
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:DGCR2-CLTCL1 (FusionGDB2 ID:22400) |
Fusion Gene Summary for DGCR2-CLTCL1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: DGCR2-CLTCL1 | Fusion gene ID: 22400 | Hgene | Tgene | Gene symbol | DGCR2 | CLTCL1 | Gene ID | 9993 | 8218 |
| Gene name | DiGeorge syndrome critical region gene 2 | clathrin heavy chain like 1 | |
| Synonyms | DGS-C|IDD|LAN|SEZ-12 | CHC22|CLH22|CLTCL|CLTD | |
| Cytomap | 22q11.21 | 22q11.21 | |
| Type of gene | protein-coding | protein-coding | |
| Description | integral membrane protein DGCR2/IDDDiGeorge syndrome critical region protein 2integral membrane protein deleted in DiGeorge syndrome | clathrin heavy chain 2CLH-22Clathrin, heavy polypeptide Dclathrin heavy chain on chromosome 22clathrin, heavy polypeptide-like 1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | P98153 | P53675 | |
| Ensembl transtripts involved in fusion gene | ENST00000545799, ENST00000263196, ENST00000537045, ENST00000473832, | ENST00000263200, ENST00000353891, ENST00000427926, ENST00000442042, | |
| Fusion gene scores | * DoF score | 16 X 15 X 8=1920 | 9 X 7 X 7=441 |
| # samples | 21 | 12 | |
| ** MAII score | log2(21/1920*10)=-3.1926450779424 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(12/441*10)=-1.877744249949 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: DGCR2 [Title/Abstract] AND CLTCL1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | DGCR2(19076881)-CLTCL1(19263353), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | DGCR2-CLTCL1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. DGCR2-CLTCL1 seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. DGCR2-CLTCL1 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | CLTCL1 | GO:0000278 | mitotic cell cycle | 19509056 |
| Tgene | CLTCL1 | GO:0000278 | mitotic cell cycle | 20065094 |
| Tgene | CLTCL1 | GO:0006898 | receptor-mediated endocytosis | 19509056 |
 Fusion gene breakpoints across DGCR2 (5'-gene) Fusion gene breakpoints across DGCR2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
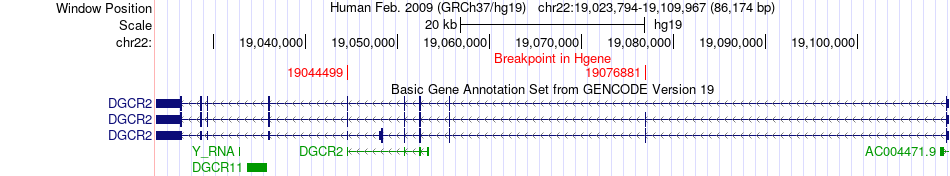 |
 Fusion gene breakpoints across CLTCL1 (3'-gene) Fusion gene breakpoints across CLTCL1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
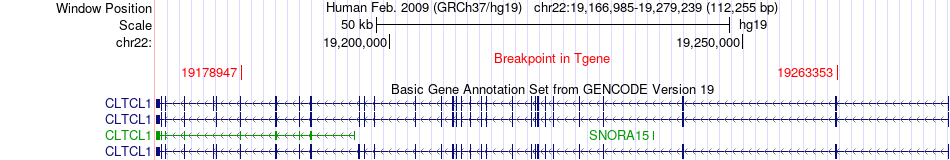 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-A7-A26G-01A | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| ChimerDB4 | BRCA | TCGA-A7-A4SC-01A | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| ChimerDB4 | STAD | TCGA-CD-A489-01A | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
Top |
Fusion Gene ORF analysis for DGCR2-CLTCL1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 3UTR-3CDS | ENST00000545799 | ENST00000263200 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| 3UTR-3CDS | ENST00000545799 | ENST00000353891 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| 3UTR-3CDS | ENST00000545799 | ENST00000427926 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| 3UTR-intron | ENST00000545799 | ENST00000442042 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| 5CDS-5UTR | ENST00000263196 | ENST00000442042 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| 5CDS-5UTR | ENST00000545799 | ENST00000442042 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| 5CDS-intron | ENST00000263196 | ENST00000442042 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| 5CDS-intron | ENST00000263196 | ENST00000442042 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| 5CDS-intron | ENST00000537045 | ENST00000442042 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| 5CDS-intron | ENST00000545799 | ENST00000442042 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| Frame-shift | ENST00000263196 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| Frame-shift | ENST00000263196 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| Frame-shift | ENST00000263196 | ENST00000263200 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| Frame-shift | ENST00000263196 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| Frame-shift | ENST00000263196 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| Frame-shift | ENST00000263196 | ENST00000353891 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| Frame-shift | ENST00000263196 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| Frame-shift | ENST00000263196 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| Frame-shift | ENST00000263196 | ENST00000427926 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| Frame-shift | ENST00000537045 | ENST00000263200 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| Frame-shift | ENST00000537045 | ENST00000353891 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| Frame-shift | ENST00000537045 | ENST00000427926 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| In-frame | ENST00000545799 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| In-frame | ENST00000545799 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| In-frame | ENST00000545799 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| In-frame | ENST00000545799 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| In-frame | ENST00000545799 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| In-frame | ENST00000545799 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| intron-3CDS | ENST00000473832 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-3CDS | ENST00000473832 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| intron-3CDS | ENST00000473832 | ENST00000263200 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-3CDS | ENST00000473832 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-3CDS | ENST00000473832 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| intron-3CDS | ENST00000473832 | ENST00000353891 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-3CDS | ENST00000473832 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-3CDS | ENST00000473832 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| intron-3CDS | ENST00000473832 | ENST00000427926 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-3CDS | ENST00000537045 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-3CDS | ENST00000537045 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| intron-3CDS | ENST00000537045 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-3CDS | ENST00000537045 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| intron-3CDS | ENST00000537045 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-3CDS | ENST00000537045 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| intron-5UTR | ENST00000473832 | ENST00000442042 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| intron-5UTR | ENST00000537045 | ENST00000442042 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - |
| intron-intron | ENST00000473832 | ENST00000442042 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-intron | ENST00000473832 | ENST00000442042 | DGCR2 | chr22 | 19044499 | - | CLTCL1 | chr22 | 19263353 | - |
| intron-intron | ENST00000537045 | ENST00000442042 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000545799 | DGCR2 | chr22 | 19076881 | - | ENST00000353891 | CLTCL1 | chr22 | 19263353 | - | 5630 | 403 | 256 | 5112 | 1618 |
