
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:CYP3A4-ITGAV (FusionGDB2 ID:21072) |
Fusion Gene Summary for CYP3A4-ITGAV |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: CYP3A4-ITGAV | Fusion gene ID: 21072 | Hgene | Tgene | Gene symbol | CYP3A4 | ITGAV | Gene ID | 1576 | 3685 |
| Gene name | cytochrome P450 family 3 subfamily A member 4 | integrin subunit alpha V | |
| Synonyms | CP33|CP34|CYP3A|CYP3A3|CYPIIIA3|CYPIIIA4|HLP|NF-25|P450C3|P450PCN1 | CD51|MSK8|VNRA|VTNR | |
| Cytomap | 7q22.1 | 2q32.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | cytochrome P450 3A41,4-cineole 2-exo-monooxygenase1,8-cineole 2-exo-monooxygenaseP450-III, steroid induciblealbendazole monooxygenasealbendazole monooxygenase (sulfoxide-forming)albendazole sulfoxidasecholesterol 25-hydroxylasecytochrome P450 3A3 | integrin alpha-Vantigen identified by monoclonal antibody L230integrin alphaVbeta3integrin, alpha V (vitronectin receptor, alpha polypeptide, antigen CD51)vitronectin receptor subunit alpha | |
| Modification date | 20200327 | 20200313 | |
| UniProtAcc | P08684 | P06756 | |
| Ensembl transtripts involved in fusion gene | ENST00000336411, ENST00000354593, | ENST00000474571, ENST00000261023, ENST00000374907, ENST00000433736, | |
| Fusion gene scores | * DoF score | 2 X 2 X 2=8 | 9 X 8 X 6=432 |
| # samples | 2 | 9 | |
| ** MAII score | log2(2/8*10)=1.32192809488736 | log2(9/432*10)=-2.26303440583379 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: CYP3A4 [Title/Abstract] AND ITGAV [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | CYP3A4(99381634)-ITGAV(187540331), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | CYP3A4 | GO:0002933 | lipid hydroxylation | 14559847 |
| Hgene | CYP3A4 | GO:0008202 | steroid metabolic process | 14559847 |
| Hgene | CYP3A4 | GO:0008210 | estrogen metabolic process | 11555828|12865317|14559847 |
| Hgene | CYP3A4 | GO:0009822 | alkaloid catabolic process | 15039299 |
| Hgene | CYP3A4 | GO:0016098 | monoterpenoid metabolic process | 16401082 |
| Hgene | CYP3A4 | GO:0017144 | drug metabolic process | 15327587|19219744 |
| Hgene | CYP3A4 | GO:0042572 | retinol metabolic process | 10681376 |
| Hgene | CYP3A4 | GO:0042573 | retinoic acid metabolic process | 11093772 |
| Hgene | CYP3A4 | GO:0042737 | drug catabolic process | 15039299 |
| Hgene | CYP3A4 | GO:0042738 | exogenous drug catabolic process | 18619574 |
| Hgene | CYP3A4 | GO:0046483 | heterocycle metabolic process | 15327587 |
| Hgene | CYP3A4 | GO:0055114 | oxidation-reduction process | 16401082|19219744 |
| Hgene | CYP3A4 | GO:0070989 | oxidative demethylation | 15039299|18619574 |
| Tgene | ITGAV | GO:0007155 | cell adhesion | 10218736 |
| Tgene | ITGAV | GO:0008284 | positive regulation of cell proliferation | 19578119 |
| Tgene | ITGAV | GO:0033627 | cell adhesion mediated by integrin | 12807887|17158881 |
| Tgene | ITGAV | GO:0034446 | substrate adhesion-dependent cell spreading | 24658351 |
| Tgene | ITGAV | GO:0045785 | positive regulation of cell adhesion | 10708943 |
| Tgene | ITGAV | GO:0050764 | regulation of phagocytosis | 10570297 |
| Tgene | ITGAV | GO:0070588 | calcium ion transmembrane transport | 18395422 |
| Tgene | ITGAV | GO:1901388 | regulation of transforming growth factor beta activation | 22278742 |
| Tgene | ITGAV | GO:2000536 | negative regulation of entry of bacterium into host cell | 10570297 |
 Fusion gene breakpoints across CYP3A4 (5'-gene) Fusion gene breakpoints across CYP3A4 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
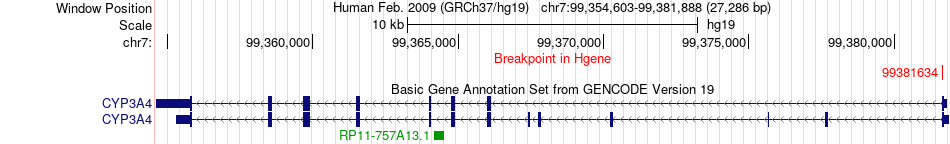 |
 Fusion gene breakpoints across ITGAV (3'-gene) Fusion gene breakpoints across ITGAV (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
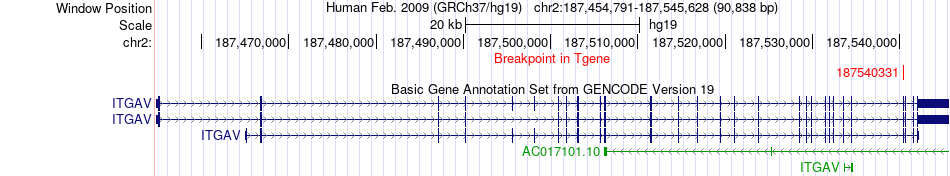 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | ERR315457 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + |
Top |
Fusion Gene ORF analysis for CYP3A4-ITGAV |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000336411 | ENST00000474571 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + |
| 5CDS-intron | ENST00000354593 | ENST00000474571 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + |
| In-frame | ENST00000336411 | ENST00000261023 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + |
| In-frame | ENST00000336411 | ENST00000374907 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + |
| In-frame | ENST00000336411 | ENST00000433736 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + |
| In-frame | ENST00000354593 | ENST00000261023 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + |
| In-frame | ENST00000354593 | ENST00000374907 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + |
| In-frame | ENST00000354593 | ENST00000433736 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000354593 | CYP3A4 | chr7 | 99381634 | - | ENST00000261023 | ITGAV | chr2 | 187540331 | + | 4225 | 175 | 136 | 615 | 159 |
