
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:EYS-CAMSAP3 (FusionGDB2 ID:28080) |
Fusion Gene Summary for EYS-CAMSAP3 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: EYS-CAMSAP3 | Fusion gene ID: 28080 | Hgene | Tgene | Gene symbol | EYS | CAMSAP3 | Gene ID | 346007 | 57662 |
| Gene name | eyes shut homolog | calmodulin regulated spectrin associated protein family member 3 | |
| Synonyms | C6orf178|C6orf179|C6orf180|EGFL10|EGFL11|RP25|SPAM|bA166P24.2|bA307F22.3|bA74E24.1|dJ1018A4.2|dJ22I17.2|dJ303F19.1 | KIAA1543|NEZHA|PPP1R80 | |
| Cytomap | 6q12 | 19p13.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein eyes shut homologEGF-like-domain, multiple 10EGF-like-domain, multiple 11epidermal growth factor-like protein 10epidermal growth factor-like protein 11protein spacemaker homolog | calmodulin-regulated spectrin-associated protein 3protein phosphatase 1, regulatory subunit 80 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q5T1H1 | Q9P1Y5 | |
| Ensembl transtripts involved in fusion gene | ENST00000370621, ENST00000503581, ENST00000370616, ENST00000342421, ENST00000370618, ENST00000393380, ENST00000486069, | ENST00000160298, ENST00000446248, | |
| Fusion gene scores | * DoF score | 11 X 11 X 4=484 | 3 X 3 X 3=27 |
| # samples | 10 | 3 | |
| ** MAII score | log2(10/484*10)=-2.27500704749987 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(3/27*10)=0.15200309344505 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: EYS [Title/Abstract] AND CAMSAP3 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | EYS(65300116)-CAMSAP3(7670112), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | EYS-CAMSAP3 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | CAMSAP3 | GO:0000226 | microtubule cytoskeleton organization | 24486153|27693509 |
| Tgene | CAMSAP3 | GO:0010923 | negative regulation of phosphatase activity | 19389623 |
| Tgene | CAMSAP3 | GO:0030334 | regulation of cell migration | 27693509 |
| Tgene | CAMSAP3 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity | 26715742|27802168 |
| Tgene | CAMSAP3 | GO:0031113 | regulation of microtubule polymerization | 24486153 |
| Tgene | CAMSAP3 | GO:0045198 | establishment of epithelial cell apical/basal polarity | 26715742|27802168 |
| Tgene | CAMSAP3 | GO:0051893 | regulation of focal adhesion assembly | 27693509 |
| Tgene | CAMSAP3 | GO:1903358 | regulation of Golgi organization | 28089391 |
 Fusion gene breakpoints across EYS (5'-gene) Fusion gene breakpoints across EYS (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
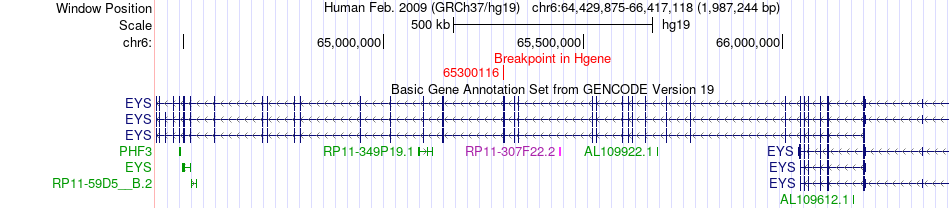 |
 Fusion gene breakpoints across CAMSAP3 (3'-gene) Fusion gene breakpoints across CAMSAP3 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
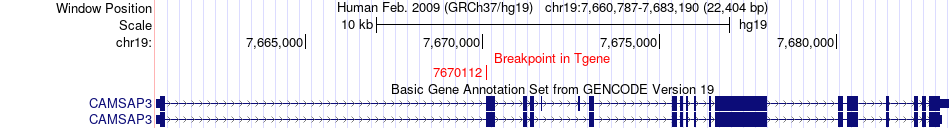 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LGG | TCGA-TM-A7CF | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
Top |
Fusion Gene ORF analysis for EYS-CAMSAP3 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000370621 | ENST00000160298 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| Frame-shift | ENST00000370621 | ENST00000446248 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| Frame-shift | ENST00000503581 | ENST00000160298 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| Frame-shift | ENST00000503581 | ENST00000446248 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| In-frame | ENST00000370616 | ENST00000160298 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| In-frame | ENST00000370616 | ENST00000446248 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| intron-3CDS | ENST00000342421 | ENST00000160298 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| intron-3CDS | ENST00000342421 | ENST00000446248 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| intron-3CDS | ENST00000370618 | ENST00000160298 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| intron-3CDS | ENST00000370618 | ENST00000446248 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| intron-3CDS | ENST00000393380 | ENST00000160298 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| intron-3CDS | ENST00000393380 | ENST00000446248 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| intron-3CDS | ENST00000486069 | ENST00000160298 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
| intron-3CDS | ENST00000486069 | ENST00000446248 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000370616 | EYS | chr6 | 65300116 | - | ENST00000160298 | CAMSAP3 | chr19 | 7670112 | + | 9284 | 5644 | 0 | 9245 | 3081 |
| ENST00000370616 | EYS | chr6 | 65300116 | - | ENST00000446248 | CAMSAP3 | chr19 | 7670112 | + | 9574 | 5644 | 0 | 9326 | 3108 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000370616 | ENST00000160298 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + | 0.001044544 | 0.9989555 |
| ENST00000370616 | ENST00000446248 | EYS | chr6 | 65300116 | - | CAMSAP3 | chr19 | 7670112 | + | 0.000967962 | 0.999032 |
Top |
Fusion Genomic Features for EYS-CAMSAP3 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| EYS | chr6 | 65300115 | - | CAMSAP3 | chr19 | 7670111 | + | 8.86E-07 | 0.99999917 |
| EYS | chr6 | 65300115 | - | CAMSAP3 | chr19 | 7670111 | + | 8.86E-07 | 0.99999917 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
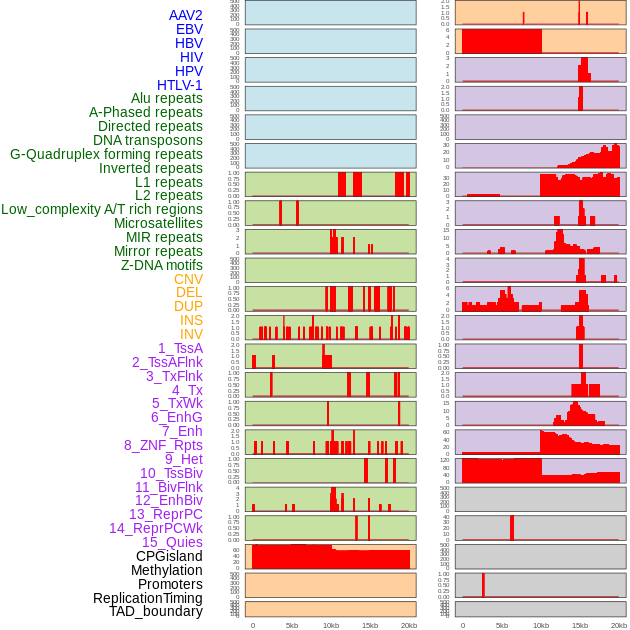 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
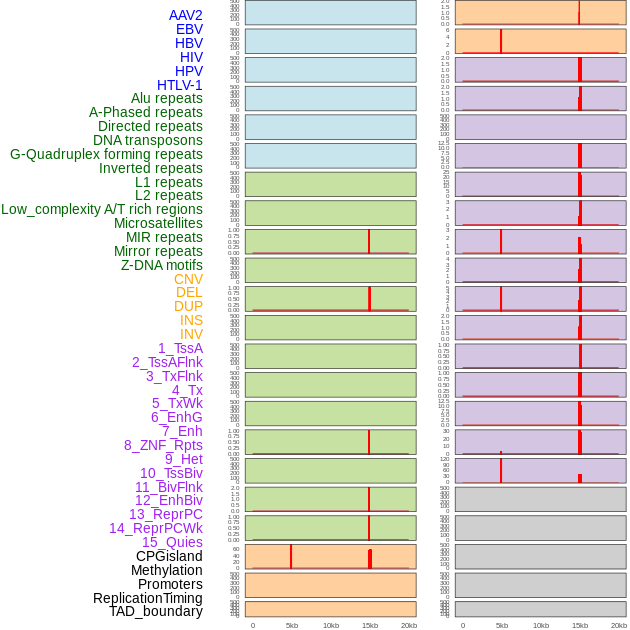 |
Top |
Fusion Protein Features for EYS-CAMSAP3 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr6:65300116/chr19:7670112) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| EYS | CAMSAP3 |
| FUNCTION: Required to maintain the integrity of photoreceptor cells (PubMed:18836446). Specifically required for normal morphology of the photoreceptor ciliary pocket, and might thus facilitate protein trafficking between the photoreceptor inner and outer segments via the transition zone (By similarity). {ECO:0000250|UniProtKB:B8JI71, ECO:0000269|PubMed:18836446}. | FUNCTION: Key microtubule-organizing protein that specifically binds the minus-end of non-centrosomal microtubules and regulates their dynamics and organization (PubMed:19041755, PubMed:23169647). Specifically recognizes growing microtubule minus-ends and autonomously decorates and stabilizes microtubule lattice formed by microtubule minus-end polymerization (PubMed:24486153). Acts on free microtubule minus-ends that are not capped by microtubule-nucleating proteins or other factors and protects microtubule minus-ends from depolymerization (PubMed:24486153). In addition, it also reduces the velocity of microtubule polymerization (PubMed:24486153). Required for the biogenesis and the maintenance of zonula adherens by anchoring the minus-end of microtubules to zonula adherens and by recruiting the kinesin KIFC3 to those junctional sites (PubMed:19041755). Required for orienting the apical-to-basal polarity of microtubules