
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:EPS15L1-EPN1 (FusionGDB2 ID:27073) |
Fusion Gene Summary for EPS15L1-EPN1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: EPS15L1-EPN1 | Fusion gene ID: 27073 | Hgene | Tgene | Gene symbol | EPS15L1 | EPN1 | Gene ID | 58513 | 29924 |
| Gene name | epidermal growth factor receptor pathway substrate 15 like 1 | epsin 1 | |
| Synonyms | EPS15R | - | |
| Cytomap | 19p13.11 | 19q13.42 | |
| Type of gene | protein-coding | protein-coding | |
| Description | epidermal growth factor receptor substrate 15-like 1epidermal growth factor receptor substrate EPS15Reps15-related protein | epsin-1EH domain-binding mitotic phosphoproteinEPS-15-interacting protein 1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q9UBC2 | Q9Y6I3 | |
| Ensembl transtripts involved in fusion gene | ENST00000248070, ENST00000455140, ENST00000535753, ENST00000594975, ENST00000597937, ENST00000602009, | ENST00000591743, ENST00000085079, ENST00000270460, ENST00000411543, | |
| Fusion gene scores | * DoF score | 23 X 17 X 14=5474 | 7 X 5 X 6=210 |
| # samples | 34 | 8 | |
| ** MAII score | log2(34/5474*10)=-4.00898878322726 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(8/210*10)=-1.39231742277876 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: EPS15L1 [Title/Abstract] AND EPN1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | EPS15L1(16582723)-EPN1(56196761), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across EPS15L1 (5'-gene) Fusion gene breakpoints across EPS15L1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
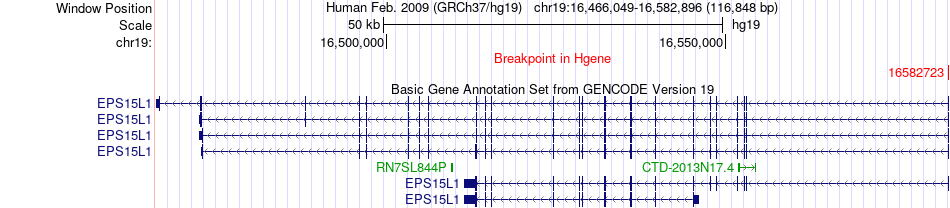 |
 Fusion gene breakpoints across EPN1 (3'-gene) Fusion gene breakpoints across EPN1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
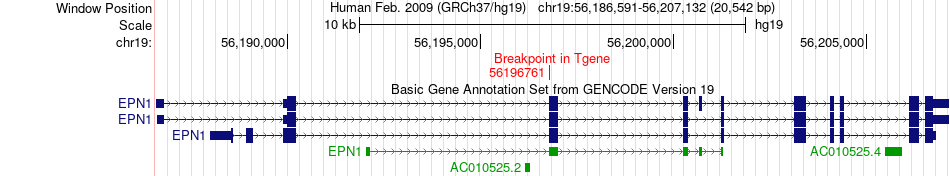 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | 5381N | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
Top |
Fusion Gene ORF analysis for EPS15L1-EPN1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000248070 | ENST00000591743 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| 5CDS-3UTR | ENST00000455140 | ENST00000591743 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| 5CDS-3UTR | ENST00000535753 | ENST00000591743 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| 5CDS-3UTR | ENST00000594975 | ENST00000591743 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| 5CDS-3UTR | ENST00000597937 | ENST00000591743 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000248070 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000248070 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000248070 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000455140 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000455140 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000455140 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000535753 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000535753 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000535753 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000594975 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000594975 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000594975 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000597937 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000597937 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| In-frame | ENST00000597937 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| intron-3CDS | ENST00000602009 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| intron-3CDS | ENST00000602009 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| intron-3CDS | ENST00000602009 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
| intron-3UTR | ENST00000602009 | ENST00000591743 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000455140 | EPS15L1 | chr19 | 16582723 | - | ENST00000270460 | EPN1 | chr19 | 56196761 | + | 2013 | 100 | 1892 | 0 | 631 |
| ENST00000455140 | EPS15L1 | chr19 | 16582723 | - | ENST00000085079 | EPN1 | chr19 | 56196761 | + | 1935 | 100 | 1814 | 0 | 605 |
| ENST00000455140 | EPS15L1 | chr19 | 16582723 | - | ENST00000411543 | EPN1 | chr19 | 56196761 | + | 1613 | 100 | 67 | 1527 | 486 |
| ENST00000248070 | EPS15L1 | chr19 | 16582723 | - | ENST00000270460 | EPN1 | chr19 | 56196761 | + | 2086 | 173 | 1965 | 16 | 649 |
| ENST00000248070 | EPS15L1 | chr19 | 16582723 | - | ENST00000085079 | EPN1 | chr19 | 56196761 | + | 2008 | 173 | 1887 | 16 | 623 |
| ENST00000248070 | EPS15L1 | chr19 | 16582723 | - | ENST00000411543 | EPN1 | chr19 | 56196761 | + | 1686 | 173 | 2 | 1600 | 532 |
| ENST00000535753 | EPS15L1 | chr19 | 16582723 | - | ENST00000270460 | EPN1 | chr19 | 56196761 | + | 1977 | 64 | 1856 | 0 | 619 |
| ENST00000535753 | EPS15L1 | chr19 | 16582723 | - | ENST00000085079 | EPN1 | chr19 | 56196761 | + | 1899 | 64 | 1778 | 0 | 593 |
| ENST00000535753 | EPS15L1 | chr19 | 16582723 | - | ENST00000411543 | EPN1 | chr19 | 56196761 | + | 1577 | 64 | 31 | 1491 | 486 |
| ENST00000594975 | EPS15L1 | chr19 | 16582723 | - | ENST00000270460 | EPN1 | chr19 | 56196761 | + | 2086 | 173 | 1965 | 16 | 649 |
| ENST00000594975 | EPS15L1 | chr19 | 16582723 | - | ENST00000085079 | EPN1 | chr19 | 56196761 | + | 2008 | 173 | 1887 | 16 | 623 |
| ENST00000594975 | EPS15L1 | chr19 | 16582723 | - | ENST00000411543 | EPN1 | chr19 | 56196761 | + | 1686 | 173 | 2 | 1600 | 532 |
| ENST00000597937 | EPS15L1 | chr19 | 16582723 | - | ENST00000270460 | EPN1 | chr19 | 56196761 | + | 1951 | 38 | 1830 | 1 | 610 |
| ENST00000597937 | EPS15L1 | chr19 | 16582723 | - | ENST00000085079 | EPN1 | chr19 | 56196761 | + | 1873 | 38 | 1752 | 1 | 584 |
| ENST00000597937 | EPS15L1 | chr19 | 16582723 | - | ENST00000411543 | EPN1 | chr19 | 56196761 | + | 1551 | 38 | 5 | 1465 | 486 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000455140 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.028425375 | 0.9715746 |
| ENST00000455140 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.030792067 | 0.96920794 |
| ENST00000455140 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.03451399 | 0.96548605 |
| ENST00000248070 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.083810456 | 0.9161896 |
| ENST00000248070 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.09223316 | 0.9077668 |
| ENST00000248070 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.0725507 | 0.9274493 |
| ENST00000535753 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.027995424 | 0.97200453 |
| ENST00000535753 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.029482013 | 0.970518 |
| ENST00000535753 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.03374081 | 0.9662592 |
| ENST00000594975 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.083810456 | 0.9161896 |
| ENST00000594975 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.09223316 | 0.9077668 |
| ENST00000594975 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.0725507 | 0.9274493 |
| ENST00000597937 | ENST00000270460 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.028119126 | 0.97188085 |
| ENST00000597937 | ENST00000085079 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.0296521 | 0.9703479 |
| ENST00000597937 | ENST00000411543 | EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 0.03383007 | 0.9661699 |
Top |