| ENST00000354593 | CYP3A4 | chr7 | 99381634 | - | ENST00000374907 | ITGAV | chr2 | 187540331 | + | 4221 | 175 | 136 | 615 | 159 |
| ENST00000354593 | CYP3A4 | chr7 | 99381634 | - | ENST00000433736 | ITGAV | chr2 | 187540331 | + | 765 | 175 | 136 | 615 | 159 |
| ENST00000336411 | CYP3A4 | chr7 | 99381634 | - | ENST00000261023 | ITGAV | chr2 | 187540331 | + | 4305 | 255 | 216 | 695 | 159 |
| ENST00000336411 | CYP3A4 | chr7 | 99381634 | - | ENST00000374907 | ITGAV | chr2 | 187540331 | + | 4301 | 255 | 216 | 695 | 159 |
| ENST00000336411 | CYP3A4 | chr7 | 99381634 | - | ENST00000433736 | ITGAV | chr2 | 187540331 | + | 845 | 255 | 216 | 695 | 159 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000354593 | ENST00000261023 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + | 0.09854877 | 0.9014513 |
| ENST00000354593 | ENST00000374907 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + | 0.030597633 | 0.9694024 |
| ENST00000354593 | ENST00000433736 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + | 0.059244517 | 0.94075555 |
| ENST00000336411 | ENST00000261023 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + | 0.09520535 | 0.90479463 |
| ENST00000336411 | ENST00000374907 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + | 0.029533591 | 0.97046643 |
| ENST00000336411 | ENST00000433736 | CYP3A4 | chr7 | 99381634 | - | ITGAV | chr2 | 187540331 | + | 0.046907615 | 0.95309234 |
Top |
Fusion Genomic Features for CYP3A4-ITGAV |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| CYP3A4 | chr7 | 99381633 | - | ITGAV | chr2 | 187540330 | + | 0.9999975 | 2.44E-06 |
| CYP3A4 | chr7 | 99381633 | - | ITGAV | chr2 | 187540330 | + | 0.9999975 | 2.44E-06 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
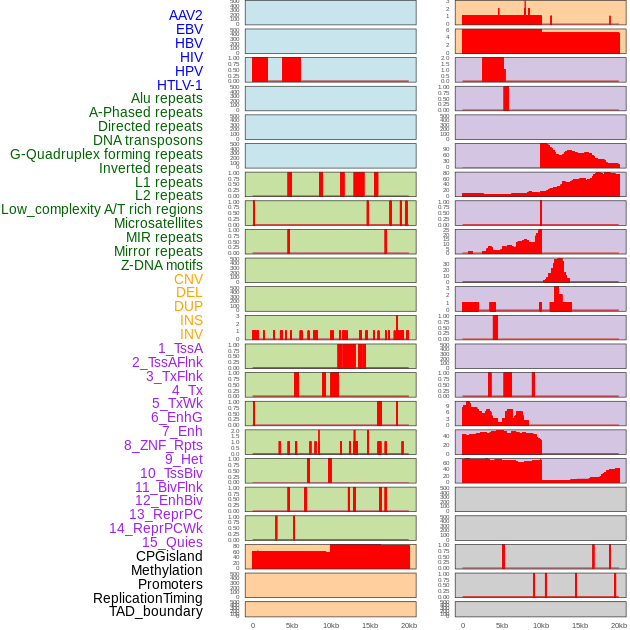 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
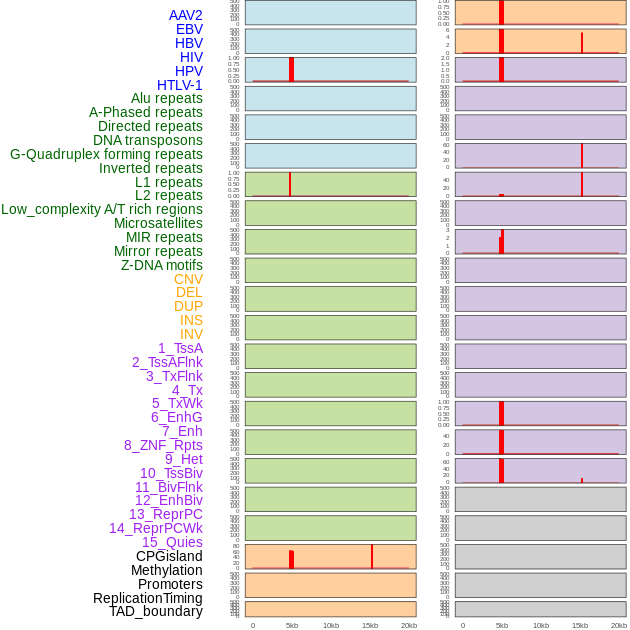 |
Top |
Fusion Protein Features for CYP3A4-ITGAV |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr7:99381634/chr2:187540331) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| CYP3A4 | ITGAV |
| FUNCTION: A cytochrome P450 monooxygenase involved in the metabolism of sterols, steroid hormones, retinoids and fatty acids (PubMed:10681376, PubMed:11093772, PubMed:11555828, PubMed:14559847, PubMed:12865317, PubMed:15373842, PubMed:15764715, PubMed:20702771, PubMed:19965576, PubMed:21490593, PubMed:21576599). Mechanistically, uses molecular oxygen inserting one oxygen atom into a substrate, and reducing the second into a water molecule, with two electrons provided by NADPH via cytochrome P450 reductase (NADPH--hemoprotein reductase). Catalyzes the hydroxylation of carbon-hydrogen bonds (PubMed:2732228, PubMed:14559847, PubMed:12865317, PubMed:15373842, PubMed:15764715, PubMed:21576599, PubMed:21490593). Exhibits high catalytic activity for the formation of hydroxyestrogens from estrone (E1) and 17beta-estradiol (E2), namely 2-hydroxy E1 and E2, as well as D-ring hydroxylated E1 and E2 at the C-16 position (PubMed:11555828, PubMed:14559847, PubMed:12865317). Plays a role in the metabolism of androgens, particularly in oxidative deactivation of testosterone (PubMed:2732228, PubMed:15373842, PubMed:15764715, PubMed:22773874). Metabolizes testosterone to less biologically active 2beta- and 6beta-hydroxytestosterones (PubMed:2732228, PubMed:15373842, PubMed:15764715). Contributes to the formation of hydroxycholesterols (oxysterols), particularly A-ring hydroxylated cholesterol at the C-4beta position, and side chain hydroxylated cholesterol at the C-25 position, likely contributing to cholesterol degradation and bile acid biosynthesis (PubMed:21576599). Catalyzes bisallylic hydroxylation of polyunsaturated fatty acids (PUFA) (PubMed:9435160). Catalyzes the epoxidation of double bonds of PUFA with a preference for the last double bond (PubMed:19965576). Metabolizes endocannabinoid arachidonoylethanolamide (anandamide) to 8,9-, 11,12-, and 14,15-epoxyeicosatrienoic acid ethanolamides (EpETrE-EAs), potentially modulating endocannabinoid system signaling (PubMed:20702771). Plays a role in the metabolism of retinoids. Displays high catalytic activity for oxidation of all-trans-retinol to all-trans-retinal, a rate-limiting step for the biosynthesis of all-trans-retinoic acid (atRA) (PubMed:10681376). Further metabolizes atRA toward 4-hydroxyretinoate and may play a role in hepatic atRA clearance (PubMed:11093772). Responsible for oxidative metabolism of xenobiotics. Acts as a 2-exo-monooxygenase for plant lipid 1,8-cineole (eucalyptol) (PubMed:11159812). Metabolizes the majority of the administered drugs. Catalyzes sulfoxidation of the anthelmintics albendazole and fenbendazole (PubMed:10759686). Hydroxylates antimalarial drug quinine (PubMed:8968357). Acts as a 1,4-cineole 2-exo-monooxygenase (PubMed:11695850). Also involved in vitamin D catabolism and calcium homeostasis. Catalyzes the inactivation of the active hormone calcitriol (1-alpha,25-dihydroxyvitamin