
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:EHF-CAT (FusionGDB2 ID:25568) |
Fusion Gene Summary for EHF-CAT |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: EHF-CAT | Fusion gene ID: 25568 | Hgene | Tgene | Gene symbol | EHF | CAT | Gene ID | 26298 | 10249 |
| Gene name | ETS homologous factor | glycine-N-acyltransferase | |
| Synonyms | ESE3|ESE3B|ESEJ | ACGNAT|CAT|GAT | |
| Cytomap | 11p13 | 11q12.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ETS homologous factorESE3 transcription factorETS domain-containing transcription factorepithelium-specific Ets transcription factor 3hEHF | glycine N-acyltransferaseAAcHRP-1(CLP)acyl-CoA:glycine N-acyltransferasearalkyl acyl-CoA N-acyltransferasearalkyl acyl-CoA:amino acid N-acyltransferasearalkyl-CoA N-acyltransferasebenzoyl-coenzyme A:glycine N-acyltransferaseepididymis secretory sp | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | Q9NZC4 | Q6ZRH7 | |
| Ensembl transtripts involved in fusion gene | ENST00000257831, ENST00000450654, ENST00000527935, ENST00000530286, ENST00000531728, ENST00000531794, ENST00000533754, ENST00000527001, | ENST00000534710, ENST00000241052, | |
| Fusion gene scores | * DoF score | 9 X 13 X 4=468 | 14 X 11 X 8=1232 |
| # samples | 12 | 18 | |
| ** MAII score | log2(12/468*10)=-1.96347412397489 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(18/1232*10)=-2.77493344436523 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: EHF [Title/Abstract] AND CAT [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | EHF(34668231)-CAT(34485652), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | EHF | GO:0045893 | positive regulation of transcription, DNA-templated | 10644770 |
| Hgene | EHF | GO:0045944 | positive regulation of transcription by RNA polymerase II | 17027647 |
| Tgene | CAT | GO:0006544 | glycine metabolic process | 22475485 |
| Tgene | CAT | GO:1901787 | benzoyl-CoA metabolic process | 22475485 |
 Fusion gene breakpoints across EHF (5'-gene) Fusion gene breakpoints across EHF (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
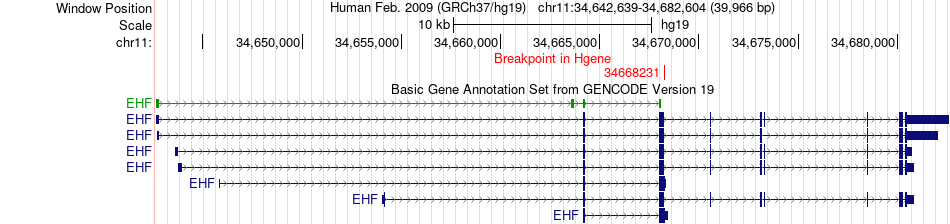 |
 Fusion gene breakpoints across CAT (3'-gene) Fusion gene breakpoints across CAT (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
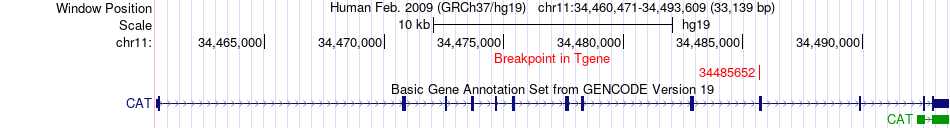 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-BH-A0B9-01A | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
Top |
