
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:FLNA-ACTG2 (FusionGDB2 ID:30627) |
Fusion Gene Summary for FLNA-ACTG2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: FLNA-ACTG2 | Fusion gene ID: 30627 | Hgene | Tgene | Gene symbol | FLNA | ACTG2 | Gene ID | 2316 | 72 |
| Gene name | filamin A | actin gamma 2, smooth muscle | |
| Synonyms | ABP-280|ABPX|CSBS|CVD1|FGS2|FLN|FLN-A|FLN1|FMD|MNS|NHBP|OPD|OPD1|OPD2|XLVD|XMVD | ACT|ACTA3|ACTE|ACTL3|ACTSG|VSCM | |
| Cytomap | Xq28 | 2p13.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | filamin-Aactin binding protein 280alpha-filaminendothelial actin-binding proteinepididymis secretory sperm binding proteinfilamin A, alphafilamin-1non-muscle filamin | actin, gamma-enteric smooth muscleactin, gamma 2, smooth muscle, entericactin-like proteinalpha-actin-3 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | P21333 | P63267 | |
| Ensembl transtripts involved in fusion gene | ENST00000344736, ENST00000360319, ENST00000369850, ENST00000422373, ENST00000369856, ENST00000498491, | ENST00000409624, ENST00000409918, ENST00000345517, ENST00000409731, | |
| Fusion gene scores | * DoF score | 24 X 29 X 9=6264 | 12 X 15 X 8=1440 |
| # samples | 34 | 20 | |
| ** MAII score | log2(34/6264*10)=-4.20347756115334 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(20/1440*10)=-2.84799690655495 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: FLNA [Title/Abstract] AND ACTG2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ACTG2(74146651)-FLNA(153590628), # samples:1 FLNA(153595765)-ACTG2(74140612), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | ACTG2-FLNA seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. ACTG2-FLNA seems lost the major protein functional domain in Tgene partner, which is a CGC due to the frame-shifted ORF. ACTG2-FLNA seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. ACTG2-FLNA seems lost the major protein functional domain in Tgene partner, which is a tumor suppressor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | FLNA | GO:0016479 | negative regulation of transcription by RNA polymerase I | 22307607 |
| Hgene | FLNA | GO:0030334 | regulation of cell migration | 16291724 |
| Hgene | FLNA | GO:0031532 | actin cytoskeleton reorganization | 10051605 |
| Hgene | FLNA | GO:0034394 | protein localization to cell surface | 18322202 |
| Hgene | FLNA | GO:0043113 | receptor clustering | 10692483 |
| Hgene | FLNA | GO:0043433 | negative regulation of DNA-binding transcription factor activity | 15684392 |
| Hgene | FLNA | GO:0044319 | wound healing, spreading of cells | 16291724 |
| Hgene | FLNA | GO:0045184 | establishment of protein localization | 18322202 |
| Hgene | FLNA | GO:0051764 | actin crosslink formation | 10051605 |
| Hgene | FLNA | GO:0072659 | protein localization to plasma membrane | 24951510 |
| Hgene | FLNA | GO:0090307 | mitotic spindle assembly | 18548008 |
| Hgene | FLNA | GO:1901381 | positive regulation of potassium ion transmembrane transport | 24951510 |
 Fusion gene breakpoints across FLNA (5'-gene) Fusion gene breakpoints across FLNA (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
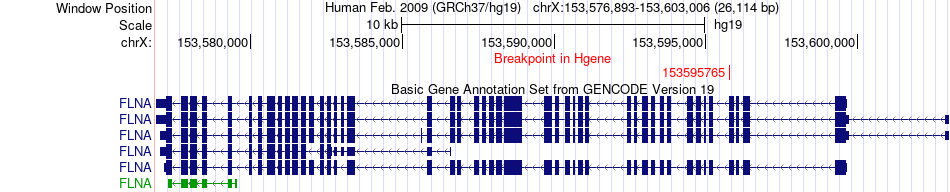 |
 Fusion gene breakpoints across ACTG2 (3'-gene) Fusion gene breakpoints across ACTG2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
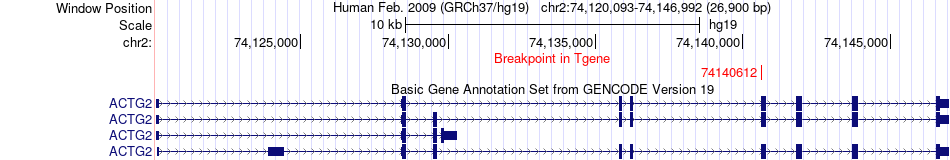 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | PRAD | TCGA-H9-A6BX-01A | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
Top |