D(3)) (PubMed:29461981). {ECO:0000269|PubMed:10681376, ECO:0000269|PubMed:10759686, ECO:0000269|PubMed:11093772, ECO:0000269|PubMed:11159812, ECO:0000269|PubMed:11555828, ECO:0000269|PubMed:11695850, ECO:0000269|PubMed:12865317, ECO:0000269|PubMed:14559847, ECO:0000269|PubMed:15373842, ECO:0000269|PubMed:15764715, ECO:0000269|PubMed:19965576, ECO:0000269|PubMed:20702771, ECO:0000269|PubMed:21490593, ECO:0000269|PubMed:21576599, ECO:0000269|PubMed:22773874, ECO:0000269|PubMed:2732228, ECO:0000269|PubMed:29461981, ECO:0000269|PubMed:8968357, ECO:0000269|PubMed:9435160}. | FUNCTION: The alpha-V (ITGAV) integrins are receptors for vitronectin, cytotactin, fibronectin, fibrinogen, laminin, matrix metalloproteinase-2, osteopontin, osteomodulin, prothrombin, thrombospondin and vWF. They recognize the sequence R-G-D in a wide array of ligands. ITGAV:ITGB3 binds to fractalkine (CX3CL1) and may act as its coreceptor in CX3CR1-dependent fractalkine signaling (PubMed:23125415). ITGAV:ITGB3 binds to NRG1 (via EGF domain) and this binding is essential for NRG1-ERBB signaling (PubMed:20682778). ITGAV:ITGB3 binds to FGF1 and this binding is essential for FGF1 signaling (PubMed:18441324). ITGAV:ITGB3 binds to FGF2 and this binding is essential for FGF2 signaling (PubMed:28302677). ITGAV:ITGB3 binds to IGF1 and this binding is essential for IGF1 signaling (PubMed:19578119). ITGAV:ITGB3 binds to IGF2 and this binding is essential for IGF2 signaling (PubMed:28873464). ITGAV:ITGB3 binds to IL1B and this binding is essential for IL1B signaling (PubMed:29030430). ITGAV:ITGB3 binds to PLA2G2A via a site (site 2) which is distinct from the classical ligand-binding site (site 1) and this induces integrin conformational changes and enhanced ligand binding to site 1 (PubMed:18635536, PubMed:25398877). ITGAV:ITGB3 and ITGAV:ITGB6 act as a receptor for fibrillin-1 (FBN1) and mediate R-G-D-dependent cell adhesion to FBN1 (PubMed:12807887, PubMed:17158881). Integrin alpha-V/beta-6 or alpha-V/beta-8 (ITGAV:ITGB6 or ITGAV:ITGB8) mediates R-G-D-dependent release of transforming growth factor beta-1 (TGF-beta-1) from regulatory Latency-associated peptide (LAP), thereby playing a key role in TGF-beta-1 activation (PubMed:15184403, PubMed:22278742, PubMed:28117447). ITGAV:ITGB3 act as a receptor for CD40LG (PubMed:31331973). {ECO:0000269|PubMed:12807887, ECO:0000269|PubMed:15184403, ECO:0000269|PubMed:17158881, ECO:0000269|PubMed:18441324, ECO:0000269|PubMed:18635536, ECO:0000269|PubMed:19578119, ECO:0000269|PubMed:20682778, ECO:0000269|PubMed:22278742, ECO:0000269|PubMed:23125415, ECO:0000269|PubMed:25398877, ECO:0000269|PubMed:28117447, ECO:0000269|PubMed:28302677, ECO:0000269|PubMed:28873464, ECO:0000269|PubMed:29030430, ECO:0000269|PubMed:31331973}.; FUNCTION: (Microbial infection) Integrin ITGAV:ITGB5 acts as a receptor for Adenovirus type C. {ECO:0000269|PubMed:20615244}.; FUNCTION: (Microbial infection) Integrin ITGAV:ITGB5 and ITGAV:ITGB3 act as receptors for Coxsackievirus A9 and B1. {ECO:0000269|PubMed:15194773, ECO:0000269|PubMed:7519807, ECO:0000269|PubMed:9426447}.; FUNCTION: (Microbial infection) Integrin ITGAV:ITGB3 acts as a receptor for Herpes virus 8/HHV-8. {ECO:0000269|PubMed:18045938}.; FUNCTION: (Microbial infection) Integrin ITGAV:ITGB6 acts as a receptor for herpes simplex 1/HHV-1. {ECO:0000269|PubMed:24367260}.; FUNCTION: (Microbial infection) Integrin ITGAV:ITGB3 acts as a receptor for Human parechovirus 1. {ECO:0000269|PubMed:11160695}.; FUNCTION: (Microbial infection) Integrin ITGAV:ITGB3 acts as a receptor for West nile virus. {ECO:0000269|PubMed:23658209}.; FUNCTION: (Microbial infection) In case of HIV-1 infection, the interaction with extracellular viral Tat protein seems to enhance angiogenesis in Kaposi's sarcoma lesions. {ECO:0000269|PubMed:10397733}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | CYP3A4 | chr7:99381634 | chr2:187540331 | ENST00000336411 | - | 1 | 13 | 2_22 | 23 | 504.0 | Transmembrane | Helical |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 1019_1023 | 902 | 1049.0 | Motif | Note=GFFKR motif | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 1019_1023 | 866 | 1013.0 | Motif | Note=GFFKR motif | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 1019_1023 | 856 | 1003.0 | Motif | Note=GFFKR motif | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 1017_1048 | 902 | 1049.0 | Topological domain | Cytoplasmic | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 1017_1048 | 866 | 1013.0 | Topological domain | Cytoplasmic | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 1017_1048 | 856 | 1003.0 | Topological domain | Cytoplasmic | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 993_1016 | 902 | 1049.0 | Transmembrane | Helical | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 993_1016 | 866 | 1013.0 | Transmembrane | Helical | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 993_1016 | 856 | 1003.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 260_268 | 902 | 1049.