Fusion Gene ORF analysis for EHF-CAT |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000257831 | ENST00000534710 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| 5CDS-intron | ENST00000450654 | ENST00000534710 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| 5CDS-intron | ENST00000527935 | ENST00000534710 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| 5CDS-intron | ENST00000530286 | ENST00000534710 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| 5CDS-intron | ENST00000531728 | ENST00000534710 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| 5CDS-intron | ENST00000531794 | ENST00000534710 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| 5CDS-intron | ENST00000533754 | ENST00000534710 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| In-frame | ENST00000257831 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| In-frame | ENST00000450654 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| In-frame | ENST00000527935 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| In-frame | ENST00000530286 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| In-frame | ENST00000531728 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| In-frame | ENST00000531794 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| In-frame | ENST00000533754 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| intron-3CDS | ENST00000527001 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
| intron-intron | ENST00000527001 | ENST00000534710 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000257831 | EHF | chr11 | 34668231 | + | ENST00000241052 | CAT | chr11 | 34485652 | + | 1482 | 464 | 121 | 852 | 243 |
| ENST00000450654 | EHF | chr11 | 34668231 | + | ENST00000241052 | CAT | chr11 | 34485652 | + | 1458 | 440 | 97 | 828 | 243 |
| ENST00000530286 | EHF | chr11 | 34668231 | + | ENST00000241052 | CAT | chr11 | 34485652 | + | 1501 | 483 | 140 | 871 | 243 |
| ENST00000533754 | EHF | chr11 | 34668231 | + | ENST00000241052 | CAT | chr11 | 34485652 | + | 1578 | 560 | 208 | 948 | 246 |
| ENST00000531728 | EHF | chr11 | 34668231 | + | ENST00000241052 | CAT | chr11 | 34485652 | + | 1257 | 239 | 214 | 627 | 137 |
| ENST00000531794 | EHF | chr11 | 34668231 | + | ENST00000241052 | CAT | chr11 | 34485652 | + | 1553 | 535 | 6 | 923 | 305 |
| ENST00000527935 | EHF | chr11 | 34668231 | + | ENST00000241052 | CAT | chr11 | 34485652 | + | 1329 | 311 | 370 | 699 | 109 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000257831 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + | 0.002548717 | 0.99745125 |
| ENST00000450654 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + | 0.002579209 | 0.9974208 |
| ENST00000530286 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + | 0.002532126 | 0.9974679 |
| ENST00000533754 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + | 0.001630996 | 0.99836904 |
| ENST00000531728 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + | 0.00490205 | 0.99509794 |
| ENST00000531794 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + | 0.001380737 | 0.9986193 |
