
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:FAM168B-HMCES (FusionGDB2 ID:28678) |
Fusion Gene Summary for FAM168B-HMCES |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: FAM168B-HMCES | Fusion gene ID: 28678 | Hgene | Tgene | Gene symbol | FAM168B | HMCES | Gene ID | 130074 | 56941 |
| Gene name | family with sequence similarity 168 member B | 5-hydroxymethylcytosine binding, ES cell specific | |
| Synonyms | MANI | C3orf37|DC12|SRAPD1 | |
| Cytomap | 2q21.1 | 3q21.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | myelin-associated neurite-outgrowth inhibitorprotein FAM168B | abasic site processing protein HMCES5-hydroxymethylcytosine (hmC) binding, ES cell-specificES cell-specific 5hmC-binding proteinSOS response associated peptidase domain containing 1SRAP domain-containing protein 1UPF0361 protein C3orf37embryonic ste | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | A1KXE4 | Q96FZ2 | |
| Ensembl transtripts involved in fusion gene | ENST00000389915, ENST00000409185, | ENST00000383463, ENST00000389735, ENST00000417226, ENST00000502878, | |
| Fusion gene scores | * DoF score | 8 X 7 X 5=280 | 6 X 5 X 4=120 |
| # samples | 9 | 7 | |
| ** MAII score | log2(9/280*10)=-1.63742992061529 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/120*10)=-0.777607578663552 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: FAM168B [Title/Abstract] AND HMCES [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | FAM168B(131840150)-HMCES(129017197), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | HMCES | GO:0006974 | cellular response to DNA damage stimulus | 30554877 |
 Fusion gene breakpoints across FAM168B (5'-gene) Fusion gene breakpoints across FAM168B (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
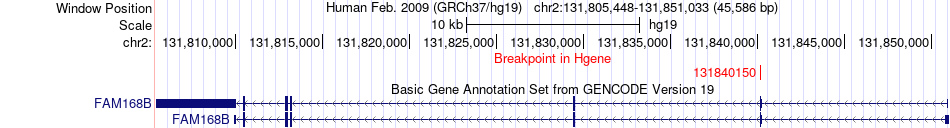 |
 Fusion gene breakpoints across HMCES (3'-gene) Fusion gene breakpoints across HMCES (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
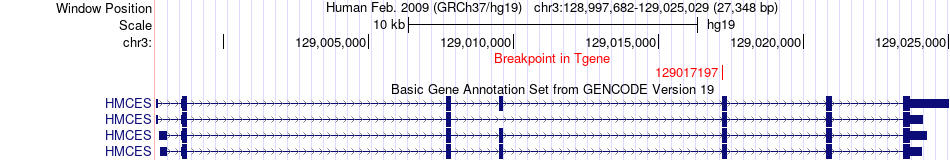 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-BR-A4QI-01A | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + |
Top |
Fusion Gene ORF analysis for FAM168B-HMCES |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000389915 | ENST00000383463 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + |
| In-frame | ENST00000389915 | ENST00000389735 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + |
| In-frame | ENST00000389915 | ENST00000417226 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + |
| In-frame | ENST00000389915 | ENST00000502878 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + |
| In-frame | ENST00000409185 | ENST00000383463 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + |
| In-frame | ENST00000409185 | ENST00000389735 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + |
| In-frame | ENST00000409185 | ENST00000417226 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + |
| In-frame | ENST00000409185 | ENST00000502878 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000409185 | FAM168B | chr2 | 131840150 | - | ENST00000383463 | HMCES | chr3 | 129017197 | + | 2151 | 178 | 115 | 789 | 224 |
| ENST00000409185 | FAM168B | chr2 | 131840150 | - | ENST00000417226 | HMCES | chr3 | 129017197 | + | 1258 | 178 | 115 | 789 | 224 |