Fusion Genomic Features for EPS15L1-EPN1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 1.48E-13 | 1 |
| EPS15L1 | chr19 | 16582723 | - | EPN1 | chr19 | 56196761 | + | 1.48E-13 | 1 |
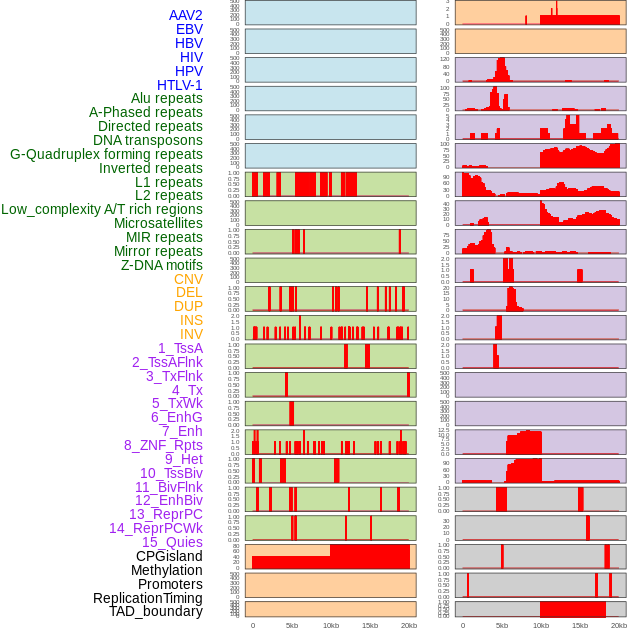
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
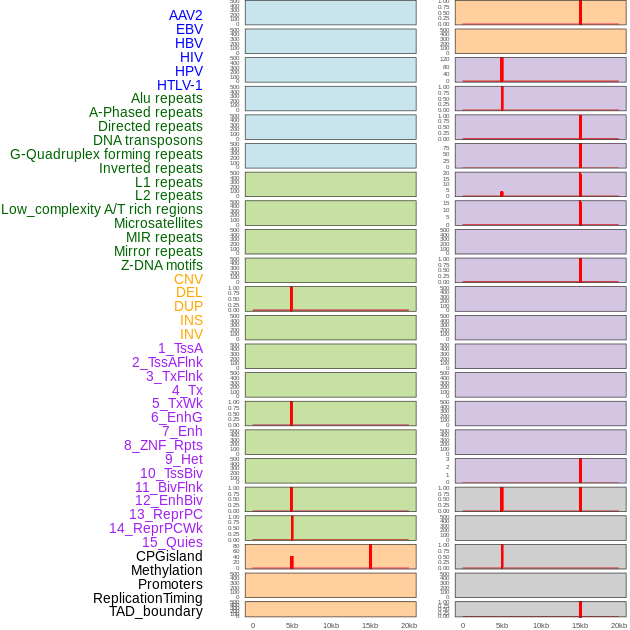
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for EPS15L1-EPN1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:16582723/chr19:56196761) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| EPS15L1 | EPN1 |
| FUNCTION: Seems to be a constitutive component of clathrin-coated pits that is required for receptor-mediated endocytosis. Involved in endocytosis of integrin beta-1 (ITGB1) and transferrin receptor (TFR); internalization of ITGB1 as DAB2-dependent cargo but not TFR seems to require association with DAB2. {ECO:0000269|PubMed:22648170, ECO:0000269|PubMed:9407958}. | FUNCTION: Binds to membranes enriched in phosphatidylinositol 4,5-bisphosphate (PtdIns(4,5)P2). Modifies membrane curvature and facilitates the formation of clathrin-coated invaginations (By similarity). Regulates receptor-mediated endocytosis. {ECO:0000250, ECO:0000269|PubMed:10557078}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 267_573 | 76 | 551.0 | Compositional bias | Note=Ala/Gly/Pro-rich | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 267_573 | 76 | 577.0 | Compositional bias | Note=Ala/Gly/Pro-rich | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 267_573 | 187 | 663.0 | Compositional bias | Note=Ala/Gly/Pro-rich | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 183_202 | 76 | 551.0 | Domain | UIM 1 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 208_227 | 76 | 551.0 | Domain | UIM 2 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 233_252 | 76 | 551.0 | Domain | UIM 3 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 183_202 | 76 | 577.0 | Domain | UIM 1 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 208_227 | 76 | 577.0 | Domain | UIM 2 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 233_252 | 76 | 577.0 | Domain | UIM 3 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 208_227 | 187 | 663.0 | Domain | UIM 2 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 233_252 | 187 | 663.0 | Domain | UIM 3 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 402_411 | 76 | 551.0 | Motif | Note=[DE]-X(1%2C2)-F-X-X-[FL]-X-X-X-R motif | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 402_411 | 76 | 577.0 | Motif | Note=[DE]-X(1%2C2)-F-X-X-[FL]-X-X-X-R motif | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 402_411 | 187 | 663.0 | Motif | Note=[DE]-X(1%2C2)-F-X-X-[FL]-X-X-X-R motif | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 274_379 | 76 | 551.0 | Region | Note=8 X 3 AA repeats of [ED]-P-W | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 502_574 | 76 | 551.0 | Region | Note=3 X 3 AA repeats of N-P-F | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 274_379 | 76 | 577.0 | Region | Note=8 X 3 AA repeats of [ED]-P-W | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 502_574 | 76 | 577.0 | Region | Note=3 X 3 AA repeats of N-P-F | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 274_379 | 187 | 663.0 | Region | Note=8 X 3 AA repeats of [ED]-P-W | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 502_574 | 187 | 663.0 | Region | Note=3 X 3 AA repeats of N-P-F | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 274_276 | 76 | 551.0 | Repeat | Note=1 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 294_296 | 76 | 551.0 | Repeat | Note=2 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 306_308 | 76 | 551.0 | Repeat | Note=3 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 319_321 | 76 | 551.0 | Repeat | Note=4 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 332_334 | 76 | 551.0 | Repeat | Note=5 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 349_351 | 76 | 551.0 | Repeat | Note=6 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 367_369 | 76 | 551.0 | Repeat | Note=7 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 377_379 | 76 | 551.0 | Repeat | Note=8 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 502_504 | 76 | 551.0 | Repeat | Note=1 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 518_520 | 76 | 551.0 | Repeat | Note=2 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 572_574 | 76 | 551.0 | Repeat | Note=3 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 274_276 | 76 | 577.0 | Repeat | Note=1 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 294_296 | 76 | 577.0 | Repeat | Note=2 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 306_308 | 76 | 577.0 | Repeat | Note=3 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 319_321 | 76 | 577.0 | Repeat | Note=4 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 332_334 | 76 | 577.0 | Repeat | Note=5 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 349_351 | 76 | 577.0 | Repeat | Note=6 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 367_369 | 76 | 577.0 | Repeat | Note=7 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 377_379 | 76 | 577.0 | Repeat | Note=8 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 502_504 | 76 | 577.0 | Repeat | Note=1 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 518_520 | 76 | 577.0 | Repeat | Note=2 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 572_574 | 76 | 577.0 | Repeat | Note=3 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 274_276 | 187 | 663.0 | Repeat | Note=1 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 294_296 | 187 | 663.0 | Repeat | Note=2 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 306_308 | 187 | 663.0 | Repeat | Note=3 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 319_321 | 187 | 663.0 | Repeat | Note=4 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 332_334 | 187 | 663.0 | Repeat | Note=5 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 349_351 | 187 | 663.0 | Repeat | Note=6 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 367_369 | 187 | 663.0 | Repeat | Note=7 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 377_379 | 187 | 663.0 | Repeat | Note=8 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 502_504 | 187 | 663.0 | Repeat | Note=1 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 518_520 | 187 | 663.0 | Repeat | Note=2 | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 572_574 | 187 | 663.0 | Repeat | Note=3 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 172_184 | 11 | 865.0 | Calcium binding | Ontology_term=ECO:0000255 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 172_184 | 11 | 911.0 | Calcium binding | Ontology_term=ECO:0000255 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 172_184 | 11 | 755.0 | Calcium binding | Ontology_term=ECO:0000255 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 172_184 | 11 | 602.0 | Calcium binding | Ontology_term=ECO:0000255 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 386_553 | 11 | 865.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 386_553 | 11 | 911.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 386_553 | 11 | 755.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 386_553 | 11 | 602.