
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ESRP1-NDUFAF6 (FusionGDB2 ID:27565) |
Fusion Gene Summary for ESRP1-NDUFAF6 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ESRP1-NDUFAF6 | Fusion gene ID: 27565 | Hgene | Tgene | Gene symbol | ESRP1 | NDUFAF6 | Gene ID | 54845 | 137682 |
| Gene name | epithelial splicing regulatory protein 1 | NADH:ubiquinone oxidoreductase complex assembly factor 6 | |
| Synonyms | DFNB109|RBM35A|RMB35A | C8orf38|MC1DN17 | |
| Cytomap | 8q22.1 | 8q22.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | epithelial splicing regulatory protein 1RNA-binding motif protein 35ARNA-binding protein 35A | NADH dehydrogenase (ubiquinone) complex I, assembly factor 6UPF0551 protein C8orf38, mitochondrialputative phytoene synthase | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q6NXG1 | Q330K2 | |
| Ensembl transtripts involved in fusion gene | ENST00000358397, ENST00000423620, ENST00000433389, ENST00000454170, ENST00000523347, | ENST00000286687, ENST00000396111, ENST00000396124, ENST00000521840, ENST00000542894, ENST00000396113, | |
| Fusion gene scores | * DoF score | 9 X 7 X 6=378 | 7 X 10 X 6=420 |
| # samples | 9 | 11 | |
| ** MAII score | log2(9/378*10)=-2.0703893278914 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(11/420*10)=-1.93288580414146 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: ESRP1 [Title/Abstract] AND NDUFAF6 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ESRP1(95680478)-NDUFAF6(96047682), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | ESRP1 | GO:0043484 | regulation of RNA splicing | 19285943 |
 Fusion gene breakpoints across ESRP1 (5'-gene) Fusion gene breakpoints across ESRP1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
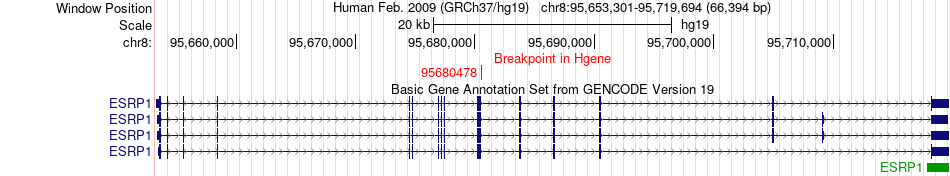 |
 Fusion gene breakpoints across NDUFAF6 (3'-gene) Fusion gene breakpoints across NDUFAF6 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
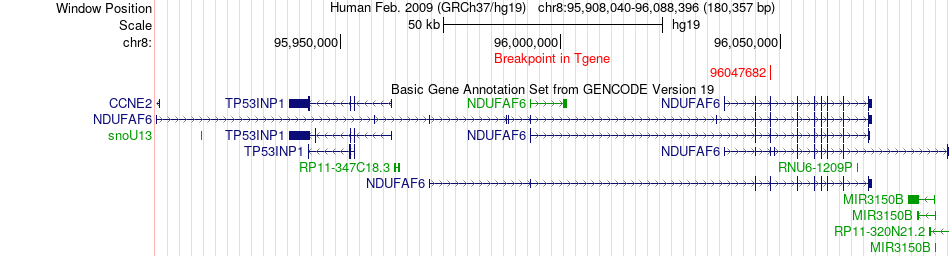 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | UCEC | TCGA-EY-A212-01A | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
Top |
Fusion Gene ORF analysis for ESRP1-NDUFAF6 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000358397 | ENST00000286687 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-5UTR | ENST00000358397 | ENST00000396111 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-5UTR | ENST00000423620 | ENST00000286687 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-5UTR | ENST00000423620 | ENST00000396111 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-5UTR | ENST00000433389 | ENST00000286687 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-5UTR | ENST00000433389 | ENST00000396111 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-5UTR | ENST00000454170 | ENST00000286687 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-5UTR | ENST00000454170 | ENST00000396111 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000358397 | ENST00000396124 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000358397 | ENST00000521840 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000358397 | ENST00000542894 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000423620 | ENST00000396124 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000423620 | ENST00000521840 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000423620 | ENST00000542894 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000433389 | ENST00000396124 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000433389 | ENST00000521840 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000433389 | ENST00000542894 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000454170 | ENST00000396124 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000454170 | ENST00000521840 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| 5CDS-intron | ENST00000454170 | ENST00000542894 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| In-frame | ENST00000358397 | ENST00000396113 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| In-frame | ENST00000423620 | ENST00000396113 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| In-frame | ENST00000433389 | ENST00000396113 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| In-frame | ENST00000454170 | ENST00000396113 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| intron-3CDS | ENST00000523347 | ENST00000396113 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| intron-5UTR | ENST00000523347 | ENST00000286687 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| intron-5UTR | ENST00000523347 | ENST00000396111 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| intron-intron | ENST00000523347 | ENST00000396124 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| intron-intron | ENST00000523347 | ENST00000521840 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
| intron-intron | ENST00000523347 | ENST00000542894 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000423620 | ESRP1 | chr8 | 95680478 | + | ENST00000396113 | NDUFAF6 | chr8 | 96047682 | + | 2956 | 1478 | 20 | 2182 | 720 |
| ENST00000433389 | ESRP1 | chr8 | 95680478 | + | ENST00000396113 | NDUFAF6 | chr8 | 96047682 | + | 2901 | 1423 | 34 | 2127 | 697 |
| ENST00000358397 | ESRP1 | chr8 | 95680478 | + | ENST00000396113 | NDUFAF6 | chr8 | 96047682 | + | 2856 | 1378 | 145 | 2082 | 645 |
| ENST00000454170 | ESRP1 | chr8 | 95680478 | + | ENST00000396113 | NDUFAF6 | chr8 | 96047682 | + | 2831 | 1353 | 120 | 2057 | 645 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000423620 | ENST00000396113 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + | 0.002690304 | 0.9973097 |
| ENST00000433389 | ENST00000396113 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + | 0.002793678 | 0.9972063 |
| ENST00000358397 | ENST00000396113 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + | 0.003052832 | 0.9969472 |
| ENST00000454170 | ENST00000396113 | ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047682 | + | 0.0025172 | 0.99748284 |
Top |
Fusion Genomic Features for ESRP1-NDUFAF6 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047681 | + | 0.001381867 | 0.9986181 |
