 | ENST00000397356 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000564326 | ENST00000397353 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000564326 | ENST00000397356 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000566206 | ENST00000397353 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000566206 | ENST00000397356 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000568704 | ENST00000397353 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000568704 | ENST00000397356 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000569418 | ENST00000397353 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| Frame-shift | ENST00000569418 | ENST00000397356 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| In-frame | ENST00000311559 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| In-frame | ENST00000398568 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| In-frame | ENST00000427738 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| In-frame | ENST00000540145 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| In-frame | ENST00000564326 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| In-frame | ENST00000566206 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| In-frame | ENST00000568704 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
| In-frame | ENST00000569418 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000569418 | CYLD | chr16 | 50788335 | + | ENST00000455446 | HAGH | chr16 | 1859834 | - | 1782 | 1191 | 278 | 1300 | 340 |
| ENST00000540145 | CYLD | chr16 | 50788335 | + | ENST00000455446 | HAGH | chr16 | 1859834 | - | 1919 | 1328 | 415 | 1437 | 340 |
| ENST00000311559 | CYLD | chr16 | 50788335 | + | ENST00000455446 | HAGH | chr16 | 1859834 | - | 1895 | 1304 | 391 | 1413 | 340 |
| ENST00000564326 | CYLD | chr16 | 50788335 | + | ENST00000455446 | HAGH | chr16 | 1859834 | - | 1698 | 1107 | 194 | 1216 | 340 |
| ENST00000566206 | CYLD | chr16 | 50788335 | + | ENST00000455446 | HAGH | chr16 | 1859834 | - | 1754 | 1163 | 250 | 1272 | 340 |
| ENST00000398568 | CYLD | chr16 | 50788335 | + | ENST00000455446 | HAGH | chr16 | 1859834 | - | 1804 | 1213 | 300 | 1322 | 340 |
| ENST00000427738 | CYLD | chr16 | 50788335 | + | ENST00000455446 | HAGH | chr16 | 1859834 | - | 1709 | 1118 | 205 | 1227 | 340 |
| ENST00000568704 | CYLD | chr16 | 50788335 | + | ENST00000455446 | HAGH | chr16 | 1859834 | - | 1654 | 1063 | 150 | 1172 | 340 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000569418 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - | 0.000356486 | 0.99964356 |
| ENST00000540145 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - | 0.000215028 | 0.999785 |
| ENST00000311559 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - | 0.000212641 | 0.9997874 |
| ENST00000564326 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - | 0.000403401 | 0.99959666 |
| ENST00000566206 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - | 0.000362295 | 0.9996377 |
| ENST00000398568 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - | 0.000342777 | 0.9996573 |
| ENST00000427738 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - | 0.000357053 | 0.99964297 |
| ENST00000568704 | ENST00000455446 | CYLD | chr16 | 50788335 | + | HAGH | chr16 | 1859834 | - | 0.000379031 | 0.9996209 |
Top |
Fusion Genomic Features for CYLD-HAGH |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
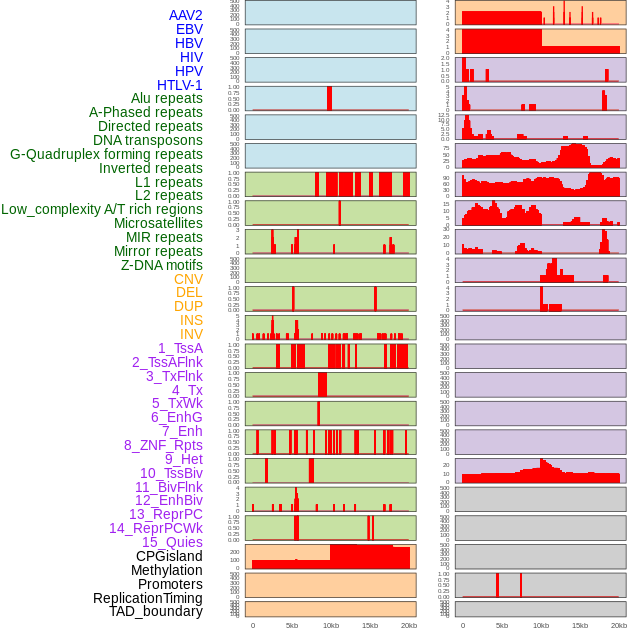 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for CYLD-HAGH |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr16:50788335/chr16:1859834) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CYLD | HAGH |