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 750_816 | 11 | 865.0 | Compositional bias | Note=Pro-rich |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 750_816 | 11 | 911.0 | Compositional bias | Note=Pro-rich |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 750_816 | 11 | 755.0 | Compositional bias | Note=Pro-rich |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 750_816 | 11 | 602.0 | Compositional bias | Note=Pro-rich |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 127_215 | 11 | 865.0 | Domain | EH 2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 159_194 | 11 | 865.0 | Domain | EF-hand |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 15_104 | 11 | 865.0 | Domain | EH 1 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 275_365 | 11 | 865.0 | Domain | EH 3 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 127_215 | 11 | 911.0 | Domain | EH 2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 159_194 | 11 | 911.0 | Domain | EF-hand |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 15_104 | 11 | 911.0 | Domain | EH 1 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 275_365 | 11 | 911.0 | Domain | EH 3 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 127_215 | 11 | 755.0 | Domain | EH 2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 159_194 | 11 | 755.0 | Domain | EF-hand |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 15_104 | 11 | 755.0 | Domain | EH 1 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 275_365 | 11 | 755.0 | Domain | EH 3 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 127_215 | 11 | 602.0 | Domain | EH 2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 159_194 | 11 | 602.0 | Domain | EF-hand |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 15_104 | 11 | 602.0 | Domain | EH 1 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 275_365 | 11 | 602.0 | Domain | EH 3 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 618_835 | 11 | 865.0 | Region | Note=15 X 3 AA repeats of D-P-F |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 618_835 | 11 | 911.0 | Region | Note=15 X 3 AA repeats of D-P-F |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 618_835 | 11 | 755.0 | Region | Note=15 X 3 AA repeats of D-P-F |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 618_835 | 11 | 602.0 | Region | Note=15 X 3 AA repeats of D-P-F |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 618_620 | 11 | 865.0 | Repeat | Note=1 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 640_642 | 11 | 865.0 | Repeat | Note=2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 645_647 | 11 | 865.0 | Repeat | Note=3 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 650_652 | 11 | 865.0 | Repeat | Note=4 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 656_658 | 11 | 865.0 | Repeat | Note=5 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 661_663 | 11 | 865.0 | Repeat | Note=6 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 667_669 | 11 | 865.0 | Repeat | Note=7 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 685_687 | 11 | 865.0 | Repeat | Note=8 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 690_692 | 11 | 865.0 | Repeat | Note=9 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 709_711 | 11 | 865.0 | Repeat | Note=10 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 728_730 | 11 | 865.0 | Repeat | Note=11 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 754_756 | 11 | 865.0 | Repeat | Note=12 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 806_808 | 11 | 865.0 | Repeat | Note=13 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 812_814 | 11 | 865.0 | Repeat | Note=14 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 833_835 | 11 | 865.0 | Repeat | Note=15 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 618_620 | 11 | 911.0 | Repeat | Note=1 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 640_642 | 11 | 911.0 | Repeat | Note=2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 645_647 | 11 | 911.0 | Repeat | Note=3 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 650_652 | 11 | 911.0 | Repeat | Note=4 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 656_658 | 11 | 911.0 | Repeat | Note=5 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 661_663 | 11 | 911.0 | Repeat | Note=6 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 667_669 | 11 | 911.0 | Repeat | Note=7 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 685_687 | 11 | 911.0 | Repeat | Note=8 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 690_692 | 11 | 911.0 | Repeat | Note=9 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 709_711 | 11 | 911.0 | Repeat | Note=10 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 728_730 | 11 | 911.0 | Repeat | Note=11 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 754_756 | 11 | 911.0 | Repeat | Note=12 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 806_808 | 11 | 911.0 | Repeat | Note=13 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 812_814 | 11 | 911.0 | Repeat | Note=14 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 833_835 | 11 | 911.0 | Repeat | Note=15 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 618_620 | 11 | 755.0 | Repeat | Note=1 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 640_642 | 11 | 755.0 | Repeat | Note=2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 645_647 | 11 | 755.0 | Repeat | Note=3 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 650_652 | 11 | 755.0 | Repeat | Note=4 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 656_658 | 11 | 755.0 | Repeat | Note=5 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 661_663 | 11 | 755.0 | Repeat | Note=6 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 667_669 | 11 | 755.0 | Repeat | Note=7 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 685_687 | 11 | 755.0 | Repeat | Note=8 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 690_692 | 11 | 755.0 | Repeat | Note=9 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 709_711 | 11 | 755.0 | Repeat | Note=10 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 728_730 | 11 | 755.0 | Repeat | Note=11 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 754_756 | 11 | 755.0 | Repeat | Note=12 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 806_808 | 11 | 755.0 | Repeat | Note=13 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 812_814 | 11 | 755.0 | Repeat | Note=14 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 833_835 | 11 | 755.0 | Repeat | Note=15 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 618_620 | 11 | 602.0 | Repeat | Note=1 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 640_642 | 11 | 602.0 | Repeat | Note=2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 645_647 | 11 | 602.0 | Repeat | Note=3 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 650_652 | 11 | 602.0 | Repeat | Note=4 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 656_658 | 11 | 602.0 | Repeat | Note=5 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 661_663 | 11 | 602.0 | Repeat | Note=6 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 667_669 | 11 | 602.0 | Repeat | Note=7 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 685_687 | 11 | 602.0 | Repeat | Note=8 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 690_692 | 11 | 602.0 | Repeat | Note=9 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 709_711 | 11 | 602.0 | Repeat | Note=10 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 728_730 | 11 | 602.0 | Repeat | Note=11 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 754_756 | 11 | 602.0 | Repeat | Note=12 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 806_808 | 11 | 602.0 | Repeat | Note=13 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 812_814 | 11 | 602.0 | Repeat | Note=14 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 833_835 | 11 | 602.0 | Repeat | Note=15 |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000085079 | 1 | 10 | 12_144 | 76 | 551.0 | Domain | ENTH | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000270460 | 1 | 11 | 12_144 | 76 | 577.0 | Domain | ENTH | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 12_144 | 187 | 663.0 | Domain | ENTH | |