| ENST00000527935 | ENST00000241052 | EHF | chr11 | 34668231 | + | CAT | chr11 | 34485652 | + | 0.7868189 | 0.21318108 |
Top |
Fusion Genomic Features for EHF-CAT |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| EHF | chr11 | 34668231 | + | CAT | chr11 | 34485651 | + | 0.001270443 | 0.9987295 |
| EHF | chr11 | 34668231 | + | CAT | chr11 | 34485651 | + | 0.001270443 | 0.9987295 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
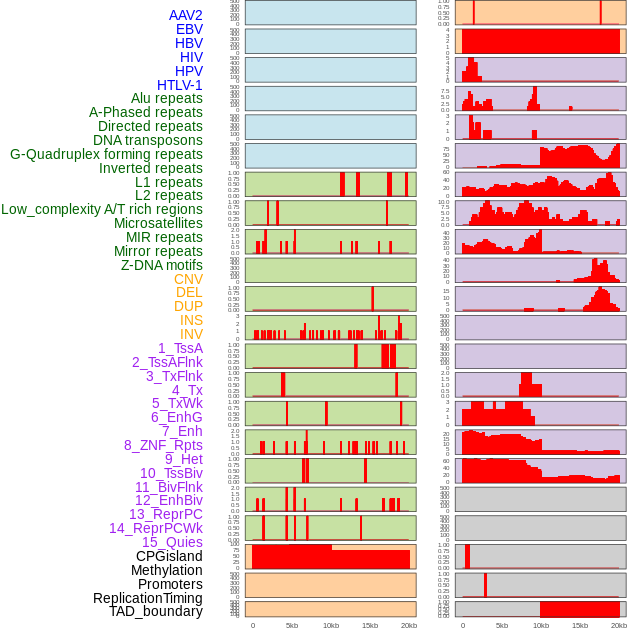 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
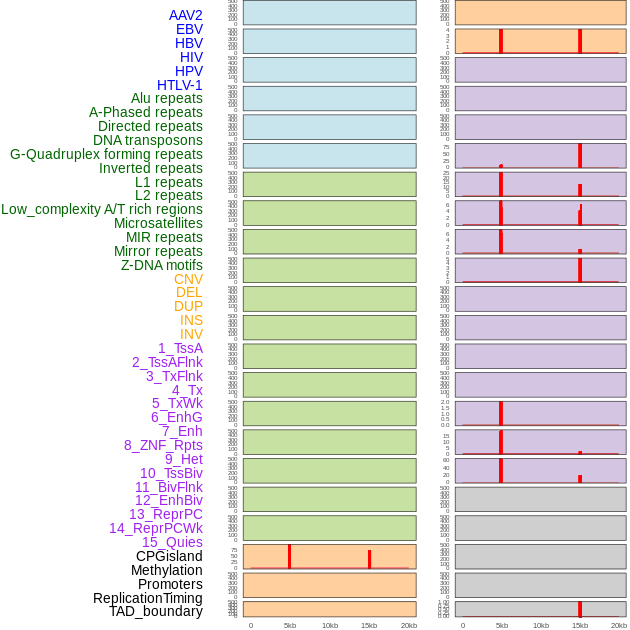 |
Top |
Fusion Protein Features for EHF-CAT |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr11:34668231/chr11:34485652) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| EHF | CAT |
| FUNCTION: Transcriptional activator that may play a role in regulating epithelial cell differentiation and proliferation. May act as a repressor for a specific subset of ETS/AP-1-responsive genes and as a modulator of the nuclear response to mitogen-activated protein kinase signaling cascades. Binds to DNA sequences containing the consensus nucleotide core sequence GGAA. Involved in regulation of TNFRSF10B/DR5 expression through Ets-binding sequences on the TNFRSF10B/DR5 promoter. May contribute to development and carcinogenesis by acting as a tumor suppressor gene or anti-oncogene. {ECO:0000269|PubMed:10527851, ECO:0000269|PubMed:10644770, ECO:0000269|PubMed:11259407, ECO:0000269|PubMed:12444029, ECO:0000269|PubMed:17027647}. | FUNCTION: Probably involved in sperm cell hyperactivation via its association with CATSPER1. Sperm cell hyperactivation is needed for sperm motility which is essential late in the preparation of sperm for fertilization. {ECO:0000250|UniProtKB:C6KI89}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000257831 | + | 3 | 9 | 29_115 | 114 | 301.0 | Domain | PNT |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000450654 | + | 3 | 8 | 29_115 | 114 | 278.0 | Domain | PNT |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000530286 | + | 3 | 9 | 29_115 | 114 | 301.0 | Domain | PNT |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000531794 | + | 3 | 9 | 29_115 | 136 | 323.0 | Domain | PNT |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000533754 | + | 3 | 9 | 29_115 | 114 | 301.0 | Domain | PNT |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000257831 | + | 3 | 9 | 207_289 | 114 | 301.0 | DNA binding | ETS |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000450654 | + | 3 | 8 | 207_289 | 114 | 278.0 | DNA binding | ETS |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000530286 | + | 3 | 9 | 207_289 | 114 | 301.0 | DNA binding | ETS |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000531794 | + | 3 | 9 | 207_289 | 136 | 323.0 | DNA binding | ETS |
