
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:DNAJB6-AGT (FusionGDB2 ID:23431) |
Fusion Gene Summary for DNAJB6-AGT |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: DNAJB6-AGT | Fusion gene ID: 23431 | Hgene | Tgene | Gene symbol | DNAJB6 | AGT | Gene ID | 10049 | 189 |
| Gene name | DnaJ heat shock protein family (Hsp40) member B6 | alanine--glyoxylate and serine--pyruvate aminotransferase | |
| Synonyms | DJ4|DnaJ|HHDJ1|HSJ-2|HSJ2|LGMD1D|LGMD1E|LGMDD1|MRJ|MSJ-1 | AGT|AGT1|AGXT1|PH1|SPAT|SPT|TLH6 | |
| Cytomap | 7q36.3 | 2q37.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | dnaJ homolog subfamily B member 6DnaJ (Hsp40) homolog, subfamily B, member 6DnaJ-like 2 proteinheat shock protein J2 | serine--pyruvate aminotransferaseL-alanine: glyoxylate aminotransferase 1alanine-glyoxylate aminotransferasehepatic peroxisomal alanine:glyoxylate aminotransferase | |
| Modification date | 20200328 | 20200313 | |
| UniProtAcc | O75190 | Q6RW13 | |
| Ensembl transtripts involved in fusion gene | ENST00000429029, ENST00000262177, ENST00000452797, ENST00000443280, ENST00000494267, | ENST00000366667, | |
| Fusion gene scores | * DoF score | 7 X 5 X 7=245 | 2 X 2 X 3=12 |
| # samples | 9 | 3 | |
| ** MAII score | log2(9/245*10)=-1.4447848426729 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(3/12*10)=1.32192809488736 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: DNAJB6 [Title/Abstract] AND AGT [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | DNAJB6(157178305)-AGT(230841946), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | DNAJB6-AGT seems lost the major protein functional domain in Tgene partner, which is a cell metabolism gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | DNAJB6 | GO:0006457 | protein folding | 11896048 |
| Hgene | DNAJB6 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process | 11896048 |
| Hgene | DNAJB6 | GO:0045109 | intermediate filament organization | 10954706 |
| Hgene | DNAJB6 | GO:0090084 | negative regulation of inclusion body assembly | 21231916 |
| Tgene | AGT | GO:0009436 | glyoxylate catabolic process | 22198249 |
| Tgene | AGT | GO:0019265 | glycine biosynthetic process, by transamination of glyoxylate | 22198249 |
| Tgene | AGT | GO:0019448 | L-cysteine catabolic process | 18492492 |
| Tgene | AGT | GO:0042853 | L-alanine catabolic process | 17696873|18492492|22198249 |
 Fusion gene breakpoints across DNAJB6 (5'-gene) Fusion gene breakpoints across DNAJB6 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