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 314_322 | 902 | 1049.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 379_387 | 902 | 1049.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 443_451 | 902 | 1049.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 260_268 | 866 | 1013.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 314_322 | 866 | 1013.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 379_387 | 866 | 1013.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 443_451 | 866 | 1013.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 260_268 | 856 | 1003.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 314_322 | 856 | 1003.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 379_387 | 856 | 1003.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 443_451 | 856 | 1003.0 | Calcium binding | Ontology_term=ECO:0000255,ECO:0000269 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 109_170 | 902 | 1049.0 | Repeat | FG-GAP 2 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 173_225 | 902 | 1049.0 | Repeat | FG-GAP 3 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 237_291 | 902 | 1049.0 | Repeat | FG-GAP 4 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 292_357 | 902 | 1049.0 | Repeat | FG-GAP 5 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 32_98 | 902 | 1049.0 | Repeat | FG-GAP 1 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 358_415 | 902 | 1049.0 | Repeat | FG-GAP 6 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 419_482 | 902 | 1049.0 | Repeat | FG-GAP 7 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 109_170 | 866 | 1013.0 | Repeat | FG-GAP 2 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 173_225 | 866 | 1013.0 | Repeat | FG-GAP 3 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 237_291 | 866 | 1013.0 | Repeat | FG-GAP 4 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 292_357 | 866 | 1013.0 | Repeat | FG-GAP 5 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 32_98 | 866 | 1013.0 | Repeat | FG-GAP 1 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 358_415 | 866 | 1013.0 | Repeat | FG-GAP 6 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 419_482 | 866 | 1013.0 | Repeat | FG-GAP 7 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 109_170 | 856 | 1003.0 | Repeat | FG-GAP 2 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 173_225 | 856 | 1003.0 | Repeat | FG-GAP 3 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 237_291 | 856 | 1003.0 | Repeat | FG-GAP 4 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 292_357 | 856 | 1003.0 | Repeat | FG-GAP 5 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 32_98 | 856 | 1003.0 | Repeat | FG-GAP 1 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 358_415 | 856 | 1003.0 | Repeat | FG-GAP 6 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 419_482 | 856 | 1003.0 | Repeat | FG-GAP 7 | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000261023 | 25 | 30 | 31_992 | 902 | 1049.0 | Topological domain | Extracellular | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000374907 | 23 | 28 | 31_992 | 866 | 1013.0 | Topological domain | Extracellular | |
| Tgene | ITGAV | chr7:99381634 | chr2:187540331 | ENST00000433736 | 25 | 30 | 31_992 | 856 | 1003.0 | Topological domain | Extracellular |
Top |
Fusion Gene Sequence for CYP3A4-ITGAV |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >21072_21072_1_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000336411_ITGAV_chr2_187540331_ENST00000261023_length(transcript)=4305nt_BP=255nt ACAGGTGGCCCTGCTACTGGCTGCAGCTCCAGCCCTGCCTCCTTCTCTAGCATATAAACAATCCAACAGCCTCACTGAATCACTGCTGTG CAGGGCAGGAAAGCTCCATGCACATAGCCCAGCAAAGAGCAACACAGAGCTGAAAGGAAGACTCAGAGGAGAGAGATAAGTAAGGAAAGT AGTGATGGCTCTCATCCCAGACTTGGCCATGGAAACCTGGCTTCTCCTGGCTGTCAGCCTGGTGCTCCTCTATCTGGTTGTGGAGTTGCT CAGTGCTTGAAGATTGTCTGCCAAGTTGGGAGATTAGACAGAGGAAAGAGTGCAATCTTGTACGTAAAGTCATTACTGTGGACTGAGACT TTTATGAATAAAGAAAATCAGAATCATTCCTATTCTCTGAAGTCGTCTGCTTCATTTAATGTCATAGAGTTTCCTTATAAGAATCTTCCA ATTGAGGATATCACCAACTCCACATTGGTTACCACTAATGTCACCTGGGGCATTCAGCCAGCGCCCATGCCTGTGCCTGTGTGGGTGATC ATTTTAGCAGTTCTAGCAGGATTGTTGCTACTGGCTGTTTTGGTATTTGTAATGTACAGGATGGGCTTTTTTAAACGGGTCCGGCCACCT CAAGAAGAACAAGAAAGGGAGCAGCTTCAACCTCATGAAAATGGTGAAGGAAACTCAGAAACTTAACTGCAGTTTTTAAGTTATGCTACA TCTTGACCCACTAGAATTAGCAACTTTATTATAGATTTAAACTTTCTTCATGAGGAGTAAAAATCCAAGGCTTTACTGCTGATAGTGCTA ATTGGCATTAACCACAAAATGAGAATTATATTTGTCAACCTTCTCCTTATAAATAAGTTCAGACATACATTTAATAACATAGGGTGACTT GTGTTTTTAGGTATTTAAATAATAAAATTTCAAGGGATAGTTTTTATTCAATGTATATAAGACAGGTAGTGCCTGATTTACTACTTTATA TAAAATAGTACCTCCTTCAGTTACTGTTTCTGATTTAATGTACGGAACTTTATTTGTTGTTGTTGTTGTTGTTGTTGTTGTTGTTTTAAA GCAGTCCAAATTTGGACCTTAGCAATCATGTCTTTTGTATAGGTACTTAATGTTAATACATATTACACTACAGTTTACTTTTCAGAATAC TAAAGACTTTATAACTGCATGAACTTGGATTTTTTTAATCACTCATATGGTAGAATTTTATAAACACATACATGATACCATCCAAATTCT TGCTTTTAATAACAAAGGTACAATATTTTGTTTTAGTATGAAAATCTGGTAGATCCTATTACACTTCTGTTTATATTAAATCCACAATAT