
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ESYT3-RPN1 (FusionGDB2 ID:27616) |
Fusion Gene Summary for ESYT3-RPN1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ESYT3-RPN1 | Fusion gene ID: 27616 | Hgene | Tgene | Gene symbol | ESYT3 | RPN1 | Gene ID | 83850 | 6184 |
| Gene name | extended synaptotagmin 3 | ribophorin I | |
| Synonyms | CHR3SYT|E-Syt3|FAM62C | OST1|RBPH1 | |
| Cytomap | 3q22.3 | 3q21.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | extended synaptotagmin-3chr3 synaptotagminextended synaptotagmin protein 3family with sequence similarity 62 (C2 domain containing), member C | dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 1RPN-Idolichyl-diphosphooligosaccharide-protein glycosyltransferase 67 kDa subunitoligosaccharyltransferase 1 homologoligosaccharyltransferase complex subunit (non-catalytic)ribop | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | A0FGR9 | P04843 | |
| Ensembl transtripts involved in fusion gene | ENST00000289135, ENST00000389567, ENST00000460133, | ENST00000497289, ENST00000490166, ENST00000296255, | |
| Fusion gene scores | * DoF score | 3 X 2 X 3=18 | 11 X 7 X 7=539 |
| # samples | 3 | 11 | |
| ** MAII score | log2(3/18*10)=0.736965594166206 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(11/539*10)=-2.29278174922785 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: ESYT3 [Title/Abstract] AND RPN1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ESYT3(138153967)-RPN1(128363826), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across ESYT3 (5'-gene) Fusion gene breakpoints across ESYT3 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
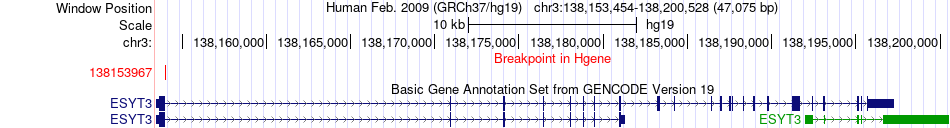 |
 Fusion gene breakpoints across RPN1 (3'-gene) Fusion gene breakpoints across RPN1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
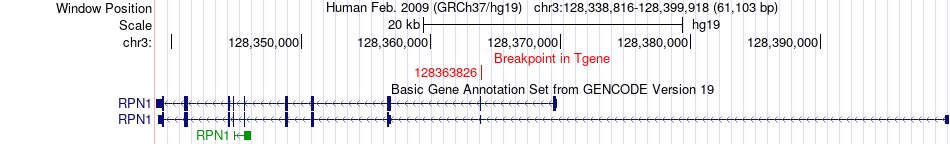 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LUSC | TCGA-66-2756-01A | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
| ChimerDB4 | LUSC | TCGA-66-2756 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
Top |
Fusion Gene ORF analysis for ESYT3-RPN1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000289135 | ENST00000497289 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