in epithelial cells: acts by tethering non-centrosomal microtubules to the apical cortex, leading to their longitudinal orientation (PubMed:27802168, PubMed:26715742). Plays a key role in early embryos, which lack centrosomes: accumulates at the microtubule bridges that connect pairs of cells and enables the formation of a non-centrosomal microtubule-organizing center that directs intracellular transport in the early embryo (By similarity). Couples non-centrosomal microtubules with actin: interaction with MACF1 at the minus ends of non-centrosomal microtubules, tethers the microtubules to actin filaments, regulating focal adhesion size and cell migration (PubMed:27693509). Plays a key role in the generation of non-centrosomal microtubules by accumulating in the pericentrosomal region and cooperating with KATNA1 to release non-centrosomal microtubules from the centrosome (PubMed:28386021). Through the microtubule cytoskeleton, also regulates the organization of cellular organelles including the Golgi and the early endosomes (PubMed:28089391). Through interaction with AKAP9, involved in translocation of Golgi vesicles in epithelial cells, where microtubules are mainly non-centrosomal (PubMed:28089391). Plays an important role in motile cilia function by facilitatating proper orientation of basal bodies and formation of central microtubule pairs in motile cilia (By similarity). {ECO:0000250|UniProtKB:Q80VC9, ECO:0000269|PubMed:19041755, ECO:0000269|PubMed:23169647, ECO:0000269|PubMed:24486153, ECO:0000269|PubMed:26715742, ECO:0000269|PubMed:27693509, ECO:0000269|PubMed:27802168, ECO:0000269|PubMed:28089391, ECO:0000269|PubMed:28386021}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 1004_1040 | 1881 | 3166.0 | Domain | EGF-like 16%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 1042_1077 | 1881 | 3166.0 | Domain | EGF-like 17 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 1079_1115 | 1881 | 3166.0 | Domain | EGF-like 18 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 1117_1159 | 1881 | 3166.0 | Domain | EGF-like 19 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 1161_1197 | 1881 | 3166.0 | Domain | EGF-like 20%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 170_212 | 1881 | 3166.0 | Domain | EGF-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 213_254 | 1881 | 3166.0 | Domain | EGF-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 256_292 | 1881 | 3166.0 | Domain | EGF-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 332_368 | 1881 | 3166.0 | Domain | EGF-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 370_406 | 1881 | 3166.0 | Domain | EGF-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 567_602 | 1881 | 3166.0 | Domain | EGF-like 6 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 643_679 | 1881 | 3166.0 | Domain | EGF-like 7 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 681_720 | 1881 | 3166.0 | Domain | EGF-like 8%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 733_769 | 1881 | 3166.0 | Domain | EGF-like 9%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 771_807 | 1881 | 3166.0 | Domain | EGF-like 10%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 809_847 | 1881 | 3166.0 | Domain | EGF-like 11 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 849_888 | 1881 | 3166.0 | Domain | EGF-like 12 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 890_926 | 1881 | 3166.0 | Domain | EGF-like 13 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 928_964 | 1881 | 3166.0 | Domain | EGF-like 14%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 966_1002 | 1881 | 3166.0 | Domain | EGF-like 15 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 1004_1040 | 1881 | 3166.0 | Domain | EGF-like 16%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 1042_1077 | 1881 | 3166.0 | Domain | EGF-like 17 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 1079_1115 | 1881 | 3166.0 | Domain | EGF-like 18 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 1117_1159 | 1881 | 3166.0 | Domain | EGF-like 19 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 1161_1197 | 1881 | 3166.0 | Domain | EGF-like 20%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 170_212 | 1881 | 3166.0 | Domain | EGF-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 213_254 | 1881 | 3166.0 | Domain | EGF-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 256_292 | 1881 | 3166.0 | Domain | EGF-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 332_368 | 1881 | 3166.0 | Domain | EGF-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 370_406 | 1881 | 3166.0 | Domain | EGF-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 567_602 | 1881 | 3166.0 | Domain | EGF-like 6 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 643_679 | 1881 | 3166.0 | Domain | EGF-like 7 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 681_720 | 1881 | 3166.0 | Domain | EGF-like 8%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 733_769 | 1881 | 3166.0 | Domain | EGF-like 9%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 771_807 | 1881 | 3166.0 | Domain | EGF-like 10%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 809_847 | 1881 | 3166.0 | Domain | EGF-like 11 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 849_888 | 1881 | 3166.0 | Domain | EGF-like 12 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 890_926 | 1881 | 3166.0 | Domain | EGF-like 13 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 928_964 | 1881 | 3166.0 | Domain | EGF-like 14%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 966_1002 | 1881 | 3166.0 | Domain | EGF-like 15 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 1004_1040 | 1881 | 3145.0 | Domain | EGF-like 16%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 1042_1077 | 1881 | 3145.0 | Domain | EGF-like 17 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 1079_1115 | 1881 | 3145.0 | Domain | EGF-like 18 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 1117_1159 | 1881 | 3145.0 | Domain | EGF-like 19 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 1161_1197 | 1881 | 3145.0 | Domain | EGF-like 20%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 170_212 | 1881 | 3145.0 | Domain | EGF-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 213_254 | 1881 | 3145.0 | Domain | EGF-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 256_292 | 1881 | 3145.0 | Domain | EGF-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 332_368 | 1881 | 3145.0 | Domain | EGF-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 370_406 | 1881 | 3145.0 | Domain | EGF-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 567_602 | 1881 | 3145.0 | Domain | EGF-like 6 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 643_679 | 1881 | 3145.0 | Domain | EGF-like 7 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 681_720 | 1881 | 3145.0 | Domain | EGF-like 8%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 733_769 | 1881 | 3145.0 | Domain | EGF-like 9%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 771_807 | 1881 | 3145.0 | Domain | EGF-like 10%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 809_847 | 1881 | 3145.0 | Domain | EGF-like 11 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 849_888 | 1881 | 3145.0 | Domain | EGF-like 12 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 890_926 | 1881 | 3145.0 | Domain | EGF-like 13 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 928_964 | 1881 | 3145.0 | Domain | EGF-like 14%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 966_1002 | 1881 | 3145.0 | Domain | EGF-like 15 |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000160298 | 0 | 17 | 594_628 | 49 | 1250.