TTTATTACATTTTTAACTTGTATAAATTTTAGGTCAAATCCTTCAAGCCAACCTATACTAAAAATTAGTTCCATAATCACAAATGGCTCT TTTGTGTAATTGTTTAATTTCACCTGAATATCATAATGCTTAAAGCCATATGGAGTTGGAAATTATTTCCAAAGCATATTTATTCCATTG TTTTAGTCTGGCTATTTACAGTATAAAAAAAGCATTTTTATTAAAATACTGTGTAGTTCTTTGAGATAGTTGCTTATGCATATAGTAAGT ATTACATTCTTAGAGTAGAGCAGAGTTTTTAGTTAGTATTAATTTATTTTCCTCCATTCATGTACTTTTCCTTATATTTCCAAAACTGTT ACTGAGAATGGGTCAAGATCAGTGAGAAATCTTTACAGTTGACAGGAACCTGGACCCCTTACCCCAACTTTATGAGTAATGCTTGGAATA AAAACTCTTAAGGCAACTCACTGATTTACTTCTAGCAATAGCATGATGTTACAGGAATATTACCTCTGTTTAAGCAAGGTAATGTGTAAA ATCAGTCTCGGCTGTCAGAATAACTTCTAAAAGGTATTTTTATAAGCAGTTCAAGTTACTGAAAACCTTTTAAACCTTTCTGAAGTTCGT TAGTATAAATTACTTTTCTAGGATTATTAATAAAAGCCACATAGGTGGCAAGTTGTAGTTTTATATGGCTCTGTAGAGTGGTGAACCTTC TAGAGGAATATATGATTTATTCACAGTTCCTCAAGGCCTGGGGATGATGATCAGTTATACCTATTTTTGTGCAATTACATCATGTTGTAC ATTAGAAATGGAGAGTTTAATAGCTCTTTAACTGCTGTCCTCATTAGGTAATGATAAATATTTCCCTTAAATAATTGACTATTTTGCTGT GTTTTAAAAATGATTGAAATTTATCTTGCCATATCTCATAATTTCATGCACAAGTTGACTGAGCTAATCTTGAGAATATATTCGTAAAAT AGGAGCACATTTAGTTGAGGTATACAAGGTAGGACTCTAGACAAAACCTTCTATTTTAGCTTTAGTGAATTTCAAAAGTAATGGGTCTTG GAGTATAGATTTTTATTAGTAGCTTGAAAGAGCTTAATCATATGCAGTAAGTATTTTTATTACCAATAAATTTAAAATTTTTTAAGAAAA ATATTTTTATCCTAGGGCCAAGTGTTGCCTGCCACCAATCAGTAAGTTAGTCTATAACAAATTTTACCCTAACAGTTTTACCACCTAGTA ACAGTCATTTCTGAAAATATGTTGGATAGAAAGTCACTCTTTGGCAAAAGTGTTAGAATTTGCTTTTGTGCCATCTATTCCTTTTATGGC ATCTATCTTGAAAGTAATCTTGTATTGGAGATTGAAAGATGCTGTAATTTAGAAATTAACATGATATCTTAAATTACCTTTATGAAATAT AGTTTTGTATAATAGCATAGATTTTCCTTCAAAAAATGAACATTTATATATCTACAAAAATATGGAGAAGAGTAATTTGAAAGCCTACTT TCTGAAGAAAATGGTGGGATTTTTTTTTATCATGATTAAATATCAAAAAATTGCCCTATGAAAACTTTAAATCTCTAAAACATTTGAAAT ACTACCATATTTGTGATTTATTGAGAATAAAAATCCATTTTGAAATGTAAAATTTTTATGATCTGATTCAGTTTTAAGAAAACATGAATG AACTAGAAGATATTAAAAACATTTGACATTGGTAAGAAATATTGATACTGATATTGATTTTTATATAGGTATTTATTTCAGAATTGATAT TTTGAGAAAAATACATGTGAGTCATTTTTTCTGTTTCTCTTTTCTCTTAACGATTATCACTGTAATTCTGAATCTGAAAGGTAAAACAAT TAGTCAAAATATTATTGCCATCATTCTACCTGTGTTATGAAACTACTTATTCATAGTTAATTCTCATTAACACTTACATTTCCATAAAGA AAACTCAAGTATTAATAAAAGAGACTTTACTGGCTTAAGAGGGCTGTGAAAGATTTTTGATAGTGAATCATGACCCTAAGGGAGAGATTT GTGTGATAAAAGTATTGTATATAATAGATCAGCGATTTTTGTAAGGCAAACAGAATTTGTAAGTTGGCAGATCTTCCTAAGTTGCAAAAT GTAATGATGAGCTTGGTGGAGAAGAATGAGTCGTTCTTGGAATACCTATGTGCAGCCACTACCCATCTCAATGTCACCTTGTTTGCATTC TTGGATAGCTTGTATATGTAGTAGTTTGATGAATAATTTAAAGAAAAACACCTAAAATTTGAAAAATGATTGTAGGATCAAAAAAGGCAG ATGAAATTACTTAATACTCAGTGTTTTGGAGAGTATTCCTTTTAGTTTGTTGGTTGGCTGGTTTGAACGATAGAAATATGCAGCATGCAA TATATGCTTATATTTCATTTTAATTTCTGATATATAATGAACTTCTTGGGAGAGGTACTGAATCTTTGATGTTTTTTGTCATTGTTCTCA AGTGCAATATAACAATGTAACCAAATCTAGATAATTTCAAAGTTGTCATTAATTTAGTAAGCCTAATATAAACAAATATTTGTATTATTT TTGTTAGCAGGAAAGAGTGATTAAGTGAGGTTATTTACCCCTAAATGGTCCATTCTGCATTGTATTTCAGGCTGGAAATGAATTATTCTT TACCAGTTTTGAAACACTTTGAAATATCCTAAGGTAACTTGGAAGCTGTGTAGTATATCAAATTAATTTGCTACCTAATAACATAGAAAG TAAATATCTTTGTGGTCACCCACATTGGGTGAGACAGAAAATGAATCTGTTCTAAAATTTGTAATTTGCTAACTTGATTTGAGTTAGTGA >21072_21072_1_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000336411_ITGAV_chr2_187540331_ENST00000261023_length(amino acids)=159AA_BP=13 MASPGCQPGAPLSGCGVAQCLKIVCQVGRLDRGKSAILYVKSLLWTETFMNKENQNHSYSLKSSASFNVIEFPYKNLPIEDITNSTLVTT -------------------------------------------------------------- >21072_21072_2_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000336411_ITGAV_chr2_187540331_ENST00000374907_length(transcript)=4301nt_BP=255nt ACAGGTGGCCCTGCTACTGGCTGCAGCTCCAGCCCTGCCTCCTTCTCTAGCATATAAACAATCCAACAGCCTCACTGAATCACTGCTGTG CAGGGCAGGAAAGCTCCATGCACATAGCCCAGCAAAGAGCAACACAGAGCTGAAAGGAAGACTCAGAGGAGAGAGATAAGTAAGGAAAGT AGTGATGGCTCTCATCCCAGACTTGGCCATGGAAACCTGGCTTCTCCTGGCTGTCAGCCTGGTGCTCCTCTATCTGGTTGTGGAGTTGCT CAGTGCTTGAAGATTGTCTGCCAAGTTGGGAGATTAGACAGAGGAAAGAGTGCAATCTTGTACGTAAAGTCATTACTGTGGACTGAGACT TTTATGAATAAAGAAAATCAGAATCATTCCTATTCTCTGAAGTCGTCTGCTTCATTTAATGTCATAGAGTTTCCTTATAAGAATCTTCCA ATTGAGGATATCACCAACTCCACATTGGTTACCACTAATGTCACCTGGGGCATTCAGCCAGCGCCCATGCCTGTGCCTGTGTGGGTGATC ATTTTAGCAGTTCTAGCAGGATTGTTGCTACTGGCTGTTTTGGTATTTGTAATGTACAGGATGGGCTTTTTTAAACGGGTCCGGCCACCT CAAGAAGAACAAGAAAGGGAGCAGCTTCAACCTCATGAAAATGGTGAAGGAAACTCAGAAACTTAACTGCAGTTTTTAAGTTATGCTACA TCTTGACCCACTAGAATTAGCAACTTTATTATAGATTTAAACTTTCTTCATGAGGAGTAAAAATCCAAGGCTTTACTGCTGATAGTGCTA ATTGGCATTAACCACAAAATGAGAATTATATTTGTCAACCTTCTCCTTATAAATAAGTTCAGACATACATTTAATAACATAGGGTGACTT GTGTTTTTAGGTATTTAAATAATAAAATTTCAAGGGATAGTTTTTATTCAATGTATATAAGACAGGTAGTGCCTGATTTACTACTTTATA TAAAATAGTACCTCCTTCAGTTACTGTTTCTGATTTAATGTACGGAACTTTATTTGTTGTTGTTGTTGTTGTTGTTGTTGTTGTTTTAAA GCAGTCCAAATTTGGACCTTAGCAATCATGTCTTTTGTATAGGTACTTAATGTTAATACATATTACACTACAGTTTACTTTTCAGAATAC TAAAGACTTTATAACTGCATGAACTTGGATTTTTTTAATCACTCATATGGTAGAATTTTATAAACACATACATGATACCATCCAAATTCT TGCTTTTAATAACAAAGGTACAATATTTTGTTTTAGTATGAAAATCTGGTAGATCCTATTACACTTCTGTTTATATTAAATCCACAATAT TTTATTACATTTTTAACTTGTATAAATTTTAGGTCAAATCCTTCAAGCCAACCTATACTAAAAATTAGTTCCATAATCACAAATGGCTCT TTTGTGTAATTGTTTAATTTCACCTGAATATCATAATGCTTAAAGCCATATGGAGTTGGAAATTATTTCCAAAGCATATTTATTCCATTG TTTTAGTCTGGCTATTTACAGTATAAAAAAAGCATTTTTATTAAAATACTGTGTAGTTCTTTGAGATAGTTGCTTATGCATATAGTAAGT