| 5CDS-5UTR | ENST00000389567 | ENST00000497289 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
| 5CDS-intron | ENST00000289135 | ENST00000490166 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
| 5CDS-intron | ENST00000389567 | ENST00000490166 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
| In-frame | ENST00000289135 | ENST00000296255 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
| In-frame | ENST00000389567 | ENST00000296255 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
| intron-3CDS | ENST00000460133 | ENST00000296255 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
| intron-5UTR | ENST00000460133 | ENST00000497289 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
| intron-intron | ENST00000460133 | ENST00000490166 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000389567 | ESYT3 | chr3 | 138153967 | + | ENST00000296255 | RPN1 | chr3 | 128363826 | - | 2513 | 513 | 186 | 2075 | 629 |
| ENST00000289135 | ESYT3 | chr3 | 138153967 | + | ENST00000296255 | RPN1 | chr3 | 128363826 | - | 2501 | 501 | 174 | 2063 | 629 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000389567 | ENST00000296255 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - | 0.001010584 | 0.9989894 |
| ENST00000289135 | ENST00000296255 | ESYT3 | chr3 | 138153967 | + | RPN1 | chr3 | 128363826 | - | 0.000991383 | 0.9990086 |
Top |
Fusion Genomic Features for ESYT3-RPN1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
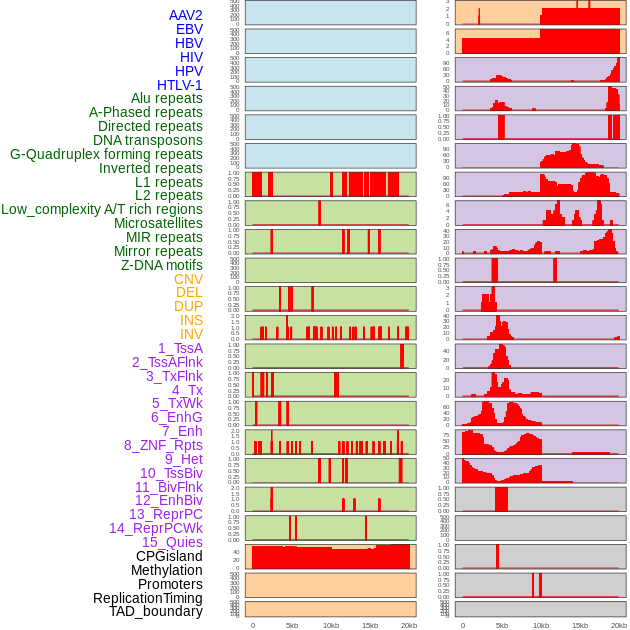 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for ESYT3-RPN1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr3:138153967/chr3:128363826) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| ESYT3 | RPN1 |
| FUNCTION: Binds glycerophospholipids in a barrel-like domain and may play a role in cellular lipid transport (By similarity). Tethers the endoplasmic reticulum to the cell membrane and promotes the formation of appositions between the endoplasmic reticulum and the cell membrane. {ECO:0000250, ECO:0000269|PubMed:23791178}. | FUNCTION: Subunit of the oligosaccharyl transferase (OST) complex that catalyzes the initial transfer of a defined glycan (Glc(3)Man(9)GlcNAc(2) in eukaryotes) from the lipid carrier dolichol-pyrophosphate to an asparagine residue within an Asn-X-Ser/Thr consensus