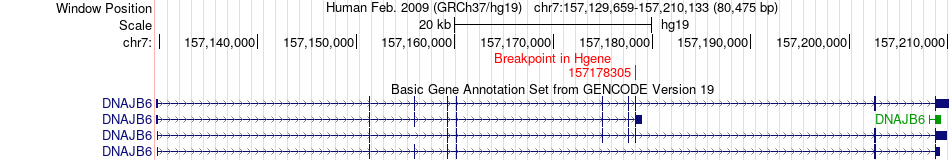 |
 Fusion gene breakpoints across AGT (3'-gene) Fusion gene breakpoints across AGT (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
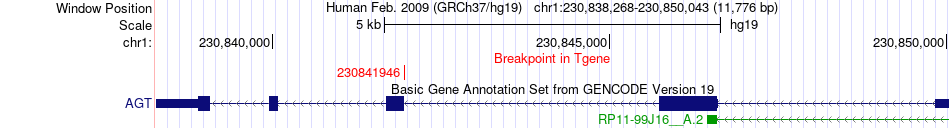 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LUSC | TCGA-98-A53H-01A | DNAJB6 | chr7 | 157178305 | + | AGT | chr1 | 230841946 | - |
| ChimerDB4 | LUSC | TCGA-98-A53H | DNAJB6 | chr7 | 157178305 | + | AGT | chr1 | 230841946 | - |
Top |
Fusion Gene ORF analysis for DNAJB6-AGT |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000429029 | ENST00000366667 | DNAJB6 | chr7 | 157178305 | + | AGT | chr1 | 230841946 | - |
| In-frame | ENST00000262177 | ENST00000366667 | DNAJB6 | chr7 | 157178305 | + | AGT | chr1 | 230841946 | - |
| In-frame | ENST00000452797 | ENST00000366667 | DNAJB6 | chr7 | 157178305 | + | AGT | chr1 | 230841946 | - |
| intron-3CDS | ENST00000443280 | ENST00000366667 | DNAJB6 | chr7 | 157178305 | + | AGT | chr1 | 230841946 | - |
| intron-3CDS | ENST00000494267 | ENST00000366667 | DNAJB6 | chr7 | 157178305 | + | AGT | chr1 | 230841946 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000262177 | DNAJB6 | chr7 | 157178305 | + | ENST00000366667 | AGT | chr1 | 230841946 | - | 2116 | 896 | 205 | 1497 | 430 |
| ENST00000452797 | DNAJB6 | chr7 | 157178305 | + | ENST00000366667 | AGT | chr1 | 230841946 | - | 1956 | 736 | 192 | 1337 | 381 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000262177 | ENST00000366667 | DNAJB6 | chr7 | 157178305 | + | AGT | chr1 | 230841946 | - | 0.002139063 | 0.99786097 |
| ENST00000452797 | ENST00000366667 | DNAJB6 | chr7 | 157178305 | + | AGT | chr1 | 230841946 | - | 0.002674623 | 0.9973253 |
Top |
Fusion Genomic Features for DNAJB6-AGT |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
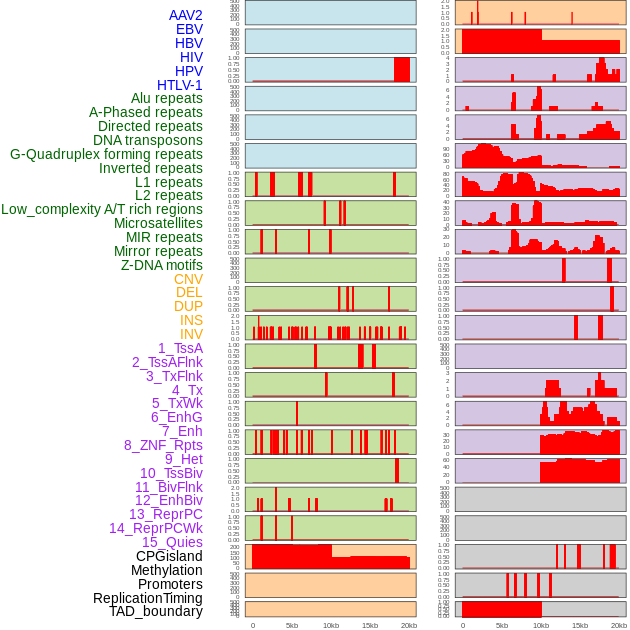 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for DNAJB6-AGT |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr7:157178305/chr1:230841946) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| DNAJB6 | AGT |
| FUNCTION: Plays an indispensable role in the organization of KRT8/KRT18 filaments. Acts as an endogenous molecular chaperone for neuronal proteins including huntingtin. Suppresses aggregation and toxicity of polyglutamine-containing, aggregation-prone proteins. Isoform B but not isoform A inhibits huntingtin aggregation. Has a stimulatory effect on the ATPase activity of HSP70 in a dose-dependent and time-dependent manner