
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:FAM162A-CASR (FusionGDB2 ID:28630) |
Fusion Gene Summary for FAM162A-CASR |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: FAM162A-CASR | Fusion gene ID: 28630 | Hgene | Tgene | Gene symbol | FAM162A | CASR | Gene ID | 26355 | 846 |
| Gene name | family with sequence similarity 162 member A | calcium sensing receptor | |
| Synonyms | C3orf28|E2IG5|HGTD-P | CAR|EIG8|FHH|FIH|GPRC2A|HHC|HHC1|HYPOC1|NSHPT|PCAR1|hCasR | |
| Cytomap | 3q21.1 | 3q13.33-q21.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | protein FAM162AE2-induced gene 5 proteinHIF-1 alpha-responsive proapoptotic moleculegrowth and transformation-dependent protein | extracellular calcium-sensing receptorparathyroid Ca(2+)-sensing receptor 1parathyroid cell calcium-sensing receptor 1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q96A26 | P41180 | |
| Ensembl transtripts involved in fusion gene | ENST00000469967, ENST00000477892, ENST00000232125, | ENST00000490131, ENST00000498619, ENST00000296154, | |
| Fusion gene scores | * DoF score | 6 X 5 X 5=150 | 8 X 9 X 7=504 |
| # samples | 10 | 15 | |
| ** MAII score | log2(10/150*10)=-0.584962500721156 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(15/504*10)=-1.74846123300404 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: FAM162A [Title/Abstract] AND CASR [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | FAM162A(122103146)-CASR(121980375), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | FAM162A-CASR seems lost the major protein functional domain in Tgene partner, which is a IUPHAR drug target due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | FAM162A | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process | 15082785 |
| Hgene | FAM162A | GO:0043065 | positive regulation of apoptotic process | 15082785 |
| Hgene | FAM162A | GO:0071456 | cellular response to hypoxia | 15082785 |
| Hgene | FAM162A | GO:0090200 | positive regulation of release of cytochrome c from mitochondria | 15082785 |
| Tgene | CASR | GO:0005513 | detection of calcium ion | 27434672 |
| Tgene | CASR | GO:0006874 | cellular calcium ion homeostasis | 27434672 |
| Tgene | CASR | GO:0007186 | G protein-coupled receptor signaling pathway | 27434672 |
| Tgene | CASR | GO:0070509 | calcium ion import | 20846291 |
 Fusion gene breakpoints across FAM162A (5'-gene) Fusion gene breakpoints across FAM162A (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Fusion gene breakpoints across CASR (3'-gene) Fusion gene breakpoints across CASR (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BLCA | TCGA-K4-A3WV-01A | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| ChimerDB4 | LUAD | TCGA-91-8499-01A | FAM162A | chr3 | 122103146 | - | CASR | chr3 | 121980375 | + |
| ChimerDB4 | LUAD | TCGA-91-8499-01A | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| ChimerDB4 | LUAD | TCGA-91-8499 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| ChimerDB4 | STAD | TCGA-VQ-A91K-01A | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| ChimerDB4 | STAD | TCGA-VQ-A91K | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
Top |
Fusion Gene ORF analysis for FAM162A-CASR |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000469967 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| 5CDS-5UTR | ENST00000469967 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| 5CDS-5UTR | ENST00000477892 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| 5CDS-5UTR | ENST00000477892 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| 5CDS-intron | ENST00000469967 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| 5CDS-intron | ENST00000477892 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| 5UTR-3CDS | ENST00000232125 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
| 5UTR-5UTR | ENST00000232125 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| 5UTR-5UTR | ENST00000232125 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| 5UTR-intron | ENST00000232125 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972795 | + |
| Frame-shift | ENST00000469967 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| Frame-shift | ENST00000469967 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| Frame-shift | ENST00000469967 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| Frame-shift | ENST00000469967 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
| Frame-shift | ENST00000469967 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| Frame-shift | ENST00000469967 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| Frame-shift | ENST00000469967 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| Frame-shift | ENST00000469967 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