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000160298 | 0 | 17 | 696_729 | 49 | 1250.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000160298 | 0 | 17 | 896_935 | 49 | 1250.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000446248 | 0 | 19 | 594_628 | 49 | 1277.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000446248 | 0 | 19 | 696_729 | 49 | 1277.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000446248 | 0 | 19 | 896_935 | 49 | 1277.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000160298 | 0 | 17 | 437_548 | 49 | 1250.0 | Compositional bias | Note=Pro-rich | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000160298 | 0 | 17 | 730_841 | 49 | 1250.0 | Compositional bias | Note=Pro-rich | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000446248 | 0 | 19 | 437_548 | 49 | 1277.0 | Compositional bias | Note=Pro-rich | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000446248 | 0 | 19 | 730_841 | 49 | 1277.0 | Compositional bias | Note=Pro-rich | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000160298 | 0 | 17 | 1109_1243 | 49 | 1250.0 | Domain | CKK | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000160298 | 0 | 17 | 203_312 | 49 | 1250.0 | Domain | Calponin-homology (CH) | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000446248 | 0 | 19 | 1109_1243 | 49 | 1277.0 | Domain | CKK | |
| Tgene | CAMSAP3 | chr6:65300116 | chr19:7670112 | ENST00000446248 | 0 | 19 | 203_312 | 49 | 1277.0 | Domain | Calponin-homology (CH) |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 1004_1040 | 0 | 595.0 | Domain | EGF-like 16%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 1042_1077 | 0 | 595.0 | Domain | EGF-like 17 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 1079_1115 | 0 | 595.0 | Domain | EGF-like 18 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 1117_1159 | 0 | 595.0 | Domain | EGF-like 19 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 1161_1197 | 0 | 595.0 | Domain | EGF-like 20%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 170_212 | 0 | 595.0 | Domain | EGF-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 1883_2063 | 0 | 595.0 | Domain | Laminin G-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2099_2140 | 0 | 595.0 | Domain | EGF-like 21 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 213_254 | 0 | 595.0 | Domain | EGF-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2145_2339 | 0 | 595.0 | Domain | Laminin G-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2335_2368 | 0 | 595.0 | Domain | EGF-like 22 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2371_2408 | 0 | 595.0 | Domain | EGF-like 23 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2419_2609 | 0 | 595.0 | Domain | Laminin G-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 256_292 | 0 | 595.0 | Domain | EGF-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2610_2646 | 0 | 595.0 | Domain | EGF-like 24 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2648_2689 | 0 | 595.0 | Domain | EGF-like 25 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2717_2895 | 0 | 595.0 | Domain | Laminin G-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2896_2932 | 0 | 595.0 | Domain | EGF-like 26 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2933_2970 | 0 | 595.0 | Domain | EGF-like 27 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 2975_3165 | 0 | 595.0 | Domain | Laminin G-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 332_368 | 0 | 595.0 | Domain | EGF-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 370_406 | 0 | 595.0 | Domain | EGF-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 567_602 | 0 | 595.0 | Domain | EGF-like 6 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 643_679 | 0 | 595.0 | Domain | EGF-like 7 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 681_720 | 0 | 595.0 | Domain | EGF-like 8%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 733_769 | 0 | 595.0 | Domain | EGF-like 9%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 771_807 | 0 | 595.0 | Domain | EGF-like 10%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 809_847 | 0 | 595.0 | Domain | EGF-like 11 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 849_888 | 0 | 595.0 | Domain | EGF-like 12 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 890_926 | 0 | 595.0 | Domain | EGF-like 13 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 928_964 | 0 | 595.0 | Domain | EGF-like 14%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000342421 | - | 1 | 9 | 966_1002 | 0 | 595.0 | Domain | EGF-like 15 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 1883_2063 | 1881 | 3166.0 | Domain | Laminin G-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2099_2140 | 1881 | 3166.0 | Domain | EGF-like 21 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2145_2339 | 1881 | 3166.0 | Domain | Laminin G-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2335_2368 | 1881 | 3166.0 | Domain | EGF-like 22 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2371_2408 | 1881 | 3166.0 | Domain | EGF-like 23 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2419_2609 | 1881 | 3166.0 | Domain | Laminin G-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2610_2646 | 1881 | 3166.0 | Domain | EGF-like 24 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2648_2689 | 1881 | 3166.0 | Domain | EGF-like 25 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2717_2895 | 1881 | 3166.0 | Domain | Laminin G-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2896_2932 | 1881 | 3166.0 | Domain | EGF-like 26 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2933_2970 | 1881 | 3166.0 | Domain | EGF-like 27 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370616 | - | 23 | 41 | 2975_3165 | 1881 | 3166.0 | Domain | Laminin G-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 1004_1040 | 0 | 595.0 | Domain | EGF-like 16%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 1042_1077 | 0 | 595.0 | Domain | EGF-like 17 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 1079_1115 | 0 | 595.0 | Domain | EGF-like 