motif in nascent polypeptide chains, the first step in protein N-glycosylation (PubMed:31831667). N-glycosylation occurs cotranslationally and the complex associates with the Sec61 complex at the channel-forming translocon complex that mediates protein translocation across the endoplasmic reticulum (ER). All subunits are required for a maximal enzyme activity. {ECO:0000250|UniProtKB:E2RQ08, ECO:0000269|PubMed:31831667}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ESYT3 | chr3:138153967 | chr3:128363826 | ENST00000389567 | + | 1 | 23 | 1_29 | 109 | 887.0 | Topological domain | Cytoplasmic |
| Hgene | ESYT3 | chr3:138153967 | chr3:128363826 | ENST00000389567 | + | 1 | 23 | 30_50 | 109 | 887.0 | Transmembrane | Helical |
| Hgene | ESYT3 | chr3:138153967 | chr3:128363826 | ENST00000389567 | + | 1 | 23 | 51_71 | 109 | 887.0 | Transmembrane | Helical |
| Tgene | RPN1 | chr3:138153967 | chr3:128363826 | ENST00000296255 | 0 | 10 | 458_607 | 87 | 608.0 | Topological domain | Cytoplasmic | |
| Tgene | RPN1 | chr3:138153967 | chr3:128363826 | ENST00000296255 | 0 | 10 | 439_457 | 87 | 608.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ESYT3 | chr3:138153967 | chr3:128363826 | ENST00000389567 | + | 1 | 23 | 114_291 | 109 | 887.0 | Domain | SMP-LTD |
| Hgene | ESYT3 | chr3:138153967 | chr3:128363826 | ENST00000389567 | + | 1 | 23 | 291_408 | 109 | 887.0 | Domain | C2 1 |
| Hgene | ESYT3 | chr3:138153967 | chr3:128363826 | ENST00000389567 | + | 1 | 23 | 426_566 | 109 | 887.0 | Domain | C2 2 |
| Hgene | ESYT3 | chr3:138153967 | chr3:128363826 | ENST00000389567 | + | 1 | 23 | 754_876 | 109 | 887.0 | Domain | C2 3 |
| Hgene | ESYT3 | chr3:138153967 | chr3:128363826 | ENST00000389567 | + | 1 | 23 | 801_808 | 109 | 887.0 | Region | Required for phosphatidylinositol 4%2C5-bisphosphate-dependent location at the cell membrane |
| Hgene | ESYT3 | chr3:138153967 | chr3:128363826 | ENST00000389567 | + | 1 | 23 | 72_886 | 109 | 887.0 | Topological domain | Cytoplasmic |
| Tgene | RPN1 | chr3:138153967 | chr3:128363826 | ENST00000296255 | 0 | 10 | 24_438 | 87 | 608.0 | Topological domain | Lumenal |
Top |