and hence acts as a co-chaperone of HSP70. Also reduces cellular toxicity and caspase-3 activity. {ECO:0000269|PubMed:10954706, ECO:0000269|PubMed:11896048, ECO:0000269|PubMed:20159555, ECO:0000269|PubMed:22366786, ECO:0000269|PubMed:28233300}. | FUNCTION: Appears to be a negative regulator of type-1 angiotensin II receptor-mediated signaling by regulating receptor internalisation as well as mechanism of receptor desensitization such as phosphorylation. Induces also a decrease in cell proliferation and angiotensin II-stimulated transcriptional activity. {ECO:0000269|PubMed:12960423}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000262177 | + | 8 | 10 | 155_194 | 230 | 327.0 | Compositional bias | Note=Ser-rich |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000262177 | + | 8 | 10 | 83_172 | 230 | 327.0 | Compositional bias | Note=Gly/Phe-rich |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000262177 | + | 8 | 10 | 2_69 | 230 | 327.0 | Domain | J |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000429029 | + | 1 | 8 | 155_194 | 0 | 242.0 | Compositional bias | Note=Ser-rich |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000429029 | + | 1 | 8 | 83_172 | 0 | 242.0 | Compositional bias | Note=Gly/Phe-rich |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000429029 | + | 1 | 8 | 2_69 | 0 | 242.0 | Domain | J |
Top |
Fusion Gene Sequence for DNAJB6-AGT |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >23431_23431_1_DNAJB6-AGT_DNAJB6_chr7_157178305_ENST00000262177_AGT_chr1_230841946_ENST00000366667_length(transcript)=2116nt_BP=896nt AGTTGACGCCGCCGTGCCGCCGCCGCGTGACGCATTTCCTGTTTGTTGTTGGAGAAAGGAGAGAAAGGAAAGCGCGAGGAGCCGCCGCCA CCACCAGCGCAGCAGTCCTGGAGCTGTGAGGAGATTCGGGCCGTCACCCTGCCTCCCCTGCGTCCCGCCACCGGCCGCTTCTGTCCTCGG ACCCATTCCAACAATCTCGTAAAACATGGTGGATTACTATGAAGTTCTAGGCGTGCAGAGACATGCCTCACCCGAGGATATTAAAAAGGC ATATCGGAAACTGGCACTGAAGTGGCATCCAGATAAAAATCCTGAGAATAAAGAAGAAGCAGAGAGAAAATTCAAGCAAGTAGCGGAGGC ATATGAAGTGCTGTCGGATGCTAAGAAACGGGACATCTATGACAAATATGGCAAAGAAGGATTAAATGGTGGAGGAGGAGGTGGAAGTCA TTTTGACAGTCCATTTGAATTTGGCTTCACATTCCGTAACCCAGATGATGTCTTCAGGGAATTTTTTGGTGGAAGGGACCCATTTTCATT TGACTTCTTTGAAGACCCTTTTGAGGACTTCTTTGGGAATCGAAGGGGTCCCCGAGGAAGCAGAAGCCGAGGGACGGGGTCGTTTTTCTC TGCGTTCAGTGGATTTCCGTCTTTTGGAAGTGGATTTTCTTCTTTTGATACAGGATTTACTTCATTTGGGTCACTAGGTCACGGGGGCCT CACTTCATTCTCTTCCACGTCATTTGGTGGTAGTGGCATGGGCAACTTCAAATCGATATCAACTTCAACTAAAATGGTTAATGGCAGAAA AATCACTACAAAGAGAATTGTCGAGAACGGTCAAGAAAGAGTAGAAGTTGAAGAAGATGGCCAGTTAAAGTCCTTAACAATAAATGGGAA