| FUNCTION: Deubiquitinase that specifically cleaves 'Lys-63'- and linear 'Met-1'-linked polyubiquitin chains and is involved in NF-kappa-B activation and TNF-alpha-induced necroptosis (PubMed:18636086, PubMed:26670046, PubMed:27458237, PubMed:26997266, PubMed:27591049, PubMed:29291351, PubMed:18313383). Plays an important role in the regulation of pathways leading to NF-kappa-B activation (PubMed:12917689, PubMed:12917691). Contributes to the regulation of cell survival, proliferation and differentiation via its effects on NF-kappa-B activation (PubMed:12917690). Negative regulator of Wnt signaling (PubMed:20227366). Inhibits HDAC6 and thereby promotes acetylation of alpha-tubulin and stabilization of microtubules (PubMed:19893491). Plays a role in the regulation of microtubule dynamics, and thereby contributes to the regulation of cell proliferation, cell polarization, cell migration, and angiogenesis (PubMed:18222923, PubMed:20194890). Required for normal cell cycle progress and normal cytokinesis (PubMed:17495026, PubMed:19893491). Inhibits nuclear translocation of NF-kappa-B (PubMed:18636086). Plays a role in the regulation of inflammation and the innate immune response, via its effects on NF-kappa-B activation (PubMed:18636086). Dispensable for the maturation of intrathymic natural killer cells, but required for the continued survival of immature natural killer cells (By similarity). Negatively regulates TNFRSF11A signaling and osteoclastogenesis (By similarity). Involved in the regulation of ciliogenesis, allowing ciliary basal bodies to migrate and dock to the plasma membrane; this process does not depend on NF-kappa-B activation (By similarity). Ability to remove linear ('Met-1'-linked) polyubiquitin chains regulates innate immunity and TNF-alpha-induced necroptosis: recruited to the LUBAC complex via interaction with SPATA2 and restricts linear polyubiquitin formation on target proteins (PubMed:26997266, PubMed:26670046, PubMed:27458237, PubMed:27591049). Regulates innate immunity by restricting linear polyubiquitin formation on RIPK2 in response to NOD2 stimulation (PubMed:26997266). Involved in TNF-alpha-induced necroptosis by removing linear ('Met-1'-linked) polyubiquitin chains from RIPK1, thereby regulating the kinase activity of RIPK1 (By similarity). Removes 'Lys-63' linked polyubiquitin chain of MAP3K7, which inhibits phosphorylation and blocks downstream activation of the JNK-p38 kinase cascades (PubMed:29291351). {ECO:0000250|UniProtKB:Q80TQ2, ECO:0000269|PubMed:12917689, ECO:0000269|PubMed:12917690, ECO:0000269|PubMed:12917691, ECO:0000269|PubMed:17495026, ECO:0000269|PubMed:18222923, ECO:0000269|PubMed:18313383, ECO:0000269|PubMed:18636086, ECO:0000269|PubMed:19893491, ECO:0000269|PubMed:20194890, ECO:0000269|PubMed:20227366, ECO:0000269|PubMed:26670046, ECO:0000269|PubMed:26997266, ECO:0000269|PubMed:27458237, ECO:0000269|PubMed:27591049, ECO:0000269|PubMed:29291351}. | FUNCTION: Hydrolase acting on ester bonds. {ECO:0000305}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000311559 | + | 6 | 20 | 153_198 | 304 | 957.0 | Domain | CAP-Gly 1 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000311559 | + | 6 | 20 | 253_286 | 304 | 957.0 | Domain | CAP-Gly 2 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000398568 | + | 5 | 18 | 153_198 | 304 | 954.0 | Domain | CAP-Gly 1 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000398568 | + | 5 | 18 | 253_286 | 304 | 954.0 | Domain | CAP-Gly 2 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000427738 | + | 4 | 18 | 153_198 | 304 | 957.0 | Domain | CAP-Gly 1 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000427738 | + | 4 | 18 | 253_286 | 304 | 957.0 | Domain | CAP-Gly 2 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000564326 | + | 4 | 17 | 153_198 | 304 | 954.0 | Domain | CAP-Gly 1 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000564326 | + | 4 | 17 | 253_286 | 304 | 954.0 | Domain | CAP-Gly 2 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000569418 | + | 5 | 18 | 153_198 | 304 | 954.0 | Domain | CAP-Gly 1 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000569418 | + | 5 | 18 | 253_286 | 304 | 954.0 | Domain | CAP-Gly 2 |