| ESRP1 | chr8 | 95680478 | + | NDUFAF6 | chr8 | 96047681 | + | 0.001381867 | 0.9986181 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
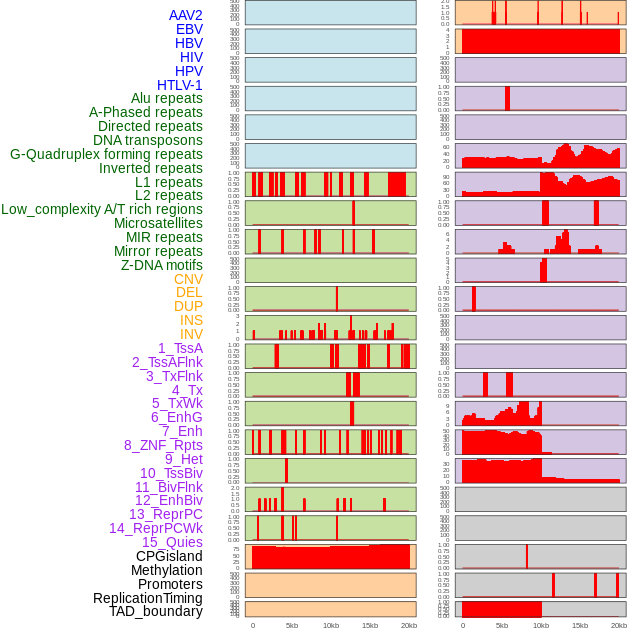 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
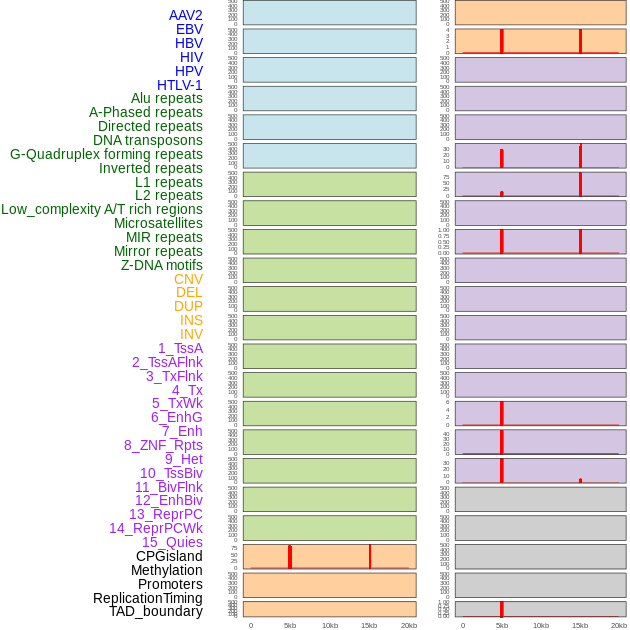 |
Top |
Fusion Protein Features for ESRP1-NDUFAF6 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr8:95680478/chr8:96047682) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| ESRP1 | NDUFAF6 |
| FUNCTION: mRNA splicing factor that regulates the formation of epithelial cell-specific isoforms. Specifically regulates the expression of FGFR2-IIIb, an epithelial cell-specific isoform of FGFR2. Also regulates the splicing of CD44, CTNND1, ENAH, 3 transcripts that undergo changes in splicing during the epithelial-to-mesenchymal transition (EMT). Acts by directly binding specific sequences in mRNAs. Binds the GU-rich sequence motifs in the ISE/ISS-3, a cis-element regulatory region present in the mRNA of FGFR2 (PubMed:19285943). Regulates splicing and expression of genes involved in inner ear development, auditory hair cell differentiation, and cell fate specification in the cochlear epithelium (By similarity). {ECO:0000250|UniProtKB:Q3US41, ECO:0000269|PubMed:19285943}. | FUNCTION: Involved in the assembly of mitochondrial NADH:ubiquinone oxidoreductase complex (complex I) at early stages. May play a role in the biogenesis of MT-ND1. {ECO:0000269|PubMed:18614015, ECO:0000269|PubMed:22019594}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000358397 | + | 10 | 16 | 225_302 | 411 | 1099.6666666666667 | Domain | RRM 1 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000358397 | + | 10 | 16 | 326_406 | 411 | 1099.6666666666667 | Domain | RRM 2 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000423620 | + | 10 | 15 | 225_302 | 411 | 660.0 | Domain | RRM 1 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000423620 | + | 10 | 15 | 326_406 | 411 | 660.0 | Domain | RRM 2 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000433389 | + | 10 | 16 | 225_302 | 411 | 1074.3333333333333 | Domain | RRM 1 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000433389 | + | 10 | 16 | 326_406 | 411 | 1074.3333333333333 | Domain | RRM 2 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000454170 | + | 10 | 14 | 225_302 | 411 | 609.0 | Domain | RRM 1 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000454170 | + | 10 | 14 | 326_406 | 411 | 609.0 | Domain | RRM 2 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000358397 | + | 10 | 16 | 445_525 | 411 | 1099.6666666666667 | Domain | RRM 3 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000423620 | + | 10 | 15 | 445_525 | 411 | 660.0 | Domain | RRM 3 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000433389 | + | 10 | 16 | 445_525 | 411 | 1074.3333333333333 | Domain | RRM 3 |
| Hgene | ESRP1 | chr8:95680478 | chr8:96047682 | ENST00000454170 | + | 10 | 14 | 445_525 | 411 | 609.0 | Domain | RRM 3 |
Top |