18 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 1117_1159 | 0 | 595.0 | Domain | EGF-like 19 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 1161_1197 | 0 | 595.0 | Domain | EGF-like 20%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 170_212 | 0 | 595.0 | Domain | EGF-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 1883_2063 | 0 | 595.0 | Domain | Laminin G-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2099_2140 | 0 | 595.0 | Domain | EGF-like 21 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 213_254 | 0 | 595.0 | Domain | EGF-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2145_2339 | 0 | 595.0 | Domain | Laminin G-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2335_2368 | 0 | 595.0 | Domain | EGF-like 22 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2371_2408 | 0 | 595.0 | Domain | EGF-like 23 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2419_2609 | 0 | 595.0 | Domain | Laminin G-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 256_292 | 0 | 595.0 | Domain | EGF-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2610_2646 | 0 | 595.0 | Domain | EGF-like 24 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2648_2689 | 0 | 595.0 | Domain | EGF-like 25 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2717_2895 | 0 | 595.0 | Domain | Laminin G-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2896_2932 | 0 | 595.0 | Domain | EGF-like 26 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2933_2970 | 0 | 595.0 | Domain | EGF-like 27 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 2975_3165 | 0 | 595.0 | Domain | Laminin G-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 332_368 | 0 | 595.0 | Domain | EGF-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 370_406 | 0 | 595.0 | Domain | EGF-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 567_602 | 0 | 595.0 | Domain | EGF-like 6 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 643_679 | 0 | 595.0 | Domain | EGF-like 7 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 681_720 | 0 | 595.0 | Domain | EGF-like 8%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 733_769 | 0 | 595.0 | Domain | EGF-like 9%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 771_807 | 0 | 595.0 | Domain | EGF-like 10%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 809_847 | 0 | 595.0 | Domain | EGF-like 11 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 849_888 | 0 | 595.0 | Domain | EGF-like 12 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 890_926 | 0 | 595.0 | Domain | EGF-like 13 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 928_964 | 0 | 595.0 | Domain | EGF-like 14%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370618 | - | 1 | 11 | 966_1002 | 0 | 595.0 | Domain | EGF-like 15 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 1883_2063 | 1881 | 3166.0 | Domain | Laminin G-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2099_2140 | 1881 | 3166.0 | Domain | EGF-like 21 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2145_2339 | 1881 | 3166.0 | Domain | Laminin G-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2335_2368 | 1881 | 3166.0 | Domain | EGF-like 22 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2371_2408 | 1881 | 3166.0 | Domain | EGF-like 23 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2419_2609 | 1881 | 3166.0 | Domain | Laminin G-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2610_2646 | 1881 | 3166.0 | Domain | EGF-like 24 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2648_2689 | 1881 | 3166.0 | Domain | EGF-like 25 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2717_2895 | 1881 | 3166.0 | Domain | Laminin G-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2896_2932 | 1881 | 3166.0 | Domain | EGF-like 26 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2933_2970 | 1881 | 3166.0 | Domain | EGF-like 27 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000370621 | - | 26 | 44 | 2975_3165 | 1881 | 3166.0 | Domain | Laminin G-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 1004_1040 | 0 | 620.0 | Domain | EGF-like 16%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 1042_1077 | 0 | 620.0 | Domain | EGF-like 17 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 1079_1115 | 0 | 620.0 | Domain | EGF-like 18 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 1117_1159 | 0 | 620.0 | Domain | EGF-like 19 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 1161_1197 | 0 | 620.0 | Domain | EGF-like 20%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 170_212 | 0 | 620.0 | Domain | EGF-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 1883_2063 | 0 | 620.0 | Domain | Laminin G-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2099_2140 | 0 | 620.0 | Domain | EGF-like 21 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 213_254 | 0 | 620.0 | Domain | EGF-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2145_2339 | 0 | 620.0 | Domain | Laminin G-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2335_2368 | 0 | 620.0 | Domain | EGF-like 22 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2371_2408 | 0 | 620.0 | Domain | EGF-like 23 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2419_2609 | 0 | 620.0 | Domain | Laminin G-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 256_292 | 0 | 620.0 | Domain | EGF-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2610_2646 | 0 | 620.0 | Domain | EGF-like 24 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2648_2689 | 0 | 620.0 | Domain | EGF-like 25 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2717_2895 | 0 | 620.0 | Domain | Laminin G-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2896_2932 | 0 | 620.0 | Domain | EGF-like 26 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2933_2970 | 0 | 620.0 | Domain | EGF-like 27 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 2975_3165 | 0 | 620.0 | Domain | Laminin G-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 332_368 | 0 | 620.0 | Domain | EGF-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 370_406 | 0 | 620.0 | Domain | EGF-like 5 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 567_602 | 0 | 620.0 | Domain | EGF-like 6 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 643_679 | 0 | 620.0 | Domain | EGF-like 7 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 681_720 | 0 | 620.0 | Domain | EGF-like 8%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 733_769 | 0 | 620.0 | Domain | EGF-like 9%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 771_807 | 0 | 620.0 | Domain | EGF-like 10%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 809_847 | 0 | 620.0 | Domain | EGF-like 11 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 849_888 | 0 | 620.0 | Domain | EGF-like 12 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 890_926 | 0 | 620.0 | Domain | EGF-like 