ATTACATTCTTAGAGTAGAGCAGAGTTTTTAGTTAGTATTAATTTATTTTCCTCCATTCATGTACTTTTCCTTATATTTCCAAAACTGTT ACTGAGAATGGGTCAAGATCAGTGAGAAATCTTTACAGTTGACAGGAACCTGGACCCCTTACCCCAACTTTATGAGTAATGCTTGGAATA AAAACTCTTAAGGCAACTCACTGATTTACTTCTAGCAATAGCATGATGTTACAGGAATATTACCTCTGTTTAAGCAAGGTAATGTGTAAA ATCAGTCTCGGCTGTCAGAATAACTTCTAAAAGGTATTTTTATAAGCAGTTCAAGTTACTGAAAACCTTTTAAACCTTTCTGAAGTTCGT TAGTATAAATTACTTTTCTAGGATTATTAATAAAAGCCACATAGGTGGCAAGTTGTAGTTTTATATGGCTCTGTAGAGTGGTGAACCTTC TAGAGGAATATATGATTTATTCACAGTTCCTCAAGGCCTGGGGATGATGATCAGTTATACCTATTTTTGTGCAATTACATCATGTTGTAC ATTAGAAATGGAGAGTTTAATAGCTCTTTAACTGCTGTCCTCATTAGGTAATGATAAATATTTCCCTTAAATAATTGACTATTTTGCTGT GTTTTAAAAATGATTGAAATTTATCTTGCCATATCTCATAATTTCATGCACAAGTTGACTGAGCTAATCTTGAGAATATATTCGTAAAAT AGGAGCACATTTAGTTGAGGTATACAAGGTAGGACTCTAGACAAAACCTTCTATTTTAGCTTTAGTGAATTTCAAAAGTAATGGGTCTTG GAGTATAGATTTTTATTAGTAGCTTGAAAGAGCTTAATCATATGCAGTAAGTATTTTTATTACCAATAAATTTAAAATTTTTTAAGAAAA ATATTTTTATCCTAGGGCCAAGTGTTGCCTGCCACCAATCAGTAAGTTAGTCTATAACAAATTTTACCCTAACAGTTTTACCACCTAGTA ACAGTCATTTCTGAAAATATGTTGGATAGAAAGTCACTCTTTGGCAAAAGTGTTAGAATTTGCTTTTGTGCCATCTATTCCTTTTATGGC ATCTATCTTGAAAGTAATCTTGTATTGGAGATTGAAAGATGCTGTAATTTAGAAATTAACATGATATCTTAAATTACCTTTATGAAATAT AGTTTTGTATAATAGCATAGATTTTCCTTCAAAAAATGAACATTTATATATCTACAAAAATATGGAGAAGAGTAATTTGAAAGCCTACTT TCTGAAGAAAATGGTGGGATTTTTTTTTATCATGATTAAATATCAAAAAATTGCCCTATGAAAACTTTAAATCTCTAAAACATTTGAAAT ACTACCATATTTGTGATTTATTGAGAATAAAAATCCATTTTGAAATGTAAAATTTTTATGATCTGATTCAGTTTTAAGAAAACATGAATG AACTAGAAGATATTAAAAACATTTGACATTGGTAAGAAATATTGATACTGATATTGATTTTTATATAGGTATTTATTTCAGAATTGATAT TTTGAGAAAAATACATGTGAGTCATTTTTTCTGTTTCTCTTTTCTCTTAACGATTATCACTGTAATTCTGAATCTGAAAGGTAAAACAAT TAGTCAAAATATTATTGCCATCATTCTACCTGTGTTATGAAACTACTTATTCATAGTTAATTCTCATTAACACTTACATTTCCATAAAGA AAACTCAAGTATTAATAAAAGAGACTTTACTGGCTTAAGAGGGCTGTGAAAGATTTTTGATAGTGAATCATGACCCTAAGGGAGAGATTT GTGTGATAAAAGTATTGTATATAATAGATCAGCGATTTTTGTAAGGCAAACAGAATTTGTAAGTTGGCAGATCTTCCTAAGTTGCAAAAT GTAATGATGAGCTTGGTGGAGAAGAATGAGTCGTTCTTGGAATACCTATGTGCAGCCACTACCCATCTCAATGTCACCTTGTTTGCATTC TTGGATAGCTTGTATATGTAGTAGTTTGATGAATAATTTAAAGAAAAACACCTAAAATTTGAAAAATGATTGTAGGATCAAAAAAGGCAG ATGAAATTACTTAATACTCAGTGTTTTGGAGAGTATTCCTTTTAGTTTGTTGGTTGGCTGGTTTGAACGATAGAAATATGCAGCATGCAA TATATGCTTATATTTCATTTTAATTTCTGATATATAATGAACTTCTTGGGAGAGGTACTGAATCTTTGATGTTTTTTGTCATTGTTCTCA AGTGCAATATAACAATGTAACCAAATCTAGATAATTTCAAAGTTGTCATTAATTTAGTAAGCCTAATATAAACAAATATTTGTATTATTT TTGTTAGCAGGAAAGAGTGATTAAGTGAGGTTATTTACCCCTAAATGGTCCATTCTGCATTGTATTTCAGGCTGGAAATGAATTATTCTT TACCAGTTTTGAAACACTTTGAAATATCCTAAGGTAACTTGGAAGCTGTGTAGTATATCAAATTAATTTGCTACCTAATAACATAGAAAG TAAATATCTTTGTGGTCACCCACATTGGGTGAGACAGAAAATGAATCTGTTCTAAAATTTGTAATTTGCTAACTTGATTTGAGTTAGTGA >21072_21072_2_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000336411_ITGAV_chr2_187540331_ENST00000374907_length(amino acids)=159AA_BP=13 MASPGCQPGAPLSGCGVAQCLKIVCQVGRLDRGKSAILYVKSLLWTETFMNKENQNHSYSLKSSASFNVIEFPYKNLPIEDITNSTLVTT -------------------------------------------------------------- >21072_21072_3_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000336411_ITGAV_chr2_187540331_ENST00000433736_length(transcript)=845nt_BP=255nt ACAGGTGGCCCTGCTACTGGCTGCAGCTCCAGCCCTGCCTCCTTCTCTAGCATATAAACAATCCAACAGCCTCACTGAATCACTGCTGTG CAGGGCAGGAAAGCTCCATGCACATAGCCCAGCAAAGAGCAACACAGAGCTGAAAGGAAGACTCAGAGGAGAGAGATAAGTAAGGAAAGT AGTGATGGCTCTCATCCCAGACTTGGCCATGGAAACCTGGCTTCTCCTGGCTGTCAGCCTGGTGCTCCTCTATCTGGTTGTGGAGTTGCT CAGTGCTTGAAGATTGTCTGCCAAGTTGGGAGATTAGACAGAGGAAAGAGTGCAATCTTGTACGTAAAGTCATTACTGTGGACTGAGACT TTTATGAATAAAGAAAATCAGAATCATTCCTATTCTCTGAAGTCGTCTGCTTCATTTAATGTCATAGAGTTTCCTTATAAGAATCTTCCA ATTGAGGATATCACCAACTCCACATTGGTTACCACTAATGTCACCTGGGGCATTCAGCCAGCGCCCATGCCTGTGCCTGTGTGGGTGATC ATTTTAGCAGTTCTAGCAGGATTGTTGCTACTGGCTGTTTTGGTATTTGTAATGTACAGGATGGGCTTTTTTAAACGGGTCCGGCCACCT CAAGAAGAACAAGAAAGGGAGCAGCTTCAACCTCATGAAAATGGTGAAGGAAACTCAGAAACTTAACTGCAGTTTTTAAGTTATGCTACA TCTTGACCCACTAGAATTAGCAACTTTATTATAGATTTAAACTTTCTTCATGAGGAGTAAAAATCCAAGGCTTTACTGCTGATAGTGCTA >21072_21072_3_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000336411_ITGAV_chr2_187540331_ENST00000433736_length(amino acids)=159AA_BP=13 MASPGCQPGAPLSGCGVAQCLKIVCQVGRLDRGKSAILYVKSLLWTETFMNKENQNHSYSLKSSASFNVIEFPYKNLPIEDITNSTLVTT -------------------------------------------------------------- >21072_21072_4_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000354593_ITGAV_chr2_187540331_ENST00000261023_length(transcript)=4225nt_BP=175nt CACTGCTGTGCAGGGCAGGAAAGCTCCATGCACATAGCCCAGCAAAGAGCAACACAGAGCTGAAAGGAAGACTCAGAGGAGAGAGATAAG TAAGGAAAGTAGTGATGGCTCTCATCCCAGACTTGGCCATGGAAACCTGGCTTCTCCTGGCTGTCAGCCTGGTGCTCCTCTATCTGGTTG TGGAGTTGCTCAGTGCTTGAAGATTGTCTGCCAAGTTGGGAGATTAGACAGAGGAAAGAGTGCAATCTTGTACGTAAAGTCATTACTGTG GACTGAGACTTTTATGAATAAAGAAAATCAGAATCATTCCTATTCTCTGAAGTCGTCTGCTTCATTTAATGTCATAGAGTTTCCTTATAA GAATCTTCCAATTGAGGATATCACCAACTCCACATTGGTTACCACTAATGTCACCTGGGGCATTCAGCCAGCGCCCATGCCTGTGCCTGT GTGGGTGATCATTTTAGCAGTTCTAGCAGGATTGTTGCTACTGGCTGTTTTGGTATTTGTAATGTACAGGATGGGCTTTTTTAAACGGGT