Fusion Gene Sequence for ESRP1-NDUFAF6 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >27565_27565_1_ESRP1-NDUFAF6_ESRP1_chr8_95680478_ENST00000358397_NDUFAF6_chr8_96047682_ENST00000396113_length(transcript)=2856nt_BP=1378nt CTTTTGCACTAGCAGTAGCAAGGAAGGGGGGTGGGCGCTCTTTCTTTTTCTCTTAGAAGAGGGTTTAGCACAGGTTTTTTCGTTCTCACT TCCACACCACCTTACCGCCTCCCGACCCCCCCTCTCCCCCTCCCCACCTATCGTCATGACGGCCTCTCCGGATTACTTGGTGGTGCTTTT TGGGATCACTGCTGGGGCCACCGGGGCCAAGCTAGGCTCGGATGAGAAGGAGTTGATCCTGCTGTTCTGGAAAGTCGTGGATCTGGCCAA CAAGAAGGTGGGACAGTTGCACGAAGTGCTAGTTAGACCGGATCAGTTGGAACTGACGGAGGACTGCAAAGAAGAAACTAAAATAGACGT CGAAAGCCTGTCCTCGGCGTCGCAGCTGGACCAAGCCCTCCGACAGTTTAACCAGTCAGTGAGCAATGAACTGAATATTGGAGTAGGGAC TTCCTTCTGTCTCTGTACTGATGGGCAGCTTCATGTCAGGCAAATCCTGCATCCTGAGGCTTCCAAGAAGAATGTACTATTACCTGAATG CTTCTATTCCTTTTTTGATCTTCGAAAAGAATTCAAGAAATGTTGCCCTGGTTCACCTGATATTGACAAACTGGACGTTGCCACAATGAC AGAGTATTTAAATTTTGAGAAGAGTAGTTCAGTCTCTCGATATGGAGCCTCTCAAGTTGAAGATATGGGGAATATAATTTTAGCAATGAT TTCAGAGCCTTATAATCACAGGTTTTCAGATCCAGAGAGAGTGAATTACAAGTTTGAAAGTGGAACTTGCAGCAAGATGGAACTTATTGA TGATAACACCGTAGTCAGGGCACGAGGTTTACCATGGCAGTCTTCAGATCAAGATATTGCAAGATTCTTCAAAGGACTCAATATTGCCAA GGGAGGTGCAGCACTTTGTCTGAATGCTCAGGGTCGAAGGAACGGAGAAGCTCTGGTTAGGTTTGTAAGTGAGGAGCACCGAGACCTAGC ACTACAGAGGCACAAACATCACATGGGGACCCGGTATATTGAGGTTTACAAAGCAACAGGTGAAGATTTCCTTAAAATTGCTGGTGGTAC TTCCAATGAGGTAGCCCAGTTTCTCTCCAAGGAAAATCAAGTCATTGTCCGCATGCGGGGGCTCCCTTTCACGGCCACAGCTGAAGAAGT GGTGGCCTTCTTTGGACAGCATTGCCCTATTACTGGGGGAAAGGAAGGCATCCTCTTTGTCACCTACCCAGATGGTAGGCCAACAGGGGA CGCTTTTGTCCTCTTTGCCTGTGAGGAATATGCACAGAATGCGTTGAGGAAGCATAAAGACTTGTTGGGTAAAAGATACATTGAACTCTT CAGGAGCACAGCAGCTGAAGTTCAGCAGGTTAAAGACTCAGTCTCTGAGAAAACAATTGGACTGATGCGAATGCAGTTTTGGAAAAAAAC TGTGGAAGATATATACTGTGACAATCCACCACATCAGCCTGTGGCCATTGAACTATGGAAGGCTGTTAAAAGACATAATCTGACTAAAAG ATGGCTTATGAAAATCGTCGATGAAAGAGAAAAAAATCTGGATGACAAAGCATATCGTAATATCAAGGAACTGGAAAATTATGCTGAAAA CACACAGAGCTCTCTTCTTTACTTAACACTAGAAATATTGGGTATAAAGGATCTTCATGCAGATCATGCTGCAAGTCATATTGGAAAAGC ACAAGGCATTGTCACTTGCTTGAGAGCAACACCATATCATGGGAGCAGAAGAAAGGTGTTCCTTCCCATGGATATTTGTATGCTGCATGG TGTTTCACAAGAGGACTTTCTACGGAGGAACCAAGATAAAAATGTGAGAGATGTAATATATGACATTGCCAGTCAAGCACACTTGCACCT AAAGCATGCTAGGTCCTTTCACAAAACTGTTCCTGTGAAAGCATTTCCTGCTTTTCTTCAGACGGTTTCTCTAGAGGACTTTCTAAAGAA AATTCAGCGAGTGGATTTTGATATATTCCACCCATCTTTACAGCAGAAGAATACATTACTTCCATTATATTTGTATATTCAGTCATGGAG AAAAACATATTAAAATAATTTCATGGCATTGATGTTAATTCTAGTCTATTAGTTTTATAAAAGTTAGGATTCTTATTTAGGAACAACAGG AAATGACTGTTAAGGAGAAAATGAATTTATTGAATGGGATGTCAAGTAGCTCACAAAATTGAGAAGTTGCCGTGGGACTAGGGTGCCAAG ACTGCTGAGATTCTGGCAGAGGCACTGCCATTGGCACCCCTGGCCCAGACAGTGTCTGCTGTGCTGTCCCTGGAAGCTGGAGCTCCAACA GGGAATATTGGTGTAGCTGTGCATAGGCCATAAGCCACACTTGAGCCAGTGGGCAGGGAGAAGGAATGTCATTCTCCCTTCACCACTGGG GAAGGAAAGGGGAAAGGGCTCAAGTTATCATAGGGGCTTACATGCTGAGCAGCTATAACCTAGGTGTTCAGAACACCTGGAAGATGCATT TATTTAGGGGTCACAAATAGTGAAATTAATATTTGAGGAATATCAGTCACGTGTTGTGGGCTTTTATCCTTAACGTTTAATTTTTACAAT CTTGTGTAAAAATTTATCCATGATAATAACATAACTTCTTTAAGGTTGGAGTGGCTGGCAAGAGCAGAGCCTTAATTTGACTTTAGCTCT GGCTGGTCTGGAAGCTTTTACTTTTTTGTTAATCCCACTTCCTCACTCTGTCATTCAGCATTATCATGATCGTGAACAAAATTGTACTGA >27565_27565_1_ESRP1-NDUFAF6_ESRP1_chr8_95680478_ENST00000358397_NDUFAF6_chr8_96047682_ENST00000396113_length(amino acids)=645AA_BP=0 MTASPDYLVVLFGITAGATGAKLGSDEKELILLFWKVVDLANKKVGQLHEVLVRPDQLELTEDCKEETKIDVESLSSASQLDQALRQFNQ SVSNELNIGVGTSFCLCTDGQLHVRQILHPEASKKNVLLPECFYSFFDLRKEFKKCCPGSPDIDKLDVATMTEYLNFEKSSSVSRYGASQ VEDMGNIILAMISEPYNHRFSDPERVNYKFESGTCSKMELIDDNTVVRARGLPWQSSDQDIARFFKGLNIAKGGAALCLNAQGRRNGEAL VRFVSEEHRDLALQRHKHHMGTRYIEVYKATGEDFLKIAGGTSNEVAQFLSKENQVIVRMRGLPFTATAEEVVAFFGQHCPITGGKEGIL FVTYPDGRPTGDAFVLFACEEYAQNALRKHKDLLGKRYIELFRSTAAEVQQVKDSVSEKTIGLMRMQFWKKTVEDIYCDNPPHQPVAIEL WKAVKRHNLTKRWLMKIVDEREKNLDDKAYRNIKELENYAENTQSSLLYLTLEILGIKDLHADHAASHIGKAQGIVTCLRATPYHGSRRK VFLPMDICMLHGVSQEDFLRRNQDKNVRDVIYDIASQAHLHLKHARSFHKTVPVKAFPAFLQTVSLEDFLKKIQRVDFDIFHPSLQQKNT -------------------------------------------------------------- >27565_27565_2_ESRP1-NDUFAF6_ESRP1_chr8_95680478_ENST00000423620_NDUFAF6_chr8_96047682_ENST00000396113_length(transcript)=2956nt_BP=1478nt ATACAAAAAGGGCAGGCCTCCTGCGCCCTCCTTCCCACCCCCCTTCCTGCCCTGGGCGTGGAGCACCGACCAGGTGTGGGCTTGGGTGGT TGGTTACCGCCTTTTGCACTAGCAGTAGCAAGGAAGGGGGGTGGGCGCTCTTTCTTTTTCTCTTAGAAGAGGGTTTAGCACAGGTTTTTT CGTTCTCACTTCCACACCACCTTACCGCCTCCCGACCCCCCCTCTCCCCCTCCCCACCTATCGTCATGACGGCCTCTCCGGATTACTTGG TGGTGCTTTTTGGGATCACTGCTGGGGCCACCGGGGCCAAGCTAGGCTCGGATGAGAAGGAGTTGATCCTGCTGTTCTGGAAAGTCGTGG ATCTGGCCAACAAGAAGGTGGGACAGTTGCACGAAGTGCTAGTTAGACCGGATCAGTTGGAACTGACGGAGGACTGCAAAGAAGAAACTA AAATAGACGTCGAAAGCCTGTCCTCGGCGTCGCAGCTGGACCAAGCCCTCCGACAGTTTAACCAGTCAGTGAGCAATGAACTGAATATTG GAGTAGGGACTTCCTTCTGTCTCTGTACTGATGGGCAGCTTCATGTCAGGCAAATCCTGCATCCTGAGGCTTCCAAGAAGAATGTACTAT TACCTGAATGCTTCTATTCCTTTTTTGATCTTCGAAAAGAATTCAAGAAATGTTGCCCTGGTTCACCTGATATTGACAAACTGGACGTTG CCACAATGACAGAGTATTTAAATTTTGAGAAGAGTAGTTCAGTCTCTCGATATGGAGCCTCTCAAGTTGAAGATATGGGGAATATAATTT TAGCAATGATTTCAGAGCCTTATAATCACAGGTTTTCAGATCCAGAGAGAGTGAATTACAAGTTTGAAAGTGGAACTTGCAGCAAGATGG AACTTATTGATGATAACACCGTAGTCAGGGCACGAGGTTTACCATGGCAGTCTTCAGATCAAGATATTGCAAGATTCTTCAAAGGACTCA ATATTGCCAAGGGAGGTGCAGCACTTTGTCTGAATGCTCAGGGTCGAAGGAACGGAGAAGCTCTGGTTAGGTTTGTAAGTGAGGAGCACC GAGACCTAGCACTACAGAGGCACAAACATCACATGGGGACCCGGTATATTGAGGTTTACAAAGCAACAGGTGAAGATTTCCTTAAAATTG CTGGTGGTACTTCCAATGAGGTAGCCCAGTTTCTCTCCAAGGAAAATCAAGTCATTGTCCGCATGCGGGGGCTCCCTTTCACGGCCACAG CTGAAGAAGTGGTGGCCTTCTTTGGACAGCATTGCCCTATTACTGGGGGAAAGGAAGGCATCCTCTTTGTCACCTACCCAGATGGTAGGC CAACAGGGGACGCTTTTGTCCTCTTTGCCTGTGAGGAATATGCACAGAATGCGTTGAGGAAGCATAAAGACTTGTTGGGTAAAAGATACA TTGAACTCTTCAGGAGCACAGCAGCTGAAGTTCAGCAGGTTAAAGACTCAGTCTCTGAGAAAACAATTGGACTGATGCGAATGCAGTTTT GGAAAAAAACTGTGGAAGATATATACTGTGACAATCCACCACATCAGCCTGTGGCCATTGAACTATGGAAGGCTGTTAAAAGACATAATC TGACTAAAAGATGGCTTATGAAAATCGTCGATGAAAGAGAAAAAAATCTGGATGACAAAGCATATCGTAATATCAAGGAACTGGAAAATT ATGCTGAAAACACACAGAGCTCTCTTCTTTACTTAACACTAGAAATATTGGGTATAAAGGATCTTCATGCAGATCATGCTGCAAGTCATA TTGGAAAAGCACAAGGCATTGTCACTTGCTTGAGAGCAACACCATATCATGGGAGCAGAAGAAAGGTGTTCCTTCCCATGGATATTTGTA TGCTGCATGGTGTTTCACAAGAGGACTTTCTACGGAGGAACCAAGATAAAAATGTGAGAGATGTAATATATGACATTGCCAGTCAAGCAC ACTTGCACCTAAAGCATGCTAGGTCCTTTCACAAAACTGTTCCTGTGAAAGCATTTCCTGCTTTTCTTCAGACGGTTTCTCTAGAGGACT TTCTAAAGAAAATTCAGCGAGTGGATTTTGATATATTCCACCCATCTTTACAGCAGAAGAATACATTACTTCCATTATATTTGTATATTC AGTCATGGAGAAAAACATATTAAAATAATTTCATGGCATTGATGTTAATTCTAGTCTATTAGTTTTATAAAAGTTAGGATTCTTATTTAG GAACAACAGGAAATGACTGTTAAGGAGAAAATGAATTTATTGAATGGGATGTCAAGTAGCTCACAAAATTGAGAAGTTGCCGTGGGACTA