| ENST00000409185 | FAM168B | chr2 | 131840150 | - | ENST00000502878 | HMCES | chr3 | 129017197 | + | 1406 | 178 | 115 | 789 | 224 |
| ENST00000409185 | FAM168B | chr2 | 131840150 | - | ENST00000389735 | HMCES | chr3 | 129017197 | + | 1227 | 178 | 115 | 789 | 224 |
| ENST00000389915 | FAM168B | chr2 | 131840150 | - | ENST00000383463 | HMCES | chr3 | 129017197 | + | 2301 | 328 | 265 | 939 | 224 |
| ENST00000389915 | FAM168B | chr2 | 131840150 | - | ENST00000417226 | HMCES | chr3 | 129017197 | + | 1408 | 328 | 265 | 939 | 224 |
| ENST00000389915 | FAM168B | chr2 | 131840150 | - | ENST00000502878 | HMCES | chr3 | 129017197 | + | 1556 | 328 | 265 | 939 | 224 |
| ENST00000389915 | FAM168B | chr2 | 131840150 | - | ENST00000389735 | HMCES | chr3 | 129017197 | + | 1377 | 328 | 265 | 939 | 224 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000409185 | ENST00000383463 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + | 0.003598353 | 0.99640167 |
| ENST00000409185 | ENST00000417226 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + | 0.003691127 | 0.99630886 |
| ENST00000409185 | ENST00000502878 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + | 0.003354837 | 0.99664515 |
| ENST00000409185 | ENST00000389735 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + | 0.004125617 | 0.99587435 |
| ENST00000389915 | ENST00000383463 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + | 0.002971971 | 0.99702805 |
| ENST00000389915 | ENST00000417226 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + | 0.003576608 | 0.9964234 |
| ENST00000389915 | ENST00000502878 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + | 0.002914815 | 0.99708515 |
| ENST00000389915 | ENST00000389735 | FAM168B | chr2 | 131840150 | - | HMCES | chr3 | 129017197 | + | 0.004024156 | 0.9959759 |
Top |
Fusion Genomic Features for FAM168B-HMCES |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| FAM168B | chr2 | 131840149 | - | HMCES | chr3 | 129017196 | + | 8.56E-08 | 0.9999999 |
| FAM168B | chr2 | 131840149 | - | HMCES | chr3 | 129017196 | + | 8.56E-08 | 0.9999999 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
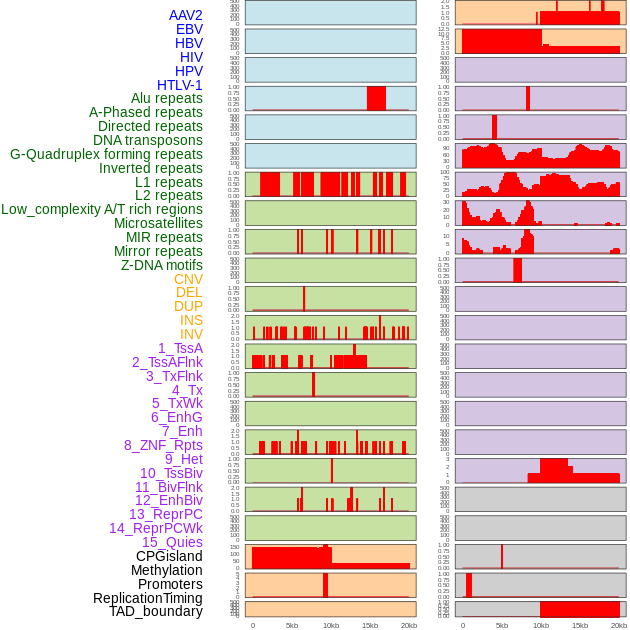 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
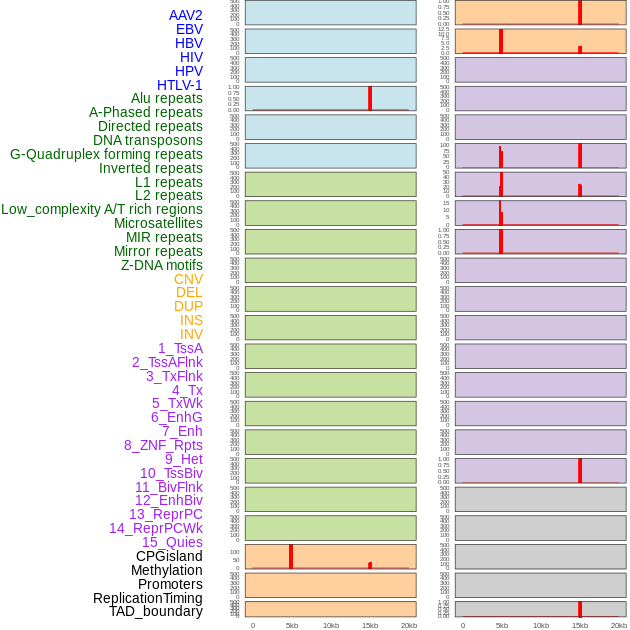 |
Top |
Fusion Protein Features for FAM168B-HMCES |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr2:131840150/chr3:129017197) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| FAM168B | HMCES |
| FUNCTION: Inhibitor of neuronal axonal outgrowth. Acts as a negative regulator of CDC42 and STAT3 and a positive regulator of STMN2. Positive regulator of CDC27. {ECO:0000250|UniProtKB:D4AEP3}. | FUNCTION: Sensor of abasic sites in single-stranded DNA (ssDNA) required to preserve genome integrity by promoting error-free repair of abasic sites (PubMed:30554877, PubMed:31235915, PubMed:31235913). Acts as an enzyme that recognizes and binds abasic sites in ssDNA at replication forks and chemically modifies the lesion by forming a covalent cross-link with DNA: forms a stable thiazolidine linkage between a ring-opened abasic site and the alpha-amino and sulfhydryl substituents of its N-terminal catalytic cysteine residue (PubMed:30554877, PubMed:31235913). The HMCES DNA-protein cross-link is then degraded by the proteasome (PubMed:30554877). Promotes error-free repair of abasic sites by acting as a 'suicide' enzyme that is degraded, thereby protecting abasic sites from translesion synthesis (TLS) polymerases and endonucleases that are error-prone and would generate mutations and double-strand breaks (PubMed:30554877). Has preference for ssDNA, but can also accommodate double-stranded DNA with 3' or 5' overhang (dsDNA), and dsDNA-ssDNA 3' junction (PubMed:31235915, PubMed:31806351). Also involved in class switch recombination (CSR) in B-cells independently of the formation of a DNA-protein cross-link: acts by binding and protecting ssDNA overhangs to promote DNA double-strand break repair through the microhomology-mediated alternative-end-joining (Alt-EJ) pathway (By similarity). Acts as a protease: mediates autocatalytic processing of its N-terminal methionine in order to expose the catalytic cysteine (By similarity). {ECO:0000250|UniProtKB:Q8R1M0, ECO:0000269|PubMed:30554877, ECO:0000269|PubMed:31235913, ECO:0000269|PubMed:31235915, ECO:0000269|PubMed:31806351}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000389915 | - | 2 | 7 | 1_18 | 23 | 157.0 | Topological domain | Cytoplasmic |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000409185 | - | 2 | 7 | 1_18 | 23 | 1704.0 | Topological domain | Cytoplasmic |
| Tgene | HMCES | chr2:131840150 | chr3:129017197 | ENST00000383463 | 3 | 7 | 332_338 | 151 | 355.0 | Motif | PIP-box | |
| Tgene | HMCES | chr2:131840150 | chr3:129017197 | ENST00000389735 | 3 | 7 | 332_338 | 151 | 355.0 | Motif | PIP-box | |
| Tgene | HMCES | chr2:131840150 | chr3:129017197 | ENST00000502878 | 3 | 7 | 332_338 | 151 | 355.0 | Motif | PIP-box |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000389915 | - | 2 | 7 | 165_195 | 23 | 157.0 | Topological domain | Cytoplasmic |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000389915 | - | 2 | 7 | 43_142 | 23 | 157.0 | Topological domain | Extracellular |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000409185 | - | 2 | 7 | 165_195 | 23 | 1704.0 | Topological domain | Cytoplasmic |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000409185 | - | 2 | 7 | 43_142 | 23 | 1704.0 | Topological domain | Extracellular |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000389915 | - | 2 | 7 | 143_164 | 23 | 157.0 | Transmembrane | Helical |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000389915 | - | 2 | 7 | 19_42 | 23 | 157.0 | Transmembrane | Helical |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000409185 | - | 2 | 7 | 143_164 | 23 | 1704.0 | Transmembrane | Helical |
| Hgene | FAM168B | chr2:131840150 | chr3:129017197 | ENST00000409185 | - | 2 | 7 | 19_42 | 23 | 1704.0 | Transmembrane | Helical |
Top |
Fusion Gene Sequence for FAM168B-HMCES |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >28678_28678_1_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000389915_HMCES_chr3_129017197_ENST00000383463_length(transcript)=2301nt_BP=328nt GTCGGCGGCTCCGGTGGCTGCGCAGCGTCGGCGGGGAGCGCGGGCTGCGGAGAGGCGGGCCGGGCCAAGCGGAGCCGAGCGAGCGGGAGC GCGGCGTCCGGGAGGCGGCGGAGACGCGGGGCTCGGAGGGTCAGCCTCTTATCGTAGCAGGTCTCCTCGGCACGCCCCCCTTGTTTCGCC CCACGGCCAAGCCCGCCGCGGGCCGGCGTGCGCTGGTCACTGAGGCCCAGGTCGCCGCCGCGGCGCGTTTTTGAAATCATGAATCCTGTT