CCGGCCACCTCAAGAAGAACAAGAAAGGGAGCAGCTTCAACCTCATGAAAATGGTGAAGGAAACTCAGAAACTTAACTGCAGTTTTTAAG TTATGCTACATCTTGACCCACTAGAATTAGCAACTTTATTATAGATTTAAACTTTCTTCATGAGGAGTAAAAATCCAAGGCTTTACTGCT GATAGTGCTAATTGGCATTAACCACAAAATGAGAATTATATTTGTCAACCTTCTCCTTATAAATAAGTTCAGACATACATTTAATAACAT AGGGTGACTTGTGTTTTTAGGTATTTAAATAATAAAATTTCAAGGGATAGTTTTTATTCAATGTATATAAGACAGGTAGTGCCTGATTTA CTACTTTATATAAAATAGTACCTCCTTCAGTTACTGTTTCTGATTTAATGTACGGAACTTTATTTGTTGTTGTTGTTGTTGTTGTTGTTG TTGTTTTAAAGCAGTCCAAATTTGGACCTTAGCAATCATGTCTTTTGTATAGGTACTTAATGTTAATACATATTACACTACAGTTTACTT TTCAGAATACTAAAGACTTTATAACTGCATGAACTTGGATTTTTTTAATCACTCATATGGTAGAATTTTATAAACACATACATGATACCA TCCAAATTCTTGCTTTTAATAACAAAGGTACAATATTTTGTTTTAGTATGAAAATCTGGTAGATCCTATTACACTTCTGTTTATATTAAA TCCACAATATTTTATTACATTTTTAACTTGTATAAATTTTAGGTCAAATCCTTCAAGCCAACCTATACTAAAAATTAGTTCCATAATCAC AAATGGCTCTTTTGTGTAATTGTTTAATTTCACCTGAATATCATAATGCTTAAAGCCATATGGAGTTGGAAATTATTTCCAAAGCATATT TATTCCATTGTTTTAGTCTGGCTATTTACAGTATAAAAAAAGCATTTTTATTAAAATACTGTGTAGTTCTTTGAGATAGTTGCTTATGCA TATAGTAAGTATTACATTCTTAGAGTAGAGCAGAGTTTTTAGTTAGTATTAATTTATTTTCCTCCATTCATGTACTTTTCCTTATATTTC CAAAACTGTTACTGAGAATGGGTCAAGATCAGTGAGAAATCTTTACAGTTGACAGGAACCTGGACCCCTTACCCCAACTTTATGAGTAAT GCTTGGAATAAAAACTCTTAAGGCAACTCACTGATTTACTTCTAGCAATAGCATGATGTTACAGGAATATTACCTCTGTTTAAGCAAGGT AATGTGTAAAATCAGTCTCGGCTGTCAGAATAACTTCTAAAAGGTATTTTTATAAGCAGTTCAAGTTACTGAAAACCTTTTAAACCTTTC TGAAGTTCGTTAGTATAAATTACTTTTCTAGGATTATTAATAAAAGCCACATAGGTGGCAAGTTGTAGTTTTATATGGCTCTGTAGAGTG GTGAACCTTCTAGAGGAATATATGATTTATTCACAGTTCCTCAAGGCCTGGGGATGATGATCAGTTATACCTATTTTTGTGCAATTACAT CATGTTGTACATTAGAAATGGAGAGTTTAATAGCTCTTTAACTGCTGTCCTCATTAGGTAATGATAAATATTTCCCTTAAATAATTGACT ATTTTGCTGTGTTTTAAAAATGATTGAAATTTATCTTGCCATATCTCATAATTTCATGCACAAGTTGACTGAGCTAATCTTGAGAATATA TTCGTAAAATAGGAGCACATTTAGTTGAGGTATACAAGGTAGGACTCTAGACAAAACCTTCTATTTTAGCTTTAGTGAATTTCAAAAGTA ATGGGTCTTGGAGTATAGATTTTTATTAGTAGCTTGAAAGAGCTTAATCATATGCAGTAAGTATTTTTATTACCAATAAATTTAAAATTT TTTAAGAAAAATATTTTTATCCTAGGGCCAAGTGTTGCCTGCCACCAATCAGTAAGTTAGTCTATAACAAATTTTACCCTAACAGTTTTA CCACCTAGTAACAGTCATTTCTGAAAATATGTTGGATAGAAAGTCACTCTTTGGCAAAAGTGTTAGAATTTGCTTTTGTGCCATCTATTC CTTTTATGGCATCTATCTTGAAAGTAATCTTGTATTGGAGATTGAAAGATGCTGTAATTTAGAAATTAACATGATATCTTAAATTACCTT TATGAAATATAGTTTTGTATAATAGCATAGATTTTCCTTCAAAAAATGAACATTTATATATCTACAAAAATATGGAGAAGAGTAATTTGA AAGCCTACTTTCTGAAGAAAATGGTGGGATTTTTTTTTATCATGATTAAATATCAAAAAATTGCCCTATGAAAACTTTAAATCTCTAAAA CATTTGAAATACTACCATATTTGTGATTTATTGAGAATAAAAATCCATTTTGAAATGTAAAATTTTTATGATCTGATTCAGTTTTAAGAA AACATGAATGAACTAGAAGATATTAAAAACATTTGACATTGGTAAGAAATATTGATACTGATATTGATTTTTATATAGGTATTTATTTCA GAATTGATATTTTGAGAAAAATACATGTGAGTCATTTTTTCTGTTTCTCTTTTCTCTTAACGATTATCACTGTAATTCTGAATCTGAAAG GTAAAACAATTAGTCAAAATATTATTGCCATCATTCTACCTGTGTTATGAAACTACTTATTCATAGTTAATTCTCATTAACACTTACATT TCCATAAAGAAAACTCAAGTATTAATAAAAGAGACTTTACTGGCTTAAGAGGGCTGTGAAAGATTTTTGATAGTGAATCATGACCCTAAG GGAGAGATTTGTGTGATAAAAGTATTGTATATAATAGATCAGCGATTTTTGTAAGGCAAACAGAATTTGTAAGTTGGCAGATCTTCCTAA GTTGCAAAATGTAATGATGAGCTTGGTGGAGAAGAATGAGTCGTTCTTGGAATACCTATGTGCAGCCACTACCCATCTCAATGTCACCTT GTTTGCATTCTTGGATAGCTTGTATATGTAGTAGTTTGATGAATAATTTAAAGAAAAACACCTAAAATTTGAAAAATGATTGTAGGATCA AAAAAGGCAGATGAAATTACTTAATACTCAGTGTTTTGGAGAGTATTCCTTTTAGTTTGTTGGTTGGCTGGTTTGAACGATAGAAATATG CAGCATGCAATATATGCTTATATTTCATTTTAATTTCTGATATATAATGAACTTCTTGGGAGAGGTACTGAATCTTTGATGTTTTTTGTC ATTGTTCTCAAGTGCAATATAACAATGTAACCAAATCTAGATAATTTCAAAGTTGTCATTAATTTAGTAAGCCTAATATAAACAAATATT TGTATTATTTTTGTTAGCAGGAAAGAGTGATTAAGTGAGGTTATTTACCCCTAAATGGTCCATTCTGCATTGTATTTCAGGCTGGAAATG AATTATTCTTTACCAGTTTTGAAACACTTTGAAATATCCTAAGGTAACTTGGAAGCTGTGTAGTATATCAAATTAATTTGCTACCTAATA ACATAGAAAGTAAATATCTTTGTGGTCACCCACATTGGGTGAGACAGAAAATGAATCTGTTCTAAAATTTGTAATTTGCTAACTTGATTT >21072_21072_4_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000354593_ITGAV_chr2_187540331_ENST00000261023_length(amino acids)=159AA_BP=13 MASPGCQPGAPLSGCGVAQCLKIVCQVGRLDRGKSAILYVKSLLWTETFMNKENQNHSYSLKSSASFNVIEFPYKNLPIEDITNSTLVTT -------------------------------------------------------------- >21072_21072_5_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000354593_ITGAV_chr2_187540331_ENST00000374907_length(transcript)=4221nt_BP=175nt CACTGCTGTGCAGGGCAGGAAAGCTCCATGCACATAGCCCAGCAAAGAGCAACACAGAGCTGAAAGGAAGACTCAGAGGAGAGAGATAAG TAAGGAAAGTAGTGATGGCTCTCATCCCAGACTTGGCCATGGAAACCTGGCTTCTCCTGGCTGTCAGCCTGGTGCTCCTCTATCTGGTTG TGGAGTTGCTCAGTGCTTGAAGATTGTCTGCCAAGTTGGGAGATTAGACAGAGGAAAGAGTGCAATCTTGTACGTAAAGTCATTACTGTG GACTGAGACTTTTATGAATAAAGAAAATCAGAATCATTCCTATTCTCTGAAGTCGTCTGCTTCATTTAATGTCATAGAGTTTCCTTATAA GAATCTTCCAATTGAGGATATCACCAACTCCACATTGGTTACCACTAATGTCACCTGGGGCATTCAGCCAGCGCCCATGCCTGTGCCTGT GTGGGTGATCATTTTAGCAGTTCTAGCAGGATTGTTGCTACTGGCTGTTTTGGTATTTGTAATGTACAGGATGGGCTTTTTTAAACGGGT CCGGCCACCTCAAGAAGAACAAGAAAGGGAGCAGCTTCAACCTCATGAAAATGGTGAAGGAAACTCAGAAACTTAACTGCAGTTTTTAAG TTATGCTACATCTTGACCCACTAGAATTAGCAACTTTATTATAGATTTAAACTTTCTTCATGAGGAGTAAAAATCCAAGGCTTTACTGCT GATAGTGCTAATTGGCATTAACCACAAAATGAGAATTATATTTGTCAACCTTCTCCTTATAAATAAGTTCAGACATACATTTAATAACAT AGGGTGACTTGTGTTTTTAGGTATTTAAATAATAAAATTTCAAGGGATAGTTTTTATTCAATGTATATAAGACAGGTAGTGCCTGATTTA