13 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 928_964 | 0 | 620.0 | Domain | EGF-like 14%3B calcium-binding |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000393380 | - | 1 | 12 | 966_1002 | 0 | 620.0 | Domain | EGF-like 15 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 1883_2063 | 1881 | 3145.0 | Domain | Laminin G-like 1 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2099_2140 | 1881 | 3145.0 | Domain | EGF-like 21 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2145_2339 | 1881 | 3145.0 | Domain | Laminin G-like 2 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2335_2368 | 1881 | 3145.0 | Domain | EGF-like 22 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2371_2408 | 1881 | 3145.0 | Domain | EGF-like 23 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2419_2609 | 1881 | 3145.0 | Domain | Laminin G-like 3 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2610_2646 | 1881 | 3145.0 | Domain | EGF-like 24 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2648_2689 | 1881 | 3145.0 | Domain | EGF-like 25 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2717_2895 | 1881 | 3145.0 | Domain | Laminin G-like 4 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2896_2932 | 1881 | 3145.0 | Domain | EGF-like 26 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2933_2970 | 1881 | 3145.0 | Domain | EGF-like 27 |
| Hgene | EYS | chr6:65300116 | chr19:7670112 | ENST00000503581 | - | 26 | 43 | 2975_3165 | 1881 | 3145.0 | Domain | Laminin G-like 5 |
Top |
Fusion Gene Sequence for EYS-CAMSAP3 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >28080_28080_1_EYS-CAMSAP3_EYS_chr6_65300116_ENST00000370616_CAMSAP3_chr19_7670112_ENST00000160298_length(transcript)=9284nt_BP=5644nt ATGACTGACAAATCAATCGTCATTCTGAGCCTGATGGTTTTTCACAGCTCTTTCATAAATGGAAAAACATGTAGACGGCAATTGGTGGAA GAATGGCATCCACAACCCTCATCATATGTGGTAAATTGGACACTAACAGAAAACATCTGCTTGGACTTCTACAGAGATTGCTGGTTTTTG GGTGTAAACACTAAAATAGATACTTCAGGCAATCAAGCTGTTCCCCAGATTTGCCCTTTGCAAATTCAATTAGGAGATATCCTTGTAATT TCTTCTGAACCATCTCTTCAATTTCCAGAAATAAATTTGATGAATGTTTCTGAAACATCTTTCGTTGGCTGTGTGCAAAATACCACAACG GAAGATCAGTTACTTTTTGGCTGCAGACTAAAAGGAATGCACACTGTTAATTCTAAGTGGCTGAGTGTTGGGACACATTATTTTATCACA GTTATGGCAAGTGGTCCATCACCTTGTCCACTGGGACTTCGACTAAATGTGACAGTGAAACAGCAGTTCTGCCAGGAATCTCTGAGTTCA GAATTTTGCTCTGGTCATGGTAAATGTCTTAGTGAAGCTTGGAGCAAGACATATAGCTGCCATTGCCAGCCTCCATTTTCTGGAAAATAC TGCCAGGAACTTGATGCATGTTCTTTTAAACCATGTAAAAATAATGGCAGTTGCATTAATAAAAGAGAAAATTGGGATGAGCAAGCATAT GAATGTGTCTGTCACCCACCATTTACAGGAAAGAATTGCTCAGAAATAATTGGCCAGTGTCAACCACATGTCTGTTTCCATGGAAACTGC AGCAATATTACTTCAAATAGTTTCATTTGTGAATGTGATGAGCAATTTTCAGGTCCATTCTGTGAGGTGTCAGCAAAACCTTGTGTTTCT CTGCTTTTTTGGAAAAGAGGAATTTGCCCAAATAGCAGTTCTGCTTATACTTATGAATGCCCAAAAGGATCTTCCAGCCAAAATGGTGAA ACTGATGTCAGTGAGTTTTCATTAGTACCATGTCAGAATGGTACTGACTGCATCAAAATATCAAATGATGTTATGTGCATCTGTTCACCA ATATTTACAGATTTGCTTTGTAAGAGCATTCAAACATCATGTGAGTCATTTCCTTTGAGGAATAATGCTACATGTAAGAAATGTGAGAAA GATTATCCTTGCAGCTGTATATCAGGATTTACTGAAAAAAACTGTGAGAAAGCAATTGACCACTGTAAACTGCTCAGCATCAACTGTCTG AATGAAGAATGGTGTTTCAATATAATTGGAAGATTCAAATATGTATGCATTCCAGGGTGCACAAAAAATCCATGTTGGTTTTTGAAGAAT GTTTACCTAATTCATCAACACCTCTGCTACTGTGGAGTCACCTTCCATGGTATTTGCCAAGATAAAGGTCCTGCTCAATTTGAATATGTG TGGCAATTGGGATTTGCAGGATCTGAAGGCGAAAAGTGCCAAGGGGTTATTGATGCCTATTTCTTTCTGGCTGCAAACTGCACTGAAGAT GCAACCTATGTGAACGATCCTGAAGATAATAATTCTTCATGTTGGTTCCCACATGAAGGCACAAAAGAGATTTGTGCAAATGGATGCAGT TGTTTGAGTGAAGAAGACAGTCAGGAATATCGGTATCTATGTTTTCTCAGATGGGCTGGCAACATGTATCTGGAAAATACAACTGATGAT CAAGAAAATGAGTGTCAACATGAAGCTGTTTGTAAAGATGAAATTAATAGACCCAGATGCAGCTGTTCTCTTAGTTACATTGGCAGATTG TGTGTTGTCAATGTTGACTATTGCTTAGGGAACCACAGTATATCAGTGCATGGCCTCTGCCTGGCCCTTTCGCACAATTGTAACTGTAGC GGTCTGCAAAGATATGAAAGGAACATCTGTGAGATAGATACTGAAGACTGCAAATCTGCGTCCTGCAAAAATGGAACAACTAGTACACAT TTAAGGGGATATTTCTTCCGCAAGTGTGTCCCAGGATTTAAAGGTACGCAATGTGAAATTGATATAGATGAGTGTGCTTCACATCCCTGC AAAAATGGAGCCACCTGCATTGACCAACCTGGTAATTACTTCTGCCAGTGTGTGCCTCCATTTAAAGTGGTTGATGGCTTTTCCTGCTTA TGCAATCCAGGCTATGTTGGGATAAGATGTGAACAGGACATTGATGACTGCATCCTGAATGCCTGTGAGCACAATTCTACCTGCAAAGAC CTGCATCTCAGCTACCAGTGTGTGTGCCTATCTGATTGGGAAGGAAATTTTTGTGAACAAGAATCCAATGAGTGTAAAATGAATCCTTGC AAGAACAATTCCACCTGTACTGACCTTTACAAAAGCTATCGATGTGAGTGTACATCTGGATGGACTGGACAGAACTGTAGTGAAGAAATA AATGAATGCGACTCTGATCCATGCATGAATGGAGGTCTTTGTCATGAATCTACCATCCCTGGACAATTTGTATGTCTGTGCCCACCCCTT TATACTGGACAATTTTGCCACCAACGCTATAACCTTTGTGACCTACTTCATAACCCTTGCAGAAACAACTCAACATGCTTAGCTTTGGTA GACGCAAATCAGCATTGTATTTGTAGAGAAGAATTTGAAGGTAAAAACTGTGAAATTGATGTGAAAGACTGCCTCTTCCTTTCCTGCCAG GATTATGGTGACTGTGAAGATATGGTCAACAATTTCAGGTGTATTTGCAGACCTGGGTTTTCTGGATCTCTGTGTGAAATTGAAATTAAT GAATGTTCCTCTGAACCTTGCAAAAATAATGGAACATGTGTGGATCTGACAAACAGATTTTTTTGTAATTGTGAACCTGAGTACCATGGG CCCTTCTGTGAACTTGATGTAAATAAATGTAAAATCTCACCTTGTCTAGATGAAGAAAATTGTGTCTACAGGACTGATGGATACAACTGC CTCTGTGCCCCTGGTTATACAGGCATCAACTGTGAAATAAATCTAGATGAATGCCTATCAGAGCCCTGTCTCCATGATGGAGTTTGTATC GATGGCATCAATCATTATACCTGTGACTGCAAGAGTGGGTTTTTTGGAACACACTGTGAAACAAACGCCAATGATTGCCTTTCAAATCCT TGTCTACATGGAAGGTGCACAGAACTTATTAATGAATATCCATGTTCATGTGATGCAGATGGGACTAGCACACAATGTAAGATCAAAATT AATGACTGCACATCAATCCCTTGTATGAATGAAGGCTTCTGTCAGAAGTCAGCACATGGATTTACTTGCATTTGCCCACGTGGATACACT GGTGCATACTGTGAAAAAAGCATTGATAATTGTGCTGAGCCTGAACTTAATTCAGTCATCTGTCTTAATGGAGGGATCTGTGTTGATGGG CCTGGACATACTTTTGACTGCAGATGTCTTCCTGGATTTTCTGGTCAATTTTGTGAAATTAATATAAATGAATGCTCTTCATCACCATGT CTACATGGTGCAGACTGTGAAGATCACATCAATGGATATGTTTGCAAATGCCAACCAGGATGGTCTGGACACCACTGTGAGAATGAGCTT GAGTGCATTCCCAACTCATGTGTTCATGAACTCTGCATGGAGAATGAACCTGGCTCGACATGTTTATGCACACCTGGATTTATGACCTGC TCCATTGGGCTTCTTTGTGGTGATGAAATAAGGAGAATTACCTGTTTAACTCCCATCTTTCAAAGAACAGATCCCATTTCCACACAGACA TATACAATTCCCCCTTCTGAGACTTTGGTCAGCAGCTTTCCATCTATAAAGGCTACTAGAATACCAGCCATAATGGACACTTACCCAGTT GATCAAGGTCCAAAACAGACAGGCATTGTCAAGCACGACATTCTCCCAACCACTGGTTTGGCAACATTAAGAATTAGCACACCCTTGGAA AGCTACTTACTCCAAGAACTGATTGTCACTAGAGAGCTTTCAGCAAAACACAGTCTTCTTTCTTCCGCAGATGTTTCCTCTTCTCGATTC CTGAATTTTGGTATTCGTGACCCAGCACAAATTGTCCAGGACAAAACATCGGTATCACATATGCCTATTCGAACTTCAGCAGCCACACTA GGTTTCTTTTTTCCTGATAGGAGAGCAAGGACCCCATTTATCATGTCTTCCTTAATGTCAGATTTTATTTTTCCTACACAATCTTTATTA TTTGAGAACTGTCAGACTGTTGCTTTATCTGCTACCCCAACGACTTCAGTAATTAGAAGCATTCCAGGGGCTGATATTGAGCTAAACAGG CAGTCATTACTCTCCCGTGGATTCCTGCTTATAGCTGCCTCCATAAGTGCAACTCCAGTTGTCTCTAGGGGGGCTCAAGAGGATATTGAA GAATATTCAGCTGATTCTTTAATTTCAAGAAGAGAGCACTGGAGATTGCTCAGCCCCTCGATGTCTCCCATTTTTCCTGCTAAGGTAATT ATATCTAAACAGGTAACCATCTTAAACTCATCAGCTCTGCACCGGTTCAGTACAAAAGCCTTCAATCCCAGTGAATATCAGGCTATTACT GAGGCTTCAAGCAACCAGAGACTCACAAACATCAAATCACAGGCTGCTGATTCTTTAAGAGAATTAAGCCAAACATGTGCAACATGTTCT ATGACTGAAATAAAATCCTCTCGTGAATTCTCAGATCAAGTTTTGCATAGCAAACAGTCCCACTTTTATGAGACATTCTGGATGAACTCA GCGATATTAGCCAGCTGGTATGCACTAATGGGAGCTCAAACTATCACTTCTGGGCATTCATTTTCTTCTGCTACTGAAATAACACCATCA GTGGCATTCACAGAAGTGCCATCTTTATTTCCTTCTAAAAAGAGTGCAAAAAGAACAATTTTATCCTCATCCTTGGAAGAATCCATTACC CTATCAAGTAATTTGGATGTTAATTTATGTTTGGATAAGACTTGTTTATCCATTGTCCCTTCACAAACTATCTCTTCAGACTTGATGAAT TCTGATTTGACTTCAAAAATGACTACTGATGAACTGTCAGTATCAGAAAACATTTTAAAACTATTGAAAATAAGACAATATGGCATAACT