| Tgene | HAGH | chr16:50788335 | chr16:1859834 | ENST00000397353 | 7 | 10 | 221_223 | 201 | 261.0 | Region | Substrate binding | |
| Tgene | HAGH | chr16:50788335 | chr16:1859834 | ENST00000397353 | 7 | 10 | 297_300 | 201 | 261.0 | Region | Substrate binding | |
| Tgene | HAGH | chr16:50788335 | chr16:1859834 | ENST00000397356 | 6 | 9 | 297_300 | 249 | 309.0 | Region | Substrate binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000311559 | + | 6 | 20 | 492_535 | 304 | 957.0 | Domain | CAP-Gly 3 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000311559 | + | 6 | 20 | 592_950 | 304 | 957.0 | Domain | Note=USP |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000398568 | + | 5 | 18 | 492_535 | 304 | 954.0 | Domain | CAP-Gly 3 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000398568 | + | 5 | 18 | 592_950 | 304 | 954.0 | Domain | Note=USP |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000427738 | + | 4 | 18 | 492_535 | 304 | 957.0 | Domain | CAP-Gly 3 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000427738 | + | 4 | 18 | 592_950 | 304 | 957.0 | Domain | Note=USP |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000564326 | + | 4 | 17 | 492_535 | 304 | 954.0 | Domain | CAP-Gly 3 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000564326 | + | 4 | 17 | 592_950 | 304 | 954.0 | Domain | Note=USP |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000569418 | + | 5 | 18 | 492_535 | 304 | 954.0 | Domain | CAP-Gly 3 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000569418 | + | 5 | 18 | 592_950 | 304 | 954.0 | Domain | Note=USP |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000311559 | + | 6 | 20 | 781_833 | 304 | 957.0 | Region | B-box |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000398568 | + | 5 | 18 | 781_833 | 304 | 954.0 | Region | B-box |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000427738 | + | 4 | 18 | 781_833 | 304 | 957.0 | Region | B-box |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000564326 | + | 4 | 17 | 781_833 | 304 | 954.0 | Region | B-box |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000569418 | + | 5 | 18 | 781_833 | 304 | 954.0 | Region | B-box |
| Tgene | HAGH | chr16:50788335 | chr16:1859834 | ENST00000397353 | 7 | 10 | 191_193 | 201 | 261.0 | Region | Substrate binding | |
| Tgene | HAGH | chr16:50788335 | chr16:1859834 | ENST00000397356 | 6 | 9 | 191_193 | 249 | 309.0 | Region | Substrate binding | |
| Tgene | HAGH | chr16:50788335 | chr16:1859834 | ENST00000397356 | 6 | 9 | 221_223 | 249 | 309.0 | Region | Substrate binding |
Top |
Fusion Gene Sequence for CYLD-HAGH |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >20992_20992_1_CYLD-HAGH_CYLD_chr16_50788335_ENST00000311559_HAGH_chr16_1859834_ENST00000455446_length(transcript)=1895nt_BP=1304nt GGGGGCGGGCCCAGGTAGCAGGTTTGGCTGCGCGGGGGCCGCGCGTCGGAGTTTCCCCCTTTCTAGGGTGAGGATGGTTCTACACAGCCA CCCGGAGTTCCTTAGTTGAAAGGTGCGCCCTGCTGTGACAGAATGTGGTAATTGTAATCTTTAACATTTTCATGTAAAACATATTTCCTG ATCATCTTTCCATTGTCTTCATGGAAAATTGATAAATATTTGTGCCTTCCAACTCTCGTCTTGGTTGAATGACTTCATCTTAATACAACA TGGACACCACGTTGCTGAAAACATGCTTTGGGACTGCCACTGAATTTATCTTTTGCGGTTTTATGACAAAGTTATTAGTAGTTTCCCTTT TTTGAATTAGTATTTTGAAGTTAATATCACAATGAGTTCAGGCTTATGGAGCCAAGAAAAAGTCACTTCACCCTACTGGGAAGAGCGGAT TTTTTACTTGCTTCTTCAAGAATGCAGCGTTACAGACAAACAAACACAAAAGCTCCTTAAAGTACCGAAGGGAAGTATAGGACAGTATAT TCAAGATCGTTCTGTGGGGCATTCAAGGATTCCTTCTGCAAAAGGCAAGAAAAATCAGATTGGATTAAAAATTCTAGAGCAACCTCATGC AGTTCTCTTTGTTGATGAAAAGGATGTTGTAGAGATAAATGAAAAGTTCACAGAGTTACTTTTGGCAATTACCAATTGTGAGGAGAGGTT CAGCCTGTTTAAAAACAGAAACAGACTAAGTAAAGGCCTCCAAATAGACGTGGGCTGTCCTGTGAAAGTACAGCTGAGATCTGGGGAAGA AAAATTTCCTGGAGTTGTACGCTTCAGAGGACCCCTGTTAGCAGAGAGGACAGTCTCCGGAATATTCTTTGGAGTTGAATTGCTGGAAGA AGGTCGTGGTCAAGGTTTCACTGACGGGGTGTACCAAGGGAAACAGCTTTTTCAGTGTGATGAAGATTGTGGCGTGTTTGTTGCATTGGA CAAGCTAGAACTCATAGAAGATGATGACACTGCATTGGAAAGTGATTACGCAGGTCCTGGGGACACAATGCAGGTCGAACTTCCTCCTTT