Fusion Gene ORF analysis for FLNA-ACTG2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000344736 | ENST00000409624 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| 5CDS-intron | ENST00000344736 | ENST00000409918 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| 5CDS-intron | ENST00000360319 | ENST00000409624 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| 5CDS-intron | ENST00000360319 | ENST00000409918 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| 5CDS-intron | ENST00000369850 | ENST00000409624 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| 5CDS-intron | ENST00000369850 | ENST00000409918 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| 5CDS-intron | ENST00000422373 | ENST00000409624 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| 5CDS-intron | ENST00000422373 | ENST00000409918 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| In-frame | ENST00000344736 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| In-frame | ENST00000344736 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| In-frame | ENST00000360319 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| In-frame | ENST00000360319 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| In-frame | ENST00000369850 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| In-frame | ENST00000369850 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| In-frame | ENST00000422373 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| In-frame | ENST00000422373 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| intron-3CDS | ENST00000369856 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| intron-3CDS | ENST00000369856 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| intron-3CDS | ENST00000498491 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| intron-3CDS | ENST00000498491 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| intron-intron | ENST00000369856 | ENST00000409624 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| intron-intron | ENST00000369856 | ENST00000409918 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| intron-intron | ENST00000498491 | ENST00000409624 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
| intron-intron | ENST00000498491 | ENST00000409918 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000360319 | FLNA | chrX | 153595765 | - | ENST00000409731 | ACTG2 | chr2 | 74140612 | + | 1876 | 906 | 2 | 1585 | 527 |
| ENST00000360319 | FLNA | chrX | 153595765 | - | ENST00000345517 | ACTG2 | chr2 | 74140612 | + | 1876 | 906 | 2 | 1585 | 527 |
| ENST00000422373 | FLNA | chrX | 153595765 | - | ENST00000409731 | ACTG2 | chr2 | 74140612 | + | 2087 | 1117 | 213 | 1796 | 527 |
| ENST00000422373 | FLNA | chrX | 153595765 | - | ENST00000345517 | ACTG2 | chr2 | 74140612 | + | 2087 | 1117 | 213 | 1796 | 527 |
| ENST00000369850 | FLNA | chrX | 153595765 | - | ENST00000409731 | ACTG2 | chr2 | 74140612 | + | 2075 | 1105 | 201 | 1784 | 527 |
| ENST00000369850 | FLNA | chrX | 153595765 | - | ENST00000345517 | ACTG2 | chr2 | 74140612 | + | 2075 | 1105 | 201 | 1784 | 527 |
| ENST00000344736 | FLNA | chrX | 153595765 | - | ENST00000409731 | ACTG2 | chr2 | 74140612 | + | 1881 | 911 | 7 | 1590 | 527 |
| ENST00000344736 | FLNA | chrX | 153595765 | - | ENST00000345517 | ACTG2 | chr2 | 74140612 | + | 1881 | 911 | 7 | 1590 | 527 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000360319 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + | 0.014363902 | 0.9856361 |
| ENST00000360319 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + | 0.014363902 | 0.9856361 |
| ENST00000422373 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + | 0.016290585 | 0.98370945 |
| ENST00000422373 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + | 0.016290585 | 0.98370945 |
| ENST00000369850 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + | 0.01616069 | 0.9838394 |
| ENST00000369850 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + | 0.01616069 | 0.9838394 |
| ENST00000344736 | ENST00000409731 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + | 0.014432815 | 0.9855672 |
| ENST00000344736 | ENST00000345517 | FLNA | chrX | 153595765 | - | ACTG2 | chr2 | 74140612 | + | 0.014432815 | 0.9855672 |
Top |
Fusion Genomic Features for FLNA-ACTG2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| FLNA | chrX | 153595764 | - | ACTG2 | chr2 | 74140611 | + | 0.001036899 | 0.9989631 |