Fusion Gene Sequence for ESYT3-RPN1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >27616_27616_1_ESYT3-RPN1_ESYT3_chr3_138153967_ENST00000289135_RPN1_chr3_128363826_ENST00000296255_length(transcript)=2501nt_BP=501nt GGCGCTGCTCTCCGTCGCAGAGAACCCTGAGCTCGGCGCGCCGAGAGTCCCAGCAGGGCAAGGGGGCGCGGCGTCCTGGTCCTCGAGCTT GGGAGACAGATGCGCATGGGCGTGGGGGCATGCGGACCTAAGCTCGGGTGAAGCTCTCGGGAAGGGCAAGACTGCGGCGACGAGATGCGA GCAGAGGAGCCCTGCGCCCCCGGGGCCCCCAGCGCCCTGGGAGCCCAGCGCACGCCGGGCCCCGAGCTGCGCCTGTCCAGCCAGCTGCTG CCCGAGCTCTGTACCTTCGTGGTGCGCGTGCTGTTCTACCTGGGGCCTGTCTACCTAGCTGGCTACCTGGGGCTCAGCATAACCTGGTTG CTGCTCGGCGCCCTGCTGTGGATGTGGTGGCGCAGGAACCGCCGCGGGAAGCTTGGGCGCCTGGCCGCCGCCTTCGAATTCCTTGACAAT GAACGCGAGTTCATCAGCCGCGAGCTGCGGGGCCAGCACCTGCCAGCCTGGGTAAAGGGAGAAGATGAGGAAGAGAACAATTTGGAAGTA CGTGAAACCAAAATTAAGGGTAAAAGTGGGAGATTCTTCACAGTCAAGCTCCCAGTTGCTCTTGATCCTGGGGCCAAGATTTCAGTCATT GTGGAAACAGTCTACACCCATGTGCTTCATCCGTATCCAACCCAGATCACCCAGTCAGAGAAACAGTTTGTGGTGTTTGAGGGGAACCAT TATTTCTACTCTCCCTATCCAACGAAGACACAAACCATGCGTGTGAAGCTTGCCTCTCGAAATGTGGAGAGCTACACCAAGCTGGGGAAC CCCACGCGCTCTGAGGACCTACTGGATTATGGGCCTTTCAGAGATGTGCCTGCCTATAGTCAGGATACTTTTAAAGTACATTATGAGAAC AACAGCCCTTTCCTGACCATCACCAGCATGACCCGAGTCATTGAAGTCTCTCACTGGGGTAATATTGCTGTGGAAGAAAATGTGGACTTA AAGCACACAGGAGCTGTGCTTAAGGGGCCTTTCTCACGCTATGATTACCAGAGACAGCCAGATAGTGGAATATCCTCCATCCGTTCTTTT AAGACCATCCTTCCTGCTGCTGCCCAGGATGTTTATTACCGGGATGAGATTGGCAATGTTTCTACCAGCCACCTCCTTATTTTGGATGAC TCTGTAGAGATGGAAATCCGGCCTCGCTTCCCTCTCTTTGGCGGGTGGAAGACCCATTACATCGTTGGCTACAACCTCCCAAGCTATGAG TACCTCTATAATTTGGGTGACCAGTATGCACTGAAGATGAGGTTTGTGGACCATGTGTTTGATGAACAAGTGATAGATTCTCTGACTGTG AAGATCATCCTGCCTGAAGGAGCCAAGAACATTGAAATTGATAGTCCCTATGAAATCAGCCGTGCCCCAGATGAGCTGCACTACACCTAT CTGGACACATTTGGCCGCCCTGTGATTGTTGCCTACAAGAAAAATCTGGTAGAACAGCACATTCAGGACATTGTGGTCCACTACACGTTC AACAAGGTGCTCATGCTGCAGGAGCCCCTGCTGGTGGTGGCGGCCTTCTACATCCTGTTCTTCACCGTTATCATCTATGTTCGGCTGGAC TTCTCCATCACCAAGGATCCAGCCGCAGAAGCCAGGATGAAGGTAGCCTGCATCACAGAGCAGGTCTTGACCCTGGTCAACAAGAGAATA GGCCTTTACCGTCACTTTGACGAGACCGTCAATAGGTACAAGCAATCCCGGGACATCTCCACCCTCAACAGTGGCAAGAAGAGCCTGGAG ACTGAACACAAGGCCTTGACCAGTGAGATTGCACTGCTGCAGTCCAGGCTGAAGACAGAGGGCTCTGATCTGTGCGACAGAGTGAGCGAA ATGCAGAAGCTGGATGCACAGGTCAAGGAGCTGGTGCTGAAGTCGGCGGTGGAGGCTGAGCGCCTGGTGGCTGGCAAGCTCAAGAAAGAC ACGTACATTGAGAATGAGAAGCTCATCTCAGGAAAGCGCCAGGAGCTGGTCACCAAGATCGACCACATCCTGGATGCCCTGTAGCCCCTG CCCGCATCCTCCAGGGGGCCCAGGGTGCCTGCACTTTGCTGTGGCAGGCAGATTGGGTGGTAGTGGGAGGTTGTGCATGGAGGCCAGTGA AAGCTGACATCTGTAAAAGGCCTTCAAGGAAGAGAAACCAGGCCCTGCGTCAGGCAGTGTGAGTTTGCCGTTTGTCCTTAACTTTCTTTT