GGAAATAAACTCCAGAGTTTCTTTGAAGGTTGGAGAAACAATAGAATCTGGAACAGTTATATTCTGTGATGTTTTGCCAGGAAAAGAAAG CTTAGGATATTTTGTTGGTGTGGACATGGATAACCCTATTGGCAACTGGGATGGAAGATTTGATGGAGTGCAGCTTTGTAGTTTTGCGTG TGTTGAAAGTACAATTCTATTGCACATCAATGATATCATCCCAGGAGAAGTACAGCATCGGGGAGCCCACAGTGCCATCCACCCTGGCAG AGGAGTTTACCTACAACCCCTTCATGAGAGTGAGGGAGAAGACGGTGCAGCAGCACGCAGGTGAGACGGACCCGGTGACCACCATGCGGG CCGTGCGCAGGGAGAAGGACCAGTTCAAGATGCCCCGGGACTGAGGCCGCCCTGCACCTTCAGCGGATTTGGGGATTAGGCTCTTTTAGG TAACTGGCTTTCCTGCTGGTCCGTGCGGGAAATTCAGTCTTGATTTAACCTTAATTTTACAGCCCTTGGCTTGTGTTATCGGACATTCTA ATGCATATTTATAAGAGAAGTTTAACAAGTATTTATTCCCATAACTTCCCCTTGCAGACTGTTTTCTTCTGGCCTTTAGGCCTCTGCTTG TCCCCTCGGGCCTCCTTCCTCCTCCGTGAGCCACTGCAGTCCCCCCACCCACGCTTCAGGTCAGGCGTGAGCCGGTGCCTTTGCTGTGGC CTTGGCTGGGTGCTGATGGGACTGTCAGGCCCACTGCTGCGGTCGGTACCTTTGCAGCCTGTGCTTGTGGTCTGATCTGCTCAGGCCCCC >20992_20992_1_CYLD-HAGH_CYLD_chr16_50788335_ENST00000311559_HAGH_chr16_1859834_ENST00000455446_length(amino acids)=340AA_BP=304 MSSGLWSQEKVTSPYWEERIFYLLLQECSVTDKQTQKLLKVPKGSIGQYIQDRSVGHSRIPSAKGKKNQIGLKILEQPHAVLFVDEKDVV EINEKFTELLLAITNCEERFSLFKNRNRLSKGLQIDVGCPVKVQLRSGEEKFPGVVRFRGPLLAERTVSGIFFGVELLEEGRGQGFTDGV YQGKQLFQCDEDCGVFVALDKLELIEDDDTALESDYAGPGDTMQVELPPLEINSRVSLKVGETIESGTVIFCDVLPGKESLGYFVGVDMD -------------------------------------------------------------- >20992_20992_2_CYLD-HAGH_CYLD_chr16_50788335_ENST00000398568_HAGH_chr16_1859834_ENST00000455446_length(transcript)=1804nt_BP=1213nt CGGAGGTAAATACTAGGGCGGTGGGTGTGGGGAGCCGGGGCCGGCCCGGGACGCGGGCTGGGGAGCCGGGGCGAGGGGCGACGCCCCGCC GCCCGAGTTTCCCCCTTTCTAGGGTGAGGATGGTTCTACACAGCCACCCGGAGTTCCTTAGTTGAAAGGTGCGCCCTGCTGTGACAGCAT GGACACCACGTTGCTGAAAACATGCTTTGGGACTGCCACTGAATTTATCTTTTGCGGTTTTATGACAAAGTTATTAGTAGTTTCCCTTTT TTGAATTAGTATTTTGAAGTTAATATCACAATGAGTTCAGGCTTATGGAGCCAAGAAAAAGTCACTTCACCCTACTGGGAAGAGCGGATT TTTTACTTGCTTCTTCAAGAATGCAGCGTTACAGACAAACAAACACAAAAGCTCCTTAAAGTACCGAAGGGAAGTATAGGACAGTATATT CAAGATCGTTCTGTGGGGCATTCAAGGATTCCTTCTGCAAAAGGCAAGAAAAATCAGATTGGATTAAAAATTCTAGAGCAACCTCATGCA GTTCTCTTTGTTGATGAAAAGGATGTTGTAGAGATAAATGAAAAGTTCACAGAGTTACTTTTGGCAATTACCAATTGTGAGGAGAGGTTC AGCCTGTTTAAAAACAGAAACAGACTAAGTAAAGGCCTCCAAATAGACGTGGGCTGTCCTGTGAAAGTACAGCTGAGATCTGGGGAAGAA AAATTTCCTGGAGTTGTACGCTTCAGAGGACCCCTGTTAGCAGAGAGGACAGTCTCCGGAATATTCTTTGGAGTTGAATTGCTGGAAGAA GGTCGTGGTCAAGGTTTCACTGACGGGGTGTACCAAGGGAAACAGCTTTTTCAGTGTGATGAAGATTGTGGCGTGTTTGTTGCATTGGAC AAGCTAGAACTCATAGAAGATGATGACACTGCATTGGAAAGTGATTACGCAGGTCCTGGGGACACAATGCAGGTCGAACTTCCTCCTTTG GAAATAAACTCCAGAGTTTCTTTGAAGGTTGGAGAAACAATAGAATCTGGAACAGTTATATTCTGTGATGTTTTGCCAGGAAAAGAAAGC TTAGGATATTTTGTTGGTGTGGACATGGATAACCCTATTGGCAACTGGGATGGAAGATTTGATGGAGTGCAGCTTTGTAGTTTTGCGTGT GTTGAAAGTACAATTCTATTGCACATCAATGATATCATCCCAGGAGAAGTACAGCATCGGGGAGCCCACAGTGCCATCCACCCTGGCAGA GGAGTTTACCTACAACCCCTTCATGAGAGTGAGGGAGAAGACGGTGCAGCAGCACGCAGGTGAGACGGACCCGGTGACCACCATGCGGGC CGTGCGCAGGGAGAAGGACCAGTTCAAGATGCCCCGGGACTGAGGCCGCCCTGCACCTTCAGCGGATTTGGGGATTAGGCTCTTTTAGGT AACTGGCTTTCCTGCTGGTCCGTGCGGGAAATTCAGTCTTGATTTAACCTTAATTTTACAGCCCTTGGCTTGTGTTATCGGACATTCTAA TGCATATTTATAAGAGAAGTTTAACAAGTATTTATTCCCATAACTTCCCCTTGCAGACTGTTTTCTTCTGGCCTTTAGGCCTCTGCTTGT CCCCTCGGGCCTCCTTCCTCCTCCGTGAGCCACTGCAGTCCCCCCACCCACGCTTCAGGTCAGGCGTGAGCCGGTGCCTTTGCTGTGGCC TTGGCTGGGTGCTGATGGGACTGTCAGGCCCACTGCTGCGGTCGGTACCTTTGCAGCCTGTGCTTGTGGTCTGATCTGCTCAGGCCCCCG >20992_20992_2_CYLD-HAGH_CYLD_chr16_50788335_ENST00000398568_HAGH_chr16_1859834_ENST00000455446_length(amino acids)=340AA_BP=304 MSSGLWSQEKVTSPYWEERIFYLLLQECSVTDKQTQKLLKVPKGSIGQYIQDRSVGHSRIPSAKGKKNQIGLKILEQPHAVLFVDEKDVV EINEKFTELLLAITNCEERFSLFKNRNRLSKGLQIDVGCPVKVQLRSGEEKFPGVVRFRGPLLAERTVSGIFFGVELLEEGRGQGFTDGV YQGKQLFQCDEDCGVFVALDKLELIEDDDTALESDYAGPGDTMQVELPPLEINSRVSLKVGETIESGTVIFCDVLPGKESLGYFVGVDMD -------------------------------------------------------------- >20992_20992_3_CYLD-HAGH_CYLD_chr16_50788335_ENST00000427738_HAGH_chr16_1859834_ENST00000455446_length(transcript)=1709nt_BP=1118nt AGTTTCCCCCTTTCTAGGGTGAGGATGGTTCTACACAGCCACCCGGAGTTCCTTAGTTGAAAGGTGCGCCCTGCTGTGACAGCATGGACA