ATGGGACCCACTGAGGTACTAAATCAAGAGAGCTTATTGGACATGGAAAAAAGTAAAGGATCTCATACACTGTTCAAACTTCACCCAAGT GATAGTTCTCTGGATTTTGAGTTAAACTTACAAATTTATCCGGATGTTACTTTAAAGACATATTCAGAAATTACACATGCAAATGACTTC AAAAATAATCTGCCACCATTGACAGGCTCAGTGCCTGATTTTTCAGAAGTCACCACCAATGTTGCATTTTATACAGTTTCAGCAACTCCA GCACTTTCAATACAGACGTCTTCCTCCATGTCTGTAATTAGGCCAGATTGGCCATATTTTACAGATTATATGACCTCTCTTAAAAAAGAG GTCAAGACTTCTTCAGAATGGTCCAAATGGGAACTTCAGCCTAGTGTGCAATATCAGGAATTTCCCACTGCAAGCCGGCATCTTCCCTTC ACTAGATCTCTTACTTTGTCTTCACTGGAATCCATTCTGGCACCTCAACGGCTGATGATTTCTGAGCACGTGCCCCCGGAGCTGTGGGAG CCCTTCTATACCGACCAGTACGCGCAGGAGCATGTGAAGCCCCCGGTGACACGGCTGCTGCTCTCAGCCGAGCTCTACTGCAGAGCCTGG CGCCAGGCACTGCCACAGCTTGAAACACCCCCCAACCCCTCTGCACTGCTGGCCCTGCTGGCGCGGAGGGGCACAGTGCCTGCTTTGCCC GAGCGCCCGGTGCGCGAGGCCGACCTGAGGCACCAGCCCATTCTCATGGGAGCCCACCTAGCTGTCATTGATGCCCTCATGGCTGCCTTT GCCTTCGAGTGGACAAAGACCCTGCCAGGTCCCTTGGCCCTGACCAGCTTGGAGCACAAGCTGCTTTTCTGGGTGGACACGACCGTCCGG CGGCTGCAGGAGAAGACCGAGCAGGAAGCGGCCCAGCGAGCCTCTCCAGCAGCCCCTGCAGACGGGGCGGCCCCGGCGCAGCCCTCGATC CGATACCGCAAGGACCGTGTGGTGGCGCGACGTGCCCCCTGCTTCCCGACGGTGACCAGCCTCCAGGACCTGGCCAGTGGGGCCGCGCTG GCCGCCACCATCCACTGCTATTGTCCCCAGCTGCTTCGACTTGAGGAGGTGTGCTTGAAGGACCCCATGTCTGTGGCGGACAGCCTGTAC AACCTCCAGCTCGTGCAGGATTTCTGTGCCTCTCGCCTTCCTCGTGGCTGCCCCCTGTCCCTTGAGGACTTGCTGTACGTCCCACCGCCA CTCAAGGTCAACTTGGTGGTGATGCTGGCCGAGTTGTTCATGTGTTTTGAGGTGCTCAAGCCCGACTTTGTGCAAGTGAAGGACTTGCCC GATGGTCACGCTGCCTCCCCCCGGGGCACTGAGGCCTCCCCACCTCAGAACAACAGCGGCAGTAGTTCTCCTGTCTTCACCTTCCGCCAC CCGCTTCTGTCATCTGGTGGCCCCCAGTCCCCACTCCGAGGATCCACAGGCTCCCTGAAGTCTTCCCCGTCCATGTCCCATATGGAGGCC CTGGGCAAGGCCTGGAACCGGCAGCTCAGCCGTCCCCTCTCCCAGGCTGTGTCATTCAGCACCCCCTTTGGCCTGGACAGCGACGTGGAT GTCGTCATGGGAGACCCTGTGCTCCTCCGCTCTGTGAGCTCGGACAGCCTGGGCCCCCCGCGTCCCGCGCCGGCCAGGACCCCCACCCAG CCACCCCCGGAGCCTGGTGACCTGCCCACCATCGAGGAAGCTCTGCAGATCATCCACAGTGCCGAGCCCCGGCTCCTCCCAGATGGGGCG GCCGACGGCAGCTTCTACCTCCACTCCCCTGAGGGGCCCTCCAAGCCATCCCTGGCCTCCCCCTACCTGCCCGAGGGGACCTCCAAACCA CTGTCCGACAGGCCCACCAAAGCACCAGTGTACATGCCACACCCCGAGACCCCCTCGAAACCATCTCCCTGTCTGGTGGGGGAGGCATCG AAACCGCCAGCCCCATCCGAGGGGTCCCCGAAGGCGGTGGCTTCGTCCCCAGCAGCCACCAACTCCGAGGTGAAAATGACCAGCTTTGCA GAACGCAAGAAACAGCTGGTGAAGGCAGAGGCTGAGGCCGGAGCGGGGTCCCCCACGTCCACTCCGGCCCCGCCGGAGGCCCTGAGCTCG GAGATGAGTGAGCTCAGCGCCCGGCTGGAGGAGAAACGCAGAGCCATCGAGGCTCAGAAGCGACGGATTGAGGCCATATTCGCCAAGCAC CGCCAGCGGCTGGGCAAAAGCGCCTTCCTGCAGGTGCAGCCGCGGGAAGCCTCTGGGGAGGCGGAAGCAGAGGCGGAGGAGGCCGATTCC GGTCCAGTCCCTGGTGGGGAGCGGCCCGCAGGCGAGGGCCAGGGTGAGCCAACCTCACGGCCCAAGGCAGTGACCTTCTCGCCAGACCTG GGCCCGGTGCCCCACGAGGGGCTGGGGGAATACAATCGAGCGGTCAGCAAGCTGAGTGCCGCCTTGAGCTCGCTGCAGCGGGACATGCAG AGGCTCACGGACCAGCAGCAGCGGCTCCTGGCCCCGCCCGAGGCCCCCGGATCCGCCCCACCACCTGCTGCGTGGGTCATCCCTGGCCCC ACGACGGGGCCCAAAGCTGCATCCCCCAGCCCCGCCCGGCGAGTCCCGGCCACCCGGCGCAGCCCTGGGCCCGGGCCCAGCCAGTCACCC CGCAGCCCGAAACACACGCGGCCAGCGGAGCTGCGGCTGGCACCCTTGACCAGGGTGCTTACGCCACCCCACGACGTAGACAGCCTCCCC CACCTGCGCAAGTTCTCGCCGAGCCAGGTGCCCGTGCAGACGCGCTCTTCCATCCTCCTGGCGGAGGAGACGCCCCCCGAGGAGCCAGCC GCCCGGCCGGGCCTCATCGAGATCCCGCTGGGCAGCCTGGCAGATCCCGCCGCCGAGGACGAGGGAGACGGGAGCCCCGCTGGTGCTGAG GATTCCTTGGAGGAGGAGGCGTCTTCGGAGGGGGAGCCCCGGGTGGGGCTGGGGTTCTTCTACAAGGATGAAGACAAGCCTGAGGACGAG ATGGCCCAAAAGCGGGCCAGCCTGCTGGAGCGGCAGCAGCGGCGAGCAGAGGAGGCGCGGCGGCGCAAGCAGTGGCAGGAGGTGGAGAAG GAACAGCGGAGGGAGGAGGCCGCGAGGCTGGCCCAAGAGGAGGCCCCGGGCCCAGCCCCGCTTGTGTCCGCAGTCCCGATGGCGACTCCA GCCCCTGCTGCCCGGGCTCCAGCCGAGGAGGAGGTGGGCCCCCGGAAGGGGGACTTCACGCGGCAGGAGTACGAGCGCCGGGCCCAGCTG AAGCTGATGGACGACCTCGATAAGGTGCTGCGGCCCCGGGCTGCGGGGTCCGGGGGTCCAGGTCGGGGCGGGCGGAGGGCCACCCGGCCT CGCTCGGGTTGCTGTGACGACTCAGCCCTGGCACGAAGCCCAGCCCGCGGCCTGCTGGGCTCTCGGCTGAGCAAAATCTATTCCCAGTCC ACCCTGTCACTGTCCACTGTGGCCAACGAGGCCCACAATAACCTCGGGGTGAAGAGGCCCACGTCTCGGGCTCCCTCCCCGTCAGGTCTC ATGTCCCCAAGCCGCCTGCCTGGAAGCCGCGAACGGGACTGGGAAAATGGCAGCAATGCCTCCTCCCCAGCGTCAGTGCCCGAGTACACA GGTCCACGGCTGTACAAAGAACCCAGCGCCAAGTCCAACAAGTTCATCATCCACAATGCCCTATCACACTGCTGCCTGGCGGGCAAGGTG AACGAACCGCAGAAGAATCGCATTCTGGAGGAAATTGAGAAAAGCAAGGCCAACCACTTCCTGATCCTCTTTCGCGACTCGAGCTGCCAG TTCCGGGCGCTCTACACGCTGTCGGGGGAGACAGAGGAGCTGTCGCGGCTGGCAGGGTATGGGCCCCGGACCGTCACGCCCGCCATGGTG GAAGGCATCTACAAGTACAACTCGGACCGCAAGCGCTTCACCCAGATCCCCGCCAAGACCATGTCCATGAGCGTCGATGCCTTCACCATC CAGGGACACCTCTGGCAGGGCAAGAAACCCACCACTCCCAAGAAGGGCGGCGGCACCCCCAAATAGCCCCACCCGGGCGGTCCACGGGCC >28080_28080_1_EYS-CAMSAP3_EYS_chr6_65300116_ENST00000370616_CAMSAP3_chr19_7670112_ENST00000160298_length(amino acids)=3081AA_BP=1881 MTDKSIVILSLMVFHSSFINGKTCRRQLVEEWHPQPSSYVVNWTLTENICLDFYRDCWFLGVNTKIDTSGNQAVPQICPLQIQLGDILVI SSEPSLQFPEINLMNVSETSFVGCVQNTTTEDQLLFGCRLKGMHTVNSKWLSVGTHYFITVMASGPSPCPLGLRLNVTVKQQFCQESLSS EFCSGHGKCLSEAWSKTYSCHCQPPFSGKYCQELDACSFKPCKNNGSCINKRENWDEQAYECVCHPPFTGKNCSEIIGQCQPHVCFHGNC SNITSNSFICECDEQFSGPFCEVSAKPCVSLLFWKRGICPNSSSAYTYECPKGSSSQNGETDVSEFSLVPCQNGTDCIKISNDVMCICSP IFTDLLCKSIQTSCESFPLRNNATCKKCEKDYPCSCISGFTEKNCEKAIDHCKLLSINCLNEEWCFNIIGRFKYVCIPGCTKNPCWFLKN VYLIHQHLCYCGVTFHGICQDKGPAQFEYVWQLGFAGSEGEKCQGVIDAYFFLAANCTEDATYVNDPEDNNSSCWFPHEGTKEICANGCS CLSEEDSQEYRYLCFLRWAGNMYLENTTDDQENECQHEAVCKDEINRPRCSCSLSYIGRLCVVNVDYCLGNHSISVHGLCLALSHNCNCS GLQRYERNICEIDTEDCKSASCKNGTTSTHLRGYFFRKCVPGFKGTQCEIDIDECASHPCKNGATCIDQPGNYFCQCVPPFKVVDGFSCL CNPGYVGIRCEQDIDDCILNACEHNSTCKDLHLSYQCVCLSDWEGNFCEQESNECKMNPCKNNSTCTDLYKSYRCECTSGWTGQNCSEEI NECDSDPCMNGGLCHESTIPGQFVCLCPPLYTGQFCHQRYNLCDLLHNPCRNNSTCLALVDANQHCICREEFEGKNCEIDVKDCLFLSCQ DYGDCEDMVNNFRCICRPGFSGSLCEIEINECSSEPCKNNGTCVDLTNRFFCNCEPEYHGPFCELDVNKCKISPCLDEENCVYRTDGYNC LCAPGYTGINCEINLDECLSEPCLHDGVCIDGINHYTCDCKSGFFGTHCETNANDCLSNPCLHGRCTELINEYPCSCDADGTSTQCKIKI NDCTSIPCMNEGFCQKSAHGFTCICPRGYTGAYCEKSIDNCAEPELNSVICLNGGICVDGPGHTFDCRCLPGFSGQFCEININECSSSPC LHGADCEDHINGYVCKCQPGWSGHHCENELECIPNSCVHELCMENEPGSTCLCTPGFMTCSIGLLCGDEIRRITCLTPIFQRTDPISTQT YTIPPSETLVSSFPSIKATRIPAIMDTYPVDQGPKQTGIVKHDILPTTGLATLRISTPLESYLLQELIVTRELSAKHSLLSSADVSSSRF LNFGIRDPAQIVQDKTSVSHMPIRTSAATLGFFFPDRRARTPFIMSSLMSDFIFPTQSLLFENCQTVALSATPTTSVIRSIPGADIELNR QSLLSRGFLLIAASISATPVVSRGAQEDIEEYSADSLISRREHWRLLSPSMSPIFPAKVIISKQVTILNSSALHRFSTKAFNPSEYQAIT EASSNQRLTNIKSQAADSLRELSQTCATCSMTEIKSSREFSDQVLHSKQSHFYETFWMNSAILASWYALMGAQTITSGHSFSSATEITPS VAFTEVPSLFPSKKSAKRTILSSSLEESITLSSNLDVNLCLDKTCLSIVPSQTISSDLMNSDLTSKMTTDELSVSENILKLLKIRQYGIT MGPTEVLNQESLLDMEKSKGSHTLFKLHPSDSSLDFELNLQIYPDVTLKTYSEITHANDFKNNLPPLTGSVPDFSEVTTNVAFYTVSATP ALSIQTSSSMSVIRPDWPYFTDYMTSLKKEVKTSSEWSKWELQPSVQYQEFPTASRHLPFTRSLTLSSLESILAPQRLMISEHVPPELWE PFYTDQYAQEHVKPPVTRLLLSAELYCRAWRQALPQLETPPNPSALLALLARRGTVPALPERPVREADLRHQPILMGAHLAVIDALMAAF AFEWTKTLPGPLALTSLEHKLLFWVDTTVRRLQEKTEQEAAQRASPAAPADGAAPAQPSIRYRKDRVVARRAPCFPTVTSLQDLASGAAL AATIHCYCPQLLRLEEVCLKDPMSVADSLYNLQLVQDFCASRLPRGCPLSLEDLLYVPPPLKVNLVVMLAELFMCFEVLKPDFVQVKDLP DGHAASPRGTEASPPQNNSGSSSPVFTFRHPLLSSGGPQSPLRGSTGSLKSSPSMSHMEALGKAWNRQLSRPLSQAVSFSTPFGLDSDVD VVMGDPVLLRSVSSDSLGPPRPAPARTPTQPPPEPGDLPTIEEALQIIHSAEPRLLPDGAADGSFYLHSPEGPSKPSLASPYLPEGTSKP LSDRPTKAPVYMPHPETPSKPSPCLVGEASKPPAPSEGSPKAVASSPAATNSEVKMTSFAERKKQLVKAEAEAGAGSPTSTPAPPEALSS EMSELSARLEEKRRAIEAQKRRIEAIFAKHRQRLGKSAFLQVQPREASGEAEAEAEEADSGPVPGGERPAGEGQGEPTSRPKAVTFSPDL