TATAGTCCTGGATCTTCTGGGGTTCCCTATGCAAATGCCAAAGGAATTGGTTATCCAGTCAGGTAGCATTGGTGCTGCAGATAGTCCTGA GAACTGGGAGAAAGTCTGGGACAACTGGAGGCTGCTGACAATGGCCGGGATCTTTGACTGCTGGGAGCCCCCAGAGGGAGGAGATGTCCT GTATTCCTATACCATCATCACAGTGGATTCCTGCAAAGGCTTGAGTGACATCCACCACAGGATGCCTGCCATATTAGATGGAGAGGAGGC AGTTTCTAAATGGCTTGACTTTGGTGAAGTCTCAACTCAGGAAGCTCTGAAATTAATCCACCCAACAGAGAACATCACCTTCCATGCAGT CTCTTCTGTGGTGAACAACTCGCGAAACAACACTCCTGAGTGTCTGGCTCCTGTCGACTTGGTGGTCAAAAAGGAGCTCAGGGCAAGTGG CAGTAGCCAGAGGATGTTGCAGTGGTTGGCCACAAAGTCACCCAAAAAGGAAGACTCAAAAACACCTCAAAAGGAAGAGTCAGATGTTCC CCAGTGGTCCAGTCAGTTCCTGCAGAAGAGTCCACTCCCCACCAAGAGAGGCACTGCAGGACTCCTAGAGCAATGGCTGAAGCGGGAGAA GGAGGAGGAACCTGTGGCCAAGCGTCCTTACAGCCAGTGACACAGGACTTTCAGAGACCAAGGCCAGGGTCTGCTGCACTGCTGTTCTGA TAATAGGTTCTTAACATTGTATGTATATGTGTTTGCTTTGGGAGGAGGTGGCACTGTGTTAGTTGACAGTTGTGGGCTCATGTAGTCTTT TTTGCCATGAGTAGGAGCCCCTAGTGGGGCTGGTGGACAGCTTTGGAAGAGGTGTCCTGCTGCTGTTACCAGCCATGTGGGCCCCATAGG GGCACTGCGCCTGCTGCCCTTTCCTGGCAGGGCTGGTGGAGTCTTCCCTCAAAGCATGCCTTACCCAGCTGGGAAGTCTCTGCCCTGATC TGGTACTCCTTGTAGTAAGCTGTTTTCTGCTCAGCCACTGGGCTCTTTCACTTTTTTAGTTCTTAAAAATTTATTTTTAAGTTCTAAAAT AAAATAAAAATAAGTTCTTAAAATTTATTTTTTTCCTGAATAAATTGTATTTGGTAAACTTCTGCCTACATTTTGGAAAGTGATGCTGGT GGGGAAAGTTCTAGATCTTACTTGGTTTCTTCTAGAATCAGTCTTCAGGAATGGATTTTGTCACAAATGGGGCATGGGGGCTTTCTGAGG AAATAACTACAAGTCTTGGTGGTGGGCTCCTTATTATGTTTCTTTTTCTTTCATTCTTGATACTTGGAAGTCGTCTGAATCCTTTAGCTT CAAACCAGCCTGAGTTTGAGTGCTTGCCGTAGCAGAAACTATCCTTACCACAGGTGGGAAGGAAAGGACCAGTTTCTAGCAGTGTCGGGC CACTCCTCTTTCGAACATCCCTAAGGGAGGCATTCACAAAAGCTGTCCCAAGCAGCTGGAAGAAAACAGCTTCCGAGATGACCAGGAGGA CTGGGCGGCGCCGAGCCCAGAACGCTCCTGGCGCAGCACCGTTGGCGTTGGCCGATTGCTGCTGGTGGGGGGCGGGGGTGCAGGCCCCAG TCTCTATGCAAATCAGGGATCAGAAGATCGGAATTTCCACCAATCAGCGGGAAGCCTCGGCCCTGTAACTGCTAATGGGAGACAGCAGCG CCACGCCACAGGCTTTTCCCCTGGTTTCGGGAGGGGTGGGGAGCCAGGTGGGGCTCCCGCCCAGACCCTTTCCCGAGGTCCGCCCTCTCC GCCTTTTCTCTAAATTCCTCTTTTGAGTGCCCTCCCTTCCGGTTGAGAGGCGGGGGTTGGCCCGTAGTTGTACACTCAGTCACCCTGCAC TGTGGAGGCGGGGGCCTCCCTTGTGGACTGATTTGCGTGGGATTTGGTTGTTTTATTAAGAGATTTAAAAAATTCAGATGACTTACTAGT >28678_28678_1_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000389915_HMCES_chr3_129017197_ENST00000383463_length(amino acids)=224AA_BP=20 MFIVLDLLGFPMQMPKELVIQSGSIGAADSPENWEKVWDNWRLLTMAGIFDCWEPPEGGDVLYSYTIITVDSCKGLSDIHHRMPAILDGE EAVSKWLDFGEVSTQEALKLIHPTENITFHAVSSVVNNSRNNTPECLAPVDLVVKKELRASGSSQRMLQWLATKSPKKEDSKTPQKEESD -------------------------------------------------------------- >28678_28678_2_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000389915_HMCES_chr3_129017197_ENST00000389735_length(transcript)=1377nt_BP=328nt GTCGGCGGCTCCGGTGGCTGCGCAGCGTCGGCGGGGAGCGCGGGCTGCGGAGAGGCGGGCCGGGCCAAGCGGAGCCGAGCGAGCGGGAGC GCGGCGTCCGGGAGGCGGCGGAGACGCGGGGCTCGGAGGGTCAGCCTCTTATCGTAGCAGGTCTCCTCGGCACGCCCCCCTTGTTTCGCC CCACGGCCAAGCCCGCCGCGGGCCGGCGTGCGCTGGTCACTGAGGCCCAGGTCGCCGCCGCGGCGCGTTTTTGAAATCATGAATCCTGTT TATAGTCCTGGATCTTCTGGGGTTCCCTATGCAAATGCCAAAGGAATTGGTTATCCAGTCAGGTAGCATTGGTGCTGCAGATAGTCCTGA GAACTGGGAGAAAGTCTGGGACAACTGGAGGCTGCTGACAATGGCCGGGATCTTTGACTGCTGGGAGCCCCCAGAGGGAGGAGATGTCCT GTATTCCTATACCATCATCACAGTGGATTCCTGCAAAGGCTTGAGTGACATCCACCACAGGATGCCTGCCATATTAGATGGAGAGGAGGC AGTTTCTAAATGGCTTGACTTTGGTGAAGTCTCAACTCAGGAAGCTCTGAAATTAATCCACCCAACAGAGAACATCACCTTCCATGCAGT CTCTTCTGTGGTGAACAACTCGCGAAACAACACTCCTGAGTGTCTGGCTCCTGTCGACTTGGTGGTCAAAAAGGAGCTCAGGGCAAGTGG