| ENST00000545799 | DGCR2 | chr22 | 19076881 | - | ENST00000263200 | CLTCL1 | chr22 | 19263353 | - | 5801 | 403 | 256 | 5283 | 1675 |
| ENST00000545799 | DGCR2 | chr22 | 19076881 | - | ENST00000427926 | CLTCL1 | chr22 | 19263353 | - | 5798 | 403 | 256 | 5283 | 1675 |
| ENST00000545799 | DGCR2 | chr22 | 19076881 | - | ENST00000353891 | CLTCL1 | chr22 | 19178947 | - | 1481 | 403 | 256 | 963 | 235 |
| ENST00000545799 | DGCR2 | chr22 | 19076881 | - | ENST00000263200 | CLTCL1 | chr22 | 19178947 | - | 1652 | 403 | 256 | 1134 | 292 |
| ENST00000545799 | DGCR2 | chr22 | 19076881 | - | ENST00000427926 | CLTCL1 | chr22 | 19178947 | - | 1649 | 403 | 256 | 1134 | 292 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000545799 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - | 0.003844786 | 0.9961552 |
| ENST00000545799 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - | 0.003822658 | 0.9961773 |
| ENST00000545799 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19263353 | - | 0.003831981 | 0.99616796 |
| ENST00000545799 | ENST00000353891 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - | 0.005450898 | 0.9945491 |
| ENST00000545799 | ENST00000263200 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - | 0.005775743 | 0.99422425 |
| ENST00000545799 | ENST00000427926 | DGCR2 | chr22 | 19076881 | - | CLTCL1 | chr22 | 19178947 | - | 0.005736781 | 0.99426323 |
Top |
Fusion Genomic Features for DGCR2-CLTCL1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
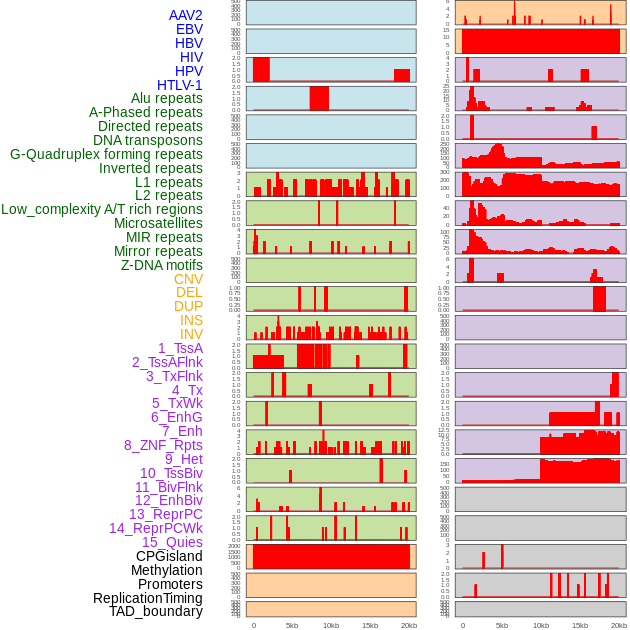 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for DGCR2-CLTCL1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr22:19076881/chr22:19263353) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| DGCR2 | CLTCL1 |
| FUNCTION: Putative adhesion receptor, that could be involved in cell-cell or cell-matrix interactions required for normal cell differentiation and migration. | FUNCTION: Clathrin is the major protein of the polyhedral coat of coated pits and vesicles. Two different adapter protein complexes link the clathrin lattice either to the plasma membrane or to the trans-Golgi network (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000263196 | - | 2 | 10 | 28_68 | 67 | 551.0 | Domain | LDL-receptor class A |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000263196 | - | 2 | 10 | 28_68 | 67 | 551.0 | Domain | LDL-receptor class A |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 1551_1640 | 1397 | 1712.3333333333333 | Region | Trimerization | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 1551_1640 | 1397 | 1665.0 | Region | Trimerization | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 1551_1640 | 1397 | 1710.6666666666667 | Region | Trimerization | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 108_149 | 14 | 1712.3333333333333 | Region | Note=WD40-like repeat 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 1213_1522 | 14 | 1712.3333333333333 | Region | Involved in binding clathrin light chain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 150_195 | 14 | 1712.3333333333333 | Region | Note=WD40-like repeat 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 1551_1640 | 14 | 1712.3333333333333 | Region | Trimerization | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 196_257 | 14 | 1712.3333333333333 | Region | Note=WD40-like repeat 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 24_67 | 14 | 1712.3333333333333 | Region | Note=WD40-like repeat 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 258_301 | 14 | 1712.3333333333333 | Region | Note=WD40-like repeat 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 302_330 | 14 | 1712.3333333333333 | Region | Note=WD40-like repeat 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 449_465 | 14 | 1712.3333333333333 | Region | Binding site for the uncoating ATPase%2C involved in lattice disassembly | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 480_523 | 14 | 1712.3333333333333 | Region | Note=Flexible linker | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 524_1640 | 14 | 1712.3333333333333 | Region | Note=Heavy chain arm | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 524_634 | 14 | 1712.3333333333333 | Region | Note=Distal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 639_1640 | 14 | 1712.3333333333333 | Region | Note=Proximal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 68_107 | 14 | 1712.3333333333333 | Region | Note=WD40-like repeat 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 108_149 | 14 | 1665.0 | Region | Note=WD40-like repeat 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 1213_1522 | 14 | 1665.0 | Region | Involved in binding clathrin light chain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 150_195 | 14 | 1665.0 | Region | Note=WD40-like repeat 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 1551_1640 | 14 | 1665.0 | Region | Trimerization | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 196_257 | 14 | 1665.0 | Region | Note=WD40-like repeat 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 24_67 | 14 | 1665.0 | Region | Note=WD40-like repeat 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 258_301 | 14 | 1665.0 | Region | Note=WD40-like repeat 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 302_330 | 14 | 1665.0 | Region | Note=WD40-like repeat 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 449_465 | 14 | 1665.0 | Region | Binding site for the uncoating ATPase%2C involved in lattice disassembly | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 480_523 | 14 | 1665.0 | Region | Note=Flexible linker | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 524_1640 | 14 | 1665.0 | Region | Note=Heavy chain arm | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 524_634 | 14 | 1665.0 | Region | Note=Distal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 639_1640 | 14 | 1665.0 | Region | Note=Proximal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 68_107 | 14 | 1665.0 | Region | Note=WD40-like repeat 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 108_149 | 14 | 1710.6666666666667 | Region | Note=WD40-like repeat 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 1213_1522 | 14 | 1710.6666666666667 | Region | Involved in binding clathrin light chain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 150_195 | 14 | 1710.6666666666667 | Region | Note=WD40-like repeat 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 1551_1640 | 14 | 1710.6666666666667 | Region | Trimerization | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 196_257 | 14 | 1710.6666666666667 | Region | Note=WD40-like repeat 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 24_67 | 14 | 1710.6666666666667 | Region | Note=WD40-like repeat 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 258_301 | 14 | 1710.6666666666667 | Region | Note=WD40-like repeat 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 302_330 | 14 | 1710.6666666666667 | Region | Note=WD40-like repeat 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 449_465 | 14 | 1710.6666666666667 | Region | Binding site for the uncoating ATPase%2C involved in lattice disassembly | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 480_523 | 14 | 1710.6666666666667 | Region | Note=Flexible linker | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 524_1640 | 14 | 1710.6666666666667 | Region | Note=Heavy chain arm | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 524_634 | 14 | 1710.6666666666667 | Region | Note=Distal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 639_1640 | 14 | 1710.6666666666667 | Region | Note=Proximal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 68_107 | 14 | 1710.6666666666667 | Region | Note=WD40-like repeat 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 1423_1566 | 1397 | 1712.3333333333333 | Repeat | Note=CHCR 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 1423_1566 | 1397 | 1665.0 | Repeat | Note=CHCR 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 1423_1566 | 1397 | 1710.6666666666667 | Repeat | Note=CHCR 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 1128_1269 | 14 | 1712.3333333333333 | Repeat | Note=CHCR 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 1274_1420 | 14 | 1712.3333333333333 | Repeat | Note=CHCR 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 1423_1566 | 14 | 1712.3333333333333 | Repeat | Note=CHCR 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 537_683 | 14 | 1712.3333333333333 | Repeat | Note=CHCR 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 686_828 | 14 | 1712.3333333333333 | Repeat | Note=CHCR 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 833_972 | 14 | 1712.3333333333333 | Repeat | Note=CHCR 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 979_1124 | 14 | 1712.3333333333333 | Repeat | Note=CHCR 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 1128_1269 | 14 | 1665.0 | Repeat | Note=CHCR 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 1274_1420 | 14 | 1665.0 | Repeat | Note=CHCR 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 1423_1566 | 14 | 1665.0 | Repeat | Note=CHCR 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 537_683 | 14 | 1665.0 | Repeat | Note=CHCR 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 686_828 | 14 | 1665.0 | Repeat | Note=CHCR 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 833_972 | 14 | 1665.0 | Repeat | Note=CHCR 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 979_1124 | 14 | 1665.0 | Repeat | Note=CHCR 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 1128_1269 | 14 | 1710.6666666666667 | Repeat | Note=CHCR 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 1274_1420 | 14 | 1710.6666666666667 | Repeat | Note=CHCR 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 1423_1566 | 14 | 1710.6666666666667 | Repeat | Note=CHCR 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 537_683 | 14 | 1710.6666666666667 | Repeat | Note=CHCR 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 686_828 | 14 | 1710.6666666666667 | Repeat | Note=CHCR 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 833_972 | 14 | 1710.6666666666667 | Repeat | Note=CHCR 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 979_1124 | 14 | 1710.6666666666667 | Repeat | Note=CHCR 4 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000263196 | - | 2 | 10 | 115_241 | 67 | 551.0 | Domain | C-type lectin |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000263196 | - | 2 | 10 | 270_333 | 67 | 551.0 | Domain | Note=VWFC |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000537045 | - | 1 | 9 | 115_241 | 0 | 510.0 | Domain | C-type lectin |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000537045 | - | 1 | 9 | 270_333 | 0 | 510.0 | Domain | Note=VWFC |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000537045 | - | 1 | 9 | 28_68 | 0 | 510.0 | Domain | LDL-receptor class A |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000263196 | - | 2 | 10 | 115_241 | 67 | 551.0 | Domain | C-type lectin |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000263196 | - | 2 | 10 | 270_333 | 67 | 551.0 | Domain | Note=VWFC |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000537045 | - | 1 | 9 | 115_241 | 0 | 510.0 | Domain | C-type lectin |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000537045 | - | 1 | 9 | 270_333 | 0 | 510.0 | Domain | Note=VWFC |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000537045 | - | 1 | 9 | 28_68 | 0 | 510.0 | Domain | LDL-receptor class A |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000263196 | - | 2 | 10 | 21_349 | 67 | 551.0 | Topological domain | Extracellular |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000263196 | - | 2 | 10 | 369_550 | 67 | 551.0 | Topological domain | Cytoplasmic |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000537045 | - | 1 | 9 | 21_349 | 0 | 510.0 | Topological domain | Extracellular |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000537045 | - | 1 | 9 | 369_550 | 0 | 510.0 | Topological domain | Cytoplasmic |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000263196 | - | 2 | 10 | 21_349 | 67 | 551.0 | Topological domain | Extracellular |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000263196 | - | 2 | 10 | 369_550 | 67 | 551.0 | Topological domain | Cytoplasmic |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000537045 | - | 1 | 9 | 21_349 | 0 | 510.0 | Topological domain | Extracellular |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000537045 | - | 1 | 9 | 369_550 | 0 | 510.0 | Topological domain | Cytoplasmic |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000263196 | - | 2 | 10 | 350_368 | 67 | 551.0 | Transmembrane | Helical |
| Hgene | DGCR2 | chr22:19076881 | chr22:19178947 | ENST00000537045 | - | 1 | 9 | 350_368 | 0 | 510.0 | Transmembrane | Helical |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000263196 | - | 2 | 10 | 350_368 | 67 | 551.0 | Transmembrane | Helical |
| Hgene | DGCR2 | chr22:19076881 | chr22:19263353 | ENST00000537045 | - | 1 | 9 | 350_368 | 0 | 510.0 | Transmembrane | Helical |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 108_149 | 1397 | 1712.3333333333333 | Region | Note=WD40-like repeat 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 1213_1522 | 1397 | 1712.3333333333333 | Region | Involved in binding clathrin light chain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 150_195 | 1397 | 1712.3333333333333 | Region | Note=WD40-like repeat 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 196_257 | 1397 | 1712.3333333333333 | Region | Note=WD40-like repeat 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 24_67 | 1397 | 1712.3333333333333 | Region | Note=WD40-like repeat 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 258_301 | 1397 | 1712.3333333333333 | Region | Note=WD40-like repeat 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 2_479 | 1397 | 1712.3333333333333 | Region | Note=Globular terminal domain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 302_330 | 1397 | 1712.3333333333333 | Region | Note=WD40-like repeat 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 449_465 | 1397 | 1712.3333333333333 | Region | Binding site for the uncoating ATPase%2C involved in lattice disassembly | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 480_523 | 1397 | 1712.3333333333333 | Region | Note=Flexible linker | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 524_1640 | 1397 | 1712.3333333333333 | Region | Note=Heavy chain arm | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 524_634 | 1397 | 1712.3333333333333 | Region | Note=Distal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 639_1640 | 1397 | 1712.3333333333333 | Region | Note=Proximal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 68_107 | 1397 | 1712.3333333333333 | Region | Note=WD40-like repeat 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 108_149 | 1397 | 1665.0 | Region | Note=WD40-like repeat 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 1213_1522 | 1397 | 1665.0 | Region | Involved in binding clathrin light chain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 150_195 | 1397 | 1665.0 | Region | Note=WD40-like repeat 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 196_257 | 1397 | 1665.0 | Region | Note=WD40-like repeat 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 24_67 | 1397 | 1665.0 | Region | Note=WD40-like repeat 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 258_301 | 1397 | 1665.0 | Region | Note=WD40-like repeat 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 2_479 | 1397 | 1665.0 | Region | Note=Globular terminal domain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 302_330 | 1397 | 1665.0 | Region | Note=WD40-like repeat 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 449_465 | 1397 | 1665.0 | Region | Binding site for the uncoating ATPase%2C involved in lattice disassembly | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 480_523 | 1397 | 1665.0 | Region | Note=Flexible linker | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 524_1640 | 1397 | 1665.0 | Region | Note=Heavy chain arm | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 524_634 | 1397 | 1665.0 | Region | Note=Distal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 639_1640 | 1397 | 1665.0 | Region | Note=Proximal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 68_107 | 1397 | 1665.0 | Region | Note=WD40-like repeat 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 108_149 | 1397 | 1710.6666666666667 | Region | Note=WD40-like repeat 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 1213_1522 | 1397 | 1710.6666666666667 | Region | Involved in binding clathrin light chain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 150_195 | 1397 | 1710.6666666666667 | Region | Note=WD40-like repeat 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 196_257 | 1397 | 1710.6666666666667 | Region | Note=WD40-like repeat 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 24_67 | 1397 | 1710.6666666666667 | Region | Note=WD40-like repeat 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 258_301 | 1397 | 1710.6666666666667 | Region | Note=WD40-like repeat 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 2_479 | 1397 | 1710.6666666666667 | Region | Note=Globular terminal domain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 302_330 | 1397 | 1710.6666666666667 | Region | Note=WD40-like repeat 7 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 449_465 | 1397 | 1710.6666666666667 | Region | Binding site for the uncoating ATPase%2C involved in lattice disassembly | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 480_523 | 1397 | 1710.6666666666667 | Region | Note=Flexible linker | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 524_1640 | 1397 | 1710.6666666666667 | Region | Note=Heavy chain arm | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 524_634 | 1397 | 1710.6666666666667 | Region | Note=Distal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 639_1640 | 1397 | 1710.6666666666667 | Region | Note=Proximal segment | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 68_107 | 1397 | 1710.6666666666667 | Region | Note=WD40-like repeat 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000263200 | 0 | 33 | 2_479 | 14 | 1712.3333333333333 | Region | Note=Globular terminal domain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000353891 | 0 | 32 | 2_479 | 14 | 1665.0 | Region | Note=Globular terminal domain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19263353 | ENST00000427926 | 0 | 34 | 2_479 | 14 | 1710.6666666666667 | Region | Note=Globular terminal domain | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 1128_1269 | 1397 | 1712.3333333333333 | Repeat | Note=CHCR 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 1274_1420 | 1397 | 1712.3333333333333 | Repeat | Note=CHCR 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 537_683 | 1397 | 1712.3333333333333 | Repeat | Note=CHCR 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 686_828 | 1397 | 1712.3333333333333 | Repeat | Note=CHCR 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 833_972 | 1397 | 1712.3333333333333 | Repeat | Note=CHCR 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000263200 | 25 | 33 | 979_1124 | 1397 | 1712.3333333333333 | Repeat | Note=CHCR 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 1128_1269 | 1397 | 1665.0 | Repeat | Note=CHCR 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 1274_1420 | 1397 | 1665.0 | Repeat | Note=CHCR 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 537_683 | 1397 | 1665.0 | Repeat | Note=CHCR 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 686_828 | 1397 | 1665.0 | Repeat | Note=CHCR 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 833_972 | 1397 | 1665.0 | Repeat | Note=CHCR 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000353891 | 25 | 32 | 979_1124 | 1397 | 1665.0 | Repeat | Note=CHCR 4 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 1128_1269 | 1397 | 1710.6666666666667 | Repeat | Note=CHCR 5 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 1274_1420 | 1397 | 1710.6666666666667 | Repeat | Note=CHCR 6 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 537_683 | 1397 | 1710.6666666666667 | Repeat | Note=CHCR 1 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 686_828 | 1397 | 1710.6666666666667 | Repeat | Note=CHCR 2 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 833_972 | 1397 | 1710.6666666666667 | Repeat | Note=CHCR 3 | |
| Tgene | CLTCL1 | chr22:19076881 | chr22:19178947 | ENST00000427926 | 26 | 34 | 979_1124 | 1397 | 1710.6666666666667 | Repeat | Note=CHCR 4 |
Top |