CCACGTTGCTGAAAACATGCTTTGGGACTGCCACTGAATTTATCTTTTGCGGTTTTATGACAAAGTTATTAGTAGTTTCCCTTTTTTGAA TTAGTATTTTGAAGTTAATATCACAATGAGTTCAGGCTTATGGAGCCAAGAAAAAGTCACTTCACCCTACTGGGAAGAGCGGATTTTTTA CTTGCTTCTTCAAGAATGCAGCGTTACAGACAAACAAACACAAAAGCTCCTTAAAGTACCGAAGGGAAGTATAGGACAGTATATTCAAGA TCGTTCTGTGGGGCATTCAAGGATTCCTTCTGCAAAAGGCAAGAAAAATCAGATTGGATTAAAAATTCTAGAGCAACCTCATGCAGTTCT CTTTGTTGATGAAAAGGATGTTGTAGAGATAAATGAAAAGTTCACAGAGTTACTTTTGGCAATTACCAATTGTGAGGAGAGGTTCAGCCT GTTTAAAAACAGAAACAGACTAAGTAAAGGCCTCCAAATAGACGTGGGCTGTCCTGTGAAAGTACAGCTGAGATCTGGGGAAGAAAAATT TCCTGGAGTTGTACGCTTCAGAGGACCCCTGTTAGCAGAGAGGACAGTCTCCGGAATATTCTTTGGAGTTGAATTGCTGGAAGAAGGTCG TGGTCAAGGTTTCACTGACGGGGTGTACCAAGGGAAACAGCTTTTTCAGTGTGATGAAGATTGTGGCGTGTTTGTTGCATTGGACAAGCT AGAACTCATAGAAGATGATGACACTGCATTGGAAAGTGATTACGCAGGTCCTGGGGACACAATGCAGGTCGAACTTCCTCCTTTGGAAAT AAACTCCAGAGTTTCTTTGAAGGTTGGAGAAACAATAGAATCTGGAACAGTTATATTCTGTGATGTTTTGCCAGGAAAAGAAAGCTTAGG ATATTTTGTTGGTGTGGACATGGATAACCCTATTGGCAACTGGGATGGAAGATTTGATGGAGTGCAGCTTTGTAGTTTTGCGTGTGTTGA AAGTACAATTCTATTGCACATCAATGATATCATCCCAGGAGAAGTACAGCATCGGGGAGCCCACAGTGCCATCCACCCTGGCAGAGGAGT TTACCTACAACCCCTTCATGAGAGTGAGGGAGAAGACGGTGCAGCAGCACGCAGGTGAGACGGACCCGGTGACCACCATGCGGGCCGTGC GCAGGGAGAAGGACCAGTTCAAGATGCCCCGGGACTGAGGCCGCCCTGCACCTTCAGCGGATTTGGGGATTAGGCTCTTTTAGGTAACTG GCTTTCCTGCTGGTCCGTGCGGGAAATTCAGTCTTGATTTAACCTTAATTTTACAGCCCTTGGCTTGTGTTATCGGACATTCTAATGCAT ATTTATAAGAGAAGTTTAACAAGTATTTATTCCCATAACTTCCCCTTGCAGACTGTTTTCTTCTGGCCTTTAGGCCTCTGCTTGTCCCCT CGGGCCTCCTTCCTCCTCCGTGAGCCACTGCAGTCCCCCCACCCACGCTTCAGGTCAGGCGTGAGCCGGTGCCTTTGCTGTGGCCTTGGC >20992_20992_3_CYLD-HAGH_CYLD_chr16_50788335_ENST00000427738_HAGH_chr16_1859834_ENST00000455446_length(amino acids)=340AA_BP=304 MSSGLWSQEKVTSPYWEERIFYLLLQECSVTDKQTQKLLKVPKGSIGQYIQDRSVGHSRIPSAKGKKNQIGLKILEQPHAVLFVDEKDVV EINEKFTELLLAITNCEERFSLFKNRNRLSKGLQIDVGCPVKVQLRSGEEKFPGVVRFRGPLLAERTVSGIFFGVELLEEGRGQGFTDGV YQGKQLFQCDEDCGVFVALDKLELIEDDDTALESDYAGPGDTMQVELPPLEINSRVSLKVGETIESGTVIFCDVLPGKESLGYFVGVDMD -------------------------------------------------------------- >20992_20992_4_CYLD-HAGH_CYLD_chr16_50788335_ENST00000540145_HAGH_chr16_1859834_ENST00000455446_length(transcript)=1919nt_BP=1328nt GTGCGGTTCGGAGGCGGGGCAGGTGGGGGCGGGCCCAGGTAGCAGGTTTGGCTGCGCGGGGGCCGCGCGTCGGAGTTTCCCCCTTTCTAG GGTGAGGATGGTTCTACACAGCCACCCGGAGTTCCTTAGTTGAAAGGTGCGCCCTGCTGTGACAGAATGTGGTAATTGTAATCTTTAACA TTTTCATGTAAAACATATTTCCTGATCATCTTTCCATTGTCTTCATGGAAAATTGATAAATATTTGTGCCTTCCAACTCTCGTCTTGGTT GAATGACTTCATCTTAATACAACATGGACACCACGTTGCTGAAAACATGCTTTGGGACTGCCACTGAATTTATCTTTTGCGGTTTTATGA CAAAGTTATTAGTAGTTTCCCTTTTTTGAATTAGTATTTTGAAGTTAATATCACAATGAGTTCAGGCTTATGGAGCCAAGAAAAAGTCAC TTCACCCTACTGGGAAGAGCGGATTTTTTACTTGCTTCTTCAAGAATGCAGCGTTACAGACAAACAAACACAAAAGCTCCTTAAAGTACC GAAGGGAAGTATAGGACAGTATATTCAAGATCGTTCTGTGGGGCATTCAAGGATTCCTTCTGCAAAAGGCAAGAAAAATCAGATTGGATT AAAAATTCTAGAGCAACCTCATGCAGTTCTCTTTGTTGATGAAAAGGATGTTGTAGAGATAAATGAAAAGTTCACAGAGTTACTTTTGGC AATTACCAATTGTGAGGAGAGGTTCAGCCTGTTTAAAAACAGAAACAGACTAAGTAAAGGCCTCCAAATAGACGTGGGCTGTCCTGTGAA AGTACAGCTGAGATCTGGGGAAGAAAAATTTCCTGGAGTTGTACGCTTCAGAGGACCCCTGTTAGCAGAGAGGACAGTCTCCGGAATATT CTTTGGAGTTGAATTGCTGGAAGAAGGTCGTGGTCAAGGTTTCACTGACGGGGTGTACCAAGGGAAACAGCTTTTTCAGTGTGATGAAGA TTGTGGCGTGTTTGTTGCATTGGACAAGCTAGAACTCATAGAAGATGATGACACTGCATTGGAAAGTGATTACGCAGGTCCTGGGGACAC AATGCAGGTCGAACTTCCTCCTTTGGAAATAAACTCCAGAGTTTCTTTGAAGGTTGGAGAAACAATAGAATCTGGAACAGTTATATTCTG TGATGTTTTGCCAGGAAAAGAAAGCTTAGGATATTTTGTTGGTGTGGACATGGATAACCCTATTGGCAACTGGGATGGAAGATTTGATGG AGTGCAGCTTTGTAGTTTTGCGTGTGTTGAAAGTACAATTCTATTGCACATCAATGATATCATCCCAGGAGAAGTACAGCATCGGGGAGC CCACAGTGCCATCCACCCTGGCAGAGGAGTTTACCTACAACCCCTTCATGAGAGTGAGGGAGAAGACGGTGCAGCAGCACGCAGGTGAGA CGGACCCGGTGACCACCATGCGGGCCGTGCGCAGGGAGAAGGACCAGTTCAAGATGCCCCGGGACTGAGGCCGCCCTGCACCTTCAGCGG ATTTGGGGATTAGGCTCTTTTAGGTAACTGGCTTTCCTGCTGGTCCGTGCGGGAAATTCAGTCTTGATTTAACCTTAATTTTACAGCCCT TGGCTTGTGTTATCGGACATTCTAATGCATATTTATAAGAGAAGTTTAACAAGTATTTATTCCCATAACTTCCCCTTGCAGACTGTTTTC TTCTGGCCTTTAGGCCTCTGCTTGTCCCCTCGGGCCTCCTTCCTCCTCCGTGAGCCACTGCAGTCCCCCCACCCACGCTTCAGGTCAGGC