CAGTAGCCAGAGGATGTTGCAGTGGTTGGCCACAAAGTCACCCAAAAAGGAAGACTCAAAAACACCTCAAAAGGAAGAGTCAGATGTTCC CCAGTGGTCCAGTCAGTTCCTGCAGAAGAGTCCACTCCCCACCAAGAGAGGCACTGCAGGACTCCTAGAGCAATGGCTGAAGCGGGAGAA GGAGGAGGAACCTGTGGCCAAGCGTCCTTACAGCCAGTGACACAGGACTTTCAGAGACCAAGGCCAGGGTCTGCTGCACTGCTGTTCTGA TAATAGGTTCTTAACATTGTATGTATATGTGTTTGCTTTGGGAGGAGGTGGCACTGTGTTAGTTGACAGTTGTGGGCTCATGTAGTCTTT TTTGCCATGAGTAGGAGCCCCTAGTGGGGCTGGTGGACAGCTTTGGAAGAGGTGTCCTGCTGCTGTTACCAGCCATGTGGGCCCCATAGG GGCACTGCGCCTGCTGCCCTTTCCTGGCAGGGCTGGTGGAGTCTTCCCTCAAAGCATGCCTTACCCAGCTGGGAAGTCTCTGCCCTGATC TGGTACTCCTTGTAGTAAGCTGTTTTCTGCTCAGCCACTGGGCTCTTTCACTTTTTTAGTTCTTAAAAATTTATTTTTAAGTTCTAAAAT >28678_28678_2_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000389915_HMCES_chr3_129017197_ENST00000389735_length(amino acids)=224AA_BP=20 MFIVLDLLGFPMQMPKELVIQSGSIGAADSPENWEKVWDNWRLLTMAGIFDCWEPPEGGDVLYSYTIITVDSCKGLSDIHHRMPAILDGE EAVSKWLDFGEVSTQEALKLIHPTENITFHAVSSVVNNSRNNTPECLAPVDLVVKKELRASGSSQRMLQWLATKSPKKEDSKTPQKEESD -------------------------------------------------------------- >28678_28678_3_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000389915_HMCES_chr3_129017197_ENST00000417226_length(transcript)=1408nt_BP=328nt GTCGGCGGCTCCGGTGGCTGCGCAGCGTCGGCGGGGAGCGCGGGCTGCGGAGAGGCGGGCCGGGCCAAGCGGAGCCGAGCGAGCGGGAGC GCGGCGTCCGGGAGGCGGCGGAGACGCGGGGCTCGGAGGGTCAGCCTCTTATCGTAGCAGGTCTCCTCGGCACGCCCCCCTTGTTTCGCC CCACGGCCAAGCCCGCCGCGGGCCGGCGTGCGCTGGTCACTGAGGCCCAGGTCGCCGCCGCGGCGCGTTTTTGAAATCATGAATCCTGTT TATAGTCCTGGATCTTCTGGGGTTCCCTATGCAAATGCCAAAGGAATTGGTTATCCAGTCAGGTAGCATTGGTGCTGCAGATAGTCCTGA GAACTGGGAGAAAGTCTGGGACAACTGGAGGCTGCTGACAATGGCCGGGATCTTTGACTGCTGGGAGCCCCCAGAGGGAGGAGATGTCCT GTATTCCTATACCATCATCACAGTGGATTCCTGCAAAGGCTTGAGTGACATCCACCACAGGATGCCTGCCATATTAGATGGAGAGGAGGC AGTTTCTAAATGGCTTGACTTTGGTGAAGTCTCAACTCAGGAAGCTCTGAAATTAATCCACCCAACAGAGAACATCACCTTCCATGCAGT CTCTTCTGTGGTGAACAACTCGCGAAACAACACTCCTGAGTGTCTGGCTCCTGTCGACTTGGTGGTCAAAAAGGAGCTCAGGGCAAGTGG CAGTAGCCAGAGGATGTTGCAGTGGTTGGCCACAAAGTCACCCAAAAAGGAAGACTCAAAAACACCTCAAAAGGAAGAGTCAGATGTTCC CCAGTGGTCCAGTCAGTTCCTGCAGAAGAGTCCACTCCCCACCAAGAGAGGCACTGCAGGACTCCTAGAGCAATGGCTGAAGCGGGAGAA GGAGGAGGAACCTGTGGCCAAGCGTCCTTACAGCCAGTGACACAGGACTTTCAGAGACCAAGGCCAGGGTCTGCTGCACTGCTGTTCTGA TAATAGGTTCTTAACATTGTATGTATATGTGTTTGCTTTGGGAGGAGGTGGCACTGTGTTAGTTGACAGTTGTGGGCTCATGTAGTCTTT TTTGCCATGAGTAGGAGCCCCTAGTGGGGCTGGTGGACAGCTTTGGAAGAGGTGTCCTGCTGCTGTTACCAGCCATGTGGGCCCCATAGG GGCACTGCGCCTGCTGCCCTTTCCTGGCAGGGCTGGTGGAGTCTTCCCTCAAAGCATGCCTTACCCAGCTGGGAAGTCTCTGCCCTGATC TGGTACTCCTTGTAGTAAGCTGTTTTCTGCTCAGCCACTGGGCTCTTTCACTTTTTTAGTTCTTAAAAATTTATTTTTAAGTTCTAAAAT >28678_28678_3_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000389915_HMCES_chr3_129017197_ENST00000417226_length(amino acids)=224AA_BP=20 MFIVLDLLGFPMQMPKELVIQSGSIGAADSPENWEKVWDNWRLLTMAGIFDCWEPPEGGDVLYSYTIITVDSCKGLSDIHHRMPAILDGE EAVSKWLDFGEVSTQEALKLIHPTENITFHAVSSVVNNSRNNTPECLAPVDLVVKKELRASGSSQRMLQWLATKSPKKEDSKTPQKEESD -------------------------------------------------------------- >28678_28678_4_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000389915_HMCES_chr3_129017197_ENST00000502878_length(transcript)=1556nt_BP=328nt GTCGGCGGCTCCGGTGGCTGCGCAGCGTCGGCGGGGAGCGCGGGCTGCGGAGAGGCGGGCCGGGCCAAGCGGAGCCGAGCGAGCGGGAGC GCGGCGTCCGGGAGGCGGCGGAGACGCGGGGCTCGGAGGGTCAGCCTCTTATCGTAGCAGGTCTCCTCGGCACGCCCCCCTTGTTTCGCC CCACGGCCAAGCCCGCCGCGGGCCGGCGTGCGCTGGTCACTGAGGCCCAGGTCGCCGCCGCGGCGCGTTTTTGAAATCATGAATCCTGTT