Fusion Gene Sequence for DGCR2-CLTCL1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >22400_22400_1_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19178947_ENST00000263200_length(transcript)=1652nt_BP=403nt GCGCGCAGCTGGAGTCGGTGCCTGAGGTTGCAGCCGAGAGTGTGCGCCAGCCCGCGGCCCAGCCGAAGCTCTTTCCCGCCGCCTCTCCGC GCCTCGCCCAGGTTCAGCTCCGCCTGACCCTCCGCTTGGCACGGTCCCCTGACCCTCAGTCCGGCCTGGTCCGGCCTTCCAGCCGCCCCG GGGACGATGAACGGAGGATAAATGGTGCCCAAGGCAGACAGCGGCGCCTTCCTGCTGCTCTTCCTGCTCGTGCTCACTGTCACCGAGCCG CTGCGGCCAGAGCTGCGGTGCAACCCTGGGCAGTTTGCGTGTCGCAGCGGCACCATCCAGTGCATCCCCCTCCCCTGGCAGTGTGACGGC TGGGCGACTTGCGAGGATGAGAGCGACGAAGCCAACTGTCCAGGTTGCCAACGTCGAGCTCTGTTACAGAGCCCTGCAGTTCTATTTGGA TTACAAACCACTGCTCATCAATGACCTGCTGCTGGTGCTTTCACCCCGGCTGGACCACACCTGGACAGTCAGTTTCTTTTCAAAGGCAGG TCAGCTGCCCCTGGTGAAGCCTTACCTGCGGTCAGTCCAGAGCCACAACAACAAGAGTGTGAATGAGGCACTCAACCACCTGCTGACAGA GGAGGAGGACTATCAGGGCTTAAGGGCATCTATCGATGCCTATGACAACTTTGACAACATCAGCCTGGCTCAGCAGCTGGAGAAGCATCA GCTGATGGAGTTCAGGTGCATTGCGGCCTATCTGTACAAGGGCAATAACTGGTGGGCCCAGAGCGTGGAGCTCTGCAAGAAGGATCATCT CTACAAGGATGCCATGCAGCATGCTGCAGAGTCGCGGGATGCTGAGCTGGCCCAGAAGTTGCTGCAGTGGTTCCTGGAGGAAGGCAAGAG GGAGTGCTTCGCAGCTTGTCTCTTCACCTGCTATGACCTGCTTCGCCCAGACATGGTGCTTGAGCTGGCCTGGAGGCACAACCTCGTGGA CTTGGCCATGCCCTACTTCATCCAGGTGATGAGGGAGTACCTGAGCAAGGTGGACAAACTGGATGCCTTGGAGAGTCTGCGCAAGCAAGA GGAGCATGTGACAGAGCCTGCCCCTCTCGTGTTTGATTTTGATGGGCATGAATGAGACCCAGCTGATTGCACTAAGCCCTGCCGTGGGCC CAGCCCCTGCCAGCTTCCCCTATGGATATGCCTCTGCTCCCAACTTCGCCAGCCTCCAATGTACAACTTCCGCGTGTAGTGGGCGTTGTC ACCACCCACCCTACCTGCAGAGTTACTAACTTCTCCAAGGAGCATGTCACTCCAGCAGCACAGGGGACGCAATGGGAGGCAGGGACACCT GGACAATATTTATTTTTGCTGAAACCCAATGACGGCAACCTCTGAGCCATCCCAGAGCCTGGGGAGGCCAGGGTAGAGGCTGACGGCGCA AGACCAGCTTTAGCCGACAACAGAGACTGGACTGTGGGCCCTCCTGCTGGAGCCAGGCCTTCCTCCTGGGCGCCTCCGACTGGCTGGAGC TGCCCCCTCCAGGCCAGTTTGAAGACTACATGAACACGTCTTGTTTGGAGGTACCGGACCTCATAAAAGGACTCTCAGCCTCTTGGCAAT >22400_22400_1_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19178947_ENST00000263200_length(amino acids)=292AA_BP=3 MSPSRCGQSCGATLGSLRVAAAPSSASPSPGSVTAGRLARMRATKPTVQVANVELCYRALQFYLDYKPLLINDLLLVLSPRLDHTWTVSF FSKAGQLPLVKPYLRSVQSHNNKSVNEALNHLLTEEEDYQGLRASIDAYDNFDNISLAQQLEKHQLMEFRCIAAYLYKGNNWWAQSVELC KKDHLYKDAMQHAAESRDAELAQKLLQWFLEEGKRECFAACLFTCYDLLRPDMVLELAWRHNLVDLAMPYFIQVMREYLSKVDKLDALES -------------------------------------------------------------- >22400_22400_2_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19178947_ENST00000353891_length(transcript)=1481nt_BP=403nt GCGCGCAGCTGGAGTCGGTGCCTGAGGTTGCAGCCGAGAGTGTGCGCCAGCCCGCGGCCCAGCCGAAGCTCTTTCCCGCCGCCTCTCCGC GCCTCGCCCAGGTTCAGCTCCGCCTGACCCTCCGCTTGGCACGGTCCCCTGACCCTCAGTCCGGCCTGGTCCGGCCTTCCAGCCGCCCCG GGGACGATGAACGGAGGATAAATGGTGCCCAAGGCAGACAGCGGCGCCTTCCTGCTGCTCTTCCTGCTCGTGCTCACTGTCACCGAGCCG CTGCGGCCAGAGCTGCGGTGCAACCCTGGGCAGTTTGCGTGTCGCAGCGGCACCATCCAGTGCATCCCCCTCCCCTGGCAGTGTGACGGC TGGGCGACTTGCGAGGATGAGAGCGACGAAGCCAACTGTCCAGGTTGCCAACGTCGAGCTCTGTTACAGAGCCCTGCAGTTCTATTTGGA TTACAAACCACTGCTCATCAATGACCTGCTGCTGGTGCTTTCACCCCGGCTGGACCACACCTGGACAGTCAGTTTCTTTTCAAAGGCAGG TCAGCTGCCCCTGGTGAAGCCTTACCTGCGGTCAGTCCAGAGCCACAACAACAAGAGTGTGAATGAGGCACTCAACCACCTGCTGACAGA GGAGGAGGACTATCAGGATGCCATGCAGCATGCTGCAGAGTCGCGGGATGCTGAGCTGGCCCAGAAGTTGCTGCAGTGGTTCCTGGAGGA AGGCAAGAGGGAGTGCTTCGCAGCTTGTCTCTTCACCTGCTATGACCTGCTTCGCCCAGACATGGTGCTTGAGCTGGCCTGGAGGCACAA CCTCGTGGACTTGGCCATGCCCTACTTCATCCAGGTGATGAGGGAGTACCTGAGCAAGGTGGACAAACTGGATGCCTTGGAGAGTCTGCG CAAGCAAGAGGAGCATGTGACAGAGCCTGCCCCTCTCGTGTTTGATTTTGATGGGCATGAATGAGACCCAGCTGATTGCACTAAGCCCTG CCGTGGGCCCAGCCCCTGCCAGCTTCCCCTATGGATATGCCTCTGCTCCCAACTTCGCCAGCCTCCAATGTACAACTTCCGCGTGTAGTG GGCGTTGTCACCACCCACCCTACCTGCAGAGTTACTAACTTCTCCAAGGAGCATGTCACTCCAGCAGCACAGGGGACGCAATGGGAGGCA GGGACACCTGGACAATATTTATTTTTGCTGAAACCCAATGACGGCAACCTCTGAGCCATCCCAGAGCCTGGGGAGGCCAGGGTAGAGGCT GACGGCGCAAGACCAGCTTTAGCCGACAACAGAGACTGGACTGTGGGCCCTCCTGCTGGAGCCAGGCCTTCCTCCTGGGCGCCTCCGACT GGCTGGAGCTGCCCCCTCCAGGCCAGTTTGAAGACTACATGAACACGTCTTGTTTGGAGGTACCGGACCTCATAAAAGGACTCTCAGCCT >22400_22400_2_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19178947_ENST00000353891_length(amino acids)=235AA_BP=3 MSPSRCGQSCGATLGSLRVAAAPSSASPSPGSVTAGRLARMRATKPTVQVANVELCYRALQFYLDYKPLLINDLLLVLSPRLDHTWTVSF FSKAGQLPLVKPYLRSVQSHNNKSVNEALNHLLTEEEDYQDAMQHAAESRDAELAQKLLQWFLEEGKRECFAACLFTCYDLLRPDMVLEL -------------------------------------------------------------- >22400_22400_3_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19178947_ENST00000427926_length(transcript)=1649nt_BP=403nt GCGCGCAGCTGGAGTCGGTGCCTGAGGTTGCAGCCGAGAGTGTGCGCCAGCCCGCGGCCCAGCCGAAGCTCTTTCCCGCCGCCTCTCCGC GCCTCGCCCAGGTTCAGCTCCGCCTGACCCTCCGCTTGGCACGGTCCCCTGACCCTCAGTCCGGCCTGGTCCGGCCTTCCAGCCGCCCCG GGGACGATGAACGGAGGATAAATGGTGCCCAAGGCAGACAGCGGCGCCTTCCTGCTGCTCTTCCTGCTCGTGCTCACTGTCACCGAGCCG CTGCGGCCAGAGCTGCGGTGCAACCCTGGGCAGTTTGCGTGTCGCAGCGGCACCATCCAGTGCATCCCCCTCCCCTGGCAGTGTGACGGC TGGGCGACTTGCGAGGATGAGAGCGACGAAGCCAACTGTCCAGGTTGCCAACGTCGAGCTCTGTTACAGAGCCCTGCAGTTCTATTTGGA TTACAAACCACTGCTCATCAATGACCTGCTGCTGGTGCTTTCACCCCGGCTGGACCACACCTGGACAGTCAGTTTCTTTTCAAAGGCAGG TCAGCTGCCCCTGGTGAAGCCTTACCTGCGGTCAGTCCAGAGCCACAACAACAAGAGTGTGAATGAGGCACTCAACCACCTGCTGACAGA GGAGGAGGACTATCAGGGCTTAAGGGCATCTATCGATGCCTATGACAACTTTGACAACATCAGCCTGGCTCAGCAGCTGGAGAAGCATCA GCTGATGGAGTTCAGGTGCATTGCGGCCTATCTGTACAAGGGCAATAACTGGTGGGCCCAGAGCGTGGAGCTCTGCAAGAAGGATCATCT CTACAAGGATGCCATGCAGCATGCTGCAGAGTCGCGGGATGCTGAGCTGGCCCAGAAGTTGCTGCAGTGGTTCCTGGAGGAAGGCAAGAG GGAGTGCTTCGCAGCTTGTCTCTTCACCTGCTATGACCTGCTTCGCCCAGACATGGTGCTTGAGCTGGCCTGGAGGCACAACCTCGTGGA CTTGGCCATGCCCTACTTCATCCAGGTGATGAGGGAGTACCTGAGCAAGGTGGACAAACTGGATGCCTTGGAGAGTCTGCGCAAGCAAGA GGAGCATGTGACAGAGCCTGCCCCTCTCGTGTTTGATTTTGATGGGCATGAATGAGACCCAGCTGATTGCACTAAGCCCTGCCGTGGGCC CAGCCCCTGCCAGCTTCCCCTATGGATATGCCTCTGCTCCCAACTTCGCCAGCCTCCAATGTACAACTTCCGCGTGTAGTGGGCGTTGTC ACCACCCACCCTACCTGCAGAGTTACTAACTTCTCCAAGGAGCATGTCACTCCAGCAGCACAGGGGACGCAATGGGAGGCAGGGACACCT GGACAATATTTATTTTTGCTGAAACCCAATGACGGCAACCTCTGAGCCATCCCAGAGCCTGGGGAGGCCAGGGTAGAGGCTGACGGCGCA AGACCAGCTTTAGCCGACAACAGAGACTGGACTGTGGGCCCTCCTGCTGGAGCCAGGCCTTCCTCCTGGGCGCCTCCGACTGGCTGGAGC TGCCCCCTCCAGGCCAGTTTGAAGACTACATGAACACGTCTTGTTTGGAGGTACCGGACCTCATAAAAGGACTCTCAGCCTCTTGGCAAT >22400_22400_3_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19178947_ENST00000427926_length(amino acids)=292AA_BP=3 MSPSRCGQSCGATLGSLRVAAAPSSASPSPGSVTAGRLARMRATKPTVQVANVELCYRALQFYLDYKPLLINDLLLVLSPRLDHTWTVSF FSKAGQLPLVKPYLRSVQSHNNKSVNEALNHLLTEEEDYQGLRASIDAYDNFDNISLAQQLEKHQLMEFRCIAAYLYKGNNWWAQSVELC KKDHLYKDAMQHAAESRDAELAQKLLQWFLEEGKRECFAACLFTCYDLLRPDMVLELAWRHNLVDLAMPYFIQVMREYLSKVDKLDALES -------------------------------------------------------------- >22400_22400_4_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19263353_ENST00000263200_length(transcript)=5801nt_BP=403nt GCGCGCAGCTGGAGTCGGTGCCTGAGGTTGCAGCCGAGAGTGTGCGCCAGCCCGCGGCCCAGCCGAAGCTCTTTCCCGCCGCCTCTCCGC GCCTCGCCCAGGTTCAGCTCCGCCTGACCCTCCGCTTGGCACGGTCCCCTGACCCTCAGTCCGGCCTGGTCCGGCCTTCCAGCCGCCCCG GGGACGATGAACGGAGGATAAATGGTGCCCAAGGCAGACAGCGGCGCCTTCCTGCTGCTCTTCCTGCTCGTGCTCACTGTCACCGAGCCG CTGCGGCCAGAGCTGCGGTGCAACCCTGGGCAGTTTGCGTGTCGCAGCGGCACCATCCAGTGCATCCCCCTCCCCTGGCAGTGTGACGGC TGGGCGACTTGCGAGGATGAGAGCGACGAAGCCAACTGTCCAGCTCCAAAACCTTGGAATTAATCCAGCTAACATTGGATTCAGCACACT GACCATGGAATCTGACAAGTTCATATGTATCCGAGAGAAAGTTGGTGAGCAGGCACAGGTCACGATCATTGACATGAGTGACCCAATGGC TCCGATCCGACGGCCTATCTCTGCAGAGAGTGCCATCATGAATCCAGCCTCTAAGGTGATAGCTCTGAAAGCTGGGAAGACACTTCAGAT CTTTAATATTGAGATGAAGAGTAAAATGAAGGCTCATACTATGGCAGAAGAAGTGATTTTCTGGAAATGGGTTTCTGTGAACACTGTTGC CTTGGTGACCGAGACCGCGGTCTACCACTGGAGCATGGAAGGTGACTCCCAGCCCATGAAGATGTTTGATAGACATACCAGTCTGGTGGG CTGCCAGGTGATTCACTACCGGACTGATGAGTACCAGAAGTGGCTGCTGCTCGTAGGCATCTCGGCTCAGCAAAACCGTGTGGTTGGAGC AATGCAGCTCTACTCTGTGGATAGGAAGGTTTCACAACCCATAGAAGGCCATGCTGCGGCTTTTGCAGAGTTCAAGATGGAGGGGAATGC CAAGCCTGCCACCCTTTTCTGCTTTGCTGTACGTAATCCCACAGGAGGCAAGTTGCACATCATTGAAGTTGGACAGCCTGCAGCGGGAAA CCAACCTTTTGTAAAGAAAGCAGTAGATGTGTTTTTTCCTCCAGAGGCACAGAATGATTTTCCAGTGGCTATGCAGATTGGAGCTAAACA TGGTGTTATTTACTTGATCACAAAGTATGGCTATCTTCATCTGTACGACCTAGAGTCTGGCGTGTGCATCTGCATGAACCGTATTAGTGC TGACACAATATTTGTCACTGCTCCACACAAACCAACCTCTGGAATTATTGGTGTCAACAAAAAGGGACAGGTACTGTCAGTTTGTGTTGA GGAAGATAACATTGTGAATTATGCAACCAACGTGCTTCAGAATCCAGACCTTGGTCTGCGTTTGGCCGTTCGTAGTAACCTGGCTGGGGC