GPVPHEGLGEYNRAVSKLSAALSSLQRDMQRLTDQQQRLLAPPEAPGSAPPPAAWVIPGPTTGPKAASPSPARRVPATRRSPGPGPSQSP RSPKHTRPAELRLAPLTRVLTPPHDVDSLPHLRKFSPSQVPVQTRSSILLAEETPPEEPAARPGLIEIPLGSLADPAAEDEGDGSPAGAE DSLEEEASSEGEPRVGLGFFYKDEDKPEDEMAQKRASLLERQQRRAEEARRRKQWQEVEKEQRREEAARLAQEEAPGPAPLVSAVPMATP APAARAPAEEEVGPRKGDFTRQEYERRAQLKLMDDLDKVLRPRAAGSGGPGRGGRRATRPRSGCCDDSALARSPARGLLGSRLSKIYSQS TLSLSTVANEAHNNLGVKRPTSRAPSPSGLMSPSRLPGSRERDWENGSNASSPASVPEYTGPRLYKEPSAKSNKFIIHNALSHCCLAGKV NEPQKNRILEEIEKSKANHFLILFRDSSCQFRALYTLSGETEELSRLAGYGPRTVTPAMVEGIYKYNSDRKRFTQIPAKTMSMSVDAFTI -------------------------------------------------------------- >28080_28080_2_EYS-CAMSAP3_EYS_chr6_65300116_ENST00000370616_CAMSAP3_chr19_7670112_ENST00000446248_length(transcript)=9574nt_BP=5644nt ATGACTGACAAATCAATCGTCATTCTGAGCCTGATGGTTTTTCACAGCTCTTTCATAAATGGAAAAACATGTAGACGGCAATTGGTGGAA GAATGGCATCCACAACCCTCATCATATGTGGTAAATTGGACACTAACAGAAAACATCTGCTTGGACTTCTACAGAGATTGCTGGTTTTTG GGTGTAAACACTAAAATAGATACTTCAGGCAATCAAGCTGTTCCCCAGATTTGCCCTTTGCAAATTCAATTAGGAGATATCCTTGTAATT TCTTCTGAACCATCTCTTCAATTTCCAGAAATAAATTTGATGAATGTTTCTGAAACATCTTTCGTTGGCTGTGTGCAAAATACCACAACG GAAGATCAGTTACTTTTTGGCTGCAGACTAAAAGGAATGCACACTGTTAATTCTAAGTGGCTGAGTGTTGGGACACATTATTTTATCACA GTTATGGCAAGTGGTCCATCACCTTGTCCACTGGGACTTCGACTAAATGTGACAGTGAAACAGCAGTTCTGCCAGGAATCTCTGAGTTCA GAATTTTGCTCTGGTCATGGTAAATGTCTTAGTGAAGCTTGGAGCAAGACATATAGCTGCCATTGCCAGCCTCCATTTTCTGGAAAATAC TGCCAGGAACTTGATGCATGTTCTTTTAAACCATGTAAAAATAATGGCAGTTGCATTAATAAAAGAGAAAATTGGGATGAGCAAGCATAT GAATGTGTCTGTCACCCACCATTTACAGGAAAGAATTGCTCAGAAATAATTGGCCAGTGTCAACCACATGTCTGTTTCCATGGAAACTGC AGCAATATTACTTCAAATAGTTTCATTTGTGAATGTGATGAGCAATTTTCAGGTCCATTCTGTGAGGTGTCAGCAAAACCTTGTGTTTCT CTGCTTTTTTGGAAAAGAGGAATTTGCCCAAATAGCAGTTCTGCTTATACTTATGAATGCCCAAAAGGATCTTCCAGCCAAAATGGTGAA ACTGATGTCAGTGAGTTTTCATTAGTACCATGTCAGAATGGTACTGACTGCATCAAAATATCAAATGATGTTATGTGCATCTGTTCACCA ATATTTACAGATTTGCTTTGTAAGAGCATTCAAACATCATGTGAGTCATTTCCTTTGAGGAATAATGCTACATGTAAGAAATGTGAGAAA GATTATCCTTGCAGCTGTATATCAGGATTTACTGAAAAAAACTGTGAGAAAGCAATTGACCACTGTAAACTGCTCAGCATCAACTGTCTG AATGAAGAATGGTGTTTCAATATAATTGGAAGATTCAAATATGTATGCATTCCAGGGTGCACAAAAAATCCATGTTGGTTTTTGAAGAAT GTTTACCTAATTCATCAACACCTCTGCTACTGTGGAGTCACCTTCCATGGTATTTGCCAAGATAAAGGTCCTGCTCAATTTGAATATGTG TGGCAATTGGGATTTGCAGGATCTGAAGGCGAAAAGTGCCAAGGGGTTATTGATGCCTATTTCTTTCTGGCTGCAAACTGCACTGAAGAT GCAACCTATGTGAACGATCCTGAAGATAATAATTCTTCATGTTGGTTCCCACATGAAGGCACAAAAGAGATTTGTGCAAATGGATGCAGT TGTTTGAGTGAAGAAGACAGTCAGGAATATCGGTATCTATGTTTTCTCAGATGGGCTGGCAACATGTATCTGGAAAATACAACTGATGAT CAAGAAAATGAGTGTCAACATGAAGCTGTTTGTAAAGATGAAATTAATAGACCCAGATGCAGCTGTTCTCTTAGTTACATTGGCAGATTG TGTGTTGTCAATGTTGACTATTGCTTAGGGAACCACAGTATATCAGTGCATGGCCTCTGCCTGGCCCTTTCGCACAATTGTAACTGTAGC GGTCTGCAAAGATATGAAAGGAACATCTGTGAGATAGATACTGAAGACTGCAAATCTGCGTCCTGCAAAAATGGAACAACTAGTACACAT TTAAGGGGATATTTCTTCCGCAAGTGTGTCCCAGGATTTAAAGGTACGCAATGTGAAATTGATATAGATGAGTGTGCTTCACATCCCTGC AAAAATGGAGCCACCTGCATTGACCAACCTGGTAATTACTTCTGCCAGTGTGTGCCTCCATTTAAAGTGGTTGATGGCTTTTCCTGCTTA TGCAATCCAGGCTATGTTGGGATAAGATGTGAACAGGACATTGATGACTGCATCCTGAATGCCTGTGAGCACAATTCTACCTGCAAAGAC CTGCATCTCAGCTACCAGTGTGTGTGCCTATCTGATTGGGAAGGAAATTTTTGTGAACAAGAATCCAATGAGTGTAAAATGAATCCTTGC AAGAACAATTCCACCTGTACTGACCTTTACAAAAGCTATCGATGTGAGTGTACATCTGGATGGACTGGACAGAACTGTAGTGAAGAAATA AATGAATGCGACTCTGATCCATGCATGAATGGAGGTCTTTGTCATGAATCTACCATCCCTGGACAATTTGTATGTCTGTGCCCACCCCTT TATACTGGACAATTTTGCCACCAACGCTATAACCTTTGTGACCTACTTCATAACCCTTGCAGAAACAACTCAACATGCTTAGCTTTGGTA GACGCAAATCAGCATTGTATTTGTAGAGAAGAATTTGAAGGTAAAAACTGTGAAATTGATGTGAAAGACTGCCTCTTCCTTTCCTGCCAG GATTATGGTGACTGTGAAGATATGGTCAACAATTTCAGGTGTATTTGCAGACCTGGGTTTTCTGGATCTCTGTGTGAAATTGAAATTAAT GAATGTTCCTCTGAACCTTGCAAAAATAATGGAACATGTGTGGATCTGACAAACAGATTTTTTTGTAATTGTGAACCTGAGTACCATGGG CCCTTCTGTGAACTTGATGTAAATAAATGTAAAATCTCACCTTGTCTAGATGAAGAAAATTGTGTCTACAGGACTGATGGATACAACTGC CTCTGTGCCCCTGGTTATACAGGCATCAACTGTGAAATAAATCTAGATGAATGCCTATCAGAGCCCTGTCTCCATGATGGAGTTTGTATC GATGGCATCAATCATTATACCTGTGACTGCAAGAGTGGGTTTTTTGGAACACACTGTGAAACAAACGCCAATGATTGCCTTTCAAATCCT TGTCTACATGGAAGGTGCACAGAACTTATTAATGAATATCCATGTTCATGTGATGCAGATGGGACTAGCACACAATGTAAGATCAAAATT AATGACTGCACATCAATCCCTTGTATGAATGAAGGCTTCTGTCAGAAGTCAGCACATGGATTTACTTGCATTTGCCCACGTGGATACACT GGTGCATACTGTGAAAAAAGCATTGATAATTGTGCTGAGCCTGAACTTAATTCAGTCATCTGTCTTAATGGAGGGATCTGTGTTGATGGG CCTGGACATACTTTTGACTGCAGATGTCTTCCTGGATTTTCTGGTCAATTTTGTGAAATTAATATAAATGAATGCTCTTCATCACCATGT CTACATGGTGCAGACTGTGAAGATCACATCAATGGATATGTTTGCAAATGCCAACCAGGATGGTCTGGACACCACTGTGAGAATGAGCTT GAGTGCATTCCCAACTCATGTGTTCATGAACTCTGCATGGAGAATGAACCTGGCTCGACATGTTTATGCACACCTGGATTTATGACCTGC TCCATTGGGCTTCTTTGTGGTGATGAAATAAGGAGAATTACCTGTTTAACTCCCATCTTTCAAAGAACAGATCCCATTTCCACACAGACA TATACAATTCCCCCTTCTGAGACTTTGGTCAGCAGCTTTCCATCTATAAAGGCTACTAGAATACCAGCCATAATGGACACTTACCCAGTT GATCAAGGTCCAAAACAGACAGGCATTGTCAAGCACGACATTCTCCCAACCACTGGTTTGGCAACATTAAGAATTAGCACACCCTTGGAA AGCTACTTACTCCAAGAACTGATTGTCACTAGAGAGCTTTCAGCAAAACACAGTCTTCTTTCTTCCGCAGATGTTTCCTCTTCTCGATTC CTGAATTTTGGTATTCGTGACCCAGCACAAATTGTCCAGGACAAAACATCGGTATCACATATGCCTATTCGAACTTCAGCAGCCACACTA GGTTTCTTTTTTCCTGATAGGAGAGCAAGGACCCCATTTATCATGTCTTCCTTAATGTCAGATTTTATTTTTCCTACACAATCTTTATTA TTTGAGAACTGTCAGACTGTTGCTTTATCTGCTACCCCAACGACTTCAGTAATTAGAAGCATTCCAGGGGCTGATATTGAGCTAAACAGG CAGTCATTACTCTCCCGTGGATTCCTGCTTATAGCTGCCTCCATAAGTGCAACTCCAGTTGTCTCTAGGGGGGCTCAAGAGGATATTGAA GAATATTCAGCTGATTCTTTAATTTCAAGAAGAGAGCACTGGAGATTGCTCAGCCCCTCGATGTCTCCCATTTTTCCTGCTAAGGTAATT ATATCTAAACAGGTAACCATCTTAAACTCATCAGCTCTGCACCGGTTCAGTACAAAAGCCTTCAATCCCAGTGAATATCAGGCTATTACT GAGGCTTCAAGCAACCAGAGACTCACAAACATCAAATCACAGGCTGCTGATTCTTTAAGAGAATTAAGCCAAACATGTGCAACATGTTCT ATGACTGAAATAAAATCCTCTCGTGAATTCTCAGATCAAGTTTTGCATAGCAAACAGTCCCACTTTTATGAGACATTCTGGATGAACTCA GCGATATTAGCCAGCTGGTATGCACTAATGGGAGCTCAAACTATCACTTCTGGGCATTCATTTTCTTCTGCTACTGAAATAACACCATCA GTGGCATTCACAGAAGTGCCATCTTTATTTCCTTCTAAAAAGAGTGCAAAAAGAACAATTTTATCCTCATCCTTGGAAGAATCCATTACC CTATCAAGTAATTTGGATGTTAATTTATGTTTGGATAAGACTTGTTTATCCATTGTCCCTTCACAAACTATCTCTTCAGACTTGATGAAT TCTGATTTGACTTCAAAAATGACTACTGATGAACTGTCAGTATCAGAAAACATTTTAAAACTATTGAAAATAAGACAATATGGCATAACT ATGGGACCCACTGAGGTACTAAATCAAGAGAGCTTATTGGACATGGAAAAAAGTAAAGGATCTCATACACTGTTCAAACTTCACCCAAGT GATAGTTCTCTGGATTTTGAGTTAAACTTACAAATTTATCCGGATGTTACTTTAAAGACATATTCAGAAATTACACATGCAAATGACTTC AAAAATAATCTGCCACCATTGACAGGCTCAGTGCCTGATTTTTCAGAAGTCACCACCAATGTTGCATTTTATACAGTTTCAGCAACTCCA GCACTTTCAATACAGACGTCTTCCTCCATGTCTGTAATTAGGCCAGATTGGCCATATTTTACAGATTATATGACCTCTCTTAAAAAAGAG GTCAAGACTTCTTCAGAATGGTCCAAATGGGAACTTCAGCCTAGTGTGCAATATCAGGAATTTCCCACTGCAAGCCGGCATCTTCCCTTC ACTAGATCTCTTACTTTGTCTTCACTGGAATCCATTCTGGCACCTCAACGGCTGATGATTTCTGAGCACGTGCCCCCGGAGCTGTGGGAG CCCTTCTATACCGACCAGTACGCGCAGGAGCATGTGAAGCCCCCGGTGACACGGCTGCTGCTCTCAGCCGAGCTCTACTGCAGAGCCTGG CGCCAGGCACTGCCACAGCTTGAAACACCCCCCAACCCCTCTGCACTGCTGGCCCTGCTGGCGCGGAGGGGCACAGTGCCTGCTTTGCCC GAGCGCCCGGTGCGCGAGGCCGACCTGAGGCACCAGCCCATTCTCATGGGAGCCCACCTAGCTGTCATTGATGCCCTCATGGCTGCCTTT GCCTTCGAGTGGACAAAGACCCTGCCAGGTCCCTTGGCCCTGACCAGCTTGGAGCACAAGCTGCTTTTCTGGGTGGACACGACCGTCCGG CGGCTGCAGGAGAAGACCGAGCAGGAAGCGGCCCAGCGAGCCTCTCCAGCAGCCCCTGCAGACGGGGCGGCCCCGGCGCAGCCCTCGTGC