TTTTTTTTTTTAAAAAAAGAAAACTTTAAAAAAACTCCCATTAAAAACAAAACATCTTTGTGTTTTGAACAAAGGAATTTTCAATATTTG ATTGGTATTCTGTTCTGAAGTCTAAGATATTTTTCAGCCTATAAAGCCCCCTGTTTTATGCCCTTCTAATTCTGATGTTTGGGTATTGTG >27616_27616_1_ESYT3-RPN1_ESYT3_chr3_138153967_ENST00000289135_RPN1_chr3_128363826_ENST00000296255_length(amino acids)=629AA_BP=109 MRAEEPCAPGAPSALGAQRTPGPELRLSSQLLPELCTFVVRVLFYLGPVYLAGYLGLSITWLLLGALLWMWWRRNRRGKLGRLAAAFEFL DNEREFISRELRGQHLPAWVKGEDEEENNLEVRETKIKGKSGRFFTVKLPVALDPGAKISVIVETVYTHVLHPYPTQITQSEKQFVVFEG NHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPTRSEDLLDYGPFRDVPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENV DLKHTGAVLKGPFSRYDYQRQPDSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDSVEMEIRPRFPLFGGWKTHYIVGYNLPS YEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVKIILPEGAKNIEIDSPYEISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHY TFNKVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARMKVACITEQVLTLVNKRIGLYRHFDETVNRYKQSRDISTLNSGKKS -------------------------------------------------------------- >27616_27616_2_ESYT3-RPN1_ESYT3_chr3_138153967_ENST00000389567_RPN1_chr3_128363826_ENST00000296255_length(transcript)=2513nt_BP=513nt GGTGCATTTCCAGGCGCTGCTCTCCGTCGCAGAGAACCCTGAGCTCGGCGCGCCGAGAGTCCCAGCAGGGCAAGGGGGCGCGGCGTCCTG GTCCTCGAGCTTGGGAGACAGATGCGCATGGGCGTGGGGGCATGCGGACCTAAGCTCGGGTGAAGCTCTCGGGAAGGGCAAGACTGCGGC GACGAGATGCGAGCAGAGGAGCCCTGCGCCCCCGGGGCCCCCAGCGCCCTGGGAGCCCAGCGCACGCCGGGCCCCGAGCTGCGCCTGTCC AGCCAGCTGCTGCCCGAGCTCTGTACCTTCGTGGTGCGCGTGCTGTTCTACCTGGGGCCTGTCTACCTAGCTGGCTACCTGGGGCTCAGC ATAACCTGGTTGCTGCTCGGCGCCCTGCTGTGGATGTGGTGGCGCAGGAACCGCCGCGGGAAGCTTGGGCGCCTGGCCGCCGCCTTCGAA TTCCTTGACAATGAACGCGAGTTCATCAGCCGCGAGCTGCGGGGCCAGCACCTGCCAGCCTGGGTAAAGGGAGAAGATGAGGAAGAGAAC AATTTGGAAGTACGTGAAACCAAAATTAAGGGTAAAAGTGGGAGATTCTTCACAGTCAAGCTCCCAGTTGCTCTTGATCCTGGGGCCAAG ATTTCAGTCATTGTGGAAACAGTCTACACCCATGTGCTTCATCCGTATCCAACCCAGATCACCCAGTCAGAGAAACAGTTTGTGGTGTTT GAGGGGAACCATTATTTCTACTCTCCCTATCCAACGAAGACACAAACCATGCGTGTGAAGCTTGCCTCTCGAAATGTGGAGAGCTACACC AAGCTGGGGAACCCCACGCGCTCTGAGGACCTACTGGATTATGGGCCTTTCAGAGATGTGCCTGCCTATAGTCAGGATACTTTTAAAGTA CATTATGAGAACAACAGCCCTTTCCTGACCATCACCAGCATGACCCGAGTCATTGAAGTCTCTCACTGGGGTAATATTGCTGTGGAAGAA AATGTGGACTTAAAGCACACAGGAGCTGTGCTTAAGGGGCCTTTCTCACGCTATGATTACCAGAGACAGCCAGATAGTGGAATATCCTCC