TATAGTCCTGGATCTTCTGGGGTTCCCTATGCAAATGCCAAAGGAATTGGTTATCCAGTCAGGTAGCATTGGTGCTGCAGATAGTCCTGA GAACTGGGAGAAAGTCTGGGACAACTGGAGGCTGCTGACAATGGCCGGGATCTTTGACTGCTGGGAGCCCCCAGAGGGAGGAGATGTCCT GTATTCCTATACCATCATCACAGTGGATTCCTGCAAAGGCTTGAGTGACATCCACCACAGGATGCCTGCCATATTAGATGGAGAGGAGGC AGTTTCTAAATGGCTTGACTTTGGTGAAGTCTCAACTCAGGAAGCTCTGAAATTAATCCACCCAACAGAGAACATCACCTTCCATGCAGT CTCTTCTGTGGTGAACAACTCGCGAAACAACACTCCTGAGTGTCTGGCTCCTGTCGACTTGGTGGTCAAAAAGGAGCTCAGGGCAAGTGG CAGTAGCCAGAGGATGTTGCAGTGGTTGGCCACAAAGTCACCCAAAAAGGAAGACTCAAAAACACCTCAAAAGGAAGAGTCAGATGTTCC CCAGTGGTCCAGTCAGTTCCTGCAGAAGAGTCCACTCCCCACCAAGAGAGGCACTGCAGGACTCCTAGAGCAATGGCTGAAGCGGGAGAA GGAGGAGGAACCTGTGGCCAAGCGTCCTTACAGCCAGTGACACAGGACTTTCAGAGACCAAGGCCAGGGTCTGCTGCACTGCTGTTCTGA TAATAGGTTCTTAACATTGTATGTATATGTGTTTGCTTTGGGAGGAGGTGGCACTGTGTTAGTTGACAGTTGTGGGCTCATGTAGTCTTT TTTGCCATGAGTAGGAGCCCCTAGTGGGGCTGGTGGACAGCTTTGGAAGAGGTGTCCTGCTGCTGTTACCAGCCATGTGGGCCCCATAGG GGCACTGCGCCTGCTGCCCTTTCCTGGCAGGGCTGGTGGAGTCTTCCCTCAAAGCATGCCTTACCCAGCTGGGAAGTCTCTGCCCTGATC TGGTACTCCTTGTAGTAAGCTGTTTTCTGCTCAGCCACTGGGCTCTTTCACTTTTTTAGTTCTTAAAAATTTATTTTTAAGTTCTAAAAT AAAATAAAAATAAGTTCTTAAAATTTATTTTTTTCCTGAATAAATTGTATTTGGTAAACTTCTGCCTACATTTTGGAAAGTGATGCTGGT GGGGAAAGTTCTAGATCTTACTTGGTTTCTTCTAGAATCAGTCTTCAGGAATGGATTTTGTCACAAATGGGGCATGGGGGCTTTCTGAGG >28678_28678_4_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000389915_HMCES_chr3_129017197_ENST00000502878_length(amino acids)=224AA_BP=20 MFIVLDLLGFPMQMPKELVIQSGSIGAADSPENWEKVWDNWRLLTMAGIFDCWEPPEGGDVLYSYTIITVDSCKGLSDIHHRMPAILDGE EAVSKWLDFGEVSTQEALKLIHPTENITFHAVSSVVNNSRNNTPECLAPVDLVVKKELRASGSSQRMLQWLATKSPKKEDSKTPQKEESD -------------------------------------------------------------- >28678_28678_5_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000409185_HMCES_chr3_129017197_ENST00000383463_length(transcript)=2151nt_BP=178nt GGGGAGCGCGGGCTGCGGAGAGGCGGGCCGGGCCAAGCGGAGCCGAGCGAGCGGGAGCGCGGCGTCCGGGAGGCGGCGGAGACGCGGGGC TCGGAGGTTTTTGAAATCATGAATCCTGTTTATAGTCCTGGATCTTCTGGGGTTCCCTATGCAAATGCCAAAGGAATTGGTTATCCAGTC AGGTAGCATTGGTGCTGCAGATAGTCCTGAGAACTGGGAGAAAGTCTGGGACAACTGGAGGCTGCTGACAATGGCCGGGATCTTTGACTG CTGGGAGCCCCCAGAGGGAGGAGATGTCCTGTATTCCTATACCATCATCACAGTGGATTCCTGCAAAGGCTTGAGTGACATCCACCACAG GATGCCTGCCATATTAGATGGAGAGGAGGCAGTTTCTAAATGGCTTGACTTTGGTGAAGTCTCAACTCAGGAAGCTCTGAAATTAATCCA CCCAACAGAGAACATCACCTTCCATGCAGTCTCTTCTGTGGTGAACAACTCGCGAAACAACACTCCTGAGTGTCTGGCTCCTGTCGACTT GGTGGTCAAAAAGGAGCTCAGGGCAAGTGGCAGTAGCCAGAGGATGTTGCAGTGGTTGGCCACAAAGTCACCCAAAAAGGAAGACTCAAA AACACCTCAAAAGGAAGAGTCAGATGTTCCCCAGTGGTCCAGTCAGTTCCTGCAGAAGAGTCCACTCCCCACCAAGAGAGGCACTGCAGG ACTCCTAGAGCAATGGCTGAAGCGGGAGAAGGAGGAGGAACCTGTGGCCAAGCGTCCTTACAGCCAGTGACACAGGACTTTCAGAGACCA AGGCCAGGGTCTGCTGCACTGCTGTTCTGATAATAGGTTCTTAACATTGTATGTATATGTGTTTGCTTTGGGAGGAGGTGGCACTGTGTT AGTTGACAGTTGTGGGCTCATGTAGTCTTTTTTGCCATGAGTAGGAGCCCCTAGTGGGGCTGGTGGACAGCTTTGGAAGAGGTGTCCTGC TGCTGTTACCAGCCATGTGGGCCCCATAGGGGCACTGCGCCTGCTGCCCTTTCCTGGCAGGGCTGGTGGAGTCTTCCCTCAAAGCATGCC TTACCCAGCTGGGAAGTCTCTGCCCTGATCTGGTACTCCTTGTAGTAAGCTGTTTTCTGCTCAGCCACTGGGCTCTTTCACTTTTTTAGT TCTTAAAAATTTATTTTTAAGTTCTAAAATAAAATAAAAATAAGTTCTTAAAATTTATTTTTTTCCTGAATAAATTGTATTTGGTAAACT TCTGCCTACATTTTGGAAAGTGATGCTGGTGGGGAAAGTTCTAGATCTTACTTGGTTTCTTCTAGAATCAGTCTTCAGGAATGGATTTTG TCACAAATGGGGCATGGGGGCTTTCTGAGGAAATAACTACAAGTCTTGGTGGTGGGCTCCTTATTATGTTTCTTTTTCTTTCATTCTTGA