| Frame-shift | ENST00000469967 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| Frame-shift | ENST00000469967 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| Frame-shift | ENST00000469967 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| Frame-shift | ENST00000469967 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
| Frame-shift | ENST00000477892 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| Frame-shift | ENST00000477892 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| Frame-shift | ENST00000477892 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| Frame-shift | ENST00000477892 | ENST00000296154 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
| Frame-shift | ENST00000477892 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| Frame-shift | ENST00000477892 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| Frame-shift | ENST00000477892 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| Frame-shift | ENST00000477892 | ENST00000490131 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
| Frame-shift | ENST00000477892 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980375 | + |
| Frame-shift | ENST00000477892 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + |
| Frame-shift | ENST00000477892 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994659 | + |
| Frame-shift | ENST00000477892 | ENST00000498619 | FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
Top |
Fusion Genomic Features for FAM162A-CASR |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972794 | + | 1.97E-06 | 0.9999981 |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + | 3.32E-08 | 1 |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + | 1.09E-08 | 1 |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + | 3.32E-08 | 1 |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + | 1.09E-08 | 1 |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121972794 | + | 1.97E-06 | 0.9999981 |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + | 3.32E-08 | 1 |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + | 1.09E-08 | 1 |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121980374 | + | 3.32E-08 | 1 |
| FAM162A | chr3 | 122103146 | + | CASR | chr3 | 121994658 | + | 1.09E-08 | 1 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
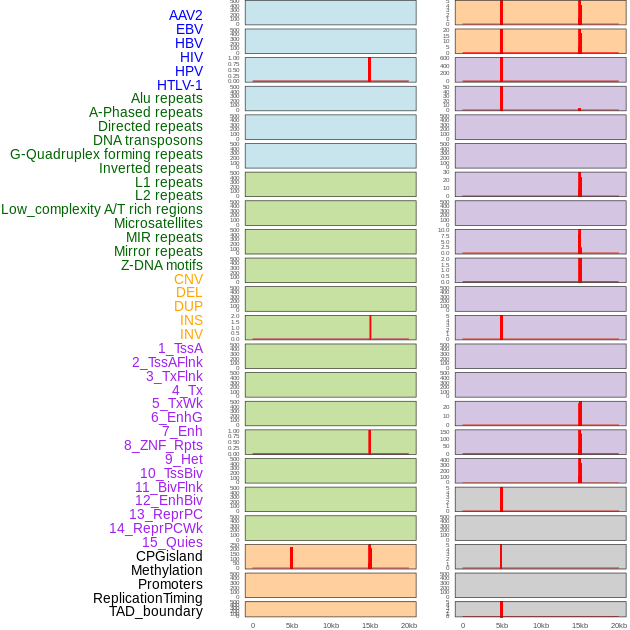 |
Top |
Fusion Protein Features for FAM162A-CASR |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/:122103146/:121980375) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| FAM162A | CASR |
| FUNCTION: Proposed to be involved in regulation of apoptosis; the exact mechanism may differ between cell types/tissues (PubMed:15082785). May be involved in hypoxia-induced cell death of transformed cells implicating cytochrome C release and caspase activation (such as CASP9) and inducing mitochondrial permeability transition (PubMed:15082785). May be involved in hypoxia-induced cell death of neuronal cells probably by promoting release of AIFM1 from mitochondria to cytoplasm and its translocation to the nucleus; however, the involvement of caspases has been reported conflictingly (By similarity). {ECO:0000250|UniProtKB:Q9D6U8, ECO:0000269|PubMed:15082785}. | FUNCTION: G-protein-coupled receptor that senses changes in the extracellular concentration of calcium ions and plays a key role in maintaining calcium homeostasis (PubMed:7759551, PubMed:8702647, PubMed:8636323, PubMed:8878438, PubMed:17555508, PubMed:19789209, PubMed:21566075, PubMed:22114145, PubMed:23966241, PubMed:25292184, PubMed:25104082, PubMed:26386835, PubMed:25766501, PubMed:22789683). Senses fluctuations in the circulating calcium concentration and modulates the production of parathyroid hormone (PTH) in parathyroid glands (By similarity). The activity of this receptor is mediated by a G-protein that activates a phosphatidylinositol-calcium second messenger system (PubMed:7759551). The G-protein-coupled receptor activity is activated by a co-agonist mechanism: aromatic amino acids, such as Trp or Phe, act concertedly with divalent cations, such as calcium or magnesium, to achieve full receptor activation (PubMed:27434672, PubMed:27386547). {ECO:0000250|UniProtKB:Q9QY96, ECO:0000269|PubMed:17555508, ECO:0000269|PubMed:19789209, ECO:0000269|PubMed:21566075, ECO:0000269|PubMed:22114145, ECO:0000269|PubMed:22789683, ECO:0000269|PubMed:23966241, ECO:0000269|PubMed:25104082, ECO:0000269|PubMed:25292184, ECO:0000269|PubMed:25766501, ECO:0000269|PubMed:26386835, ECO:0000269|PubMed:27386547, ECO:0000269|PubMed:27434672, ECO:0000269|PubMed:7759551, ECO:0000269|PubMed:8636323, ECO:0000269|PubMed:8702647, ECO:0000269|PubMed:8878438}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
Top |
Fusion Gene Sequence for FAM162A-CASR |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
Top |
Fusion Gene PPI Analysis for FAM162A-CASR |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for FAM162A-CASR |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for FAM162A-CASR |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