CTACTTTATATAAAATAGTACCTCCTTCAGTTACTGTTTCTGATTTAATGTACGGAACTTTATTTGTTGTTGTTGTTGTTGTTGTTGTTG TTGTTTTAAAGCAGTCCAAATTTGGACCTTAGCAATCATGTCTTTTGTATAGGTACTTAATGTTAATACATATTACACTACAGTTTACTT TTCAGAATACTAAAGACTTTATAACTGCATGAACTTGGATTTTTTTAATCACTCATATGGTAGAATTTTATAAACACATACATGATACCA TCCAAATTCTTGCTTTTAATAACAAAGGTACAATATTTTGTTTTAGTATGAAAATCTGGTAGATCCTATTACACTTCTGTTTATATTAAA TCCACAATATTTTATTACATTTTTAACTTGTATAAATTTTAGGTCAAATCCTTCAAGCCAACCTATACTAAAAATTAGTTCCATAATCAC AAATGGCTCTTTTGTGTAATTGTTTAATTTCACCTGAATATCATAATGCTTAAAGCCATATGGAGTTGGAAATTATTTCCAAAGCATATT TATTCCATTGTTTTAGTCTGGCTATTTACAGTATAAAAAAAGCATTTTTATTAAAATACTGTGTAGTTCTTTGAGATAGTTGCTTATGCA TATAGTAAGTATTACATTCTTAGAGTAGAGCAGAGTTTTTAGTTAGTATTAATTTATTTTCCTCCATTCATGTACTTTTCCTTATATTTC CAAAACTGTTACTGAGAATGGGTCAAGATCAGTGAGAAATCTTTACAGTTGACAGGAACCTGGACCCCTTACCCCAACTTTATGAGTAAT GCTTGGAATAAAAACTCTTAAGGCAACTCACTGATTTACTTCTAGCAATAGCATGATGTTACAGGAATATTACCTCTGTTTAAGCAAGGT AATGTGTAAAATCAGTCTCGGCTGTCAGAATAACTTCTAAAAGGTATTTTTATAAGCAGTTCAAGTTACTGAAAACCTTTTAAACCTTTC TGAAGTTCGTTAGTATAAATTACTTTTCTAGGATTATTAATAAAAGCCACATAGGTGGCAAGTTGTAGTTTTATATGGCTCTGTAGAGTG GTGAACCTTCTAGAGGAATATATGATTTATTCACAGTTCCTCAAGGCCTGGGGATGATGATCAGTTATACCTATTTTTGTGCAATTACAT CATGTTGTACATTAGAAATGGAGAGTTTAATAGCTCTTTAACTGCTGTCCTCATTAGGTAATGATAAATATTTCCCTTAAATAATTGACT ATTTTGCTGTGTTTTAAAAATGATTGAAATTTATCTTGCCATATCTCATAATTTCATGCACAAGTTGACTGAGCTAATCTTGAGAATATA TTCGTAAAATAGGAGCACATTTAGTTGAGGTATACAAGGTAGGACTCTAGACAAAACCTTCTATTTTAGCTTTAGTGAATTTCAAAAGTA ATGGGTCTTGGAGTATAGATTTTTATTAGTAGCTTGAAAGAGCTTAATCATATGCAGTAAGTATTTTTATTACCAATAAATTTAAAATTT TTTAAGAAAAATATTTTTATCCTAGGGCCAAGTGTTGCCTGCCACCAATCAGTAAGTTAGTCTATAACAAATTTTACCCTAACAGTTTTA CCACCTAGTAACAGTCATTTCTGAAAATATGTTGGATAGAAAGTCACTCTTTGGCAAAAGTGTTAGAATTTGCTTTTGTGCCATCTATTC CTTTTATGGCATCTATCTTGAAAGTAATCTTGTATTGGAGATTGAAAGATGCTGTAATTTAGAAATTAACATGATATCTTAAATTACCTT TATGAAATATAGTTTTGTATAATAGCATAGATTTTCCTTCAAAAAATGAACATTTATATATCTACAAAAATATGGAGAAGAGTAATTTGA AAGCCTACTTTCTGAAGAAAATGGTGGGATTTTTTTTTATCATGATTAAATATCAAAAAATTGCCCTATGAAAACTTTAAATCTCTAAAA CATTTGAAATACTACCATATTTGTGATTTATTGAGAATAAAAATCCATTTTGAAATGTAAAATTTTTATGATCTGATTCAGTTTTAAGAA AACATGAATGAACTAGAAGATATTAAAAACATTTGACATTGGTAAGAAATATTGATACTGATATTGATTTTTATATAGGTATTTATTTCA GAATTGATATTTTGAGAAAAATACATGTGAGTCATTTTTTCTGTTTCTCTTTTCTCTTAACGATTATCACTGTAATTCTGAATCTGAAAG GTAAAACAATTAGTCAAAATATTATTGCCATCATTCTACCTGTGTTATGAAACTACTTATTCATAGTTAATTCTCATTAACACTTACATT TCCATAAAGAAAACTCAAGTATTAATAAAAGAGACTTTACTGGCTTAAGAGGGCTGTGAAAGATTTTTGATAGTGAATCATGACCCTAAG GGAGAGATTTGTGTGATAAAAGTATTGTATATAATAGATCAGCGATTTTTGTAAGGCAAACAGAATTTGTAAGTTGGCAGATCTTCCTAA GTTGCAAAATGTAATGATGAGCTTGGTGGAGAAGAATGAGTCGTTCTTGGAATACCTATGTGCAGCCACTACCCATCTCAATGTCACCTT GTTTGCATTCTTGGATAGCTTGTATATGTAGTAGTTTGATGAATAATTTAAAGAAAAACACCTAAAATTTGAAAAATGATTGTAGGATCA AAAAAGGCAGATGAAATTACTTAATACTCAGTGTTTTGGAGAGTATTCCTTTTAGTTTGTTGGTTGGCTGGTTTGAACGATAGAAATATG CAGCATGCAATATATGCTTATATTTCATTTTAATTTCTGATATATAATGAACTTCTTGGGAGAGGTACTGAATCTTTGATGTTTTTTGTC ATTGTTCTCAAGTGCAATATAACAATGTAACCAAATCTAGATAATTTCAAAGTTGTCATTAATTTAGTAAGCCTAATATAAACAAATATT TGTATTATTTTTGTTAGCAGGAAAGAGTGATTAAGTGAGGTTATTTACCCCTAAATGGTCCATTCTGCATTGTATTTCAGGCTGGAAATG AATTATTCTTTACCAGTTTTGAAACACTTTGAAATATCCTAAGGTAACTTGGAAGCTGTGTAGTATATCAAATTAATTTGCTACCTAATA ACATAGAAAGTAAATATCTTTGTGGTCACCCACATTGGGTGAGACAGAAAATGAATCTGTTCTAAAATTTGTAATTTGCTAACTTGATTT >21072_21072_5_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000354593_ITGAV_chr2_187540331_ENST00000374907_length(amino acids)=159AA_BP=13 MASPGCQPGAPLSGCGVAQCLKIVCQVGRLDRGKSAILYVKSLLWTETFMNKENQNHSYSLKSSASFNVIEFPYKNLPIEDITNSTLVTT -------------------------------------------------------------- >21072_21072_6_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000354593_ITGAV_chr2_187540331_ENST00000433736_length(transcript)=765nt_BP=175nt CACTGCTGTGCAGGGCAGGAAAGCTCCATGCACATAGCCCAGCAAAGAGCAACACAGAGCTGAAAGGAAGACTCAGAGGAGAGAGATAAG TAAGGAAAGTAGTGATGGCTCTCATCCCAGACTTGGCCATGGAAACCTGGCTTCTCCTGGCTGTCAGCCTGGTGCTCCTCTATCTGGTTG TGGAGTTGCTCAGTGCTTGAAGATTGTCTGCCAAGTTGGGAGATTAGACAGAGGAAAGAGTGCAATCTTGTACGTAAAGTCATTACTGTG GACTGAGACTTTTATGAATAAAGAAAATCAGAATCATTCCTATTCTCTGAAGTCGTCTGCTTCATTTAATGTCATAGAGTTTCCTTATAA GAATCTTCCAATTGAGGATATCACCAACTCCACATTGGTTACCACTAATGTCACCTGGGGCATTCAGCCAGCGCCCATGCCTGTGCCTGT GTGGGTGATCATTTTAGCAGTTCTAGCAGGATTGTTGCTACTGGCTGTTTTGGTATTTGTAATGTACAGGATGGGCTTTTTTAAACGGGT CCGGCCACCTCAAGAAGAACAAGAAAGGGAGCAGCTTCAACCTCATGAAAATGGTGAAGGAAACTCAGAAACTTAACTGCAGTTTTTAAG TTATGCTACATCTTGACCCACTAGAATTAGCAACTTTATTATAGATTTAAACTTTCTTCATGAGGAGTAAAAATCCAAGGCTTTACTGCT >21072_21072_6_CYP3A4-ITGAV_CYP3A4_chr7_99381634_ENST00000354593_ITGAV_chr2_187540331_ENST00000433736_length(amino acids)=159AA_BP=13 MASPGCQPGAPLSGCGVAQCLKIVCQVGRLDRGKSAILYVKSLLWTETFMNKENQNHSYSLKSSASFNVIEFPYKNLPIEDITNSTLVTT -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for CYP3A4-ITGAV |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for CYP3A4-ITGAV |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for CYP3A4-ITGAV |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