GTGAGCCGGTGCCTTTGCTGTGGCCTTGGCTGGGTGCTGATGGGACTGTCAGGCCCACTGCTGCGGTCGGTACCTTTGCAGCCTGTGCTT >20992_20992_4_CYLD-HAGH_CYLD_chr16_50788335_ENST00000540145_HAGH_chr16_1859834_ENST00000455446_length(amino acids)=340AA_BP=304 MSSGLWSQEKVTSPYWEERIFYLLLQECSVTDKQTQKLLKVPKGSIGQYIQDRSVGHSRIPSAKGKKNQIGLKILEQPHAVLFVDEKDVV EINEKFTELLLAITNCEERFSLFKNRNRLSKGLQIDVGCPVKVQLRSGEEKFPGVVRFRGPLLAERTVSGIFFGVELLEEGRGQGFTDGV YQGKQLFQCDEDCGVFVALDKLELIEDDDTALESDYAGPGDTMQVELPPLEINSRVSLKVGETIESGTVIFCDVLPGKESLGYFVGVDMD -------------------------------------------------------------- >20992_20992_5_CYLD-HAGH_CYLD_chr16_50788335_ENST00000564326_HAGH_chr16_1859834_ENST00000455446_length(transcript)=1698nt_BP=1107nt GGCCCAGGTAGCAGGTTTGGCTGCGCGGGGGCCGCGCGTCGGAGGCATTTTGATTTCCTTTCTTTTTACAGCATGGACACCACGTTGCTG AAAACATGCTTTGGGACTGCCACTGAATTTATCTTTTGCGGTTTTATGACAAAGTTATTAGTAGTTTCCCTTTTTTGAATTAGTATTTTG AAGTTAATATCACAATGAGTTCAGGCTTATGGAGCCAAGAAAAAGTCACTTCACCCTACTGGGAAGAGCGGATTTTTTACTTGCTTCTTC AAGAATGCAGCGTTACAGACAAACAAACACAAAAGCTCCTTAAAGTACCGAAGGGAAGTATAGGACAGTATATTCAAGATCGTTCTGTGG GGCATTCAAGGATTCCTTCTGCAAAAGGCAAGAAAAATCAGATTGGATTAAAAATTCTAGAGCAACCTCATGCAGTTCTCTTTGTTGATG AAAAGGATGTTGTAGAGATAAATGAAAAGTTCACAGAGTTACTTTTGGCAATTACCAATTGTGAGGAGAGGTTCAGCCTGTTTAAAAACA GAAACAGACTAAGTAAAGGCCTCCAAATAGACGTGGGCTGTCCTGTGAAAGTACAGCTGAGATCTGGGGAAGAAAAATTTCCTGGAGTTG TACGCTTCAGAGGACCCCTGTTAGCAGAGAGGACAGTCTCCGGAATATTCTTTGGAGTTGAATTGCTGGAAGAAGGTCGTGGTCAAGGTT TCACTGACGGGGTGTACCAAGGGAAACAGCTTTTTCAGTGTGATGAAGATTGTGGCGTGTTTGTTGCATTGGACAAGCTAGAACTCATAG AAGATGATGACACTGCATTGGAAAGTGATTACGCAGGTCCTGGGGACACAATGCAGGTCGAACTTCCTCCTTTGGAAATAAACTCCAGAG TTTCTTTGAAGGTTGGAGAAACAATAGAATCTGGAACAGTTATATTCTGTGATGTTTTGCCAGGAAAAGAAAGCTTAGGATATTTTGTTG GTGTGGACATGGATAACCCTATTGGCAACTGGGATGGAAGATTTGATGGAGTGCAGCTTTGTAGTTTTGCGTGTGTTGAAAGTACAATTC TATTGCACATCAATGATATCATCCCAGGAGAAGTACAGCATCGGGGAGCCCACAGTGCCATCCACCCTGGCAGAGGAGTTTACCTACAAC CCCTTCATGAGAGTGAGGGAGAAGACGGTGCAGCAGCACGCAGGTGAGACGGACCCGGTGACCACCATGCGGGCCGTGCGCAGGGAGAAG GACCAGTTCAAGATGCCCCGGGACTGAGGCCGCCCTGCACCTTCAGCGGATTTGGGGATTAGGCTCTTTTAGGTAACTGGCTTTCCTGCT GGTCCGTGCGGGAAATTCAGTCTTGATTTAACCTTAATTTTACAGCCCTTGGCTTGTGTTATCGGACATTCTAATGCATATTTATAAGAG AAGTTTAACAAGTATTTATTCCCATAACTTCCCCTTGCAGACTGTTTTCTTCTGGCCTTTAGGCCTCTGCTTGTCCCCTCGGGCCTCCTT CCTCCTCCGTGAGCCACTGCAGTCCCCCCACCCACGCTTCAGGTCAGGCGTGAGCCGGTGCCTTTGCTGTGGCCTTGGCTGGGTGCTGAT >20992_20992_5_CYLD-HAGH_CYLD_chr16_50788335_ENST00000564326_HAGH_chr16_1859834_ENST00000455446_length(amino acids)=340AA_BP=304 MSSGLWSQEKVTSPYWEERIFYLLLQECSVTDKQTQKLLKVPKGSIGQYIQDRSVGHSRIPSAKGKKNQIGLKILEQPHAVLFVDEKDVV EINEKFTELLLAITNCEERFSLFKNRNRLSKGLQIDVGCPVKVQLRSGEEKFPGVVRFRGPLLAERTVSGIFFGVELLEEGRGQGFTDGV YQGKQLFQCDEDCGVFVALDKLELIEDDDTALESDYAGPGDTMQVELPPLEINSRVSLKVGETIESGTVIFCDVLPGKESLGYFVGVDMD -------------------------------------------------------------- >20992_20992_6_CYLD-HAGH_CYLD_chr16_50788335_ENST00000566206_HAGH_chr16_1859834_ENST00000455446_length(transcript)=1754nt_BP=1163nt AGGTAGCAGGTTTGGCTGCGCGGGGGCCGCGCGTCGGAGTTTCCCCCTTTCTAGGGTGAGGATGGTTCTACACAGCCACCCGGAGTTCCT TAGTTGAAAGGCATTTTGATTTCCTTTCTTTTTACAGCATGGACACCACGTTGCTGAAAACATGCTTTGGGACTGCCACTGAATTTATCT TTTGCGGTTTTATGACAAAGTTATTAGTAGTTTCCCTTTTTTGAATTAGTATTTTGAAGTTAATATCACAATGAGTTCAGGCTTATGGAG CCAAGAAAAAGTCACTTCACCCTACTGGGAAGAGCGGATTTTTTACTTGCTTCTTCAAGAATGCAGCGTTACAGACAAACAAACACAAAA GCTCCTTAAAGTACCGAAGGGAAGTATAGGACAGTATATTCAAGATCGTTCTGTGGGGCATTCAAGGATTCCTTCTGCAAAAGGCAAGAA AAATCAGATTGGATTAAAAATTCTAGAGCAACCTCATGCAGTTCTCTTTGTTGATGAAAAGGATGTTGTAGAGATAAATGAAAAGTTCAC AGAGTTACTTTTGGCAATTACCAATTGTGAGGAGAGGTTCAGCCTGTTTAAAAACAGAAACAGACTAAGTAAAGGCCTCCAAATAGACGT GGGCTGTCCTGTGAAAGTACAGCTGAGATCTGGGGAAGAAAAATTTCCTGGAGTTGTACGCTTCAGAGGACCCCTGTTAGCAGAGAGGAC AGTCTCCGGAATATTCTTTGGAGTTGAATTGCTGGAAGAAGGTCGTGGTCAAGGTTTCACTGACGGGGTGTACCAAGGGAAACAGCTTTT TCAGTGTGATGAAGATTGTGGCGTGTTTGTTGCATTGGACAAGCTAGAACTCATAGAAGATGATGACACTGCATTGGAAAGTGATTACGC AGGTCCTGGGGACACAATGCAGGTCGAACTTCCTCCTTTGGAAATAAACTCCAGAGTTTCTTTGAAGGTTGGAGAAACAATAGAATCTGG