GGGTGCCAAGACTGCTGAGATTCTGGCAGAGGCACTGCCATTGGCACCCCTGGCCCAGACAGTGTCTGCTGTGCTGTCCCTGGAAGCTGG AGCTCCAACAGGGAATATTGGTGTAGCTGTGCATAGGCCATAAGCCACACTTGAGCCAGTGGGCAGGGAGAAGGAATGTCATTCTCCCTT CACCACTGGGGAAGGAAAGGGGAAAGGGCTCAAGTTATCATAGGGGCTTACATGCTGAGCAGCTATAACCTAGGTGTTCAGAACACCTGG AAGATGCATTTATTTAGGGGTCACAAATAGTGAAATTAATATTTGAGGAATATCAGTCACGTGTTGTGGGCTTTTATCCTTAACGTTTAA TTTTTACAATCTTGTGTAAAAATTTATCCATGATAATAACATAACTTCTTTAAGGTTGGAGTGGCTGGCAAGAGCAGAGCCTTAATTTGA CTTTAGCTCTGGCTGGTCTGGAAGCTTTTACTTTTTTGTTAATCCCACTTCCTCACTCTGTCATTCAGCATTATCATGATCGTGAACAAA >27565_27565_2_ESRP1-NDUFAF6_ESRP1_chr8_95680478_ENST00000423620_NDUFAF6_chr8_96047682_ENST00000396113_length(amino acids)=720AA_BP=1 MRPPSHPPSCPGRGAPTRCGLGWLVTAFCTSSSKEGGWALFLFLLEEGLAQVFSFSLPHHLTASRPPLSPSPPIVMTASPDYLVVLFGIT AGATGAKLGSDEKELILLFWKVVDLANKKVGQLHEVLVRPDQLELTEDCKEETKIDVESLSSASQLDQALRQFNQSVSNELNIGVGTSFC LCTDGQLHVRQILHPEASKKNVLLPECFYSFFDLRKEFKKCCPGSPDIDKLDVATMTEYLNFEKSSSVSRYGASQVEDMGNIILAMISEP YNHRFSDPERVNYKFESGTCSKMELIDDNTVVRARGLPWQSSDQDIARFFKGLNIAKGGAALCLNAQGRRNGEALVRFVSEEHRDLALQR HKHHMGTRYIEVYKATGEDFLKIAGGTSNEVAQFLSKENQVIVRMRGLPFTATAEEVVAFFGQHCPITGGKEGILFVTYPDGRPTGDAFV LFACEEYAQNALRKHKDLLGKRYIELFRSTAAEVQQVKDSVSEKTIGLMRMQFWKKTVEDIYCDNPPHQPVAIELWKAVKRHNLTKRWLM KIVDEREKNLDDKAYRNIKELENYAENTQSSLLYLTLEILGIKDLHADHAASHIGKAQGIVTCLRATPYHGSRRKVFLPMDICMLHGVSQ EDFLRRNQDKNVRDVIYDIASQAHLHLKHARSFHKTVPVKAFPAFLQTVSLEDFLKKIQRVDFDIFHPSLQQKNTLLPLYLYIQSWRKTY -------------------------------------------------------------- >27565_27565_3_ESRP1-NDUFAF6_ESRP1_chr8_95680478_ENST00000433389_NDUFAF6_chr8_96047682_ENST00000396113_length(transcript)=2901nt_BP=1423nt GCGTGGAGCACCGACCAGGTGTGGGCTTGGGTGGTTGGTTACCGCCTTTTGCACTAGCAGTAGCAAGGAAGGGGGGTGGGCGCTCTTTCT TTTTCTCTTAGAAGAGGGTTTAGCACAGGTTTTTTCGTTCTCACTTCCACACCACCTTACCGCCTCCCGACCCCCCCTCTCCCCCTCCCC ACCTATCGTCATGACGGCCTCTCCGGATTACTTGGTGGTGCTTTTTGGGATCACTGCTGGGGCCACCGGGGCCAAGCTAGGCTCGGATGA GAAGGAGTTGATCCTGCTGTTCTGGAAAGTCGTGGATCTGGCCAACAAGAAGGTGGGACAGTTGCACGAAGTGCTAGTTAGACCGGATCA GTTGGAACTGACGGAGGACTGCAAAGAAGAAACTAAAATAGACGTCGAAAGCCTGTCCTCGGCGTCGCAGCTGGACCAAGCCCTCCGACA GTTTAACCAGTCAGTGAGCAATGAACTGAATATTGGAGTAGGGACTTCCTTCTGTCTCTGTACTGATGGGCAGCTTCATGTCAGGCAAAT CCTGCATCCTGAGGCTTCCAAGAAGAATGTACTATTACCTGAATGCTTCTATTCCTTTTTTGATCTTCGAAAAGAATTCAAGAAATGTTG CCCTGGTTCACCTGATATTGACAAACTGGACGTTGCCACAATGACAGAGTATTTAAATTTTGAGAAGAGTAGTTCAGTCTCTCGATATGG AGCCTCTCAAGTTGAAGATATGGGGAATATAATTTTAGCAATGATTTCAGAGCCTTATAATCACAGGTTTTCAGATCCAGAGAGAGTGAA TTACAAGTTTGAAAGTGGAACTTGCAGCAAGATGGAACTTATTGATGATAACACCGTAGTCAGGGCACGAGGTTTACCATGGCAGTCTTC AGATCAAGATATTGCAAGATTCTTCAAAGGACTCAATATTGCCAAGGGAGGTGCAGCACTTTGTCTGAATGCTCAGGGTCGAAGGAACGG AGAAGCTCTGGTTAGGTTTGTAAGTGAGGAGCACCGAGACCTAGCACTACAGAGGCACAAACATCACATGGGGACCCGGTATATTGAGGT TTACAAAGCAACAGGTGAAGATTTCCTTAAAATTGCTGGTGGTACTTCCAATGAGGTAGCCCAGTTTCTCTCCAAGGAAAATCAAGTCAT TGTCCGCATGCGGGGGCTCCCTTTCACGGCCACAGCTGAAGAAGTGGTGGCCTTCTTTGGACAGCATTGCCCTATTACTGGGGGAAAGGA AGGCATCCTCTTTGTCACCTACCCAGATGGTAGGCCAACAGGGGACGCTTTTGTCCTCTTTGCCTGTGAGGAATATGCACAGAATGCGTT GAGGAAGCATAAAGACTTGTTGGGTAAAAGATACATTGAACTCTTCAGGAGCACAGCAGCTGAAGTTCAGCAGGTTAAAGACTCAGTCTC TGAGAAAACAATTGGACTGATGCGAATGCAGTTTTGGAAAAAAACTGTGGAAGATATATACTGTGACAATCCACCACATCAGCCTGTGGC