
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ELANE-BSG (FusionGDB2 ID:26129) |
Fusion Gene Summary for ELANE-BSG |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ELANE-BSG | Fusion gene ID: 26129 | Hgene | Tgene | Gene symbol | ELANE | BSG | Gene ID | 1991 | 682 |
| Gene name | elastase, neutrophil expressed | basigin (Ok blood group) | |
| Synonyms | ELA2|GE|HLE|HNE|NE|PMN-E|SCN1 | 5F7|CD147|EMMPRIN|EMPRIN|OK|SLC7A11|TCSF | |
| Cytomap | 19p13.3 | 19p13.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | neutrophil elastasePMN elastasebone marrow serine proteaseelastase 2, neutrophilelastase-2granulocyte-derived elastaseleukocyte elastasemedullasinpolymorphonuclear elastase | basiginOK blood group antigencollagenase stimulatory factorextracellular matrix metalloproteinase inducerleukocyte activation antigen M6tumor cell-derived collagenase stimulatory factor | |
| Modification date | 20200313 | 20200315 | |
| UniProtAcc | P08246 | P35613 | |
| Ensembl transtripts involved in fusion gene | ENST00000263621, ENST00000590230, | ENST00000574970, ENST00000333511, ENST00000353555, ENST00000545507, ENST00000346916, | |
| Fusion gene scores | * DoF score | 4 X 2 X 2=16 | 9 X 12 X 6=648 |
| # samples | 4 | 14 | |
| ** MAII score | log2(4/16*10)=1.32192809488736 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(14/648*10)=-2.21056698593966 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: ELANE [Title/Abstract] AND BSG [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ELANE(852395)-BSG(579500), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | ELANE | GO:0002812 | biosynthetic process of antibacterial peptides active against Gram-negative bacteria | 20421939 |
| Hgene | ELANE | GO:0006508 | proteolysis | 11907569|20421939 |
| Hgene | ELANE | GO:0009411 | response to UV | 11928814 |
| Hgene | ELANE | GO:0042742 | defense response to bacterium | 14705961 |
| Hgene | ELANE | GO:0045079 | negative regulation of chemokine biosynthetic process | 12223522 |
| Hgene | ELANE | GO:0045415 | negative regulation of interleukin-8 biosynthetic process | 12223522 |
| Hgene | ELANE | GO:0045416 | positive regulation of interleukin-8 biosynthetic process | 14730209 |
| Hgene | ELANE | GO:0048661 | positive regulation of smooth muscle cell proliferation | 15010259 |
| Hgene | ELANE | GO:0070945 | neutrophil mediated killing of gram-negative bacterium | 20421939 |
 Fusion gene breakpoints across ELANE (5'-gene) Fusion gene breakpoints across ELANE (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Fusion gene breakpoints across BSG (3'-gene) Fusion gene breakpoints across BSG (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | ERR315396 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
Top |
Fusion Gene ORF analysis for ELANE-BSG |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000263621 | ENST00000574970 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
| 5CDS-3UTR | ENST00000590230 | ENST00000574970 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
| 5CDS-5UTR | ENST00000263621 | ENST00000333511 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
| 5CDS-5UTR | ENST00000263621 | ENST00000353555 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
| 5CDS-5UTR | ENST00000263621 | ENST00000545507 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
| 5CDS-5UTR | ENST00000590230 | ENST00000333511 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
| 5CDS-5UTR | ENST00000590230 | ENST00000353555 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
| 5CDS-5UTR | ENST00000590230 | ENST00000545507 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
| 5CDS-intron | ENST00000263621 | ENST00000346916 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
| 5CDS-intron | ENST00000590230 | ENST00000346916 | ELANE | chr19 | 852395 | + | BSG | chr19 | 579500 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
Top |
Fusion Genomic Features for ELANE-BSG |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| ELANE | chr19 | 852395 | + | BSG | chr19 | 579499 | + | 0.9996419 | 0.000358115 |
| ELANE | chr19 | 852395 | + | BSG | chr19 | 579499 | + | 0.9996419 | 0.000358115 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
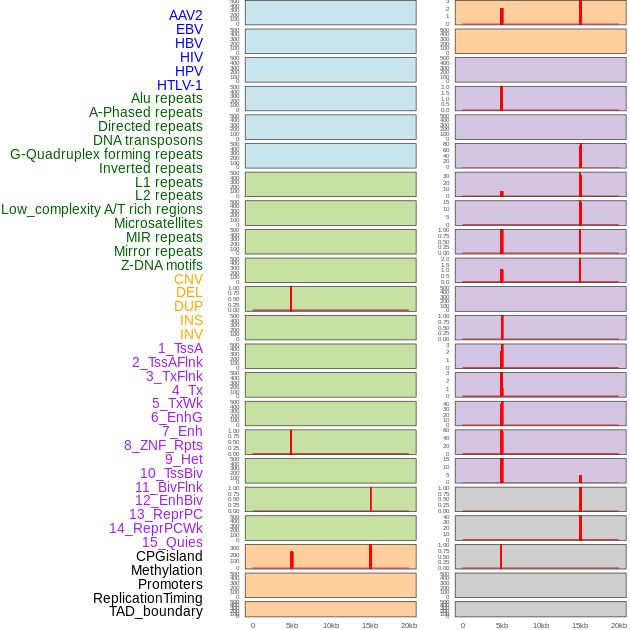 |
Top |
Fusion Protein Features for ELANE-BSG |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/:852395/:579500) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| ELANE | BSG |