| Hgene | EHF | chr11:34668231 | chr11:34485652 | ENST00000533754 | + | 3 | 9 | 207_289 | 114 | 301.0 | DNA binding | ETS |
Top |
Fusion Gene Sequence for EHF-CAT |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >25568_25568_1_EHF-CAT_EHF_chr11_34668231_ENST00000257831_CAT_chr11_34485652_ENST00000241052_length(transcript)=1482nt_BP=464nt ACTTGATAACACCCGTGGTGCCCCATCCCTATAGGAGCTGGTGAGATTGCAGCCTGCTGCCTCCCCTCCAACAGCCACAGCTATTGGATT TCCCACCCAGAATCTTTAGGTAAATGAGATCATGATTCTGGAAGGAGGTGGTGTAATGAATCTCAACCCCGGCAACAACCTCCTTCACCA GCCGCCAGCCTGGACAGACAGCTACTCCACGTGCAATGTTTCCAGTGGGTTTTTTGGAGGCCAGTGGCATGAAATTCATCCTCAGTACTG GACCAAGTACCAGGTGTGGGAGTGGCTCCAGCACCTCCTGGACACCAACCAGCTGGATGCCAATTGTATCCCTTTCCAAGAGTTCGACAT CAACGGCGAGCACCTCTGCAGCATGAGTTTGCAGGAGTTCACCCGGGCGGCAGGGACGGCGGGGCAGCTCCTCTACAGCAACTTGCAGCA TCTGAAGTGGAACGGTGGTGCTCCAAATTACTACCCCAACAGCTTTGGTGCTCCGGAACAACAGCCTTCTGCCCTGGAGCACAGCATCCA ATATTCTGGAGAAGTGCGGAGATTCAACACTGCCAATGATGATAACGTTACTCAGGTGCGGGCATTCTATGTGAACGTGCTGAATGAGGA ACAGAGGAAACGTCTGTGTGAGAACATTGCCGGCCACCTGAAGGATGCACAAATTTTCATCCAGAAGAAAGCGGTCAAGAACTTCACTGA GGTCCACCCTGACTACGGGAGCCACATCCAGGCTCTTCTGGACAAGTACAATGCTGAGAAGCCTAAGAATGCGATTCACACCTTTGTGCA GTCCGGATCTCACTTGGCGGCAAGGGAGAAGGCAAATCTGTGAGGCCGGGGCCCTGCACCTGTGCAGCGAAGCTTAGCGTTCATCCGTGT AACCCGCTCATCACTGGATGAAGATTCTCCTGTGCTAGATGTGCAAATGCAAGCTAGTGGCTTCAAAATAGAGAATCCCACTTTCTATAG CAGATTGTGTAACAATTTTAATGCTATTTCCCCAGGGGAAAATGAAGGTTAGGATTTAACAGTCATTTAAAAAAAAAATTTGTTTTGACG GATGATTGGATTATTCATTTAAAATGATTAGAAGGCAAGTTTCTAGCTAGAAATATGATTTTATTTGACAAAATTTGTTGAAATTATGTA TGTTTACATATCACCTCATGGCCTATTATATTAAAATATGGCTATAAATATATAAAAAGAAAAGATAAAGATGATCTACTCAGAAATTTT TATTTTTCTAAGGTTCTCATAGGAAAAGTACATTTAATACAGCAGTGTCATCAGAAGATAACTTGAGCACCGTCATGGCTTAATGTTTAT TCCTGATAATAATTGATCAAATTCATTTTTTTCACTGGAGTTACATTAATGTTAATTCAGCACTGATTTCACAACAGATCAATTTGTAAT >25568_25568_1_EHF-CAT_EHF_chr11_34668231_ENST00000257831_CAT_chr11_34485652_ENST00000241052_length(amino acids)=243AA_BP=114 MILEGGGVMNLNPGNNLLHQPPAWTDSYSTCNVSSGFFGGQWHEIHPQYWTKYQVWEWLQHLLDTNQLDANCIPFQEFDINGEHLCSMSL QEFTRAAGTAGQLLYSNLQHLKWNGGAPNYYPNSFGAPEQQPSALEHSIQYSGEVRRFNTANDDNVTQVRAFYVNVLNEEQRKRLCENIA -------------------------------------------------------------- >25568_25568_2_EHF-CAT_EHF_chr11_34668231_ENST00000450654_CAT_chr11_34485652_ENST00000241052_length(transcript)=1458nt_BP=440nt ATCCCTATAGGAGCTGGTGAGATTGCAGCCTGCTGCCTCCCCTCCAACAGCCACAGCTATTGGATTTCCCACCCAGAATCTTTAGGTAAA TGAGATCATGATTCTGGAAGGAGGTGGTGTAATGAATCTCAACCCCGGCAACAACCTCCTTCACCAGCCGCCAGCCTGGACAGACAGCTA CTCCACGTGCAATGTTTCCAGTGGGTTTTTTGGAGGCCAGTGGCATGAAATTCATCCTCAGTACTGGACCAAGTACCAGGTGTGGGAGTG GCTCCAGCACCTCCTGGACACCAACCAGCTGGATGCCAATTGTATCCCTTTCCAAGAGTTCGACATCAACGGCGAGCACCTCTGCAGCAT