| FLNA | chrX | 153595764 | - | ACTG2 | chr2 | 74140611 | + | 0.001036899 | 0.9989631 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
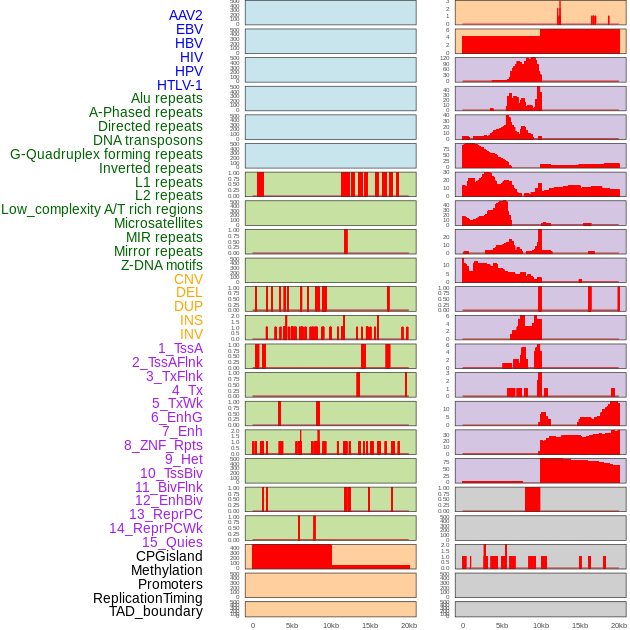 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
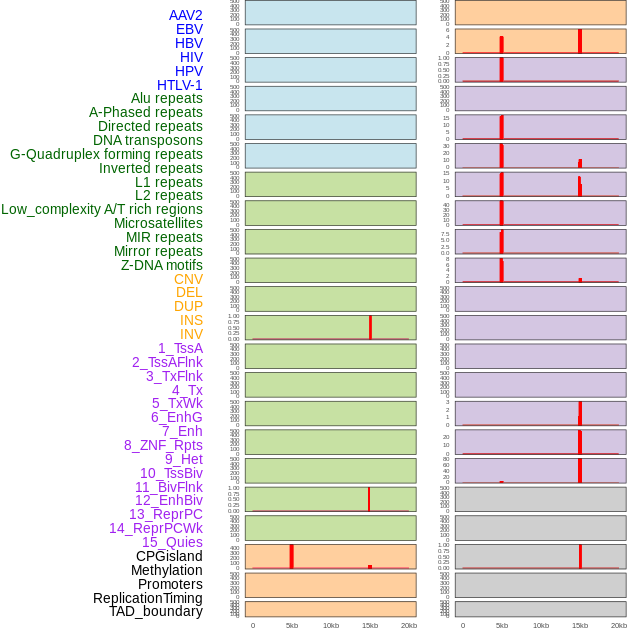 |
Top |
Fusion Protein Features for FLNA-ACTG2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chrX:74146651/chr2:153590628) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| FLNA | ACTG2 |
| FUNCTION: Promotes orthogonal branching of actin filaments and links actin filaments to membrane glycoproteins. Anchors various transmembrane proteins to the actin cytoskeleton and serves as a scaffold for a wide range of cytoplasmic signaling proteins. Interaction with FLNB may allow neuroblast migration from the ventricular zone into the cortical plate. Tethers cell surface-localized furin, modulates its rate of internalization and directs its intracellular trafficking (By similarity). Involved in ciliogenesis. Plays a role in cell-cell contacts and adherens junctions during the development of blood vessels, heart and brain organs. Plays a role in platelets morphology through interaction with SYK that regulates ITAM- and ITAM-like-containing receptor signaling, resulting in by platelet cytoskeleton organization maintenance (By similarity). During the axon guidance process, required for growth cone collapse induced by SEMA3A-mediated stimulation of neurons (PubMed:25358863). {ECO:0000250, ECO:0000250|UniProtKB:Q8BTM8, ECO:0000269|PubMed:22121117, ECO:0000269|PubMed:25358863}. | FUNCTION: Actins are highly conserved proteins that are involved in various types of cell motility and are ubiquitously expressed in all eukaryotic cells. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 166_269 | 289 | 2640.0 | Domain | Calponin-homology (CH) 2 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 43_149 | 289 | 2640.0 | Domain | Calponin-homology (CH) 1 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 166_269 | 289 | 2648.0 | Domain | Calponin-homology (CH) 2 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 43_149 | 289 | 2648.0 | Domain | Calponin-homology (CH) 1 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 166_269 | 289 | 2640.0 | Domain | Calponin-homology (CH) 2 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 43_149 | 289 | 2640.0 | Domain | Calponin-homology (CH) 1 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 2_274 | 289 | 2640.0 | Region | Note=Actin-binding |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 2_274 | 289 | 2648.0 | Region | Note=Actin-binding |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 2_274 | 289 | 2640.0 | Region | Note=Actin-binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1741_1778 | 289 | 2640.0 | Region | Note=Hinge 1 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 2517_2551 | 289 | 2640.0 | Region | Note=Hinge 2 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 2517_2647 | 289 | 2640.0 | Region | Note=Self-association site%2C tail |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1741_1778 | 289 | 2648.0 | Region | Note=Hinge 1 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 2517_2551 | 289 | 2648.0 | Region | Note=Hinge 2 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 2517_2647 | 289 | 2648.0 | Region | Note=Self-association site%2C tail |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1741_1778 | 289 | 2640.0 | Region | Note=Hinge 1 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 2517_2551 | 289 | 2640.0 | Region | Note=Hinge 2 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 2517_2647 | 289 | 2640.0 | Region | Note=Self-association site%2C tail |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1062_1154 | 289 | 2640.0 | Repeat | Note=Filamin 9 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1155_1249 | 289 | 2640.0 | Repeat | Note=Filamin 10 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1250_1349 | 289 | 2640.0 | Repeat | Note=Filamin 11 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1350_1442 | 289 | 2640.0 | Repeat | Note=Filamin 12 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1443_1539 | 289 | 2640.0 | Repeat | Note=Filamin 13 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1540_1636 | 289 | 2640.0 | Repeat | Note=Filamin 14 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1649_1740 | 289 | 2640.0 | Repeat | Note=Filamin 15 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1779_1860 | 289 | 2640.0 | Repeat | Note=Filamin 16 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1861_1950 | 289 | 2640.0 | Repeat | Note=Filamin 17 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1951_2039 | 289 | 2640.0 | Repeat | Note=Filamin 18 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 2042_2131 | 289 | 2640.0 | Repeat | Note=Filamin 19 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 2132_2230 | 289 | 2640.0 | Repeat | Note=Filamin 20 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 2233_2325 | 289 | 2640.0 | Repeat | Note=Filamin 21 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 2327_2420 | 289 | 2640.0 | Repeat | Note=Filamin 22 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 2424_2516 | 289 | 2640.0 | Repeat | Note=Filamin 23 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 2552_2646 | 289 | 2640.0 | Repeat | Note=Filamin 24 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 276_374 | 289 | 2640.0 | Repeat | Note=Filamin 1 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 376_474 | 289 | 2640.0 | Repeat | Note=Filamin 2 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 475_570 | 289 | 2640.0 | Repeat | Note=Filamin 3 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 571_663 | 289 | 2640.0 | Repeat | Note=Filamin 4 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 667_763 | 289 | 2640.0 | Repeat | Note=Filamin 5 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 764_866 | 289 | 2640.0 | Repeat | Note=Filamin 6 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 867_965 | 289 | 2640.0 | Repeat | Note=Filamin 7 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 966_1061 | 289 | 2640.0 | Repeat | Note=Filamin 8 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1062_1154 | 289 | 2648.0 | Repeat | Note=Filamin 9 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1155_1249 | 289 | 2648.0 | Repeat | Note=Filamin 10 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1250_1349 | 289 | 2648.0 | Repeat | Note=Filamin 11 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1350_1442 | 289 | 2648.0 | Repeat | Note=Filamin 12 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1443_1539 | 289 | 2648.0 | Repeat | Note=Filamin 13 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1540_1636 | 289 | 2648.0 | Repeat | Note=Filamin 14 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1649_1740 | 289 | 2648.0 | Repeat | Note=Filamin 15 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1779_1860 | 289 | 2648.0 | Repeat | Note=Filamin 16 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1861_1950 | 289 | 2648.0 | Repeat | Note=Filamin 17 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1951_2039 | 289 | 2648.0 | Repeat | Note=Filamin 18 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 2042_2131 | 289 | 2648.0 | Repeat | Note=Filamin 19 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 2132_2230 | 289 | 2648.0 | Repeat | Note=Filamin 20 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 2233_2325 | 289 | 2648.0 | Repeat | Note=Filamin 21 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 2327_2420 | 289 | 2648.0 | Repeat | Note=Filamin 22 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 2424_2516 | 289 | 2648.0 | Repeat | Note=Filamin 23 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 2552_2646 | 289 | 2648.0 | Repeat | Note=Filamin 24 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 276_374 | 