| Tgene | EPN1 | chr19:16582723 | chr19:56196761 | ENST00000411543 | 2 | 11 | 183_202 | 187 | 663.0 | Domain | UIM 1 |
Top |
Fusion Gene Sequence for EPS15L1-EPN1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >27073_27073_1_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000248070_EPN1_chr19_56196761_ENST00000085079_length(transcript)=2008nt_BP=173nt ACATGCGTGGCCCTGTCTACGGCCCCGCCCTCTCCCGGGACAGGCGGCCTGCGCACCGCCCTCCCGTCGTCCCAGTGCGCACGCGTGGCT GCAGCCCCGCCCCCGTTCCCCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGA CGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACT TCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACC GGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCC CCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGG CCGACCAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGG AGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTG CTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCA CGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGG AGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGG CCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCCGGGGGATTCGACACGGAGCCCGACGAGTTCT CTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAA GCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCACGC CCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCACGC CGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGCCTC CCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCGGGG GCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCCTGG CTCCATCCGGCTGCCCCATTCCGGCTCCCTGGGAGATCAGTGTTGTGAGTGCATGTGAAATGGGGGATCCCCACCCCCAGTGCCCTTCCC CTTCCTGGGGCCCACTCACACTACACCCTCTTCCTTTCCCACCCCACCTCCCCGGAGAGAAACTGGACATGGGGCCTGGGGAGGGGAGCT GGCCAGAGGAGGACCCCTTTCCCGTGGCATTAGAAGGGGGAGGGGTGGCTGGGGCCCCCACCCATTCCCCCTCCCTCCAAACTCCCAACC CCCAGTCAGTGTTTGAGCCTCCTCGTTCCCCTCACGCACCCGCTCACGCACCCTCGGTGAATCCTTGGTGATGATTTTGGCAACTTTGGG >27073_27073_1_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000248070_EPN1_chr19_56196761_ENST00000085079_length(amino acids)=623AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAGV GAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSWSA SSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSRSRATSCLAFSRTFTPWSLPSRSTYWK -------------------------------------------------------------- >27073_27073_2_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000248070_EPN1_chr19_56196761_ENST00000270460_length(transcript)=2086nt_BP=173nt ACATGCGTGGCCCTGTCTACGGCCCCGCCCTCTCCCGGGACAGGCGGCCTGCGCACCGCCCTCCCGTCGTCCCAGTGCGCACGCGTGGCT GCAGCCCCGCCCCCGTTCCCCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGA CGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACT TCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACC GGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCC CCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGG CCGACCAGCCCCCGTCCTGCGGCCCCGAGGACGACGCCCAGCTCCAGCTGGCCCTTAGTTTGAGCCGAGAAGAGCATGATAAGGAGGAGC GGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGGAGTCGTCCCTCATGG ACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTGCTGCCGTCCCCACGG CTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCACGCCGGCCTCTGGGG ACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGGAGGGGCCCACGCCTG ATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGGCCTTCTCAGATCCCT GGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCAGCCGGGGGATTCGACACGGAGCCCGACGAGTTCTCTGACTTTGACC GACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAAGCCCTGGGGCGT TTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCACGCCCCCCACCCGGA AGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCACGCCGCCTGGAGCCA AGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGCCTCCCGCGACGCTCA CCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCGGGGGCCCTGGCCTGC CCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCCTGGCTCCATCCGGCT GCCCCATTCCGGCTCCCTGGGAGATCAGTGTTGTGAGTGCATGTGAAATGGGGGATCCCCACCCCCAGTGCCCTTCCCCTTCCTGGGGCC CACTCACACTACACCCTCTTCCTTTCCCACCCCACCTCCCCGGAGAGAAACTGGACATGGGGCCTGGGGAGGGGAGCTGGCCAGAGGAGG ACCCCTTTCCCGTGGCATTAGAAGGGGGAGGGGTGGCTGGGGCCCCCACCCATTCCCCCTCCCTCCAAACTCCCAACCCCCAGTCAGTGT TTGAGCCTCCTCGTTCCCCTCACGCACCCGCTCACGCACCCTCGGTGAATCCTTGGTGATGATTTTGGCAACTTTGGGAATAAATGGCAA >27073_27073_2_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000248070_EPN1_chr19_56196761_ENST00000270460_length(amino acids)=649AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAG VGAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSLS CSSRLKLRASWSWASSSGPQDGGWSASSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSR SRATSCLAFSRTFTPWSLPSRSTYWKSFSVCTAYMFSLHCCDTRSEPVLMRYSISVMACWERGMSGAAIFPRTRAPSAGGTGAGLQPRVR -------------------------------------------------------------- >27073_27073_3_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000248070_EPN1_chr19_56196761_ENST00000411543_length(transcript)=1686nt_BP=173nt ACATGCGTGGCCCTGTCTACGGCCCCGCCCTCTCCCGGGACAGGCGGCCTGCGCACCGCCCTCCCGTCGTCCCAGTGCGCACGCGTGGCT GCAGCCCCGCCCCCGTTCCCCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGA CGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACT TCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACC GGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCC CCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGG CCGACCAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGG AGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTG CTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCA CGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGG AGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGG CCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCAGCCGGGGGATTCGACACGGAGCCCGACGAGT TCTCTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCC GAAGCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCA CGCCCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCA CGCCGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGC CTCCCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCG GGGGCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCC >27073_27073_3_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000248070_EPN1_chr19_56196761_ENST00000411543_length(amino acids)=532AA_BP=56 MRGPVYGPALSRDRRPAHRPPVVPVRTRGCSPAPVPPALGARVRGKMAAPLIPLSQQAMTLMEYLIKTGSERVSQQCKENMYAVQTLKDF QYVDRDGKDQGVNVREKAKQLVALLRDEDRLREERAHALKTKEKLAQTATASSAAVGSGPPPEAEQAWPQSSGEEELQLQLALAMSKEEA DQEERIRRGDDLRLQMAIEESKRETGGKEESSLMDLADVFTAPAPAPTTDPWGGPAPMAAAVPTAAPTSDPWGGPPVPPAADPWGGPAPT