TACTTGGAAGTCGTCTGAATCCTTTAGCTTCAAACCAGCCTGAGTTTGAGTGCTTGCCGTAGCAGAAACTATCCTTACCACAGGTGGGAA GGAAAGGACCAGTTTCTAGCAGTGTCGGGCCACTCCTCTTTCGAACATCCCTAAGGGAGGCATTCACAAAAGCTGTCCCAAGCAGCTGGA AGAAAACAGCTTCCGAGATGACCAGGAGGACTGGGCGGCGCCGAGCCCAGAACGCTCCTGGCGCAGCACCGTTGGCGTTGGCCGATTGCT GCTGGTGGGGGGCGGGGGTGCAGGCCCCAGTCTCTATGCAAATCAGGGATCAGAAGATCGGAATTTCCACCAATCAGCGGGAAGCCTCGG CCCTGTAACTGCTAATGGGAGACAGCAGCGCCACGCCACAGGCTTTTCCCCTGGTTTCGGGAGGGGTGGGGAGCCAGGTGGGGCTCCCGC CCAGACCCTTTCCCGAGGTCCGCCCTCTCCGCCTTTTCTCTAAATTCCTCTTTTGAGTGCCCTCCCTTCCGGTTGAGAGGCGGGGGTTGG CCCGTAGTTGTACACTCAGTCACCCTGCACTGTGGAGGCGGGGGCCTCCCTTGTGGACTGATTTGCGTGGGATTTGGTTGTTTTATTAAG >28678_28678_5_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000409185_HMCES_chr3_129017197_ENST00000383463_length(amino acids)=224AA_BP=20 MFIVLDLLGFPMQMPKELVIQSGSIGAADSPENWEKVWDNWRLLTMAGIFDCWEPPEGGDVLYSYTIITVDSCKGLSDIHHRMPAILDGE EAVSKWLDFGEVSTQEALKLIHPTENITFHAVSSVVNNSRNNTPECLAPVDLVVKKELRASGSSQRMLQWLATKSPKKEDSKTPQKEESD -------------------------------------------------------------- >28678_28678_6_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000409185_HMCES_chr3_129017197_ENST00000389735_length(transcript)=1227nt_BP=178nt GGGGAGCGCGGGCTGCGGAGAGGCGGGCCGGGCCAAGCGGAGCCGAGCGAGCGGGAGCGCGGCGTCCGGGAGGCGGCGGAGACGCGGGGC TCGGAGGTTTTTGAAATCATGAATCCTGTTTATAGTCCTGGATCTTCTGGGGTTCCCTATGCAAATGCCAAAGGAATTGGTTATCCAGTC AGGTAGCATTGGTGCTGCAGATAGTCCTGAGAACTGGGAGAAAGTCTGGGACAACTGGAGGCTGCTGACAATGGCCGGGATCTTTGACTG CTGGGAGCCCCCAGAGGGAGGAGATGTCCTGTATTCCTATACCATCATCACAGTGGATTCCTGCAAAGGCTTGAGTGACATCCACCACAG GATGCCTGCCATATTAGATGGAGAGGAGGCAGTTTCTAAATGGCTTGACTTTGGTGAAGTCTCAACTCAGGAAGCTCTGAAATTAATCCA CCCAACAGAGAACATCACCTTCCATGCAGTCTCTTCTGTGGTGAACAACTCGCGAAACAACACTCCTGAGTGTCTGGCTCCTGTCGACTT GGTGGTCAAAAAGGAGCTCAGGGCAAGTGGCAGTAGCCAGAGGATGTTGCAGTGGTTGGCCACAAAGTCACCCAAAAAGGAAGACTCAAA AACACCTCAAAAGGAAGAGTCAGATGTTCCCCAGTGGTCCAGTCAGTTCCTGCAGAAGAGTCCACTCCCCACCAAGAGAGGCACTGCAGG ACTCCTAGAGCAATGGCTGAAGCGGGAGAAGGAGGAGGAACCTGTGGCCAAGCGTCCTTACAGCCAGTGACACAGGACTTTCAGAGACCA AGGCCAGGGTCTGCTGCACTGCTGTTCTGATAATAGGTTCTTAACATTGTATGTATATGTGTTTGCTTTGGGAGGAGGTGGCACTGTGTT AGTTGACAGTTGTGGGCTCATGTAGTCTTTTTTGCCATGAGTAGGAGCCCCTAGTGGGGCTGGTGGACAGCTTTGGAAGAGGTGTCCTGC TGCTGTTACCAGCCATGTGGGCCCCATAGGGGCACTGCGCCTGCTGCCCTTTCCTGGCAGGGCTGGTGGAGTCTTCCCTCAAAGCATGCC TTACCCAGCTGGGAAGTCTCTGCCCTGATCTGGTACTCCTTGTAGTAAGCTGTTTTCTGCTCAGCCACTGGGCTCTTTCACTTTTTTAGT >28678_28678_6_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000409185_HMCES_chr3_129017197_ENST00000389735_length(amino acids)=224AA_BP=20 MFIVLDLLGFPMQMPKELVIQSGSIGAADSPENWEKVWDNWRLLTMAGIFDCWEPPEGGDVLYSYTIITVDSCKGLSDIHHRMPAILDGE EAVSKWLDFGEVSTQEALKLIHPTENITFHAVSSVVNNSRNNTPECLAPVDLVVKKELRASGSSQRMLQWLATKSPKKEDSKTPQKEESD -------------------------------------------------------------- >28678_28678_7_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000409185_HMCES_chr3_129017197_ENST00000417226_length(transcript)=1258nt_BP=178nt GGGGAGCGCGGGCTGCGGAGAGGCGGGCCGGGCCAAGCGGAGCCGAGCGAGCGGGAGCGCGGCGTCCGGGAGGCGGCGGAGACGCGGGGC TCGGAGGTTTTTGAAATCATGAATCCTGTTTATAGTCCTGGATCTTCTGGGGTTCCCTATGCAAATGCCAAAGGAATTGGTTATCCAGTC AGGTAGCATTGGTGCTGCAGATAGTCCTGAGAACTGGGAGAAAGTCTGGGACAACTGGAGGCTGCTGACAATGGCCGGGATCTTTGACTG CTGGGAGCCCCCAGAGGGAGGAGATGTCCTGTATTCCTATACCATCATCACAGTGGATTCCTGCAAAGGCTTGAGTGACATCCACCACAG