CATTGAACTATGGAAGGCTGTTAAAAGACATAATCTGACTAAAAGATGGCTTATGAAAATCGTCGATGAAAGAGAAAAAAATCTGGATGA CAAAGCATATCGTAATATCAAGGAACTGGAAAATTATGCTGAAAACACACAGAGCTCTCTTCTTTACTTAACACTAGAAATATTGGGTAT AAAGGATCTTCATGCAGATCATGCTGCAAGTCATATTGGAAAAGCACAAGGCATTGTCACTTGCTTGAGAGCAACACCATATCATGGGAG CAGAAGAAAGGTGTTCCTTCCCATGGATATTTGTATGCTGCATGGTGTTTCACAAGAGGACTTTCTACGGAGGAACCAAGATAAAAATGT GAGAGATGTAATATATGACATTGCCAGTCAAGCACACTTGCACCTAAAGCATGCTAGGTCCTTTCACAAAACTGTTCCTGTGAAAGCATT TCCTGCTTTTCTTCAGACGGTTTCTCTAGAGGACTTTCTAAAGAAAATTCAGCGAGTGGATTTTGATATATTCCACCCATCTTTACAGCA GAAGAATACATTACTTCCATTATATTTGTATATTCAGTCATGGAGAAAAACATATTAAAATAATTTCATGGCATTGATGTTAATTCTAGT CTATTAGTTTTATAAAAGTTAGGATTCTTATTTAGGAACAACAGGAAATGACTGTTAAGGAGAAAATGAATTTATTGAATGGGATGTCAA GTAGCTCACAAAATTGAGAAGTTGCCGTGGGACTAGGGTGCCAAGACTGCTGAGATTCTGGCAGAGGCACTGCCATTGGCACCCCTGGCC CAGACAGTGTCTGCTGTGCTGTCCCTGGAAGCTGGAGCTCCAACAGGGAATATTGGTGTAGCTGTGCATAGGCCATAAGCCACACTTGAG CCAGTGGGCAGGGAGAAGGAATGTCATTCTCCCTTCACCACTGGGGAAGGAAAGGGGAAAGGGCTCAAGTTATCATAGGGGCTTACATGC TGAGCAGCTATAACCTAGGTGTTCAGAACACCTGGAAGATGCATTTATTTAGGGGTCACAAATAGTGAAATTAATATTTGAGGAATATCA GTCACGTGTTGTGGGCTTTTATCCTTAACGTTTAATTTTTACAATCTTGTGTAAAAATTTATCCATGATAATAACATAACTTCTTTAAGG TTGGAGTGGCTGGCAAGAGCAGAGCCTTAATTTGACTTTAGCTCTGGCTGGTCTGGAAGCTTTTACTTTTTTGTTAATCCCACTTCCTCA CTCTGTCATTCAGCATTATCATGATCGTGAACAAAATTGTACTGAGAAAGTTTGTATAAGTGATATGAACGGTATTGTATTGAACATCCA >27565_27565_3_ESRP1-NDUFAF6_ESRP1_chr8_95680478_ENST00000433389_NDUFAF6_chr8_96047682_ENST00000396113_length(amino acids)=697AA_BP=1 MVTAFCTSSSKEGGWALFLFLLEEGLAQVFSFSLPHHLTASRPPLSPSPPIVMTASPDYLVVLFGITAGATGAKLGSDEKELILLFWKVV DLANKKVGQLHEVLVRPDQLELTEDCKEETKIDVESLSSASQLDQALRQFNQSVSNELNIGVGTSFCLCTDGQLHVRQILHPEASKKNVL LPECFYSFFDLRKEFKKCCPGSPDIDKLDVATMTEYLNFEKSSSVSRYGASQVEDMGNIILAMISEPYNHRFSDPERVNYKFESGTCSKM ELIDDNTVVRARGLPWQSSDQDIARFFKGLNIAKGGAALCLNAQGRRNGEALVRFVSEEHRDLALQRHKHHMGTRYIEVYKATGEDFLKI AGGTSNEVAQFLSKENQVIVRMRGLPFTATAEEVVAFFGQHCPITGGKEGILFVTYPDGRPTGDAFVLFACEEYAQNALRKHKDLLGKRY IELFRSTAAEVQQVKDSVSEKTIGLMRMQFWKKTVEDIYCDNPPHQPVAIELWKAVKRHNLTKRWLMKIVDEREKNLDDKAYRNIKELEN YAENTQSSLLYLTLEILGIKDLHADHAASHIGKAQGIVTCLRATPYHGSRRKVFLPMDICMLHGVSQEDFLRRNQDKNVRDVIYDIASQA -------------------------------------------------------------- >27565_27565_4_ESRP1-NDUFAF6_ESRP1_chr8_95680478_ENST00000454170_NDUFAF6_chr8_96047682_ENST00000396113_length(transcript)=2831nt_BP=1353nt GGGGGGTGGGCGCTCTTTCTTTTTCTCTTAGAAGAGGGTTTAGCACAGGTTTTTTCGTTCTCACTTCCACACCACCTTACCGCCTCCCGA CCCCCCCTCTCCCCCTCCCCACCTATCGTCATGACGGCCTCTCCGGATTACTTGGTGGTGCTTTTTGGGATCACTGCTGGGGCCACCGGG GCCAAGCTAGGCTCGGATGAGAAGGAGTTGATCCTGCTGTTCTGGAAAGTCGTGGATCTGGCCAACAAGAAGGTGGGACAGTTGCACGAA GTGCTAGTTAGACCGGATCAGTTGGAACTGACGGAGGACTGCAAAGAAGAAACTAAAATAGACGTCGAAAGCCTGTCCTCGGCGTCGCAG CTGGACCAAGCCCTCCGACAGTTTAACCAGTCAGTGAGCAATGAACTGAATATTGGAGTAGGGACTTCCTTCTGTCTCTGTACTGATGGG CAGCTTCATGTCAGGCAAATCCTGCATCCTGAGGCTTCCAAGAAGAATGTACTATTACCTGAATGCTTCTATTCCTTTTTTGATCTTCGA AAAGAATTCAAGAAATGTTGCCCTGGTTCACCTGATATTGACAAACTGGACGTTGCCACAATGACAGAGTATTTAAATTTTGAGAAGAGT AGTTCAGTCTCTCGATATGGAGCCTCTCAAGTTGAAGATATGGGGAATATAATTTTAGCAATGATTTCAGAGCCTTATAATCACAGGTTT