GATGAAGGGCTTCTCCCTGCTGGCCGAGCCCCAGGAGTTCTGGGTGGACAACAGCACCTCAGTGTCTGTTCCCATGCTCTCTGGCATGGG CACCTTCCAGCACTGGAGTGACATCCAGGACAACTTCTCGGTGACTCAAGTGCCCTTCACTGAGAGCGCCTGCCTGCTGCTGATCCAGCC TCACTATGCCTCTGACCTGGACAAGGTGGAGGGTCTCACTTTCCAGCAAAACTCCCTCAACTGGATGAAGAAACTATCTCCCCGGACCAT CCACCTGACCATGCCCCAACTGGTGCTGCAAGGATCTTATGACCTGCAGGACCTGCTCGCCCAGGCTGAGCTGCCCGCCATTCTGCACAC CGAGCTGAACCTGCAAAAATTGAGCAATGACCGCATCAGGGTGGGGGAGGTGCTGAACAGCATTTTTTTTGAGCTTGAAGCGGATGAGAG AGAGCCCACAGAGTCTACCCAACAGCTTAACAAGCCTGAGGTCTTGGAGGTGACCCTGAACCGCCCATTCCTGTTTGCTGTGTATGATCA AAGCGCCACTGCCCTGCACTTCCTGGGCCGCGTGGCCAACCCGCTGAGCACAGCATGAGGCCAGGGCCCCAGAACACAGTGCCTGGCAAG GCCTCTGCCCCTGGCCTTTGAGGCAAAGGCCAGCAGCAGATAACAACCCCGGACAAATCAGCGATGTGTCACCCCCAGTCTCCCACCTTT TCTTCTAATGAGTCGACTTTGAGCTGGAAAGCAGCCGTTTCTCCTTGGTCTAAGTGTGCTGCATGGAGTGAGCAGTAGAAGCCTGCAGCG GCACAAATGCACCTCCCAGTTTGCTGGGTTTATTTTAGAGAATGGGGGTGGGGAGGCAAGAACCAGTGTTTAGCGCGGGACTACTGTTCC AAAAAGAATTCCAACCGACCAGCTTGTTTGTGAAACAAAAAAGTGTTCCCTTTTCAAGTTGAGAACAAAAATTGGGTTTTAAAATTAAAG TATACATTTTTGCATTGCCTTCGGTTTGTATTTAGTGTCTTGAATGTAAGAACATGACCTCCGTGTAGTGTCTGTAATACCTTAGTTTTT TCCACAGATGCTTGTGATTTTTGAACAATACGTGAAAGATGCAAGCACCTGAATTTCTGTTTGAATGCGGAACCATAGCTGGTTATTTCT >23431_23431_1_DNAJB6-AGT_DNAJB6_chr7_157178305_ENST00000262177_AGT_chr1_230841946_ENST00000366667_length(amino acids)=430AA_BP=228 MVDYYEVLGVQRHASPEDIKKAYRKLALKWHPDKNPENKEEAERKFKQVAEAYEVLSDAKKRDIYDKYGKEGLNGGGGGGSHFDSPFEFG FTFRNPDDVFREFFGGRDPFSFDFFEDPFEDFFGNRRGPRGSRSRGTGSFFSAFSGFPSFGSGFSSFDTGFTSFGSLGHGGLTSFSSTSF GGSGMGNFKSISTSTKMVNGRKITTKRIVENGQERVEVEEDGQLKSLTINGKMKGFSLLAEPQEFWVDNSTSVSVPMLSGMGTFQHWSDI QDNFSVTQVPFTESACLLLIQPHYASDLDKVEGLTFQQNSLNWMKKLSPRTIHLTMPQLVLQGSYDLQDLLAQAELPAILHTELNLQKLS -------------------------------------------------------------- >23431_23431_2_DNAJB6-AGT_DNAJB6_chr7_157178305_ENST00000452797_AGT_chr1_230841946_ENST00000366667_length(transcript)=1956nt_BP=736nt GGAGAAAGGAGAGAAAGGAAAGCGCGAGGAGCCGCCGCCACCACCAGCGCAGCAGTCCTGGAGCTGTGAGGAGATTCGGGCCGTCACCCT GCCTCCCCTGCGTCCCGCCACCGGCCGCTTCTGTCCTCGGACCCATTCCAACAATCTCGTAAAACATGGTGGATTACTATGAAGTTCTAG GCGTGCAGAGACATGCCTCACCCGAGGATATTAAAAAGGCCTAAGAAACGGGACATCTATGACAAATATGGCAAAGAAGGATTAAATGGT GGAGGAGGAGGTGGAAGTCATTTTGACAGTCCATTTGAATTTGGCTTCACATTCCGTAACCCAGATGATGTCTTCAGGGAATTTTTTGGT GGAAGGGACCCATTTTCATTTGACTTCTTTGAAGACCCTTTTGAGGACTTCTTTGGGAATCGAAGGGGTCCCCGAGGAAGCAGAAGCCGA GGGACGGGGTCGTTTTTCTCTGCGTTCAGTGGATTTCCGTCTTTTGGAAGTGGATTTTCTTCTTTTGATACAGGATTTACTTCATTTGGG TCACTAGGTCACGGGGGCCTCACTTCATTCTCTTCCACGTCATTTGGTGGTAGTGGCATGGGCAACTTCAAATCGATATCAACTTCAACT