GAGTTTGCAGGAGTTCACCCGGGCGGCAGGGACGGCGGGGCAGCTCCTCTACAGCAACTTGCAGCATCTGAAGTGGAACGGTGGTGCTCC AAATTACTACCCCAACAGCTTTGGTGCTCCGGAACAACAGCCTTCTGCCCTGGAGCACAGCATCCAATATTCTGGAGAAGTGCGGAGATT CAACACTGCCAATGATGATAACGTTACTCAGGTGCGGGCATTCTATGTGAACGTGCTGAATGAGGAACAGAGGAAACGTCTGTGTGAGAA CATTGCCGGCCACCTGAAGGATGCACAAATTTTCATCCAGAAGAAAGCGGTCAAGAACTTCACTGAGGTCCACCCTGACTACGGGAGCCA CATCCAGGCTCTTCTGGACAAGTACAATGCTGAGAAGCCTAAGAATGCGATTCACACCTTTGTGCAGTCCGGATCTCACTTGGCGGCAAG GGAGAAGGCAAATCTGTGAGGCCGGGGCCCTGCACCTGTGCAGCGAAGCTTAGCGTTCATCCGTGTAACCCGCTCATCACTGGATGAAGA TTCTCCTGTGCTAGATGTGCAAATGCAAGCTAGTGGCTTCAAAATAGAGAATCCCACTTTCTATAGCAGATTGTGTAACAATTTTAATGC TATTTCCCCAGGGGAAAATGAAGGTTAGGATTTAACAGTCATTTAAAAAAAAAATTTGTTTTGACGGATGATTGGATTATTCATTTAAAA TGATTAGAAGGCAAGTTTCTAGCTAGAAATATGATTTTATTTGACAAAATTTGTTGAAATTATGTATGTTTACATATCACCTCATGGCCT ATTATATTAAAATATGGCTATAAATATATAAAAAGAAAAGATAAAGATGATCTACTCAGAAATTTTTATTTTTCTAAGGTTCTCATAGGA AAAGTACATTTAATACAGCAGTGTCATCAGAAGATAACTTGAGCACCGTCATGGCTTAATGTTTATTCCTGATAATAATTGATCAAATTC ATTTTTTTCACTGGAGTTACATTAATGTTAATTCAGCACTGATTTCACAACAGATCAATTTGTAATTGCTTACATTTTTACAATAAATAA >25568_25568_2_EHF-CAT_EHF_chr11_34668231_ENST00000450654_CAT_chr11_34485652_ENST00000241052_length(amino acids)=243AA_BP=114 MILEGGGVMNLNPGNNLLHQPPAWTDSYSTCNVSSGFFGGQWHEIHPQYWTKYQVWEWLQHLLDTNQLDANCIPFQEFDINGEHLCSMSL QEFTRAAGTAGQLLYSNLQHLKWNGGAPNYYPNSFGAPEQQPSALEHSIQYSGEVRRFNTANDDNVTQVRAFYVNVLNEEQRKRLCENIA -------------------------------------------------------------- >25568_25568_3_EHF-CAT_EHF_chr11_34668231_ENST00000527935_CAT_chr11_34485652_ENST00000241052_length(transcript)=1329nt_BP=311nt GATCATGATTCTGGAAGGAGGTGGTGTAATGAATCTCAACCCCGGCAACAACCTCCTTCACCAGCCGCCAGCCTGGACAGACAGCTACTC CACGTGCAATGGTGACTCTCTCTTTCTGTGTCTCTCCCTACCCTGCTAGGAGCCAGCTGGAAGAATAGAGATTCTGCAGATGACAGGATT CTTTGTCAGGGACAAGAAAGAGTGGTCTCACCTCCAAATTACCATGTCCCAAGACAGAGAAATGGAAGGAAGATGAAAGGACAGGATTGA AGCATAGGGTACCTGGTGTGCCTAATCCCTTATAAATGCCAGTGGTGCTCCAAATTACTACCCCAACAGCTTTGGTGCTCCGGAACAACA GCCTTCTGCCCTGGAGCACAGCATCCAATATTCTGGAGAAGTGCGGAGATTCAACACTGCCAATGATGATAACGTTACTCAGGTGCGGGC ATTCTATGTGAACGTGCTGAATGAGGAACAGAGGAAACGTCTGTGTGAGAACATTGCCGGCCACCTGAAGGATGCACAAATTTTCATCCA GAAGAAAGCGGTCAAGAACTTCACTGAGGTCCACCCTGACTACGGGAGCCACATCCAGGCTCTTCTGGACAAGTACAATGCTGAGAAGCC TAAGAATGCGATTCACACCTTTGTGCAGTCCGGATCTCACTTGGCGGCAAGGGAGAAGGCAAATCTGTGAGGCCGGGGCCCTGCACCTGT GCAGCGAAGCTTAGCGTTCATCCGTGTAACCCGCTCATCACTGGATGAAGATTCTCCTGTGCTAGATGTGCAAATGCAAGCTAGTGGCTT CAAAATAGAGAATCCCACTTTCTATAGCAGATTGTGTAACAATTTTAATGCTATTTCCCCAGGGGAAAATGAAGGTTAGGATTTAACAGT CATTTAAAAAAAAAATTTGTTTTGACGGATGATTGGATTATTCATTTAAAATGATTAGAAGGCAAGTTTCTAGCTAGAAATATGATTTTA TTTGACAAAATTTGTTGAAATTATGTATGTTTACATATCACCTCATGGCCTATTATATTAAAATATGGCTATAAATATATAAAAAGAAAA GATAAAGATGATCTACTCAGAAATTTTTATTTTTCTAAGGTTCTCATAGGAAAAGTACATTTAATACAGCAGTGTCATCAGAAGATAACT