AGAGAAGTTGTTTGTGAGAAAATTCAATACCCTCTTTGCACAGGGCAGCTATGCTGAAGCCGCCAAAGTTGCAGCGTCTGCACCAAAGGG AATCCTGCGTACCAGAGAGACGGTCCAGAAATTCCAGAGTATACCCGCTCAGTCTGGCCAGGCTTCTCCATTGCTGCAGTACTTCGGAAT CCTGCTCGACCAGGGTCAGCTCAATAAACTTGAATCCTTAGAACTTTGCCATCTGGTTCTTCAGCAGGGGCGTAAGCAACTCCTAGAGAA GTGGCTGAAAGAAGATAAGCTGGAGTGCTCAGAGGAGCTCGGAGACTTGGTCAAAACCACTGACCCCATGCTCGCTCTGAGTGTGTACCT TCGGGCAAATGTGCCAAGCAAAGTGATCCAGTGTTTTGCAGAAACAGGCCAATTCCAGAAAATTGTGCTCTATGCCAAAAAGGTTGGGTA CACCCCAGACTGGATCTTTCTGCTGAGGGGTGTAATGAAGATCAGTCCGGAACAGGGCCTGCAGTTTTCTCGAATGCTAGTGCAGGACGA GGAGCCGCTGGCCAACATTAGCCAGATTGTGGACATTTTCATGGAAAACAGTTTAATTCAGCAGTGTACTTCCTTCTTATTGGATGCCTT GAAGAATAATCGCCCAGCTGAGGGACTCCTGCAGACATGGCTGTTGGAGATGAACCTTGTTCATGCACCCCAGGTTGCAGATGCCATCCT TGGAAATAAAATGTTTACTCATTACGACCGGGCCCACATTGCCCAGCTCTGTGAGAAGGCAGGCCTCCTGCAGCAAGCACTGGAGCACTA CACCGACCTCTATGACATCAAGAGGGCTGTGGTCCACACTCACCTCCTCAATCCCGAGTGGCTTGTCAATTTCTTTGGCTCCTTATCGGT GGAGGATTCTGTGGAGTGTCTGCATGCCATGCTGTCTGCTAACATCAGACAGAACCTTCAGCTGTGTGTGCAGGTGGCCTCTAAGTACCA CGAGCAGCTGGGCACGCAGGCCCTGGTGGAGCTCTTTGAATCCTTCAAGAGTTACAAAGGCCTCTTCTACTTCCTGGGCTCAATCGTGAA CTTCAGCCAAGACCCAGATGTGCATCTGAAATACATTCAGGCTGCCTGTAAGACAGGGCAGATCAAGGAGGTGGAGAGGATATGCCGAGA GAGCAGCTGCTACAACCCAGAGCGTGTGAAGAACTTCCTGAAGGAGGCCAAGCTCACAGACCAGCTTCCCCTCATCATCGTGTGTGATCG TTTTGGCTTTGTCCATGACCTTGTCCTATATTTATACCGCAACAACCTGCAGAGGTACATTGAGATCTACGTGCAGAAGGTCAACCCTAG CCGGACCCCAGCTGTGATTGGAGGGCTGCTTGATGTGGATTGTTCTGAGGAAGTGATTAAACACTTAATCATGGCAGTGAGAGGACAGTT CTCTACTGATGAGTTGGTGGCTGAAGTAGAAAAAAGAAATAGGCTCAAGCTGCTGCTTCCCTGGCTGGAGTCCCAGATTCAGGAAGGCTG TGAGGAGCCTGCCACTCACAATGCACTGGCTAAAATCTACATCGACAGCAACAACAGCCCCGAGTGCTTCCTGAGAGAGAATGCCTACTA TGACAGCAGCGTGGTGGGCCGCTACTGTGAGAAGCGAGACCCCCATCTGGCCTGTGTTGCCTATGAGCGGGGGCAGTGTGACCTTGAGCT CATCAAGGTGTGCAATGAGAATTCTCTGTTCAAAAGCGAGGCCCGCTACCTGGTATGCAGAAAGGATCCGGAGCTCTGGGCTCACGTCCT TGAGGAGACCAACCCATCCAGGAGACAGCTAATTGACCAGGTGGTACAGACAGCATTGTCAGAAACACGGGATCCTGAAGAGATTTCGGT CACTGTCAAAGCCTTTATGACAGCCGACCTGCCTAATGAACTGATTGAACTGCTGGAGAAGATAGTTCTGGATAACTCTGTCTTCAGCGA GCACAGGAATCTACAGAATCTGTTGATCCTGACTGCCATCAAGGCAGACCGCACACGGGTCATGGAGTACATCAGCCGCCTGGACAACTA TGACGCACTGGACATCGCGAGCATCGCTGTCAGCAGCGCACTGTATGAGGAGGCCTTCACCGTTTTCCACAAGTTTGATATGAATGCCTC AGCAATCCAGGTCCTGATCGAGCACATTGGAAACCTGGACCGGGCATATGAGTTTGCGGAGAGATGCAATGAGCCTGCTGTGTGGAGTCA GCTGGCCCAAGCCCAGCTCCAGAAAGATTTGGTGAAGGAAGCCATCAACTCCTATATCAGAGGGGACGACCCTTCCTCTTACCTGGAAGT TGTTCAGTCAGCCAGCAGGAGCAACAACTGGGAGGATCTAGTTAAATTTCTGCAGATGGCCAGGAAAAAGGGCCGTGAGTCCTATATAGA GACTGAACTTATTTTTGCCTTGGCTAAAACCAGCCGTGTTTCTGAGCTAGAAGATTTTATTAATGGACCCAACAATGCCCACATCCAGCA GGTTGGAGACCGCTGTTACGAGGAGGGAATGTACGAGGCTGCCAAGCTGCTCTATAGCAATGTTTCTAACTTTGCCCGCCTGGCTTCCAC CTTGGTTCACCTCGGTGAGTATCAGGCAGCAGTGGACAACAGCCGCAAGGCCAGCAGCACCCGGACGTGGAAGGAGGTGTGCTTTGCCTG CATGGATGGACAAGAGTTCCGCTTCGCACAGCTGTGTGGTCTTCACATCGTCATTCATGCAGATGAGCTGGAGGAGCTGATGTGCTATTA CCAGGATCGTGGCTACTTTGAGGAGCTGATCTTGCTGTTGGAAGCGGCCCTGGGCCTGGAGCGGGCCCACATGGGCATGTTCACTGAGCT GGCCATCCTCTACTCCAAATTCAAGCCACAGAAGATGCTGGAGCATCTGGAGCTTTTCTGGTCCCGTGTCAACATCCCAAAGGTGCTGAG GGCTGCAGAGCAGGCACACCTGTGGGCTGAGCTGGTGTTCCTCTATGACAAGTACGAGGAGTATGACAATGCTGTGCTCACCATGATGAG CCACCCCACTGAGGCCTGGAAGGAGGGTCAGTTCAAGGACATCATTACCAAGGTTGCCAACGTCGAGCTCTGTTACAGAGCCCTGCAGTT CTATTTGGATTACAAACCACTGCTCATCAATGACCTGCTGCTGGTGCTTTCACCCCGGCTGGACCACACCTGGACAGTCAGTTTCTTTTC AAAGGCAGGTCAGCTGCCCCTGGTGAAGCCTTACCTGCGGTCAGTCCAGAGCCACAACAACAAGAGTGTGAATGAGGCACTCAACCACCT GCTGACAGAGGAGGAGGACTATCAGGGCTTAAGGGCATCTATCGATGCCTATGACAACTTTGACAACATCAGCCTGGCTCAGCAGCTGGA GAAGCATCAGCTGATGGAGTTCAGGTGCATTGCGGCCTATCTGTACAAGGGCAATAACTGGTGGGCCCAGAGCGTGGAGCTCTGCAAGAA GGATCATCTCTACAAGGATGCCATGCAGCATGCTGCAGAGTCGCGGGATGCTGAGCTGGCCCAGAAGTTGCTGCAGTGGTTCCTGGAGGA AGGCAAGAGGGAGTGCTTCGCAGCTTGTCTCTTCACCTGCTATGACCTGCTTCGCCCAGACATGGTGCTTGAGCTGGCCTGGAGGCACAA CCTCGTGGACTTGGCCATGCCCTACTTCATCCAGGTGATGAGGGAGTACCTGAGCAAGGTGGACAAACTGGATGCCTTGGAGAGTCTGCG CAAGCAAGAGGAGCATGTGACAGAGCCTGCCCCTCTCGTGTTTGATTTTGATGGGCATGAATGAGACCCAGCTGATTGCACTAAGCCCTG CCGTGGGCCCAGCCCCTGCCAGCTTCCCCTATGGATATGCCTCTGCTCCCAACTTCGCCAGCCTCCAATGTACAACTTCCGCGTGTAGTG GGCGTTGTCACCACCCACCCTACCTGCAGAGTTACTAACTTCTCCAAGGAGCATGTCACTCCAGCAGCACAGGGGACGCAATGGGAGGCA GGGACACCTGGACAATATTTATTTTTGCTGAAACCCAATGACGGCAACCTCTGAGCCATCCCAGAGCCTGGGGAGGCCAGGGTAGAGGCT GACGGCGCAAGACCAGCTTTAGCCGACAACAGAGACTGGACTGTGGGCCCTCCTGCTGGAGCCAGGCCTTCCTCCTGGGCGCCTCCGACT GGCTGGAGCTGCCCCCTCCAGGCCAGTTTGAAGACTACATGAACACGTCTTGTTTGGAGGTACCGGACCTCATAAAAGGACTCTCAGCCT >22400_22400_4_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19263353_ENST00000263200_length(amino acids)=1675AA_BP=3 MSPSRCGQSCGATLGSLRVAAAPSSASPSPGSVTAGRLARMRATKPTVQLQNLGINPANIGFSTLTMESDKFICIREKVGEQAQVTIIDM SDPMAPIRRPISAESAIMNPASKVIALKAGKTLQIFNIEMKSKMKAHTMAEEVIFWKWVSVNTVALVTETAVYHWSMEGDSQPMKMFDRH TSLVGCQVIHYRTDEYQKWLLLVGISAQQNRVVGAMQLYSVDRKVSQPIEGHAAAFAEFKMEGNAKPATLFCFAVRNPTGGKLHIIEVGQ PAAGNQPFVKKAVDVFFPPEAQNDFPVAMQIGAKHGVIYLITKYGYLHLYDLESGVCICMNRISADTIFVTAPHKPTSGIIGVNKKGQVL SVCVEEDNIVNYATNVLQNPDLGLRLAVRSNLAGAEKLFVRKFNTLFAQGSYAEAAKVAASAPKGILRTRETVQKFQSIPAQSGQASPLL QYFGILLDQGQLNKLESLELCHLVLQQGRKQLLEKWLKEDKLECSEELGDLVKTTDPMLALSVYLRANVPSKVIQCFAETGQFQKIVLYA KKVGYTPDWIFLLRGVMKISPEQGLQFSRMLVQDEEPLANISQIVDIFMENSLIQQCTSFLLDALKNNRPAEGLLQTWLLEMNLVHAPQV ADAILGNKMFTHYDRAHIAQLCEKAGLLQQALEHYTDLYDIKRAVVHTHLLNPEWLVNFFGSLSVEDSVECLHAMLSANIRQNLQLCVQV ASKYHEQLGTQALVELFESFKSYKGLFYFLGSIVNFSQDPDVHLKYIQAACKTGQIKEVERICRESSCYNPERVKNFLKEAKLTDQLPLI IVCDRFGFVHDLVLYLYRNNLQRYIEIYVQKVNPSRTPAVIGGLLDVDCSEEVIKHLIMAVRGQFSTDELVAEVEKRNRLKLLLPWLESQ IQEGCEEPATHNALAKIYIDSNNSPECFLRENAYYDSSVVGRYCEKRDPHLACVAYERGQCDLELIKVCNENSLFKSEARYLVCRKDPEL WAHVLEETNPSRRQLIDQVVQTALSETRDPEEISVTVKAFMTADLPNELIELLEKIVLDNSVFSEHRNLQNLLILTAIKADRTRVMEYIS RLDNYDALDIASIAVSSALYEEAFTVFHKFDMNASAIQVLIEHIGNLDRAYEFAERCNEPAVWSQLAQAQLQKDLVKEAINSYIRGDDPS SYLEVVQSASRSNNWEDLVKFLQMARKKGRESYIETELIFALAKTSRVSELEDFINGPNNAHIQQVGDRCYEEGMYEAAKLLYSNVSNFA RLASTLVHLGEYQAAVDNSRKASSTRTWKEVCFACMDGQEFRFAQLCGLHIVIHADELEELMCYYQDRGYFEELILLLEAALGLERAHMG MFTELAILYSKFKPQKMLEHLELFWSRVNIPKVLRAAEQAHLWAELVFLYDKYEEYDNAVLTMMSHPTEAWKEGQFKDIITKVANVELCY RALQFYLDYKPLLINDLLLVLSPRLDHTWTVSFFSKAGQLPLVKPYLRSVQSHNNKSVNEALNHLLTEEEDYQGLRASIDAYDNFDNISL AQQLEKHQLMEFRCIAAYLYKGNNWWAQSVELCKKDHLYKDAMQHAAESRDAELAQKLLQWFLEEGKRECFAACLFTCYDLLRPDMVLEL -------------------------------------------------------------- >22400_22400_5_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19263353_ENST00000353891_length(transcript)=5630nt_BP=403nt GCGCGCAGCTGGAGTCGGTGCCTGAGGTTGCAGCCGAGAGTGTGCGCCAGCCCGCGGCCCAGCCGAAGCTCTTTCCCGCCGCCTCTCCGC GCCTCGCCCAGGTTCAGCTCCGCCTGACCCTCCGCTTGGCACGGTCCCCTGACCCTCAGTCCGGCCTGGTCCGGCCTTCCAGCCGCCCCG GGGACGATGAACGGAGGATAAATGGTGCCCAAGGCAGACAGCGGCGCCTTCCTGCTGCTCTTCCTGCTCGTGCTCACTGTCACCGAGCCG CTGCGGCCAGAGCTGCGGTGCAACCCTGGGCAGTTTGCGTGTCGCAGCGGCACCATCCAGTGCATCCCCCTCCCCTGGCAGTGTGACGGC TGGGCGACTTGCGAGGATGAGAGCGACGAAGCCAACTGTCCAGCTCCAAAACCTTGGAATTAATCCAGCTAACATTGGATTCAGCACACT GACCATGGAATCTGACAAGTTCATATGTATCCGAGAGAAAGTTGGTGAGCAGGCACAGGTCACGATCATTGACATGAGTGACCCAATGGC TCCGATCCGACGGCCTATCTCTGCAGAGAGTGCCATCATGAATCCAGCCTCTAAGGTGATAGCTCTGAAAGCTGGGAAGACACTTCAGAT CTTTAATATTGAGATGAAGAGTAAAATGAAGGCTCATACTATGGCAGAAGAAGTGATTTTCTGGAAATGGGTTTCTGTGAACACTGTTGC CTTGGTGACCGAGACCGCGGTCTACCACTGGAGCATGGAAGGTGACTCCCAGCCCATGAAGATGTTTGATAGACATACCAGTCTGGTGGG CTGCCAGGTGATTCACTACCGGACTGATGAGTACCAGAAGTGGCTGCTGCTCGTAGGCATCTCGGCTCAGCAAAACCGTGTGGTTGGAGC