GATGCCTGCCATATTAGATGGAGAGGAGGCAGTTTCTAAATGGCTTGACTTTGGTGAAGTCTCAACTCAGGAAGCTCTGAAATTAATCCA CCCAACAGAGAACATCACCTTCCATGCAGTCTCTTCTGTGGTGAACAACTCGCGAAACAACACTCCTGAGTGTCTGGCTCCTGTCGACTT GGTGGTCAAAAAGGAGCTCAGGGCAAGTGGCAGTAGCCAGAGGATGTTGCAGTGGTTGGCCACAAAGTCACCCAAAAAGGAAGACTCAAA AACACCTCAAAAGGAAGAGTCAGATGTTCCCCAGTGGTCCAGTCAGTTCCTGCAGAAGAGTCCACTCCCCACCAAGAGAGGCACTGCAGG ACTCCTAGAGCAATGGCTGAAGCGGGAGAAGGAGGAGGAACCTGTGGCCAAGCGTCCTTACAGCCAGTGACACAGGACTTTCAGAGACCA AGGCCAGGGTCTGCTGCACTGCTGTTCTGATAATAGGTTCTTAACATTGTATGTATATGTGTTTGCTTTGGGAGGAGGTGGCACTGTGTT AGTTGACAGTTGTGGGCTCATGTAGTCTTTTTTGCCATGAGTAGGAGCCCCTAGTGGGGCTGGTGGACAGCTTTGGAAGAGGTGTCCTGC TGCTGTTACCAGCCATGTGGGCCCCATAGGGGCACTGCGCCTGCTGCCCTTTCCTGGCAGGGCTGGTGGAGTCTTCCCTCAAAGCATGCC TTACCCAGCTGGGAAGTCTCTGCCCTGATCTGGTACTCCTTGTAGTAAGCTGTTTTCTGCTCAGCCACTGGGCTCTTTCACTTTTTTAGT >28678_28678_7_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000409185_HMCES_chr3_129017197_ENST00000417226_length(amino acids)=224AA_BP=20 MFIVLDLLGFPMQMPKELVIQSGSIGAADSPENWEKVWDNWRLLTMAGIFDCWEPPEGGDVLYSYTIITVDSCKGLSDIHHRMPAILDGE EAVSKWLDFGEVSTQEALKLIHPTENITFHAVSSVVNNSRNNTPECLAPVDLVVKKELRASGSSQRMLQWLATKSPKKEDSKTPQKEESD -------------------------------------------------------------- >28678_28678_8_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000409185_HMCES_chr3_129017197_ENST00000502878_length(transcript)=1406nt_BP=178nt GGGGAGCGCGGGCTGCGGAGAGGCGGGCCGGGCCAAGCGGAGCCGAGCGAGCGGGAGCGCGGCGTCCGGGAGGCGGCGGAGACGCGGGGC TCGGAGGTTTTTGAAATCATGAATCCTGTTTATAGTCCTGGATCTTCTGGGGTTCCCTATGCAAATGCCAAAGGAATTGGTTATCCAGTC AGGTAGCATTGGTGCTGCAGATAGTCCTGAGAACTGGGAGAAAGTCTGGGACAACTGGAGGCTGCTGACAATGGCCGGGATCTTTGACTG CTGGGAGCCCCCAGAGGGAGGAGATGTCCTGTATTCCTATACCATCATCACAGTGGATTCCTGCAAAGGCTTGAGTGACATCCACCACAG GATGCCTGCCATATTAGATGGAGAGGAGGCAGTTTCTAAATGGCTTGACTTTGGTGAAGTCTCAACTCAGGAAGCTCTGAAATTAATCCA CCCAACAGAGAACATCACCTTCCATGCAGTCTCTTCTGTGGTGAACAACTCGCGAAACAACACTCCTGAGTGTCTGGCTCCTGTCGACTT GGTGGTCAAAAAGGAGCTCAGGGCAAGTGGCAGTAGCCAGAGGATGTTGCAGTGGTTGGCCACAAAGTCACCCAAAAAGGAAGACTCAAA AACACCTCAAAAGGAAGAGTCAGATGTTCCCCAGTGGTCCAGTCAGTTCCTGCAGAAGAGTCCACTCCCCACCAAGAGAGGCACTGCAGG ACTCCTAGAGCAATGGCTGAAGCGGGAGAAGGAGGAGGAACCTGTGGCCAAGCGTCCTTACAGCCAGTGACACAGGACTTTCAGAGACCA AGGCCAGGGTCTGCTGCACTGCTGTTCTGATAATAGGTTCTTAACATTGTATGTATATGTGTTTGCTTTGGGAGGAGGTGGCACTGTGTT AGTTGACAGTTGTGGGCTCATGTAGTCTTTTTTGCCATGAGTAGGAGCCCCTAGTGGGGCTGGTGGACAGCTTTGGAAGAGGTGTCCTGC TGCTGTTACCAGCCATGTGGGCCCCATAGGGGCACTGCGCCTGCTGCCCTTTCCTGGCAGGGCTGGTGGAGTCTTCCCTCAAAGCATGCC TTACCCAGCTGGGAAGTCTCTGCCCTGATCTGGTACTCCTTGTAGTAAGCTGTTTTCTGCTCAGCCACTGGGCTCTTTCACTTTTTTAGT TCTTAAAAATTTATTTTTAAGTTCTAAAATAAAATAAAAATAAGTTCTTAAAATTTATTTTTTTCCTGAATAAATTGTATTTGGTAAACT TCTGCCTACATTTTGGAAAGTGATGCTGGTGGGGAAAGTTCTAGATCTTACTTGGTTTCTTCTAGAATCAGTCTTCAGGAATGGATTTTG >28678_28678_8_FAM168B-HMCES_FAM168B_chr2_131840150_ENST00000409185_HMCES_chr3_129017197_ENST00000502878_length(amino acids)=224AA_BP=20 MFIVLDLLGFPMQMPKELVIQSGSIGAADSPENWEKVWDNWRLLTMAGIFDCWEPPEGGDVLYSYTIITVDSCKGLSDIHHRMPAILDGE EAVSKWLDFGEVSTQEALKLIHPTENITFHAVSSVVNNSRNNTPECLAPVDLVVKKELRASGSSQRMLQWLATKSPKKEDSKTPQKEESD -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for FAM168B-HMCES |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for FAM168B-HMCES |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for FAM168B-HMCES |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