TGAGCACCGTCATGGCTTAATGTTTATTCCTGATAATAATTGATCAAATTCATTTTTTTCACTGGAGTTACATTAATGTTAATTCAGCAC >25568_25568_3_EHF-CAT_EHF_chr11_34668231_ENST00000527935_CAT_chr11_34485652_ENST00000241052_length(amino acids)=109AA_BP= MEHSIQYSGEVRRFNTANDDNVTQVRAFYVNVLNEEQRKRLCENIAGHLKDAQIFIQKKAVKNFTEVHPDYGSHIQALLDKYNAEKPKNA -------------------------------------------------------------- >25568_25568_4_EHF-CAT_EHF_chr11_34668231_ENST00000530286_CAT_chr11_34485652_ENST00000241052_length(transcript)=1501nt_BP=483nt CTCAGCAGCAACAAATGCCTTCTGAGACAGGGATTCTTTTGATTTTGCTTGGACATTGTGGAGAAGTGTTAGCCCCAATGTGGACTGATC TGGGAACAGTGGGAAATTAACTTCTTGTTGGTAAATATCAGGCTGAGATCATGATTCTGGAAGGAGGTGGTGTAATGAATCTCAACCCCG GCAACAACCTCCTTCACCAGCCGCCAGCCTGGACAGACAGCTACTCCACGTGCAATGTTTCCAGTGGGTTTTTTGGAGGCCAGTGGCATG AAATTCATCCTCAGTACTGGACCAAGTACCAGGTGTGGGAGTGGCTCCAGCACCTCCTGGACACCAACCAGCTGGATGCCAATTGTATCC CTTTCCAAGAGTTCGACATCAACGGCGAGCACCTCTGCAGCATGAGTTTGCAGGAGTTCACCCGGGCGGCAGGGACGGCGGGGCAGCTCC TCTACAGCAACTTGCAGCATCTGAAGTGGAACGGTGGTGCTCCAAATTACTACCCCAACAGCTTTGGTGCTCCGGAACAACAGCCTTCTG CCCTGGAGCACAGCATCCAATATTCTGGAGAAGTGCGGAGATTCAACACTGCCAATGATGATAACGTTACTCAGGTGCGGGCATTCTATG TGAACGTGCTGAATGAGGAACAGAGGAAACGTCTGTGTGAGAACATTGCCGGCCACCTGAAGGATGCACAAATTTTCATCCAGAAGAAAG CGGTCAAGAACTTCACTGAGGTCCACCCTGACTACGGGAGCCACATCCAGGCTCTTCTGGACAAGTACAATGCTGAGAAGCCTAAGAATG CGATTCACACCTTTGTGCAGTCCGGATCTCACTTGGCGGCAAGGGAGAAGGCAAATCTGTGAGGCCGGGGCCCTGCACCTGTGCAGCGAA GCTTAGCGTTCATCCGTGTAACCCGCTCATCACTGGATGAAGATTCTCCTGTGCTAGATGTGCAAATGCAAGCTAGTGGCTTCAAAATAG AGAATCCCACTTTCTATAGCAGATTGTGTAACAATTTTAATGCTATTTCCCCAGGGGAAAATGAAGGTTAGGATTTAACAGTCATTTAAA AAAAAAATTTGTTTTGACGGATGATTGGATTATTCATTTAAAATGATTAGAAGGCAAGTTTCTAGCTAGAAATATGATTTTATTTGACAA AATTTGTTGAAATTATGTATGTTTACATATCACCTCATGGCCTATTATATTAAAATATGGCTATAAATATATAAAAAGAAAAGATAAAGA TGATCTACTCAGAAATTTTTATTTTTCTAAGGTTCTCATAGGAAAAGTACATTTAATACAGCAGTGTCATCAGAAGATAACTTGAGCACC GTCATGGCTTAATGTTTATTCCTGATAATAATTGATCAAATTCATTTTTTTCACTGGAGTTACATTAATGTTAATTCAGCACTGATTTCA >25568_25568_4_EHF-CAT_EHF_chr11_34668231_ENST00000530286_CAT_chr11_34485652_ENST00000241052_length(amino acids)=243AA_BP=114 MILEGGGVMNLNPGNNLLHQPPAWTDSYSTCNVSSGFFGGQWHEIHPQYWTKYQVWEWLQHLLDTNQLDANCIPFQEFDINGEHLCSMSL QEFTRAAGTAGQLLYSNLQHLKWNGGAPNYYPNSFGAPEQQPSALEHSIQYSGEVRRFNTANDDNVTQVRAFYVNVLNEEQRKRLCENIA -------------------------------------------------------------- >25568_25568_5_EHF-CAT_EHF_chr11_34668231_ENST00000531728_CAT_chr11_34485652_ENST00000241052_length(transcript)=1257nt_BP=239nt CCCTGCCTTTGTTACTGGGAAACCGAACGCGGCGCTGTGGCTTTCAGCTTGGATCATGATTCTGGAAGGAGGTGGTGTAATGAATCTCAA CCCCGGCAACAACCTCCTTCACCAGCCGCCAGCCTGGACAGACAGCTACTCCACGTGCAATGGTGACTCTCTCTTTCTGTGTCTCTCCCT ACCCTGCTAGGAGCCAGCTGGAAGAATAGAGATTCTGCAGATGACAGGATTCTTTGTCAGTGGTGCTCCAAATTACTACCCCAACAGCTT TGGTGCTCCGGAACAACAGCCTTCTGCCCTGGAGCACAGCATCCAATATTCTGGAGAAGTGCGGAGATTCAACACTGCCAATGATGATAA