CCTACGCGCTGGTACTGGAAGCTGGTTCCTCACGCAATTGCCTTCTGTTTGAAGGAGTCGGGGAGCAAACCCCCCATGATCCGATACCGC AAGGACCGTGTGGTGGCGCGACGTGCCCCCTGCTTCCCGACGGTGACCAGCCTCCAGGACCTGGCCAGTGGGGCCGCGCTGGCCGCCACC ATCCACTGCTATTGTCCCCAGCTGCTTCGACTTGAGGAGGTGTGCTTGAAGGACCCCATGTCTGTGGCGGACAGCCTGTACAACCTCCAG CTCGTGCAGGATTTCTGTGCCTCTCGCCTTCCTCGTGGCTGCCCCCTGTCCCTTGAGGACTTGCTGTACGTCCCACCGCCACTCAAGGTC AACTTGGTGGTGATGCTGGCCGAGTTGTTCATGTGTTTTGAGGTGCTCAAGCCCGACTTTGTGCAAGTGAAGGACTTGCCCGATGGTCAC GCTGCCTCCCCCCGGGGCACTGAGGCCTCCCCACCTCAGAACAACAGCGGCAGTAGTTCTCCTGTCTTCACCTTCCGCCACCCGCTTCTG TCATCTGGTGGCCCCCAGTCCCCACTCCGAGGATCCACAGGCTCCCTGAAGTCTTCCCCGTCCATGTCCCATATGGAGGCCCTGGGCAAG GCCTGGAACCGGCAGCTCAGCCGTCCCCTCTCCCAGGCTGTGTCATTCAGCACCCCCTTTGGCCTGGACAGCGACGTGGATGTCGTCATG GGAGACCCTGTGCTCCTCCGCTCTGTGAGCTCGGACAGCCTGGGCCCCCCGCGTCCCGCGCCGGCCAGGACCCCCACCCAGCCACCCCCG GAGCCTGGTGACCTGCCCACCATCGAGGAAGCTCTGCAGATCATCCACAGTGCCGAGCCCCGGCTCCTCCCAGATGGGGCGGCCGACGGC AGCTTCTACCTCCACTCCCCTGAGGGGCCCTCCAAGCCATCCCTGGCCTCCCCCTACCTGCCCGAGGGGACCTCCAAACCACTGTCCGAC AGGCCCACCAAAGCACCAGTGTACATGCCACACCCCGAGACCCCCTCGAAACCATCTCCCTGTCTGGTGGGGGAGGCATCGAAACCGCCA GCCCCATCCGAGGGGTCCCCGAAGGCGGTGGCTTCGTCCCCAGCAGCCACCAACTCCGAGGTGAAAATGACCAGCTTTGCAGAACGCAAG AAACAGCTGGTGAAGGCAGAGGCTGAGGCCGGAGCGGGGTCCCCCACGTCCACTCCGGCCCCGCCGGAGGCCCTGAGCTCGGAGATGAGT GAGCTCAGCGCCCGGCTGGAGGAGAAACGCAGAGCCATCGAGGCTCAGAAGCGACGGATTGAGGCCATATTCGCCAAGCACCGCCAGCGG CTGGGCAAAAGCGCCTTCCTGCAGGTGCAGCCGCGGGAAGCCTCTGGGGAGGCGGAAGCAGAGGCGGAGGAGGCCGATTCCGGTCCAGTC CCTGGTGGGGAGCGGCCCGCAGGCGAGGGCCAGGGTGAGCCAACCTCACGGCCCAAGGCAGTGACCTTCTCGCCAGACCTGGGCCCGGTG CCCCACGAGGGGCTGGGGGAATACAATCGAGCGGTCAGCAAGCTGAGTGCCGCCTTGAGCTCGCTGCAGCGGGACATGCAGAGGCTCACG GACCAGCAGCAGCGGCTCCTGGCCCCGCCCGAGGCCCCCGGATCCGCCCCACCACCTGCTGCGTGGGTCATCCCTGGCCCCACGACGGGG CCCAAAGCTGCATCCCCCAGCCCCGCCCGGCGAGTCCCGGCCACCCGGCGCAGCCCTGGGCCCGGGCCCAGCCAGTCACCCCGCAGCCCG AAACACACGCGGCCAGCGGAGCTGCGGCTGGCACCCTTGACCAGGGTGCTTACGCCACCCCACGACGTAGACAGCCTCCCCCACCTGCGC AAGTTCTCGCCGAGCCAGGTGCCCGTGCAGACGCGCTCTTCCATCCTCCTGGCGGAGGAGACGCCCCCCGAGGAGCCAGCCGCCCGGCCG GGCCTCATCGAGATCCCGCTGGGCAGCCTGGCAGATCCCGCCGCCGAGGACGAGGGAGACGGGAGCCCCGCTGGTGCTGAGGATTCCTTG GAGGAGGAGGCGTCTTCGGAGGGGGAGCCCCGGGTGGGGCTGGGGTTCTTCTACAAGGATGAAGACAAGCCTGAGGACGAGATGGCCCAA AAGCGGGCCAGCCTGCTGGAGCGGCAGCAGCGGCGAGCAGAGGAGGCGCGGCGGCGCAAGCAGTGGCAGGAGGTGGAGAAGGAACAGCGG AGGGAGGAGGCCGCGAGGCTGGCCCAAGAGGAGGCCCCGGGCCCAGCCCCGCTTGTGTCCGCAGTCCCGATGGCGACTCCAGCCCCTGCT GCCCGGGCTCCAGCCGAGGAGGAGGTGGGCCCCCGGAAGGGGGACTTCACGCGGCAGGAGTACGAGCGCCGGGCCCAGCTGAAGCTGATG GACGACCTCGATAAGGTGCTGCGGCCCCGGGCTGCGGGGTCCGGGGGTCCAGGTCGGGGCGGGCGGAGGGCCACCCGGCCTCGCTCGGGT TGCTGTGACGACTCAGCCCTGGCACGAAGCCCAGCCCGCGGCCTGCTGGGCTCTCGGCTGAGCAAAATCTATTCCCAGTCCACCCTGTCA CTGTCCACTGTGGCCAACGAGGCCCACAATAACCTCGGGGTGAAGAGGCCCACGTCTCGGGCTCCCTCCCCGTCAGGTCTCATGTCCCCA AGCCGCCTGCCTGGAAGCCGCGAACGGGACTGGGAAAATGGCAGCAATGCCTCCTCCCCAGCGTCAGTGCCCGAGTACACAGGTCCACGG CTGTACAAAGAACCCAGCGCCAAGTCCAACAAGTTCATCATCCACAATGCCCTATCACACTGCTGCCTGGCGGGCAAGGTGAACGAACCG CAGAAGAATCGCATTCTGGAGGAAATTGAGAAAAGCAAGGCCAACCACTTCCTGATCCTCTTTCGCGACTCGAGCTGCCAGTTCCGGGCG CTCTACACGCTGTCGGGGGAGACAGAGGAGCTGTCGCGGCTGGCAGGGTATGGGCCCCGGACCGTCACGCCCGCCATGGTGGAAGGCATC TACAAGTACAACTCGGACCGCAAGCGCTTCACCCAGATCCCCGCCAAGACCATGTCCATGAGCGTCGATGCCTTCACCATCCAGGGACAC CTCTGGCAGGGCAAGAAACCCACCACTCCCAAGAAGGGCGGCGGCACCCCCAAATAGCCCCACCCGGGCGGTCCACGGGCCGGGCCCTGT GTGCTGCGGCCGCCATCCCCTGGAGGACAGTCAGTCGGTATTCCTGGGTCCTGTCTGTCCCCAACCGTGTCTGGGTGGGGCTGGAGTCTC CACCCTCTGACTTTGAGTCCAGTCCTGCTGTGGGGGCTGAGCTGGGAGGTTCAGGGACTCAGGGCTCAGCTCAGTCCCCTTGTCTGTCCT >28080_28080_2_EYS-CAMSAP3_EYS_chr6_65300116_ENST00000370616_CAMSAP3_chr19_7670112_ENST00000446248_length(amino acids)=3108AA_BP=1881 MTDKSIVILSLMVFHSSFINGKTCRRQLVEEWHPQPSSYVVNWTLTENICLDFYRDCWFLGVNTKIDTSGNQAVPQICPLQIQLGDILVI SSEPSLQFPEINLMNVSETSFVGCVQNTTTEDQLLFGCRLKGMHTVNSKWLSVGTHYFITVMASGPSPCPLGLRLNVTVKQQFCQESLSS EFCSGHGKCLSEAWSKTYSCHCQPPFSGKYCQELDACSFKPCKNNGSCINKRENWDEQAYECVCHPPFTGKNCSEIIGQCQPHVCFHGNC SNITSNSFICECDEQFSGPFCEVSAKPCVSLLFWKRGICPNSSSAYTYECPKGSSSQNGETDVSEFSLVPCQNGTDCIKISNDVMCICSP IFTDLLCKSIQTSCESFPLRNNATCKKCEKDYPCSCISGFTEKNCEKAIDHCKLLSINCLNEEWCFNIIGRFKYVCIPGCTKNPCWFLKN VYLIHQHLCYCGVTFHGICQDKGPAQFEYVWQLGFAGSEGEKCQGVIDAYFFLAANCTEDATYVNDPEDNNSSCWFPHEGTKEICANGCS CLSEEDSQEYRYLCFLRWAGNMYLENTTDDQENECQHEAVCKDEINRPRCSCSLSYIGRLCVVNVDYCLGNHSISVHGLCLALSHNCNCS GLQRYERNICEIDTEDCKSASCKNGTTSTHLRGYFFRKCVPGFKGTQCEIDIDECASHPCKNGATCIDQPGNYFCQCVPPFKVVDGFSCL CNPGYVGIRCEQDIDDCILNACEHNSTCKDLHLSYQCVCLSDWEGNFCEQESNECKMNPCKNNSTCTDLYKSYRCECTSGWTGQNCSEEI NECDSDPCMNGGLCHESTIPGQFVCLCPPLYTGQFCHQRYNLCDLLHNPCRNNSTCLALVDANQHCICREEFEGKNCEIDVKDCLFLSCQ DYGDCEDMVNNFRCICRPGFSGSLCEIEINECSSEPCKNNGTCVDLTNRFFCNCEPEYHGPFCELDVNKCKISPCLDEENCVYRTDGYNC LCAPGYTGINCEINLDECLSEPCLHDGVCIDGINHYTCDCKSGFFGTHCETNANDCLSNPCLHGRCTELINEYPCSCDADGTSTQCKIKI NDCTSIPCMNEGFCQKSAHGFTCICPRGYTGAYCEKSIDNCAEPELNSVICLNGGICVDGPGHTFDCRCLPGFSGQFCEININECSSSPC LHGADCEDHINGYVCKCQPGWSGHHCENELECIPNSCVHELCMENEPGSTCLCTPGFMTCSIGLLCGDEIRRITCLTPIFQRTDPISTQT YTIPPSETLVSSFPSIKATRIPAIMDTYPVDQGPKQTGIVKHDILPTTGLATLRISTPLESYLLQELIVTRELSAKHSLLSSADVSSSRF LNFGIRDPAQIVQDKTSVSHMPIRTSAATLGFFFPDRRARTPFIMSSLMSDFIFPTQSLLFENCQTVALSATPTTSVIRSIPGADIELNR QSLLSRGFLLIAASISATPVVSRGAQEDIEEYSADSLISRREHWRLLSPSMSPIFPAKVIISKQVTILNSSALHRFSTKAFNPSEYQAIT EASSNQRLTNIKSQAADSLRELSQTCATCSMTEIKSSREFSDQVLHSKQSHFYETFWMNSAILASWYALMGAQTITSGHSFSSATEITPS VAFTEVPSLFPSKKSAKRTILSSSLEESITLSSNLDVNLCLDKTCLSIVPSQTISSDLMNSDLTSKMTTDELSVSENILKLLKIRQYGIT MGPTEVLNQESLLDMEKSKGSHTLFKLHPSDSSLDFELNLQIYPDVTLKTYSEITHANDFKNNLPPLTGSVPDFSEVTTNVAFYTVSATP ALSIQTSSSMSVIRPDWPYFTDYMTSLKKEVKTSSEWSKWELQPSVQYQEFPTASRHLPFTRSLTLSSLESILAPQRLMISEHVPPELWE PFYTDQYAQEHVKPPVTRLLLSAELYCRAWRQALPQLETPPNPSALLALLARRGTVPALPERPVREADLRHQPILMGAHLAVIDALMAAF AFEWTKTLPGPLALTSLEHKLLFWVDTTVRRLQEKTEQEAAQRASPAAPADGAAPAQPSCPTRWYWKLVPHAIAFCLKESGSKPPMIRYR KDRVVARRAPCFPTVTSLQDLASGAALAATIHCYCPQLLRLEEVCLKDPMSVADSLYNLQLVQDFCASRLPRGCPLSLEDLLYVPPPLKV NLVVMLAELFMCFEVLKPDFVQVKDLPDGHAASPRGTEASPPQNNSGSSSPVFTFRHPLLSSGGPQSPLRGSTGSLKSSPSMSHMEALGK AWNRQLSRPLSQAVSFSTPFGLDSDVDVVMGDPVLLRSVSSDSLGPPRPAPARTPTQPPPEPGDLPTIEEALQIIHSAEPRLLPDGAADG SFYLHSPEGPSKPSLASPYLPEGTSKPLSDRPTKAPVYMPHPETPSKPSPCLVGEASKPPAPSEGSPKAVASSPAATNSEVKMTSFAERK KQLVKAEAEAGAGSPTSTPAPPEALSSEMSELSARLEEKRRAIEAQKRRIEAIFAKHRQRLGKSAFLQVQPREASGEAEAEAEEADSGPV PGGERPAGEGQGEPTSRPKAVTFSPDLGPVPHEGLGEYNRAVSKLSAALSSLQRDMQRLTDQQQRLLAPPEAPGSAPPPAAWVIPGPTTG PKAASPSPARRVPATRRSPGPGPSQSPRSPKHTRPAELRLAPLTRVLTPPHDVDSLPHLRKFSPSQVPVQTRSSILLAEETPPEEPAARP GLIEIPLGSLADPAAEDEGDGSPAGAEDSLEEEASSEGEPRVGLGFFYKDEDKPEDEMAQKRASLLERQQRRAEEARRRKQWQEVEKEQR REEAARLAQEEAPGPAPLVSAVPMATPAPAARAPAEEEVGPRKGDFTRQEYERRAQLKLMDDLDKVLRPRAAGSGGPGRGGRRATRPRSG CCDDSALARSPARGLLGSRLSKIYSQSTLSLSTVANEAHNNLGVKRPTSRAPSPSGLMSPSRLPGSRERDWENGSNASSPASVPEYTGPR LYKEPSAKSNKFIIHNALSHCCLAGKVNEPQKNRILEEIEKSKANHFLILFRDSSCQFRALYTLSGETEELSRLAGYGPRTVTPAMVEGI -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for EYS-CAMSAP3 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for EYS-CAMSAP3 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for EYS-CAMSAP3 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