ATCCGTTCTTTTAAGACCATCCTTCCTGCTGCTGCCCAGGATGTTTATTACCGGGATGAGATTGGCAATGTTTCTACCAGCCACCTCCTT ATTTTGGATGACTCTGTAGAGATGGAAATCCGGCCTCGCTTCCCTCTCTTTGGCGGGTGGAAGACCCATTACATCGTTGGCTACAACCTC CCAAGCTATGAGTACCTCTATAATTTGGGTGACCAGTATGCACTGAAGATGAGGTTTGTGGACCATGTGTTTGATGAACAAGTGATAGAT TCTCTGACTGTGAAGATCATCCTGCCTGAAGGAGCCAAGAACATTGAAATTGATAGTCCCTATGAAATCAGCCGTGCCCCAGATGAGCTG CACTACACCTATCTGGACACATTTGGCCGCCCTGTGATTGTTGCCTACAAGAAAAATCTGGTAGAACAGCACATTCAGGACATTGTGGTC CACTACACGTTCAACAAGGTGCTCATGCTGCAGGAGCCCCTGCTGGTGGTGGCGGCCTTCTACATCCTGTTCTTCACCGTTATCATCTAT GTTCGGCTGGACTTCTCCATCACCAAGGATCCAGCCGCAGAAGCCAGGATGAAGGTAGCCTGCATCACAGAGCAGGTCTTGACCCTGGTC AACAAGAGAATAGGCCTTTACCGTCACTTTGACGAGACCGTCAATAGGTACAAGCAATCCCGGGACATCTCCACCCTCAACAGTGGCAAG AAGAGCCTGGAGACTGAACACAAGGCCTTGACCAGTGAGATTGCACTGCTGCAGTCCAGGCTGAAGACAGAGGGCTCTGATCTGTGCGAC AGAGTGAGCGAAATGCAGAAGCTGGATGCACAGGTCAAGGAGCTGGTGCTGAAGTCGGCGGTGGAGGCTGAGCGCCTGGTGGCTGGCAAG CTCAAGAAAGACACGTACATTGAGAATGAGAAGCTCATCTCAGGAAAGCGCCAGGAGCTGGTCACCAAGATCGACCACATCCTGGATGCC CTGTAGCCCCTGCCCGCATCCTCCAGGGGGCCCAGGGTGCCTGCACTTTGCTGTGGCAGGCAGATTGGGTGGTAGTGGGAGGTTGTGCAT GGAGGCCAGTGAAAGCTGACATCTGTAAAAGGCCTTCAAGGAAGAGAAACCAGGCCCTGCGTCAGGCAGTGTGAGTTTGCCGTTTGTCCT TAACTTTCTTTTTTTTTTTTTTTAAAAAAAGAAAACTTTAAAAAAACTCCCATTAAAAACAAAACATCTTTGTGTTTTGAACAAAGGAAT TTTCAATATTTGATTGGTATTCTGTTCTGAAGTCTAAGATATTTTTCAGCCTATAAAGCCCCCTGTTTTATGCCCTTCTAATTCTGATGT >27616_27616_2_ESYT3-RPN1_ESYT3_chr3_138153967_ENST00000389567_RPN1_chr3_128363826_ENST00000296255_length(amino acids)=629AA_BP=109 MRAEEPCAPGAPSALGAQRTPGPELRLSSQLLPELCTFVVRVLFYLGPVYLAGYLGLSITWLLLGALLWMWWRRNRRGKLGRLAAAFEFL DNEREFISRELRGQHLPAWVKGEDEEENNLEVRETKIKGKSGRFFTVKLPVALDPGAKISVIVETVYTHVLHPYPTQITQSEKQFVVFEG NHYFYSPYPTKTQTMRVKLASRNVESYTKLGNPTRSEDLLDYGPFRDVPAYSQDTFKVHYENNSPFLTITSMTRVIEVSHWGNIAVEENV DLKHTGAVLKGPFSRYDYQRQPDSGISSIRSFKTILPAAAQDVYYRDEIGNVSTSHLLILDDSVEMEIRPRFPLFGGWKTHYIVGYNLPS YEYLYNLGDQYALKMRFVDHVFDEQVIDSLTVKIILPEGAKNIEIDSPYEISRAPDELHYTYLDTFGRPVIVAYKKNLVEQHIQDIVVHY TFNKVLMLQEPLLVVAAFYILFFTVIIYVRLDFSITKDPAAEARMKVACITEQVLTLVNKRIGLYRHFDETVNRYKQSRDISTLNSGKKS -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for ESYT3-RPN1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ESYT3-RPN1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ESYT3-RPN1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