CGTTACTCAGGTGCGGGCATTCTATGTGAACGTGCTGAATGAGGAACAGAGGAAACGTCTGTGTGAGAACATTGCCGGCCACCTGAAGGA TGCACAAATTTTCATCCAGAAGAAAGCGGTCAAGAACTTCACTGAGGTCCACCCTGACTACGGGAGCCACATCCAGGCTCTTCTGGACAA GTACAATGCTGAGAAGCCTAAGAATGCGATTCACACCTTTGTGCAGTCCGGATCTCACTTGGCGGCAAGGGAGAAGGCAAATCTGTGAGG CCGGGGCCCTGCACCTGTGCAGCGAAGCTTAGCGTTCATCCGTGTAACCCGCTCATCACTGGATGAAGATTCTCCTGTGCTAGATGTGCA AATGCAAGCTAGTGGCTTCAAAATAGAGAATCCCACTTTCTATAGCAGATTGTGTAACAATTTTAATGCTATTTCCCCAGGGGAAAATGA AGGTTAGGATTTAACAGTCATTTAAAAAAAAAATTTGTTTTGACGGATGATTGGATTATTCATTTAAAATGATTAGAAGGCAAGTTTCTA GCTAGAAATATGATTTTATTTGACAAAATTTGTTGAAATTATGTATGTTTACATATCACCTCATGGCCTATTATATTAAAATATGGCTAT AAATATATAAAAAGAAAAGATAAAGATGATCTACTCAGAAATTTTTATTTTTCTAAGGTTCTCATAGGAAAAGTACATTTAATACAGCAG TGTCATCAGAAGATAACTTGAGCACCGTCATGGCTTAATGTTTATTCCTGATAATAATTGATCAAATTCATTTTTTTCACTGGAGTTACA >25568_25568_5_EHF-CAT_EHF_chr11_34668231_ENST00000531728_CAT_chr11_34485652_ENST00000241052_length(amino acids)=137AA_BP=9 MQMTGFFVSGAPNYYPNSFGAPEQQPSALEHSIQYSGEVRRFNTANDDNVTQVRAFYVNVLNEEQRKRLCENIAGHLKDAQIFIQKKAVK -------------------------------------------------------------- >25568_25568_6_EHF-CAT_EHF_chr11_34668231_ENST00000531794_CAT_chr11_34485652_ENST00000241052_length(transcript)=1553nt_BP=535nt AACCCACTGCTTTATTCTGCCCTGAGTGGAGATTGGTTTTGGCTCAGGCTGCTTTGTGAAACTCAGAAGCATTATCCTCTCTGCCAACTC CACGTCCTAGTCAGAGTTTTCTGTGAAGGCAAGGGCATGGGGTTGCCGGAGAGAAGAGGATTGGTCCTGCTTTTAAGCCTAGCTGAAATT CTTTTCAAGATCATGATTCTGGAAGGAGGTGGTGTAATGAATCTCAACCCCGGCAACAACCTCCTTCACCAGCCGCCAGCCTGGACAGAC AGCTACTCCACGTGCAATGTTTCCAGTGGGTTTTTTGGAGGCCAGTGGCATGAAATTCATCCTCAGTACTGGACCAAGTACCAGGTGTGG GAGTGGCTCCAGCACCTCCTGGACACCAACCAGCTGGATGCCAATTGTATCCCTTTCCAAGAGTTCGACATCAACGGCGAGCACCTCTGC AGCATGAGTTTGCAGGAGTTCACCCGGGCGGCAGGGACGGCGGGGCAGCTCCTCTACAGCAACTTGCAGCATCTGAAGTGGAACGGTGGT GCTCCAAATTACTACCCCAACAGCTTTGGTGCTCCGGAACAACAGCCTTCTGCCCTGGAGCACAGCATCCAATATTCTGGAGAAGTGCGG AGATTCAACACTGCCAATGATGATAACGTTACTCAGGTGCGGGCATTCTATGTGAACGTGCTGAATGAGGAACAGAGGAAACGTCTGTGT GAGAACATTGCCGGCCACCTGAAGGATGCACAAATTTTCATCCAGAAGAAAGCGGTCAAGAACTTCACTGAGGTCCACCCTGACTACGGG AGCCACATCCAGGCTCTTCTGGACAAGTACAATGCTGAGAAGCCTAAGAATGCGATTCACACCTTTGTGCAGTCCGGATCTCACTTGGCG GCAAGGGAGAAGGCAAATCTGTGAGGCCGGGGCCCTGCACCTGTGCAGCGAAGCTTAGCGTTCATCCGTGTAACCCGCTCATCACTGGAT GAAGATTCTCCTGTGCTAGATGTGCAAATGCAAGCTAGTGGCTTCAAAATAGAGAATCCCACTTTCTATAGCAGATTGTGTAACAATTTT AATGCTATTTCCCCAGGGGAAAATGAAGGTTAGGATTTAACAGTCATTTAAAAAAAAAATTTGTTTTGACGGATGATTGGATTATTCATT TAAAATGATTAGAAGGCAAGTTTCTAGCTAGAAATATGATTTTATTTGACAAAATTTGTTGAAATTATGTATGTTTACATATCACCTCAT GGCCTATTATATTAAAATATGGCTATAAATATATAAAAAGAAAAGATAAAGATGATCTACTCAGAAATTTTTATTTTTCTAAGGTTCTCA TAGGAAAAGTACATTTAATACAGCAGTGTCATCAGAAGATAACTTGAGCACCGTCATGGCTTAATGTTTATTCCTGATAATAATTGATCA