| FUNCTION: Modifies the functions of natural killer cells, monocytes and granulocytes. Inhibits C5a-dependent neutrophil enzyme release and chemotaxis (PubMed:15140022). Capable of killing E.coli but not S.aureus in vitro; digests outer membrane protein A (ompA) in E.coli and K.pneumoniae (PubMed:10947984). {ECO:0000269|PubMed:10947984, ECO:0000269|PubMed:15140022}. | FUNCTION: [Isoform 1]: Essential for normal retinal maturation and development (By similarity). Acts as a retinal cell surface receptor for NXNL1 and plays an important role in NXNL1-mediated survival of retinal cone photoreceptors (PubMed:25957687). In association with glucose transporter SLC16A1/GLUT1 and NXNL1, promotes retinal cone survival by enhancing aerobic glycolysis and accelerating the entry of glucose into photoreceptors (PubMed:25957687). May act as a potent stimulator of IL6 secretion in multiple cell lines that include monocytes (PubMed:21620857). {ECO:0000250|UniProtKB:P18572, ECO:0000269|PubMed:21620857, ECO:0000269|PubMed:25957687}.; FUNCTION: [Isoform 2]: Signaling receptor for cyclophilins, essential for PPIA/CYPA and PPIB/CYPB-dependent signaling related to chemotaxis and adhesion of immune cells (PubMed:11943775, PubMed:11688976). Plays an important role in targeting monocarboxylate transporters SLC16A1/GLUT1, SLC16A11 and SLC16A12 to the plasma membrane (PubMed:17127621, PubMed:21778275, PubMed:28666119). Acts as a coreceptor for vascular endothelial growth factor receptor 2 (KDR/VEGFR2) in endothelial cells enhancing its VEGFA-mediated activation and downstream signaling (PubMed:25825981). Promotes angiogenesis through EPAS1/HIF2A-mediated up-regulation of VEGFA (isoform VEGF-165 and VEGF-121) and KDR/VEGFR2 in endothelial cells (PubMed:19837976). Plays a key role in regulating tumor growth, invasion, metastasis and neoangiogenesis by stimulating the production and release of extracellular matrix metalloproteinases and KDR/VEGFR2 by both tumor cells and stromal cells (fibroblasts and endothelial cells) (PubMed:12553375, PubMed:11992541, PubMed:15833850). {ECO:0000269|PubMed:11688976, ECO:0000269|PubMed:11943775, ECO:0000269|PubMed:11992541, ECO:0000269|PubMed:12553375, ECO:0000269|PubMed:15833850, ECO:0000269|PubMed:17127621, ECO:0000269|PubMed:19837976, ECO:0000269|PubMed:21778275, ECO:0000269|PubMed:25825981, ECO:0000269|PubMed:28666119}.; FUNCTION: [Isoform 1]: (Microbial infection) Erythrocyte receptor for P.falciparum RH5 which is essential for erythrocyte invasion by the merozoite stage of P.falciparum isolates 3D7 and Dd2. {ECO:0000269|PubMed:22080952}.; FUNCTION: [Isoform 2]: (Microbial infection) Erythrocyte receptor for P.falciparum RH5 which is essential for erythrocyte invasion by the merozoite stage of P.falciparum isolates 3D7, Dd2, 7G8 and HB3 (PubMed:22080952, PubMed:26195724). Binding of P.falciparum RH5 results in BSG dimerization which triggers an increase in intracellular Ca(2+) in the erythrocyte (PubMed:28409866). This essential step leads to a rearrangement of the erythrocyte cytoskeleton required for the merozoite invasion (PubMed:28409866). {ECO:0000269|PubMed:22080952, ECO:0000269|PubMed:26195724, ECO:0000269|PubMed:28409866}.; FUNCTION: [Isoform 2]: (Microbial infection) Can facilitate human SARS coronavirus (SARS-CoV-1) infection via its interaction with virus-associated PPIA/CYPA. {ECO:0000269|PubMed:15688292}.; FUNCTION: [Isoform 2]: (Microbial infection) Can facilitate HIV-1 infection via its interaction with virus-associated PPIA/CYPA. {ECO:0000269|PubMed:11353871}.; FUNCTION: [Isoform 2]: (Microbial infection) First described as a receptor for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), it is not required for SARS-CoV-2 infection. {ECO:0000269|PubMed:33432067, ECO:0000303|PubMed:32307653}.; FUNCTION: [Isoform 2]: (Microbial infection) Acts as a receptor for measles virus. {ECO:0000269|PubMed:20147391}.; FUNCTION: [Isoform 2]: (Microbial infection) Promotes entry of pentamer-expressing human cytomegalovirus (HCMV) into epithelial and endothelial cells. {ECO:0000269|PubMed:29739904}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Gene Sequence for ELANE-BSG |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
Top |
Fusion Gene PPI Analysis for ELANE-BSG |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ELANE-BSG |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ELANE-BSG |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