AATGCAGCTCTACTCTGTGGATAGGAAGGTTTCACAACCCATAGAAGGCCATGCTGCGGCTTTTGCAGAGTTCAAGATGGAGGGGAATGC CAAGCCTGCCACCCTTTTCTGCTTTGCTGTACGTAATCCCACAGGAGGCAAGTTGCACATCATTGAAGTTGGACAGCCTGCAGCGGGAAA CCAACCTTTTGTAAAGAAAGCAGTAGATGTGTTTTTTCCTCCAGAGGCACAGAATGATTTTCCAGTGGCTATGCAGATTGGAGCTAAACA TGGTGTTATTTACTTGATCACAAAGTATGGCTATCTTCATCTGTACGACCTAGAGTCTGGCGTGTGCATCTGCATGAACCGTATTAGTGC TGACACAATATTTGTCACTGCTCCACACAAACCAACCTCTGGAATTATTGGTGTCAACAAAAAGGGACAGGTACTGTCAGTTTGTGTTGA GGAAGATAACATTGTGAATTATGCAACCAACGTGCTTCAGAATCCAGACCTTGGTCTGCGTTTGGCCGTTCGTAGTAACCTGGCTGGGGC AGAGAAGTTGTTTGTGAGAAAATTCAATACCCTCTTTGCACAGGGCAGCTATGCTGAAGCCGCCAAAGTTGCAGCGTCTGCACCAAAGGG AATCCTGCGTACCAGAGAGACGGTCCAGAAATTCCAGAGTATACCCGCTCAGTCTGGCCAGGCTTCTCCATTGCTGCAGTACTTCGGAAT CCTGCTCGACCAGGGTCAGCTCAATAAACTTGAATCCTTAGAACTTTGCCATCTGGTTCTTCAGCAGGGGCGTAAGCAACTCCTAGAGAA GTGGCTGAAAGAAGATAAGCTGGAGTGCTCAGAGGAGCTCGGAGACTTGGTCAAAACCACTGACCCCATGCTCGCTCTGAGTGTGTACCT TCGGGCAAATGTGCCAAGCAAAGTGATCCAGTGTTTTGCAGAAACAGGCCAATTCCAGAAAATTGTGCTCTATGCCAAAAAGGTTGGGTA CACCCCAGACTGGATCTTTCTGCTGAGGGGTGTAATGAAGATCAGTCCGGAACAGGGCCTGCAGTTTTCTCGAATGCTAGTGCAGGACGA GGAGCCGCTGGCCAACATTAGCCAGATTGTGGACATTTTCATGGAAAACAGTTTAATTCAGCAGTGTACTTCCTTCTTATTGGATGCCTT GAAGAATAATCGCCCAGCTGAGGGACTCCTGCAGACATGGCTGTTGGAGATGAACCTTGTTCATGCACCCCAGGTTGCAGATGCCATCCT TGGAAATAAAATGTTTACTCATTACGACCGGGCCCACATTGCCCAGCTCTGTGAGAAGGCAGGCCTCCTGCAGCAAGCACTGGAGCACTA CACCGACCTCTATGACATCAAGAGGGCTGTGGTCCACACTCACCTCCTCAATCCCGAGTGGCTTGTCAATTTCTTTGGCTCCTTATCGGT GGAGGATTCTGTGGAGTGTCTGCATGCCATGCTGTCTGCTAACATCAGACAGAACCTTCAGCTGTGTGTGCAGGTGGCCTCTAAGTACCA CGAGCAGCTGGGCACGCAGGCCCTGGTGGAGCTCTTTGAATCCTTCAAGAGTTACAAAGGCCTCTTCTACTTCCTGGGCTCAATCGTGAA CTTCAGCCAAGACCCAGATGTGCATCTGAAATACATTCAGGCTGCCTGTAAGACAGGGCAGATCAAGGAGGTGGAGAGGATATGCCGAGA GAGCAGCTGCTACAACCCAGAGCGTGTGAAGAACTTCCTGAAGGAGGCCAAGCTCACAGACCAGCTTCCCCTCATCATCGTGTGTGATCG TTTTGGCTTTGTCCATGACCTTGTCCTATATTTATACCGCAACAACCTGCAGAGGTACATTGAGATCTACGTGCAGAAGGTCAACCCTAG CCGGACCCCAGCTGTGATTGGAGGGCTGCTTGATGTGGATTGTTCTGAGGAAGTGATTAAACACTTAATCATGGCAGTGAGAGGACAGTT CTCTACTGATGAGTTGGTGGCTGAAGTAGAAAAAAGAAATAGGCTCAAGCTGCTGCTTCCCTGGCTGGAGTCCCAGATTCAGGAAGGCTG TGAGGAGCCTGCCACTCACAATGCACTGGCTAAAATCTACATCGACAGCAACAACAGCCCCGAGTGCTTCCTGAGAGAGAATGCCTACTA TGACAGCAGCGTGGTGGGCCGCTACTGTGAGAAGCGAGACCCCCATCTGGCCTGTGTTGCCTATGAGCGGGGGCAGTGTGACCTTGAGCT CATCAAGGTGTGCAATGAGAATTCTCTGTTCAAAAGCGAGGCCCGCTACCTGGTATGCAGAAAGGATCCGGAGCTCTGGGCTCACGTCCT TGAGGAGACCAACCCATCCAGGAGACAGCTAATTGACCAGGTGGTACAGACAGCATTGTCAGAAACACGGGATCCTGAAGAGATTTCGGT CACTGTCAAAGCCTTTATGACAGCCGACCTGCCTAATGAACTGATTGAACTGCTGGAGAAGATAGTTCTGGATAACTCTGTCTTCAGCGA GCACAGGAATCTACAGAATCTGTTGATCCTGACTGCCATCAAGGCAGACCGCACACGGGTCATGGAGTACATCAGCCGCCTGGACAACTA TGACGCACTGGACATCGCGAGCATCGCTGTCAGCAGCGCACTGTATGAGGAGGCCTTCACCGTTTTCCACAAGTTTGATATGAATGCCTC AGCAATCCAGGTCCTGATCGAGCACATTGGAAACCTGGACCGGGCATATGAGTTTGCGGAGAGATGCAATGAGCCTGCTGTGTGGAGTCA GCTGGCCCAAGCCCAGCTCCAGAAAGATTTGGTGAAGGAAGCCATCAACTCCTATATCAGAGGGGACGACCCTTCCTCTTACCTGGAAGT TGTTCAGTCAGCCAGCAGGAGCAACAACTGGGAGGATCTAGTTAAATTTCTGCAGATGGCCAGGAAAAAGGGCCGTGAGTCCTATATAGA GACTGAACTTATTTTTGCCTTGGCTAAAACCAGCCGTGTTTCTGAGCTAGAAGATTTTATTAATGGACCCAACAATGCCCACATCCAGCA GGTTGGAGACCGCTGTTACGAGGAGGGAATGTACGAGGCTGCCAAGCTGCTCTATAGCAATGTTTCTAACTTTGCCCGCCTGGCTTCCAC CTTGGTTCACCTCGGTGAGTATCAGGCAGCAGTGGACAACAGCCGCAAGGCCAGCAGCACCCGGACGTGGAAGGAGGTGTGCTTTGCCTG CATGGATGGACAAGAGTTCCGCTTCGCACAGCTGTGTGGTCTTCACATCGTCATTCATGCAGATGAGCTGGAGGAGCTGATGTGCTATTA CCAGGATCGTGGCTACTTTGAGGAGCTGATCTTGCTGTTGGAAGCGGCCCTGGGCCTGGAGCGGGCCCACATGGGCATGTTCACTGAGCT GGCCATCCTCTACTCCAAATTCAAGCCACAGAAGATGCTGGAGCATCTGGAGCTTTTCTGGTCCCGTGTCAACATCCCAAAGGTGCTGAG GGCTGCAGAGCAGGCACACCTGTGGGCTGAGCTGGTGTTCCTCTATGACAAGTACGAGGAGTATGACAATGCTGTGCTCACCATGATGAG CCACCCCACTGAGGCCTGGAAGGAGGGTCAGTTCAAGGACATCATTACCAAGGTTGCCAACGTCGAGCTCTGTTACAGAGCCCTGCAGTT CTATTTGGATTACAAACCACTGCTCATCAATGACCTGCTGCTGGTGCTTTCACCCCGGCTGGACCACACCTGGACAGTCAGTTTCTTTTC AAAGGCAGGTCAGCTGCCCCTGGTGAAGCCTTACCTGCGGTCAGTCCAGAGCCACAACAACAAGAGTGTGAATGAGGCACTCAACCACCT GCTGACAGAGGAGGAGGACTATCAGGATGCCATGCAGCATGCTGCAGAGTCGCGGGATGCTGAGCTGGCCCAGAAGTTGCTGCAGTGGTT CCTGGAGGAAGGCAAGAGGGAGTGCTTCGCAGCTTGTCTCTTCACCTGCTATGACCTGCTTCGCCCAGACATGGTGCTTGAGCTGGCCTG GAGGCACAACCTCGTGGACTTGGCCATGCCCTACTTCATCCAGGTGATGAGGGAGTACCTGAGCAAGGTGGACAAACTGGATGCCTTGGA GAGTCTGCGCAAGCAAGAGGAGCATGTGACAGAGCCTGCCCCTCTCGTGTTTGATTTTGATGGGCATGAATGAGACCCAGCTGATTGCAC TAAGCCCTGCCGTGGGCCCAGCCCCTGCCAGCTTCCCCTATGGATATGCCTCTGCTCCCAACTTCGCCAGCCTCCAATGTACAACTTCCG CGTGTAGTGGGCGTTGTCACCACCCACCCTACCTGCAGAGTTACTAACTTCTCCAAGGAGCATGTCACTCCAGCAGCACAGGGGACGCAA TGGGAGGCAGGGACACCTGGACAATATTTATTTTTGCTGAAACCCAATGACGGCAACCTCTGAGCCATCCCAGAGCCTGGGGAGGCCAGG GTAGAGGCTGACGGCGCAAGACCAGCTTTAGCCGACAACAGAGACTGGACTGTGGGCCCTCCTGCTGGAGCCAGGCCTTCCTCCTGGGCG CCTCCGACTGGCTGGAGCTGCCCCCTCCAGGCCAGTTTGAAGACTACATGAACACGTCTTGTTTGGAGGTACCGGACCTCATAAAAGGAC >22400_22400_5_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19263353_ENST00000353891_length(amino acids)=1618AA_BP=3 MSPSRCGQSCGATLGSLRVAAAPSSASPSPGSVTAGRLARMRATKPTVQLQNLGINPANIGFSTLTMESDKFICIREKVGEQAQVTIIDM SDPMAPIRRPISAESAIMNPASKVIALKAGKTLQIFNIEMKSKMKAHTMAEEVIFWKWVSVNTVALVTETAVYHWSMEGDSQPMKMFDRH TSLVGCQVIHYRTDEYQKWLLLVGISAQQNRVVGAMQLYSVDRKVSQPIEGHAAAFAEFKMEGNAKPATLFCFAVRNPTGGKLHIIEVGQ PAAGNQPFVKKAVDVFFPPEAQNDFPVAMQIGAKHGVIYLITKYGYLHLYDLESGVCICMNRISADTIFVTAPHKPTSGIIGVNKKGQVL SVCVEEDNIVNYATNVLQNPDLGLRLAVRSNLAGAEKLFVRKFNTLFAQGSYAEAAKVAASAPKGILRTRETVQKFQSIPAQSGQASPLL QYFGILLDQGQLNKLESLELCHLVLQQGRKQLLEKWLKEDKLECSEELGDLVKTTDPMLALSVYLRANVPSKVIQCFAETGQFQKIVLYA KKVGYTPDWIFLLRGVMKISPEQGLQFSRMLVQDEEPLANISQIVDIFMENSLIQQCTSFLLDALKNNRPAEGLLQTWLLEMNLVHAPQV ADAILGNKMFTHYDRAHIAQLCEKAGLLQQALEHYTDLYDIKRAVVHTHLLNPEWLVNFFGSLSVEDSVECLHAMLSANIRQNLQLCVQV ASKYHEQLGTQALVELFESFKSYKGLFYFLGSIVNFSQDPDVHLKYIQAACKTGQIKEVERICRESSCYNPERVKNFLKEAKLTDQLPLI IVCDRFGFVHDLVLYLYRNNLQRYIEIYVQKVNPSRTPAVIGGLLDVDCSEEVIKHLIMAVRGQFSTDELVAEVEKRNRLKLLLPWLESQ IQEGCEEPATHNALAKIYIDSNNSPECFLRENAYYDSSVVGRYCEKRDPHLACVAYERGQCDLELIKVCNENSLFKSEARYLVCRKDPEL WAHVLEETNPSRRQLIDQVVQTALSETRDPEEISVTVKAFMTADLPNELIELLEKIVLDNSVFSEHRNLQNLLILTAIKADRTRVMEYIS RLDNYDALDIASIAVSSALYEEAFTVFHKFDMNASAIQVLIEHIGNLDRAYEFAERCNEPAVWSQLAQAQLQKDLVKEAINSYIRGDDPS SYLEVVQSASRSNNWEDLVKFLQMARKKGRESYIETELIFALAKTSRVSELEDFINGPNNAHIQQVGDRCYEEGMYEAAKLLYSNVSNFA RLASTLVHLGEYQAAVDNSRKASSTRTWKEVCFACMDGQEFRFAQLCGLHIVIHADELEELMCYYQDRGYFEELILLLEAALGLERAHMG MFTELAILYSKFKPQKMLEHLELFWSRVNIPKVLRAAEQAHLWAELVFLYDKYEEYDNAVLTMMSHPTEAWKEGQFKDIITKVANVELCY RALQFYLDYKPLLINDLLLVLSPRLDHTWTVSFFSKAGQLPLVKPYLRSVQSHNNKSVNEALNHLLTEEEDYQDAMQHAAESRDAELAQK -------------------------------------------------------------- >22400_22400_6_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19263353_ENST00000427926_length(transcript)=5798nt_BP=403nt GCGCGCAGCTGGAGTCGGTGCCTGAGGTTGCAGCCGAGAGTGTGCGCCAGCCCGCGGCCCAGCCGAAGCTCTTTCCCGCCGCCTCTCCGC GCCTCGCCCAGGTTCAGCTCCGCCTGACCCTCCGCTTGGCACGGTCCCCTGACCCTCAGTCCGGCCTGGTCCGGCCTTCCAGCCGCCCCG GGGACGATGAACGGAGGATAAATGGTGCCCAAGGCAGACAGCGGCGCCTTCCTGCTGCTCTTCCTGCTCGTGCTCACTGTCACCGAGCCG CTGCGGCCAGAGCTGCGGTGCAACCCTGGGCAGTTTGCGTGTCGCAGCGGCACCATCCAGTGCATCCCCCTCCCCTGGCAGTGTGACGGC TGGGCGACTTGCGAGGATGAGAGCGACGAAGCCAACTGTCCAGCTCCAAAACCTTGGAATTAATCCAGCTAACATTGGATTCAGCACACT GACCATGGAATCTGACAAGTTCATATGTATCCGAGAGAAAGTTGGTGAGCAGGCACAGGTCACGATCATTGACATGAGTGACCCAATGGC TCCGATCCGACGGCCTATCTCTGCAGAGAGTGCCATCATGAATCCAGCCTCTAAGGTGATAGCTCTGAAAGCTGGGAAGACACTTCAGAT