AATTCATTTTTTTCACTGGAGTTACATTAATGTTAATTCAGCACTGATTTCACAACAGATCAATTTGTAATTGCTTACATTTTTACAATA >25568_25568_6_EHF-CAT_EHF_chr11_34668231_ENST00000531794_CAT_chr11_34485652_ENST00000241052_length(amino acids)=305AA_BP=176 MLYSALSGDWFWLRLLCETQKHYPLCQLHVLVRVFCEGKGMGLPERRGLVLLLSLAEILFKIMILEGGGVMNLNPGNNLLHQPPAWTDSY STCNVSSGFFGGQWHEIHPQYWTKYQVWEWLQHLLDTNQLDANCIPFQEFDINGEHLCSMSLQEFTRAAGTAGQLLYSNLQHLKWNGGAP NYYPNSFGAPEQQPSALEHSIQYSGEVRRFNTANDDNVTQVRAFYVNVLNEEQRKRLCENIAGHLKDAQIFIQKKAVKNFTEVHPDYGSH -------------------------------------------------------------- >25568_25568_7_EHF-CAT_EHF_chr11_34668231_ENST00000533754_CAT_chr11_34485652_ENST00000241052_length(transcript)=1578nt_BP=560nt ATTTTCACCGTCCATCTTTGCTGATTTACCATGCTCCCAGGATGGTGGGAGTGTGTGTTTTTAAGATGGAGAGTGTATGCTTCTGGGTTC AAGTTCACAGGTGTCTCTGCTGGTTATCTGCACTCACCTTGGTAACAGGGAGAAAGTGAGTGAATGGATTCCAAGAACTTACTGATGGAA GTCTAATTCAGGAGTTGGTTCTGCAGCCATGGAGATCATGATTCTGGAAGGAGGTGGTGTAATGAATCTCAACCCCGGCAACAACCTCCT TCACCAGCCGCCAGCCTGGACAGACAGCTACTCCACGTGCAATGTTTCCAGTGGGTTTTTTGGAGGCCAGTGGCATGAAATTCATCCTCA GTACTGGACCAAGTACCAGGTGTGGGAGTGGCTCCAGCACCTCCTGGACACCAACCAGCTGGATGCCAATTGTATCCCTTTCCAAGAGTT CGACATCAACGGCGAGCACCTCTGCAGCATGAGTTTGCAGGAGTTCACCCGGGCGGCAGGGACGGCGGGGCAGCTCCTCTACAGCAACTT GCAGCATCTGAAGTGGAACGGTGGTGCTCCAAATTACTACCCCAACAGCTTTGGTGCTCCGGAACAACAGCCTTCTGCCCTGGAGCACAG CATCCAATATTCTGGAGAAGTGCGGAGATTCAACACTGCCAATGATGATAACGTTACTCAGGTGCGGGCATTCTATGTGAACGTGCTGAA TGAGGAACAGAGGAAACGTCTGTGTGAGAACATTGCCGGCCACCTGAAGGATGCACAAATTTTCATCCAGAAGAAAGCGGTCAAGAACTT CACTGAGGTCCACCCTGACTACGGGAGCCACATCCAGGCTCTTCTGGACAAGTACAATGCTGAGAAGCCTAAGAATGCGATTCACACCTT TGTGCAGTCCGGATCTCACTTGGCGGCAAGGGAGAAGGCAAATCTGTGAGGCCGGGGCCCTGCACCTGTGCAGCGAAGCTTAGCGTTCAT CCGTGTAACCCGCTCATCACTGGATGAAGATTCTCCTGTGCTAGATGTGCAAATGCAAGCTAGTGGCTTCAAAATAGAGAATCCCACTTT CTATAGCAGATTGTGTAACAATTTTAATGCTATTTCCCCAGGGGAAAATGAAGGTTAGGATTTAACAGTCATTTAAAAAAAAAATTTGTT TTGACGGATGATTGGATTATTCATTTAAAATGATTAGAAGGCAAGTTTCTAGCTAGAAATATGATTTTATTTGACAAAATTTGTTGAAAT TATGTATGTTTACATATCACCTCATGGCCTATTATATTAAAATATGGCTATAAATATATAAAAAGAAAAGATAAAGATGATCTACTCAGA AATTTTTATTTTTCTAAGGTTCTCATAGGAAAAGTACATTTAATACAGCAGTGTCATCAGAAGATAACTTGAGCACCGTCATGGCTTAAT GTTTATTCCTGATAATAATTGATCAAATTCATTTTTTTCACTGGAGTTACATTAATGTTAATTCAGCACTGATTTCACAACAGATCAATT >25568_25568_7_EHF-CAT_EHF_chr11_34668231_ENST00000533754_CAT_chr11_34485652_ENST00000241052_length(amino acids)=246AA_BP=117 MEIMILEGGGVMNLNPGNNLLHQPPAWTDSYSTCNVSSGFFGGQWHEIHPQYWTKYQVWEWLQHLLDTNQLDANCIPFQEFDINGEHLCS MSLQEFTRAAGTAGQLLYSNLQHLKWNGGAPNYYPNSFGAPEQQPSALEHSIQYSGEVRRFNTANDDNVTQVRAFYVNVLNEEQRKRLCE -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for EHF-CAT |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for EHF-CAT |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for EHF-CAT |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