AACAGTTATATTCTGTGATGTTTTGCCAGGAAAAGAAAGCTTAGGATATTTTGTTGGTGTGGACATGGATAACCCTATTGGCAACTGGGA TGGAAGATTTGATGGAGTGCAGCTTTGTAGTTTTGCGTGTGTTGAAAGTACAATTCTATTGCACATCAATGATATCATCCCAGGAGAAGT ACAGCATCGGGGAGCCCACAGTGCCATCCACCCTGGCAGAGGAGTTTACCTACAACCCCTTCATGAGAGTGAGGGAGAAGACGGTGCAGC AGCACGCAGGTGAGACGGACCCGGTGACCACCATGCGGGCCGTGCGCAGGGAGAAGGACCAGTTCAAGATGCCCCGGGACTGAGGCCGCC CTGCACCTTCAGCGGATTTGGGGATTAGGCTCTTTTAGGTAACTGGCTTTCCTGCTGGTCCGTGCGGGAAATTCAGTCTTGATTTAACCT TAATTTTACAGCCCTTGGCTTGTGTTATCGGACATTCTAATGCATATTTATAAGAGAAGTTTAACAAGTATTTATTCCCATAACTTCCCC TTGCAGACTGTTTTCTTCTGGCCTTTAGGCCTCTGCTTGTCCCCTCGGGCCTCCTTCCTCCTCCGTGAGCCACTGCAGTCCCCCCACCCA CGCTTCAGGTCAGGCGTGAGCCGGTGCCTTTGCTGTGGCCTTGGCTGGGTGCTGATGGGACTGTCAGGCCCACTGCTGCGGTCGGTACCT >20992_20992_6_CYLD-HAGH_CYLD_chr16_50788335_ENST00000566206_HAGH_chr16_1859834_ENST00000455446_length(amino acids)=340AA_BP=304 MSSGLWSQEKVTSPYWEERIFYLLLQECSVTDKQTQKLLKVPKGSIGQYIQDRSVGHSRIPSAKGKKNQIGLKILEQPHAVLFVDEKDVV EINEKFTELLLAITNCEERFSLFKNRNRLSKGLQIDVGCPVKVQLRSGEEKFPGVVRFRGPLLAERTVSGIFFGVELLEEGRGQGFTDGV YQGKQLFQCDEDCGVFVALDKLELIEDDDTALESDYAGPGDTMQVELPPLEINSRVSLKVGETIESGTVIFCDVLPGKESLGYFVGVDMD -------------------------------------------------------------- >20992_20992_7_CYLD-HAGH_CYLD_chr16_50788335_ENST00000568704_HAGH_chr16_1859834_ENST00000455446_length(transcript)=1654nt_BP=1063nt GCATTTTGATTTCCTTTCTTTTTACAGCATGGACACCACGTTGCTGAAAACATGCTTTGGGACTGCCACTGAATTTATCTTTTGCGGTTT TATGACAAAGTTATTAGTAGTTTCCCTTTTTTGAATTAGTATTTTGAAGTTAATATCACAATGAGTTCAGGCTTATGGAGCCAAGAAAAA GTCACTTCACCCTACTGGGAAGAGCGGATTTTTTACTTGCTTCTTCAAGAATGCAGCGTTACAGACAAACAAACACAAAAGCTCCTTAAA GTACCGAAGGGAAGTATAGGACAGTATATTCAAGATCGTTCTGTGGGGCATTCAAGGATTCCTTCTGCAAAAGGCAAGAAAAATCAGATT GGATTAAAAATTCTAGAGCAACCTCATGCAGTTCTCTTTGTTGATGAAAAGGATGTTGTAGAGATAAATGAAAAGTTCACAGAGTTACTT TTGGCAATTACCAATTGTGAGGAGAGGTTCAGCCTGTTTAAAAACAGAAACAGACTAAGTAAAGGCCTCCAAATAGACGTGGGCTGTCCT GTGAAAGTACAGCTGAGATCTGGGGAAGAAAAATTTCCTGGAGTTGTACGCTTCAGAGGACCCCTGTTAGCAGAGAGGACAGTCTCCGGA ATATTCTTTGGAGTTGAATTGCTGGAAGAAGGTCGTGGTCAAGGTTTCACTGACGGGGTGTACCAAGGGAAACAGCTTTTTCAGTGTGAT GAAGATTGTGGCGTGTTTGTTGCATTGGACAAGCTAGAACTCATAGAAGATGATGACACTGCATTGGAAAGTGATTACGCAGGTCCTGGG GACACAATGCAGGTCGAACTTCCTCCTTTGGAAATAAACTCCAGAGTTTCTTTGAAGGTTGGAGAAACAATAGAATCTGGAACAGTTATA TTCTGTGATGTTTTGCCAGGAAAAGAAAGCTTAGGATATTTTGTTGGTGTGGACATGGATAACCCTATTGGCAACTGGGATGGAAGATTT GATGGAGTGCAGCTTTGTAGTTTTGCGTGTGTTGAAAGTACAATTCTATTGCACATCAATGATATCATCCCAGGAGAAGTACAGCATCGG GGAGCCCACAGTGCCATCCACCCTGGCAGAGGAGTTTACCTACAACCCCTTCATGAGAGTGAGGGAGAAGACGGTGCAGCAGCACGCAGG TGAGACGGACCCGGTGACCACCATGCGGGCCGTGCGCAGGGAGAAGGACCAGTTCAAGATGCCCCGGGACTGAGGCCGCCCTGCACCTTC AGCGGATTTGGGGATTAGGCTCTTTTAGGTAACTGGCTTTCCTGCTGGTCCGTGCGGGAAATTCAGTCTTGATTTAACCTTAATTTTACA GCCCTTGGCTTGTGTTATCGGACATTCTAATGCATATTTATAAGAGAAGTTTAACAAGTATTTATTCCCATAACTTCCCCTTGCAGACTG TTTTCTTCTGGCCTTTAGGCCTCTGCTTGTCCCCTCGGGCCTCCTTCCTCCTCCGTGAGCCACTGCAGTCCCCCCACCCACGCTTCAGGT CAGGCGTGAGCCGGTGCCTTTGCTGTGGCCTTGGCTGGGTGCTGATGGGACTGTCAGGCCCACTGCTGCGGTCGGTACCTTTGCAGCCTG >20992_20992_7_CYLD-HAGH_CYLD_chr16_50788335_ENST00000568704_HAGH_chr16_1859834_ENST00000455446_length(amino acids)=340AA_BP=304 MSSGLWSQEKVTSPYWEERIFYLLLQECSVTDKQTQKLLKVPKGSIGQYIQDRSVGHSRIPSAKGKKNQIGLKILEQPHAVLFVDEKDVV EINEKFTELLLAITNCEERFSLFKNRNRLSKGLQIDVGCPVKVQLRSGEEKFPGVVRFRGPLLAERTVSGIFFGVELLEEGRGQGFTDGV YQGKQLFQCDEDCGVFVALDKLELIEDDDTALESDYAGPGDTMQVELPPLEINSRVSLKVGETIESGTVIFCDVLPGKESLGYFVGVDMD -------------------------------------------------------------- >20992_20992_8_CYLD-HAGH_CYLD_chr16_50788335_ENST00000569418_HAGH_chr16_1859834_ENST00000455446_length(transcript)=1782nt_BP=1191nt GTGCGGTTCGGAGGCGGGGCAGGTGGGGGCGGGCCCAGGTAGCAGGTTTGGCTGCGCGGGGGCCGCGCGTCGGAGTTTCCCCCTTTCTAG GGTGAGGATGGTTCTACACAGCCACCCGGAGTTCCTTAGTTGAAAGGTGCGCCCTGCTGTGACAGCATGGACACCACGTTGCTGAAAACA TGCTTTGGGACTGCCACTGAATTTATCTTTTGCGGTTTTATGACAAAGTTATTAGTAGTTTCCCTTTTTTGAATTAGTATTTTGAAGTTA ATATCACAATGAGTTCAGGCTTATGGAGCCAAGAAAAAGTCACTTCACCCTACTGGGAAGAGCGGATTTTTTACTTGCTTCTTCAAGAAT