CTTTAATATTGAGATGAAGAGTAAAATGAAGGCTCATACTATGGCAGAAGAAGTGATTTTCTGGAAATGGGTTTCTGTGAACACTGTTGC CTTGGTGACCGAGACCGCGGTCTACCACTGGAGCATGGAAGGTGACTCCCAGCCCATGAAGATGTTTGATAGACATACCAGTCTGGTGGG CTGCCAGGTGATTCACTACCGGACTGATGAGTACCAGAAGTGGCTGCTGCTCGTAGGCATCTCGGCTCAGCAAAACCGTGTGGTTGGAGC AATGCAGCTCTACTCTGTGGATAGGAAGGTTTCACAACCCATAGAAGGCCATGCTGCGGCTTTTGCAGAGTTCAAGATGGAGGGGAATGC CAAGCCTGCCACCCTTTTCTGCTTTGCTGTACGTAATCCCACAGGAGGCAAGTTGCACATCATTGAAGTTGGACAGCCTGCAGCGGGAAA CCAACCTTTTGTAAAGAAAGCAGTAGATGTGTTTTTTCCTCCAGAGGCACAGAATGATTTTCCAGTGGCTATGCAGATTGGAGCTAAACA TGGTGTTATTTACTTGATCACAAAGTATGGCTATCTTCATCTGTACGACCTAGAGTCTGGCGTGTGCATCTGCATGAACCGTATTAGTGC TGACACAATATTTGTCACTGCTCCACACAAACCAACCTCTGGAATTATTGGTGTCAACAAAAAGGGACAGGTACTGTCAGTTTGTGTTGA GGAAGATAACATTGTGAATTATGCAACCAACGTGCTTCAGAATCCAGACCTTGGTCTGCGTTTGGCCGTTCGTAGTAACCTGGCTGGGGC AGAGAAGTTGTTTGTGAGAAAATTCAATACCCTCTTTGCACAGGGCAGCTATGCTGAAGCCGCCAAAGTTGCAGCGTCTGCACCAAAGGG AATCCTGCGTACCAGAGAGACGGTCCAGAAATTCCAGAGTATACCCGCTCAGTCTGGCCAGGCTTCTCCATTGCTGCAGTACTTCGGAAT CCTGCTCGACCAGGGTCAGCTCAATAAACTTGAATCCTTAGAACTTTGCCATCTGGTTCTTCAGCAGGGGCGTAAGCAACTCCTAGAGAA GTGGCTGAAAGAAGATAAGCTGGAGTGCTCAGAGGAGCTCGGAGACTTGGTCAAAACCACTGACCCCATGCTCGCTCTGAGTGTGTACCT TCGGGCAAATGTGCCAAGCAAAGTGATCCAGTGTTTTGCAGAAACAGGCCAATTCCAGAAAATTGTGCTCTATGCCAAAAAGGTTGGGTA CACCCCAGACTGGATCTTTCTGCTGAGGGGTGTAATGAAGATCAGTCCGGAACAGGGCCTGCAGTTTTCTCGAATGCTAGTGCAGGACGA GGAGCCGCTGGCCAACATTAGCCAGATTGTGGACATTTTCATGGAAAACAGTTTAATTCAGCAGTGTACTTCCTTCTTATTGGATGCCTT GAAGAATAATCGCCCAGCTGAGGGACTCCTGCAGACATGGCTGTTGGAGATGAACCTTGTTCATGCACCCCAGGTTGCAGATGCCATCCT TGGAAATAAAATGTTTACTCATTACGACCGGGCCCACATTGCCCAGCTCTGTGAGAAGGCAGGCCTCCTGCAGCAAGCACTGGAGCACTA CACCGACCTCTATGACATCAAGAGGGCTGTGGTCCACACTCACCTCCTCAATCCCGAGTGGCTTGTCAATTTCTTTGGCTCCTTATCGGT GGAGGATTCTGTGGAGTGTCTGCATGCCATGCTGTCTGCTAACATCAGACAGAACCTTCAGCTGTGTGTGCAGGTGGCCTCTAAGTACCA CGAGCAGCTGGGCACGCAGGCCCTGGTGGAGCTCTTTGAATCCTTCAAGAGTTACAAAGGCCTCTTCTACTTCCTGGGCTCAATCGTGAA CTTCAGCCAAGACCCAGATGTGCATCTGAAATACATTCAGGCTGCCTGTAAGACAGGGCAGATCAAGGAGGTGGAGAGGATATGCCGAGA GAGCAGCTGCTACAACCCAGAGCGTGTGAAGAACTTCCTGAAGGAGGCCAAGCTCACAGACCAGCTTCCCCTCATCATCGTGTGTGATCG TTTTGGCTTTGTCCATGACCTTGTCCTATATTTATACCGCAACAACCTGCAGAGGTACATTGAGATCTACGTGCAGAAGGTCAACCCTAG CCGGACCCCAGCTGTGATTGGAGGGCTGCTTGATGTGGATTGTTCTGAGGAAGTGATTAAACACTTAATCATGGCAGTGAGAGGACAGTT CTCTACTGATGAGTTGGTGGCTGAAGTAGAAAAAAGAAATAGGCTCAAGCTGCTGCTTCCCTGGCTGGAGTCCCAGATTCAGGAAGGCTG TGAGGAGCCTGCCACTCACAATGCACTGGCTAAAATCTACATCGACAGCAACAACAGCCCCGAGTGCTTCCTGAGAGAGAATGCCTACTA TGACAGCAGCGTGGTGGGCCGCTACTGTGAGAAGCGAGACCCCCATCTGGCCTGTGTTGCCTATGAGCGGGGGCAGTGTGACCTTGAGCT CATCAAGGTGTGCAATGAGAATTCTCTGTTCAAAAGCGAGGCCCGCTACCTGGTATGCAGAAAGGATCCGGAGCTCTGGGCTCACGTCCT TGAGGAGACCAACCCATCCAGGAGACAGCTAATTGACCAGGTGGTACAGACAGCATTGTCAGAAACACGGGATCCTGAAGAGATTTCGGT CACTGTCAAAGCCTTTATGACAGCCGACCTGCCTAATGAACTGATTGAACTGCTGGAGAAGATAGTTCTGGATAACTCTGTCTTCAGCGA GCACAGGAATCTACAGAATCTGTTGATCCTGACTGCCATCAAGGCAGACCGCACACGGGTCATGGAGTACATCAGCCGCCTGGACAACTA TGACGCACTGGACATCGCGAGCATCGCTGTCAGCAGCGCACTGTATGAGGAGGCCTTCACCGTTTTCCACAAGTTTGATATGAATGCCTC AGCAATCCAGGTCCTGATCGAGCACATTGGAAACCTGGACCGGGCATATGAGTTTGCGGAGAGATGCAATGAGCCTGCTGTGTGGAGTCA GCTGGCCCAAGCCCAGCTCCAGAAAGATTTGGTGAAGGAAGCCATCAACTCCTATATCAGAGGGGACGACCCTTCCTCTTACCTGGAAGT TGTTCAGTCAGCCAGCAGGAGCAACAACTGGGAGGATCTAGTTAAATTTCTGCAGATGGCCAGGAAAAAGGGCCGTGAGTCCTATATAGA GACTGAACTTATTTTTGCCTTGGCTAAAACCAGCCGTGTTTCTGAGCTAGAAGATTTTATTAATGGACCCAACAATGCCCACATCCAGCA GGTCGGGGACCGCTGTTACGAGGAGGGAATGTACGAGGCTGCCAAGCTGCTCTATAGCAATGTTTCTAACTTTGCCCGCCTGGCTTCCAC CTTGGTTCACCTCGGTGAGTATCAGGCAGCAGTGGACAACAGCCGCAAGGCCAGCAGCACCCGGACGTGGAAGGAGGTGTGCTTTGCCTG CATGGATGGACAAGAGTTCCGCTTCGCACAGCTGTGTGGTCTTCACATCGTCATTCATGCAGATGAGCTGGAGGAGCTGATGTGCTATTA CCAGGATCGTGGCTACTTTGAGGAGCTGATCTTGCTGTTGGAAGCGGCCCTGGGCCTGGAGCGGGCCCACATGGGCATGTTCACTGAGCT GGCCATCCTCTACTCCAAATTCAAGCCACAGAAGATGCTGGAGCATCTGGAGCTTTTCTGGTCCCGTGTCAACATCCCAAAGGTGCTGAG GGCTGCAGAGCAGGCACACCTGTGGGCTGAGCTGGTGTTCCTCTATGACAAGTACGAGGAGTATGACAATGCTGTGCTCACCATGATGAG CCACCCCACTGAGGCCTGGAAGGAGGGTCAGTTCAAGGACATCATTACCAAGGTTGCCAACGTCGAGCTCTGTTACAGAGCCCTGCAGTT CTATTTGGATTACAAACCACTGCTCATCAATGACCTGCTGCTGGTGCTTTCACCCCGGCTGGACCACACCTGGACAGTCAGTTTCTTTTC AAAGGCAGGTCAGCTGCCCCTGGTGAAGCCTTACCTGCGGTCAGTCCAGAGCCACAACAACAAGAGTGTGAATGAGGCACTCAACCACCT GCTGACAGAGGAGGAGGACTATCAGGGCTTAAGGGCATCTATCGATGCCTATGACAACTTTGACAACATCAGCCTGGCTCAGCAGCTGGA GAAGCATCAGCTGATGGAGTTCAGGTGCATTGCGGCCTATCTGTACAAGGGCAATAACTGGTGGGCCCAGAGCGTGGAGCTCTGCAAGAA GGATCATCTCTACAAGGATGCCATGCAGCATGCTGCAGAGTCGCGGGATGCTGAGCTGGCCCAGAAGTTGCTGCAGTGGTTCCTGGAGGA AGGCAAGAGGGAGTGCTTCGCAGCTTGTCTCTTCACCTGCTATGACCTGCTTCGCCCAGACATGGTGCTTGAGCTGGCCTGGAGGCACAA CCTCGTGGACTTGGCCATGCCCTACTTCATCCAGGTGATGAGGGAGTACCTGAGCAAGGTGGACAAACTGGATGCCTTGGAGAGTCTGCG CAAGCAAGAGGAGCATGTGACAGAGCCTGCCCCTCTCGTGTTTGATTTTGATGGGCATGAATGAGACCCAGCTGATTGCACTAAGCCCTG CCGTGGGCCCAGCCCCTGCCAGCTTCCCCTATGGATATGCCTCTGCTCCCAACTTCGCCAGCCTCCAATGTACAACTTCCGCGTGTAGTG GGCGTTGTCACCACCCACCCTACCTGCAGAGTTACTAACTTCTCCAAGGAGCATGTCACTCCAGCAGCACAGGGGACGCAATGGGAGGCA GGGACACCTGGACAATATTTATTTTTGCTGAAACCCAATGACGGCAACCTCTGAGCCATCCCAGAGCCTGGGGAGGCCAGGGTAGAGGCT GACGGCGCAAGACCAGCTTTAGCCGACAACAGAGACTGGACTGTGGGCCCTCCTGCTGGAGCCAGGCCTTCCTCCTGGGCGCCTCCGACT GGCTGGAGCTGCCCCCTCCAGGCCAGTTTGAAGACTACATGAACACGTCTTGTTTGGAGGTACCGGACCTCATAAAAGGACTCTCAGCCT >22400_22400_6_DGCR2-CLTCL1_DGCR2_chr22_19076881_ENST00000545799_CLTCL1_chr22_19263353_ENST00000427926_length(amino acids)=1675AA_BP=3 MSPSRCGQSCGATLGSLRVAAAPSSASPSPGSVTAGRLARMRATKPTVQLQNLGINPANIGFSTLTMESDKFICIREKVGEQAQVTIIDM SDPMAPIRRPISAESAIMNPASKVIALKAGKTLQIFNIEMKSKMKAHTMAEEVIFWKWVSVNTVALVTETAVYHWSMEGDSQPMKMFDRH TSLVGCQVIHYRTDEYQKWLLLVGISAQQNRVVGAMQLYSVDRKVSQPIEGHAAAFAEFKMEGNAKPATLFCFAVRNPTGGKLHIIEVGQ PAAGNQPFVKKAVDVFFPPEAQNDFPVAMQIGAKHGVIYLITKYGYLHLYDLESGVCICMNRISADTIFVTAPHKPTSGIIGVNKKGQVL SVCVEEDNIVNYATNVLQNPDLGLRLAVRSNLAGAEKLFVRKFNTLFAQGSYAEAAKVAASAPKGILRTRETVQKFQSIPAQSGQASPLL QYFGILLDQGQLNKLESLELCHLVLQQGRKQLLEKWLKEDKLECSEELGDLVKTTDPMLALSVYLRANVPSKVIQCFAETGQFQKIVLYA KKVGYTPDWIFLLRGVMKISPEQGLQFSRMLVQDEEPLANISQIVDIFMENSLIQQCTSFLLDALKNNRPAEGLLQTWLLEMNLVHAPQV ADAILGNKMFTHYDRAHIAQLCEKAGLLQQALEHYTDLYDIKRAVVHTHLLNPEWLVNFFGSLSVEDSVECLHAMLSANIRQNLQLCVQV ASKYHEQLGTQALVELFESFKSYKGLFYFLGSIVNFSQDPDVHLKYIQAACKTGQIKEVERICRESSCYNPERVKNFLKEAKLTDQLPLI IVCDRFGFVHDLVLYLYRNNLQRYIEIYVQKVNPSRTPAVIGGLLDVDCSEEVIKHLIMAVRGQFSTDELVAEVEKRNRLKLLLPWLESQ IQEGCEEPATHNALAKIYIDSNNSPECFLRENAYYDSSVVGRYCEKRDPHLACVAYERGQCDLELIKVCNENSLFKSEARYLVCRKDPEL WAHVLEETNPSRRQLIDQVVQTALSETRDPEEISVTVKAFMTADLPNELIELLEKIVLDNSVFSEHRNLQNLLILTAIKADRTRVMEYIS RLDNYDALDIASIAVSSALYEEAFTVFHKFDMNASAIQVLIEHIGNLDRAYEFAERCNEPAVWSQLAQAQLQKDLVKEAINSYIRGDDPS SYLEVVQSASRSNNWEDLVKFLQMARKKGRESYIETELIFALAKTSRVSELEDFINGPNNAHIQQVGDRCYEEGMYEAAKLLYSNVSNFA RLASTLVHLGEYQAAVDNSRKASSTRTWKEVCFACMDGQEFRFAQLCGLHIVIHADELEELMCYYQDRGYFEELILLLEAALGLERAHMG MFTELAILYSKFKPQKMLEHLELFWSRVNIPKVLRAAEQAHLWAELVFLYDKYEEYDNAVLTMMSHPTEAWKEGQFKDIITKVANVELCY RALQFYLDYKPLLINDLLLVLSPRLDHTWTVSFFSKAGQLPLVKPYLRSVQSHNNKSVNEALNHLLTEEEDYQGLRASIDAYDNFDNISL AQQLEKHQLMEFRCIAAYLYKGNNWWAQSVELCKKDHLYKDAMQHAAESRDAELAQKLLQWFLEEGKRECFAACLFTCYDLLRPDMVLEL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for DGCR2-CLTCL1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for DGCR2-CLTCL1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for DGCR2-CLTCL1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