TCAGATCCAGAGAGAGTGAATTACAAGTTTGAAAGTGGAACTTGCAGCAAGATGGAACTTATTGATGATAACACCGTAGTCAGGGCACGA GGTTTACCATGGCAGTCTTCAGATCAAGATATTGCAAGATTCTTCAAAGGACTCAATATTGCCAAGGGAGGTGCAGCACTTTGTCTGAAT GCTCAGGGTCGAAGGAACGGAGAAGCTCTGGTTAGGTTTGTAAGTGAGGAGCACCGAGACCTAGCACTACAGAGGCACAAACATCACATG GGGACCCGGTATATTGAGGTTTACAAAGCAACAGGTGAAGATTTCCTTAAAATTGCTGGTGGTACTTCCAATGAGGTAGCCCAGTTTCTC TCCAAGGAAAATCAAGTCATTGTCCGCATGCGGGGGCTCCCTTTCACGGCCACAGCTGAAGAAGTGGTGGCCTTCTTTGGACAGCATTGC CCTATTACTGGGGGAAAGGAAGGCATCCTCTTTGTCACCTACCCAGATGGTAGGCCAACAGGGGACGCTTTTGTCCTCTTTGCCTGTGAG GAATATGCACAGAATGCGTTGAGGAAGCATAAAGACTTGTTGGGTAAAAGATACATTGAACTCTTCAGGAGCACAGCAGCTGAAGTTCAG CAGGTTAAAGACTCAGTCTCTGAGAAAACAATTGGACTGATGCGAATGCAGTTTTGGAAAAAAACTGTGGAAGATATATACTGTGACAAT CCACCACATCAGCCTGTGGCCATTGAACTATGGAAGGCTGTTAAAAGACATAATCTGACTAAAAGATGGCTTATGAAAATCGTCGATGAA AGAGAAAAAAATCTGGATGACAAAGCATATCGTAATATCAAGGAACTGGAAAATTATGCTGAAAACACACAGAGCTCTCTTCTTTACTTA ACACTAGAAATATTGGGTATAAAGGATCTTCATGCAGATCATGCTGCAAGTCATATTGGAAAAGCACAAGGCATTGTCACTTGCTTGAGA GCAACACCATATCATGGGAGCAGAAGAAAGGTGTTCCTTCCCATGGATATTTGTATGCTGCATGGTGTTTCACAAGAGGACTTTCTACGG AGGAACCAAGATAAAAATGTGAGAGATGTAATATATGACATTGCCAGTCAAGCACACTTGCACCTAAAGCATGCTAGGTCCTTTCACAAA ACTGTTCCTGTGAAAGCATTTCCTGCTTTTCTTCAGACGGTTTCTCTAGAGGACTTTCTAAAGAAAATTCAGCGAGTGGATTTTGATATA TTCCACCCATCTTTACAGCAGAAGAATACATTACTTCCATTATATTTGTATATTCAGTCATGGAGAAAAACATATTAAAATAATTTCATG GCATTGATGTTAATTCTAGTCTATTAGTTTTATAAAAGTTAGGATTCTTATTTAGGAACAACAGGAAATGACTGTTAAGGAGAAAATGAA TTTATTGAATGGGATGTCAAGTAGCTCACAAAATTGAGAAGTTGCCGTGGGACTAGGGTGCCAAGACTGCTGAGATTCTGGCAGAGGCAC TGCCATTGGCACCCCTGGCCCAGACAGTGTCTGCTGTGCTGTCCCTGGAAGCTGGAGCTCCAACAGGGAATATTGGTGTAGCTGTGCATA GGCCATAAGCCACACTTGAGCCAGTGGGCAGGGAGAAGGAATGTCATTCTCCCTTCACCACTGGGGAAGGAAAGGGGAAAGGGCTCAAGT TATCATAGGGGCTTACATGCTGAGCAGCTATAACCTAGGTGTTCAGAACACCTGGAAGATGCATTTATTTAGGGGTCACAAATAGTGAAA TTAATATTTGAGGAATATCAGTCACGTGTTGTGGGCTTTTATCCTTAACGTTTAATTTTTACAATCTTGTGTAAAAATTTATCCATGATA ATAACATAACTTCTTTAAGGTTGGAGTGGCTGGCAAGAGCAGAGCCTTAATTTGACTTTAGCTCTGGCTGGTCTGGAAGCTTTTACTTTT TTGTTAATCCCACTTCCTCACTCTGTCATTCAGCATTATCATGATCGTGAACAAAATTGTACTGAGAAAGTTTGTATAAGTGATATGAAC >27565_27565_4_ESRP1-NDUFAF6_ESRP1_chr8_95680478_ENST00000454170_NDUFAF6_chr8_96047682_ENST00000396113_length(amino acids)=645AA_BP=0 MTASPDYLVVLFGITAGATGAKLGSDEKELILLFWKVVDLANKKVGQLHEVLVRPDQLELTEDCKEETKIDVESLSSASQLDQALRQFNQ SVSNELNIGVGTSFCLCTDGQLHVRQILHPEASKKNVLLPECFYSFFDLRKEFKKCCPGSPDIDKLDVATMTEYLNFEKSSSVSRYGASQ VEDMGNIILAMISEPYNHRFSDPERVNYKFESGTCSKMELIDDNTVVRARGLPWQSSDQDIARFFKGLNIAKGGAALCLNAQGRRNGEAL VRFVSEEHRDLALQRHKHHMGTRYIEVYKATGEDFLKIAGGTSNEVAQFLSKENQVIVRMRGLPFTATAEEVVAFFGQHCPITGGKEGIL FVTYPDGRPTGDAFVLFACEEYAQNALRKHKDLLGKRYIELFRSTAAEVQQVKDSVSEKTIGLMRMQFWKKTVEDIYCDNPPHQPVAIEL WKAVKRHNLTKRWLMKIVDEREKNLDDKAYRNIKELENYAENTQSSLLYLTLEILGIKDLHADHAASHIGKAQGIVTCLRATPYHGSRRK VFLPMDICMLHGVSQEDFLRRNQDKNVRDVIYDIASQAHLHLKHARSFHKTVPVKAFPAFLQTVSLEDFLKKIQRVDFDIFHPSLQQKNT -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for ESRP1-NDUFAF6 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ESRP1-NDUFAF6 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ESRP1-NDUFAF6 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