GCAGCGTTACAGACAAACAAACACAAAAGCTCCTTAAAGTACCGAAGGGAAGTATAGGACAGTATATTCAAGATCGTTCTGTGGGGCATT CAAGGATTCCTTCTGCAAAAGGCAAGAAAAATCAGATTGGATTAAAAATTCTAGAGCAACCTCATGCAGTTCTCTTTGTTGATGAAAAGG ATGTTGTAGAGATAAATGAAAAGTTCACAGAGTTACTTTTGGCAATTACCAATTGTGAGGAGAGGTTCAGCCTGTTTAAAAACAGAAACA GACTAAGTAAAGGCCTCCAAATAGACGTGGGCTGTCCTGTGAAAGTACAGCTGAGATCTGGGGAAGAAAAATTTCCTGGAGTTGTACGCT TCAGAGGACCCCTGTTAGCAGAGAGGACAGTCTCCGGAATATTCTTTGGAGTTGAATTGCTGGAAGAAGGTCGTGGTCAAGGTTTCACTG ACGGGGTGTACCAAGGGAAACAGCTTTTTCAGTGTGATGAAGATTGTGGCGTGTTTGTTGCATTGGACAAGCTAGAACTCATAGAAGATG ATGACACTGCATTGGAAAGTGATTACGCAGGTCCTGGGGACACAATGCAGGTCGAACTTCCTCCTTTGGAAATAAACTCCAGAGTTTCTT TGAAGGTTGGAGAAACAATAGAATCTGGAACAGTTATATTCTGTGATGTTTTGCCAGGAAAAGAAAGCTTAGGATATTTTGTTGGTGTGG ACATGGATAACCCTATTGGCAACTGGGATGGAAGATTTGATGGAGTGCAGCTTTGTAGTTTTGCGTGTGTTGAAAGTACAATTCTATTGC ACATCAATGATATCATCCCAGGAGAAGTACAGCATCGGGGAGCCCACAGTGCCATCCACCCTGGCAGAGGAGTTTACCTACAACCCCTTC ATGAGAGTGAGGGAGAAGACGGTGCAGCAGCACGCAGGTGAGACGGACCCGGTGACCACCATGCGGGCCGTGCGCAGGGAGAAGGACCAG TTCAAGATGCCCCGGGACTGAGGCCGCCCTGCACCTTCAGCGGATTTGGGGATTAGGCTCTTTTAGGTAACTGGCTTTCCTGCTGGTCCG TGCGGGAAATTCAGTCTTGATTTAACCTTAATTTTACAGCCCTTGGCTTGTGTTATCGGACATTCTAATGCATATTTATAAGAGAAGTTT AACAAGTATTTATTCCCATAACTTCCCCTTGCAGACTGTTTTCTTCTGGCCTTTAGGCCTCTGCTTGTCCCCTCGGGCCTCCTTCCTCCT CCGTGAGCCACTGCAGTCCCCCCACCCACGCTTCAGGTCAGGCGTGAGCCGGTGCCTTTGCTGTGGCCTTGGCTGGGTGCTGATGGGACT >20992_20992_8_CYLD-HAGH_CYLD_chr16_50788335_ENST00000569418_HAGH_chr16_1859834_ENST00000455446_length(amino acids)=340AA_BP=304 MSSGLWSQEKVTSPYWEERIFYLLLQECSVTDKQTQKLLKVPKGSIGQYIQDRSVGHSRIPSAKGKKNQIGLKILEQPHAVLFVDEKDVV EINEKFTELLLAITNCEERFSLFKNRNRLSKGLQIDVGCPVKVQLRSGEEKFPGVVRFRGPLLAERTVSGIFFGVELLEEGRGQGFTDGV YQGKQLFQCDEDCGVFVALDKLELIEDDDTALESDYAGPGDTMQVELPPLEINSRVSLKVGETIESGTVIFCDVLPGKESLGYFVGVDMD -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CYLD-HAGH |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000311559 | + | 6 | 20 | 470_554 | 304.3333333333333 | 957.0 | IKBKG/NEMO |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000398568 | + | 5 | 18 | 470_554 | 304.3333333333333 | 954.0 | IKBKG/NEMO |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000427738 | + | 4 | 18 | 470_554 | 304.3333333333333 | 957.0 | IKBKG/NEMO |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000564326 | + | 4 | 17 | 470_554 | 304.3333333333333 | 954.0 | IKBKG/NEMO |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000569418 | + | 5 | 18 | 470_554 | 304.3333333333333 | 954.0 | IKBKG/NEMO |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000311559 | + | 6 | 20 | 394_469 | 304.3333333333333 | 957.0 | TRAF2 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000398568 | + | 5 | 18 | 394_469 | 304.3333333333333 | 954.0 | TRAF2 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000427738 | + | 4 | 18 | 394_469 | 304.3333333333333 | 957.0 | TRAF2 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000564326 | + | 4 | 17 | 394_469 | 304.3333333333333 | 954.0 | TRAF2 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000569418 | + | 5 | 18 | 394_469 | 304.3333333333333 | 954.0 | TRAF2 |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000311559 | + | 6 | 20 | 106_593 | 304.3333333333333 | 957.0 | TRIP |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000398568 | + | 5 | 18 | 106_593 | 304.3333333333333 | 954.0 | TRIP |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000427738 | + | 4 | 18 | 106_593 | 304.3333333333333 | 957.0 | TRIP |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000564326 | + | 4 | 17 | 106_593 | 304.3333333333333 | 954.0 | TRIP |
| Hgene | CYLD | chr16:50788335 | chr16:1859834 | ENST00000569418 | + | 5 | 18 | 106_593 | 304.3333333333333 | 954.0 | TRIP |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CYLD-HAGH |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CYLD-HAGH |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