AAAATGGTTAATGGCAGAAAAATCACTACAAAGAGAATTGTCGAGAACGGTCAAGAAAGAGTAGAAGTTGAAGAAGATGGCCAGTTAAAG TCCTTAACAATAAATGGGAAGATGAAGGGCTTCTCCCTGCTGGCCGAGCCCCAGGAGTTCTGGGTGGACAACAGCACCTCAGTGTCTGTT CCCATGCTCTCTGGCATGGGCACCTTCCAGCACTGGAGTGACATCCAGGACAACTTCTCGGTGACTCAAGTGCCCTTCACTGAGAGCGCC TGCCTGCTGCTGATCCAGCCTCACTATGCCTCTGACCTGGACAAGGTGGAGGGTCTCACTTTCCAGCAAAACTCCCTCAACTGGATGAAG AAACTATCTCCCCGGACCATCCACCTGACCATGCCCCAACTGGTGCTGCAAGGATCTTATGACCTGCAGGACCTGCTCGCCCAGGCTGAG CTGCCCGCCATTCTGCACACCGAGCTGAACCTGCAAAAATTGAGCAATGACCGCATCAGGGTGGGGGAGGTGCTGAACAGCATTTTTTTT GAGCTTGAAGCGGATGAGAGAGAGCCCACAGAGTCTACCCAACAGCTTAACAAGCCTGAGGTCTTGGAGGTGACCCTGAACCGCCCATTC CTGTTTGCTGTGTATGATCAAAGCGCCACTGCCCTGCACTTCCTGGGCCGCGTGGCCAACCCGCTGAGCACAGCATGAGGCCAGGGCCCC AGAACACAGTGCCTGGCAAGGCCTCTGCCCCTGGCCTTTGAGGCAAAGGCCAGCAGCAGATAACAACCCCGGACAAATCAGCGATGTGTC ACCCCCAGTCTCCCACCTTTTCTTCTAATGAGTCGACTTTGAGCTGGAAAGCAGCCGTTTCTCCTTGGTCTAAGTGTGCTGCATGGAGTG AGCAGTAGAAGCCTGCAGCGGCACAAATGCACCTCCCAGTTTGCTGGGTTTATTTTAGAGAATGGGGGTGGGGAGGCAAGAACCAGTGTT TAGCGCGGGACTACTGTTCCAAAAAGAATTCCAACCGACCAGCTTGTTTGTGAAACAAAAAAGTGTTCCCTTTTCAAGTTGAGAACAAAA ATTGGGTTTTAAAATTAAAGTATACATTTTTGCATTGCCTTCGGTTTGTATTTAGTGTCTTGAATGTAAGAACATGACCTCCGTGTAGTG TCTGTAATACCTTAGTTTTTTCCACAGATGCTTGTGATTTTTGAACAATACGTGAAAGATGCAAGCACCTGAATTTCTGTTTGAATGCGG >23431_23431_2_DNAJB6-AGT_DNAJB6_chr7_157178305_ENST00000452797_AGT_chr1_230841946_ENST00000366667_length(amino acids)=381AA_BP=179 MPHPRILKRPKKRDIYDKYGKEGLNGGGGGGSHFDSPFEFGFTFRNPDDVFREFFGGRDPFSFDFFEDPFEDFFGNRRGPRGSRSRGTGS FFSAFSGFPSFGSGFSSFDTGFTSFGSLGHGGLTSFSSTSFGGSGMGNFKSISTSTKMVNGRKITTKRIVENGQERVEVEEDGQLKSLTI NGKMKGFSLLAEPQEFWVDNSTSVSVPMLSGMGTFQHWSDIQDNFSVTQVPFTESACLLLIQPHYASDLDKVEGLTFQQNSLNWMKKLSP RTIHLTMPQLVLQGSYDLQDLLAQAELPAILHTELNLQKLSNDRIRVGEVLNSIFFELEADEREPTESTQQLNKPEVLEVTLNRPFLFAV -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for DNAJB6-AGT |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000262177 | + | 8 | 10 | 2_146 | 230.33333333333334 | 327.0 | HSP70 |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000429029 | + | 1 | 8 | 2_146 | 0 | 242.0 | HSP70 |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000262177 | + | 8 | 10 | 119_242 | 230.33333333333334 | 327.0 | KRT18 |
| Hgene | DNAJB6 | chr7:157178305 | chr1:230841946 | ENST00000429029 | + | 1 | 8 | 119_242 | 0 | 242.0 | KRT18 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for DNAJB6-AGT |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for DNAJB6-AGT |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