PASGDPWRPAAPAGPSVDPWGGTPAPAAGEGPTPDPWGSSDGGVPVSGPSASDPWTPAPAFSDPWGGSPAKPSTNGTTAAGGFDTEPDEF SDFDRLRTALPTSGSSAGELELLAGEVPARSPGAFDMSGVRGSLAEAVGSPPPAATPTPTPPTRKTPESFLGPNAALVDLDSLVSRPGPT -------------------------------------------------------------- >27073_27073_4_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000455140_EPN1_chr19_56196761_ENST00000085079_length(transcript)=1935nt_BP=100nt AGTGCGCACGCGTGGCTGCAGCCCCGCCCCCGTTCCCCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCT CTCCCAGCAGGCCATGACGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGT GCAGACGCTGAAGGACTTCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCT GCTGCGCGACGAGGACCGGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGC AGCTGTGGGCTCAGGCCCCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGC CATGAGCAAGGAGGAGGCCGACCAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGA GACTGGGGGCAAGGAGGAGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGG CCCAGCACCCATGGCTGCTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTG GGGAGGTCCAGCCCCCACGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCC AGCCCCTGCAGCTGGGGAGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCC CTGGACACCGGCCCCGGCCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCCGGGGGATTCGACAC GGAGCCCGACGAGTTCTCTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGG AGAGGTGCCGGCCCGAAGCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGC CACACCAACTCCCACGCCCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAG CCGGCCGGGCCCCACGCCGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCC CTTCCAGCCCGCGCCTCCCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACAT CTCTCCCCTTGGCGGGGGCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAG GGCGGAAGGGGGCCTGGCTCCATCCGGCTGCCCCATTCCGGCTCCCTGGGAGATCAGTGTTGTGAGTGCATGTGAAATGGGGGATCCCCA CCCCCAGTGCCCTTCCCCTTCCTGGGGCCCACTCACACTACACCCTCTTCCTTTCCCACCCCACCTCCCCGGAGAGAAACTGGACATGGG GCCTGGGGAGGGGAGCTGGCCAGAGGAGGACCCCTTTCCCGTGGCATTAGAAGGGGGAGGGGTGGCTGGGGCCCCCACCCATTCCCCCTC CCTCCAAACTCCCAACCCCCAGTCAGTGTTTGAGCCTCCTCGTTCCCCTCACGCACCCGCTCACGCACCCTCGGTGAATCCTTGGTGATG >27073_27073_4_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000455140_EPN1_chr19_56196761_ENST00000085079_length(amino acids)=605AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAGV GAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSWSA SSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSRSRATSCLAFSRTFTPWSLPSRSTYWK -------------------------------------------------------------- >27073_27073_5_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000455140_EPN1_chr19_56196761_ENST00000270460_length(transcript)=2013nt_BP=100nt AGTGCGCACGCGTGGCTGCAGCCCCGCCCCCGTTCCCCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCT CTCCCAGCAGGCCATGACGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGT GCAGACGCTGAAGGACTTCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCT GCTGCGCGACGAGGACCGGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGC AGCTGTGGGCTCAGGCCCCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGC CATGAGCAAGGAGGAGGCCGACCAGCCCCCGTCCTGCGGCCCCGAGGACGACGCCCAGCTCCAGCTGGCCCTTAGTTTGAGCCGAGAAGA GCATGATAAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGA GGAGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGC TGCTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCC CACGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGG GGAGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCC GGCCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCAGCCGGGGGATTCGACACGGAGCCCGACGA GTTCTCTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGC CCGAAGCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCC CACGCCCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCC CACGCCGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGC GCCTCCCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGG CGGGGGCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGG CCTGGCTCCATCCGGCTGCCCCATTCCGGCTCCCTGGGAGATCAGTGTTGTGAGTGCATGTGAAATGGGGGATCCCCACCCCCAGTGCCC TTCCCCTTCCTGGGGCCCACTCACACTACACCCTCTTCCTTTCCCACCCCACCTCCCCGGAGAGAAACTGGACATGGGGCCTGGGGAGGG GAGCTGGCCAGAGGAGGACCCCTTTCCCGTGGCATTAGAAGGGGGAGGGGTGGCTGGGGCCCCCACCCATTCCCCCTCCCTCCAAACTCC CAACCCCCAGTCAGTGTTTGAGCCTCCTCGTTCCCCTCACGCACCCGCTCACGCACCCTCGGTGAATCCTTGGTGATGATTTTGGCAACT >27073_27073_5_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000455140_EPN1_chr19_56196761_ENST00000270460_length(amino acids)=631AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAG VGAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSLS CSSRLKLRASWSWASSSGPQDGGWSASSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSR SRATSCLAFSRTFTPWSLPSRSTYWKSFSVCTAYMFSLHCCDTRSEPVLMRYSISVMACWERGMSGAAIFPRTRAPSAGGTGAGLQPRVR -------------------------------------------------------------- >27073_27073_6_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000455140_EPN1_chr19_56196761_ENST00000411543_length(transcript)=1613nt_BP=100nt AGTGCGCACGCGTGGCTGCAGCCCCGCCCCCGTTCCCCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCT CTCCCAGCAGGCCATGACGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGT GCAGACGCTGAAGGACTTCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCT GCTGCGCGACGAGGACCGGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGC AGCTGTGGGCTCAGGCCCCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGC CATGAGCAAGGAGGAGGCCGACCAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGA GACTGGGGGCAAGGAGGAGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGG CCCAGCACCCATGGCTGCTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTG GGGAGGTCCAGCCCCCACGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCC AGCCCCTGCAGCTGGGGAGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCC CTGGACACCGGCCCCGGCCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCAGCCGGGGGATTCGA CACGGAGCCCGACGAGTTCTCTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGC AGGAGAGGTGCCGGCCCGAAGCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGC AGCCACACCAACTCCCACGCCCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGT GAGCCGGCCGGGCCCCACGCCGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAA CCCCTTCCAGCCCGCGCCTCCCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTA CATCTCTCCCCTTGGCGGGGGCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATC >27073_27073_6_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000455140_EPN1_chr19_56196761_ENST00000411543_length(amino acids)=486AA_BP=10 MAAPLIPLSQQAMTLMEYLIKTGSERVSQQCKENMYAVQTLKDFQYVDRDGKDQGVNVREKAKQLVALLRDEDRLREERAHALKTKEKLA QTATASSAAVGSGPPPEAEQAWPQSSGEEELQLQLALAMSKEEADQEERIRRGDDLRLQMAIEESKRETGGKEESSLMDLADVFTAPAPA PTTDPWGGPAPMAAAVPTAAPTSDPWGGPPVPPAADPWGGPAPTPASGDPWRPAAPAGPSVDPWGGTPAPAAGEGPTPDPWGSSDGGVPV SGPSASDPWTPAPAFSDPWGGSPAKPSTNGTTAAGGFDTEPDEFSDFDRLRTALPTSGSSAGELELLAGEVPARSPGAFDMSGVRGSLAE AVGSPPPAATPTPTPPTRKTPESFLGPNAALVDLDSLVSRPGPTPPGAKASNPFLPGGGPATGPSVTNPFQPAPPATLTLNQLRLSPVPP -------------------------------------------------------------- >27073_27073_7_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000535753_EPN1_chr19_56196761_ENST00000085079_length(transcript)=1899nt_BP=64nt CCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGACGCTGATGGAGTACCTCAT CAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACTTCCAGTACGTGGACCGCGA CGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACCGGCTGCGGGAAGAGCGGGC GCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCCCCCCTCCCGAGGCGGAGCA GGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGGCCGACCAGGAGGAGCGGAT CCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGGAGTCGTCCCTCATGGACCT TGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTGCTGCCGTCCCCACGGCTGC CCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCACGCCGGCCTCTGGGGACCC CTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGGAGGGGCCCACGCCTGATCC ATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGGCCTTCTCAGATCCCTGGGG AGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCCGGGGGATTCGACACGGAGCCCGACGAGTTCTCTGACTTTGACCGACTCCG CACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAAGCCCTGGGGCGTTTGACAT GAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCACGCCCCCCACCCGGAAGACGCC GGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCACGCCGCCTGGAGCCAAGGCCTC CAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGCCTCCCGCGACGCTCACCCTGAA CCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCGGGGGCCCTGGCCTGCCCCCCAT GATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCCTGGCTCCATCCGGCTGCCCCAT TCCGGCTCCCTGGGAGATCAGTGTTGTGAGTGCATGTGAAATGGGGGATCCCCACCCCCAGTGCCCTTCCCCTTCCTGGGGCCCACTCAC ACTACACCCTCTTCCTTTCCCACCCCACCTCCCCGGAGAGAAACTGGACATGGGGCCTGGGGAGGGGAGCTGGCCAGAGGAGGACCCCTT TCCCGTGGCATTAGAAGGGGGAGGGGTGGCTGGGGCCCCCACCCATTCCCCCTCCCTCCAAACTCCCAACCCCCAGTCAGTGTTTGAGCC TCCTCGTTCCCCTCACGCACCCGCTCACGCACCCTCGGTGAATCCTTGGTGATGATTTTGGCAACTTTGGGAATAAATGGCAATTCCCAC >27073_27073_7_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000535753_EPN1_chr19_56196761_ENST00000085079_length(amino acids)=593AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAGV GAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSWSA SSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSRSRATSCLAFSRTFTPWSLPSRSTYWK -------------------------------------------------------------- >27073_27073_8_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000535753_EPN1_chr19_56196761_ENST00000270460_length(transcript)=1977nt_BP=64nt CCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGACGCTGATGGAGTACCTCAT CAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACTTCCAGTACGTGGACCGCGA CGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACCGGCTGCGGGAAGAGCGGGC GCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCCCCCCTCCCGAGGCGGAGCA GGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGGCCGACCAGCCCCCGTCCTG CGGCCCCGAGGACGACGCCCAGCTCCAGCTGGCCCTTAGTTTGAGCCGAGAAGAGCATGATAAGGAGGAGCGGATCCGTCGCGGGGATGA CCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGGAGTCGTCCCTCATGGACCTTGCTGACGTCTTCAC GGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTGCTGCCGTCCCCACGGCTGCCCCCACCTCGGACCC CTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCACGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGC CCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGGAGGGGCCCACGCCTGATCCATGGGGAAGTTCCGA TGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGGCCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAA GCCCAGCACCAATGGCACAACAGCAGCCGGGGGATTCGACACGGAGCCCGACGAGTTCTCTGACTTTGACCGACTCCGCACGGCACTGCC GACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAAGCCCTGGGGCGTTTGACATGAGTGGGGTCAG GGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCACGCCCCCCACCCGGAAGACGCCGGAGTCATTCCT GGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCACGCCGCCTGGAGCCAAGGCCTCCAACCCCTTCCT GCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGCCTCCCGCGACGCTCACCCTGAACCAGCTCCGTCT CAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCGGGGGCCCTGGCCTGCCCCCCATGATGCCCCCGGG CCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCCTGGCTCCATCCGGCTGCCCCATTCCGGCTCCCTG GGAGATCAGTGTTGTGAGTGCATGTGAAATGGGGGATCCCCACCCCCAGTGCCCTTCCCCTTCCTGGGGCCCACTCACACTACACCCTCT TCCTTTCCCACCCCACCTCCCCGGAGAGAAACTGGACATGGGGCCTGGGGAGGGGAGCTGGCCAGAGGAGGACCCCTTTCCCGTGGCATT AGAAGGGGGAGGGGTGGCTGGGGCCCCCACCCATTCCCCCTCCCTCCAAACTCCCAACCCCCAGTCAGTGTTTGAGCCTCCTCGTTCCCC >27073_27073_8_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000535753_EPN1_chr19_56196761_ENST00000270460_length(amino acids)=619AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAG VGAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSLS CSSRLKLRASWSWASSSGPQDGGWSASSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSR -------------------------------------------------------------- >27073_27073_9_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000535753_EPN1_chr19_56196761_ENST00000411543_length(transcript)=1577nt_BP=64nt CCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGACGCTGATGGAGTACCTCAT CAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACTTCCAGTACGTGGACCGCGA CGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACCGGCTGCGGGAAGAGCGGGC GCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCCCCCCTCCCGAGGCGGAGCA GGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGGCCGACCAGGAGGAGCGGAT CCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGGAGTCGTCCCTCATGGACCT TGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTGCTGCCGTCCCCACGGCTGC CCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCACGCCGGCCTCTGGGGACCC CTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGGAGGGGCCCACGCCTGATCC ATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGGCCTTCTCAGATCCCTGGGG AGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCAGCCGGGGGATTCGACACGGAGCCCGACGAGTTCTCTGACTTTGACCGACT CCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAAGCCCTGGGGCGTTTGA CATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCACGCCCCCCACCCGGAAGAC GCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCACGCCGCCTGGAGCCAAGGC CTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGCCTCCCGCGACGCTCACCCT GAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCGGGGGCCCTGGCCTGCCCCC CATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCCTGGCTCCATCCGGCTGCCC >27073_27073_9_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000535753_EPN1_chr19_56196761_ENST00000411543_length(amino acids)=486AA_BP=10 MAAPLIPLSQQAMTLMEYLIKTGSERVSQQCKENMYAVQTLKDFQYVDRDGKDQGVNVREKAKQLVALLRDEDRLREERAHALKTKEKLA QTATASSAAVGSGPPPEAEQAWPQSSGEEELQLQLALAMSKEEADQEERIRRGDDLRLQMAIEESKRETGGKEESSLMDLADVFTAPAPA PTTDPWGGPAPMAAAVPTAAPTSDPWGGPPVPPAADPWGGPAPTPASGDPWRPAAPAGPSVDPWGGTPAPAAGEGPTPDPWGSSDGGVPV SGPSASDPWTPAPAFSDPWGGSPAKPSTNGTTAAGGFDTEPDEFSDFDRLRTALPTSGSSAGELELLAGEVPARSPGAFDMSGVRGSLAE AVGSPPPAATPTPTPPTRKTPESFLGPNAALVDLDSLVSRPGPTPPGAKASNPFLPGGGPATGPSVTNPFQPAPPATLTLNQLRLSPVPP -------------------------------------------------------------- >27073_27073_10_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000594975_EPN1_chr19_56196761_ENST00000085079_length(transcript)=2008nt_BP=173nt ACATGCGTGGCCCTGTCTACGGCCCCGCCCTCTCCCGGGACAGGCGGCCTGCGCACCGCCCTCCCGTCGTCCCAGTGCGCACGCGTGGCT GCAGCCCCGCCCCCGTTCCCCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGA CGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACT TCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACC GGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCC CCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGG CCGACCAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGG AGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTG CTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCA CGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGG AGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGG CCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCCGGGGGATTCGACACGGAGCCCGACGAGTTCT CTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAA GCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCACGC CCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCACGC CGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGCCTC CCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCGGGG GCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCCTGG CTCCATCCGGCTGCCCCATTCCGGCTCCCTGGGAGATCAGTGTTGTGAGTGCATGTGAAATGGGGGATCCCCACCCCCAGTGCCCTTCCC CTTCCTGGGGCCCACTCACACTACACCCTCTTCCTTTCCCACCCCACCTCCCCGGAGAGAAACTGGACATGGGGCCTGGGGAGGGGAGCT GGCCAGAGGAGGACCCCTTTCCCGTGGCATTAGAAGGGGGAGGGGTGGCTGGGGCCCCCACCCATTCCCCCTCCCTCCAAACTCCCAACC CCCAGTCAGTGTTTGAGCCTCCTCGTTCCCCTCACGCACCCGCTCACGCACCCTCGGTGAATCCTTGGTGATGATTTTGGCAACTTTGGG >27073_27073_10_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000594975_EPN1_chr19_56196761_ENST00000085079_length(amino acids)=623AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAGV GAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSWSA SSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSRSRATSCLAFSRTFTPWSLPSRSTYWK -------------------------------------------------------------- >27073_27073_11_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000594975_EPN1_chr19_56196761_ENST00000270460_length(transcript)=2086nt_BP=173nt ACATGCGTGGCCCTGTCTACGGCCCCGCCCTCTCCCGGGACAGGCGGCCTGCGCACCGCCCTCCCGTCGTCCCAGTGCGCACGCGTGGCT GCAGCCCCGCCCCCGTTCCCCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGA CGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACT TCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACC GGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCC CCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGG CCGACCAGCCCCCGTCCTGCGGCCCCGAGGACGACGCCCAGCTCCAGCTGGCCCTTAGTTTGAGCCGAGAAGAGCATGATAAGGAGGAGC GGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGGAGTCGTCCCTCATGG ACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTGCTGCCGTCCCCACGG CTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCACGCCGGCCTCTGGGG ACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGGAGGGGCCCACGCCTG ATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGGCCTTCTCAGATCCCT GGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCAGCCGGGGGATTCGACACGGAGCCCGACGAGTTCTCTGACTTTGACC GACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAAGCCCTGGGGCGT TTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCACGCCCCCCACCCGGA AGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCACGCCGCCTGGAGCCA AGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGCCTCCCGCGACGCTCA CCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCGGGGGCCCTGGCCTGC CCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCCTGGCTCCATCCGGCT GCCCCATTCCGGCTCCCTGGGAGATCAGTGTTGTGAGTGCATGTGAAATGGGGGATCCCCACCCCCAGTGCCCTTCCCCTTCCTGGGGCC CACTCACACTACACCCTCTTCCTTTCCCACCCCACCTCCCCGGAGAGAAACTGGACATGGGGCCTGGGGAGGGGAGCTGGCCAGAGGAGG ACCCCTTTCCCGTGGCATTAGAAGGGGGAGGGGTGGCTGGGGCCCCCACCCATTCCCCCTCCCTCCAAACTCCCAACCCCCAGTCAGTGT TTGAGCCTCCTCGTTCCCCTCACGCACCCGCTCACGCACCCTCGGTGAATCCTTGGTGATGATTTTGGCAACTTTGGGAATAAATGGCAA >27073_27073_11_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000594975_EPN1_chr19_56196761_ENST00000270460_length(amino acids)=649AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAG VGAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSLS CSSRLKLRASWSWASSSGPQDGGWSASSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSR SRATSCLAFSRTFTPWSLPSRSTYWKSFSVCTAYMFSLHCCDTRSEPVLMRYSISVMACWERGMSGAAIFPRTRAPSAGGTGAGLQPRVR -------------------------------------------------------------- >27073_27073_12_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000594975_EPN1_chr19_56196761_ENST00000411543_length(transcript)=1686nt_BP=173nt ACATGCGTGGCCCTGTCTACGGCCCCGCCCTCTCCCGGGACAGGCGGCCTGCGCACCGCCCTCCCGTCGTCCCAGTGCGCACGCGTGGCT GCAGCCCCGCCCCCGTTCCCCCGGCGCTCGGAGCCCGAGTCCGCGGGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGA CGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGCAGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACT TCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGCGTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACC GGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGCTGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCC CCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGGAGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGG CCGACCAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGG AGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTG CTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCA CGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGG AGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGG CCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCAGCCGGGGGATTCGACACGGAGCCCGACGAGT TCTCTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCC GAAGCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCA CGCCCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCA CGCCGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGC CTCCCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCG GGGGCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCC >27073_27073_12_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000594975_EPN1_chr19_56196761_ENST00000411543_length(amino acids)=532AA_BP=56 MRGPVYGPALSRDRRPAHRPPVVPVRTRGCSPAPVPPALGARVRGKMAAPLIPLSQQAMTLMEYLIKTGSERVSQQCKENMYAVQTLKDF QYVDRDGKDQGVNVREKAKQLVALLRDEDRLREERAHALKTKEKLAQTATASSAAVGSGPPPEAEQAWPQSSGEEELQLQLALAMSKEEA DQEERIRRGDDLRLQMAIEESKRETGGKEESSLMDLADVFTAPAPAPTTDPWGGPAPMAAAVPTAAPTSDPWGGPPVPPAADPWGGPAPT PASGDPWRPAAPAGPSVDPWGGTPAPAAGEGPTPDPWGSSDGGVPVSGPSASDPWTPAPAFSDPWGGSPAKPSTNGTTAAGGFDTEPDEF SDFDRLRTALPTSGSSAGELELLAGEVPARSPGAFDMSGVRGSLAEAVGSPPPAATPTPTPPTRKTPESFLGPNAALVDLDSLVSRPGPT -------------------------------------------------------------- >27073_27073_13_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000597937_EPN1_chr19_56196761_ENST00000085079_length(transcript)=1873nt_BP=38nt GGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGACGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGC AGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACTTCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGC GTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACCGGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGC TGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCCCCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGG AGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGGCCGACCAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGC AGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGGAGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTC CTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTGCTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCC CCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCACGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGAC CCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGGAGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCC CGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGGCCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCA ATGGCACAACAGCCGGGGGATTCGACACGGAGCCCGACGAGTTCTCTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGAGCA GCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAAGCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGGCTG AGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCACGCCCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATGCAG CCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCACGCCGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCC CAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGCCTCCCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTC CCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCGGGGGCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCCCCA ACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCCTGGCTCCATCCGGCTGCCCCATTCCGGCTCCCTGGGAGATCAGTGTTG TGAGTGCATGTGAAATGGGGGATCCCCACCCCCAGTGCCCTTCCCCTTCCTGGGGCCCACTCACACTACACCCTCTTCCTTTCCCACCCC ACCTCCCCGGAGAGAAACTGGACATGGGGCCTGGGGAGGGGAGCTGGCCAGAGGAGGACCCCTTTCCCGTGGCATTAGAAGGGGGAGGGG TGGCTGGGGCCCCCACCCATTCCCCCTCCCTCCAAACTCCCAACCCCCAGTCAGTGTTTGAGCCTCCTCGTTCCCCTCACGCACCCGCTC >27073_27073_13_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000597937_EPN1_chr19_56196761_ENST00000085079_length(amino acids)=584AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAGV GAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSWSA SSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSRSRATSCLAFSRTFTPWSLPSRSTYWK -------------------------------------------------------------- >27073_27073_14_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000597937_EPN1_chr19_56196761_ENST00000270460_length(transcript)=1951nt_BP=38nt GGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGACGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGC AGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACTTCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGC GTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACCGGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGC TGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCCCCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGG AGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGGCCGACCAGCCCCCGTCCTGCGGCCCCGAGGACGACGCCCAGCTCC AGCTGGCCCTTAGTTTGAGCCGAGAAGAGCATGATAAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGCAGATGGCAATCGAGG AGAGCAAGAGGGAGACTGGGGGCAAGGAGGAGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTCCTGCCCCGACCACAG ACCCCTGGGGGGGCCCAGCACCCATGGCTGCTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCCCCCCTGTCCCTCCAG CTGCTGATCCCTGGGGAGGTCCAGCCCCCACGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGACCCTCAGTTGACCCTT GGGGTGGGACCCCAGCCCCTGCAGCTGGGGAGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCCCGGTCAGTGGGCCCT CAGCCTCCGATCCCTGGACACCGGCCCCGGCCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCAATGGCACAACAGCAG CCGGGGGATTCGACACGGAGCCCGACGAGTTCTCTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGAGCAGCGCAGGAGAGC TGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAAGCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGGCTGAGGCTGTGGGCA GCCCCCCACCTGCAGCCACACCAACTCCCACGCCCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATGCAGCCCTCGTCGACC TGGACTCGCTGGTGAGCCGGCCGGGCCCCACGCCGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAGGCCCAGCCACTGGCC CTTCCGTCACCAACCCCTTCCAGCCCGCGCCTCCCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGCCTCCCGTCCCTGGAG CGCCACCCACGTACATCTCTCCCCTTGGCGGGGGCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCCCCAACACTAATCCCT TCCTCCTATAATCCAGGGCGGAAGGGGGCCTGGCTCCATCCGGCTGCCCCATTCCGGCTCCCTGGGAGATCAGTGTTGTGAGTGCATGTG AAATGGGGGATCCCCACCCCCAGTGCCCTTCCCCTTCCTGGGGCCCACTCACACTACACCCTCTTCCTTTCCCACCCCACCTCCCCGGAG AGAAACTGGACATGGGGCCTGGGGAGGGGAGCTGGCCAGAGGAGGACCCCTTTCCCGTGGCATTAGAAGGGGGAGGGGTGGCTGGGGCCC CCACCCATTCCCCCTCCCTCCAAACTCCCAACCCCCAGTCAGTGTTTGAGCCTCCTCGTTCCCCTCACGCACCCGCTCACGCACCCTCGG >27073_27073_14_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000597937_EPN1_chr19_56196761_ENST00000270460_length(amino acids)=610AA_BP=1 MGVWREGEWVGAPATPPPSNATGKGSSSGQLPSPGPMSSFSPGRWGGKGRGCSVSGPQEGEGHWGWGSPISHALTTLISQGAGMGQPDGA RPPSALDYRRKGLVLGAGGPGGIMGGRPGPPPRGEMYVGGAPGTGGTGLRRSWFRVSVAGGAGWKGLVTEGPVAGPPPGRKGLEALAPGG VGPGRLTSESRSTRAALGPRNDSGVFRVGGVGVGVAAGGGLPTASARDPLTPLMSNAPGLRAGTSPASSSSSPALLPEVGSAVRSRSKSE NSSGSVSNPPAAVVPLVLGLAGDPPQGSEKAGAGVQGSEAEGPLTGTPPSELPHGSGVGPSPAAGAGVPPQGSTEGPAGAAGLQGSPEAG VGAGPPQGSAAGGTGGPPQGSEVGAAVGTAAAMGAGPPQGSVVGAGAGAVKTSARSMRDDSSLPPVSLLLSSIAICSRRSSPRRIRSSLS CSSRLKLRASWSWASSSGPQDGGWSASSLLMARASWSCSSSSPLLCGHACSASGGGPEPTAADEAVAVCASFSLVLSACARSSRSRSSSR -------------------------------------------------------------- >27073_27073_15_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000597937_EPN1_chr19_56196761_ENST00000411543_length(transcript)=1551nt_BP=38nt GGAAGATGGCGGCGCCGCTCATCCCCCTCTCCCAGCAGGCCATGACGCTGATGGAGTACCTCATCAAGACCGGCTCGGAGCGCGTGTCGC AGCAGTGCAAGGAGAACATGTACGCCGTGCAGACGCTGAAGGACTTCCAGTACGTGGACCGCGACGGCAAGGACCAGGGCGTGAACGTGC GTGAGAAAGCTAAGCAGCTGGTGGCCCTGCTGCGCGACGAGGACCGGCTGCGGGAAGAGCGGGCGCACGCGCTCAAGACCAAGGAAAAGC TGGCACAGACCGCCACGGCCTCATCAGCAGCTGTGGGCTCAGGCCCCCCTCCCGAGGCGGAGCAGGCGTGGCCGCAGAGCAGCGGGGAGG AGGAGCTGCAGCTCCAGCTGGCCCTGGCCATGAGCAAGGAGGAGGCCGACCAGGAGGAGCGGATCCGTCGCGGGGATGACCTGCGGCTGC AGATGGCAATCGAGGAGAGCAAGAGGGAGACTGGGGGCAAGGAGGAGTCGTCCCTCATGGACCTTGCTGACGTCTTCACGGCCCCAGCTC CTGCCCCGACCACAGACCCCTGGGGGGGCCCAGCACCCATGGCTGCTGCCGTCCCCACGGCTGCCCCCACCTCGGACCCCTGGGGCGGCC CCCCTGTCCCTCCAGCTGCTGATCCCTGGGGAGGTCCAGCCCCCACGCCGGCCTCTGGGGACCCCTGGAGGCCTGCTGCCCCTGCAGGAC CCTCAGTTGACCCTTGGGGTGGGACCCCAGCCCCTGCAGCTGGGGAGGGGCCCACGCCTGATCCATGGGGAAGTTCCGATGGTGGGGTCC CGGTCAGTGGGCCCTCAGCCTCCGATCCCTGGACACCGGCCCCGGCCTTCTCAGATCCCTGGGGAGGGTCACCTGCCAAGCCCAGCACCA ATGGCACAACAGCAGCCGGGGGATTCGACACGGAGCCCGACGAGTTCTCTGACTTTGACCGACTCCGCACGGCACTGCCGACCTCCGGGA GCAGCGCAGGAGAGCTGGAGCTGCTGGCAGGAGAGGTGCCGGCCCGAAGCCCTGGGGCGTTTGACATGAGTGGGGTCAGGGGATCTCTGG CTGAGGCTGTGGGCAGCCCCCCACCTGCAGCCACACCAACTCCCACGCCCCCCACCCGGAAGACGCCGGAGTCATTCCTGGGGCCCAATG CAGCCCTCGTCGACCTGGACTCGCTGGTGAGCCGGCCGGGCCCCACGCCGCCTGGAGCCAAGGCCTCCAACCCCTTCCTGCCAGGCGGAG GCCCAGCCACTGGCCCTTCCGTCACCAACCCCTTCCAGCCCGCGCCTCCCGCGACGCTCACCCTGAACCAGCTCCGTCTCAGTCCTGTGC CTCCCGTCCCTGGAGCGCCACCCACGTACATCTCTCCCCTTGGCGGGGGCCCTGGCCTGCCCCCCATGATGCCCCCGGGCCCCCCGGCCC CCAACACTAATCCCTTCCTCCTATAATCCAGGGCGGAAGGGGGCCTGGCTCCATCCGGCTGCCCCATTCCGGCTCCCTGGGAGATCAGTG >27073_27073_15_EPS15L1-EPN1_EPS15L1_chr19_16582723_ENST00000597937_EPN1_chr19_56196761_ENST00000411543_length(amino acids)=486AA_BP=10 MAAPLIPLSQQAMTLMEYLIKTGSERVSQQCKENMYAVQTLKDFQYVDRDGKDQGVNVREKAKQLVALLRDEDRLREERAHALKTKEKLA QTATASSAAVGSGPPPEAEQAWPQSSGEEELQLQLALAMSKEEADQEERIRRGDDLRLQMAIEESKRETGGKEESSLMDLADVFTAPAPA PTTDPWGGPAPMAAAVPTAAPTSDPWGGPPVPPAADPWGGPAPTPASGDPWRPAAPAGPSVDPWGGTPAPAAGEGPTPDPWGSSDGGVPV SGPSASDPWTPAPAFSDPWGGSPAKPSTNGTTAAGGFDTEPDEFSDFDRLRTALPTSGSSAGELELLAGEVPARSPGAFDMSGVRGSLAE AVGSPPPAATPTPTPPTRKTPESFLGPNAALVDLDSLVSRPGPTPPGAKASNPFLPGGGPATGPSVTNPFQPAPPATLTLNQLRLSPVPP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for EPS15L1-EPN1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000248070 | - | 1 | 23 | 15_368 | 11.0 | 865.0 | DAB2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000455140 | - | 1 | 24 | 15_368 | 11.0 | 911.0 | DAB2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000535753 | - | 1 | 22 | 15_368 | 11.0 | 755.0 | DAB2 |
| Hgene | EPS15L1 | chr19:16582723 | chr19:56196761 | ENST00000597937 | - | 1 | 16 | 15_368 | 11.0 | 602.0 | DAB2 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for EPS15L1-EPN1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for EPS15L1-EPN1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