289 | 2648.0 | Repeat | Note=Filamin 1 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 376_474 | 289 | 2648.0 | Repeat | Note=Filamin 2 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 475_570 | 289 | 2648.0 | Repeat | Note=Filamin 3 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 571_663 | 289 | 2648.0 | Repeat | Note=Filamin 4 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 667_763 | 289 | 2648.0 | Repeat | Note=Filamin 5 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 764_866 | 289 | 2648.0 | Repeat | Note=Filamin 6 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 867_965 | 289 | 2648.0 | Repeat | Note=Filamin 7 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 966_1061 | 289 | 2648.0 | Repeat | Note=Filamin 8 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1062_1154 | 289 | 2640.0 | Repeat | Note=Filamin 9 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1155_1249 | 289 | 2640.0 | Repeat | Note=Filamin 10 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1250_1349 | 289 | 2640.0 | Repeat | Note=Filamin 11 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1350_1442 | 289 | 2640.0 | Repeat | Note=Filamin 12 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1443_1539 | 289 | 2640.0 | Repeat | Note=Filamin 13 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1540_1636 | 289 | 2640.0 | Repeat | Note=Filamin 14 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1649_1740 | 289 | 2640.0 | Repeat | Note=Filamin 15 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1779_1860 | 289 | 2640.0 | Repeat | Note=Filamin 16 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1861_1950 | 289 | 2640.0 | Repeat | Note=Filamin 17 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1951_2039 | 289 | 2640.0 | Repeat | Note=Filamin 18 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 2042_2131 | 289 | 2640.0 | Repeat | Note=Filamin 19 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 2132_2230 | 289 | 2640.0 | Repeat | Note=Filamin 20 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 2233_2325 | 289 | 2640.0 | Repeat | Note=Filamin 21 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 2327_2420 | 289 | 2640.0 | Repeat | Note=Filamin 22 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 2424_2516 | 289 | 2640.0 | Repeat | Note=Filamin 23 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 2552_2646 | 289 | 2640.0 | Repeat | Note=Filamin 24 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 276_374 | 289 | 2640.0 | Repeat | Note=Filamin 1 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 376_474 | 289 | 2640.0 | Repeat | Note=Filamin 2 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 475_570 | 289 | 2640.0 | Repeat | Note=Filamin 3 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 571_663 | 289 | 2640.0 | Repeat | Note=Filamin 4 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 667_763 | 289 | 2640.0 | Repeat | Note=Filamin 5 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 764_866 | 289 | 2640.0 | Repeat | Note=Filamin 6 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 867_965 | 289 | 2640.0 | Repeat | Note=Filamin 7 |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 966_1061 | 289 | 2640.0 | Repeat | Note=Filamin 8 |
Top |
Fusion Gene Sequence for FLNA-ACTG2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >30627_30627_1_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000344736_ACTG2_chr2_74140612_ENST00000345517_length(transcript)=1881nt_BP=911nt GCCGAGGCTGCAGCACCGCCCCCCCGGCTTCTCGCGCCTCAAAATGAGTAGCTCCCACTCTCGGGCGGGCCAGAGCGCAGCAGGCGCGGC TCCGGGCGGCGGCGTCGACACGCGGGACGCCGAGATGCCGGCCACCGAGAAGGACCTGGCGGAGGACGCGCCGTGGAAGAAGATCCAGCA GAACACTTTCACGCGCTGGTGCAACGAGCACCTGAAGTGCGTGAGCAAGCGCATCGCCAACCTGCAGACGGACCTGAGCGACGGGCTGCG GCTTATCGCGCTGTTGGAGGTGCTCAGCCAGAAGAAGATGCACCGCAAGCACAACCAGCGGCCCACTTTCCGCCAAATGCAGCTTGAGAA CGTGTCGGTGGCGCTCGAGTTCCTGGACCGCGAGAGCATCAAACTGGTGTCCATCGACAGCAAGGCCATCGTGGACGGGAACCTGAAGCT GATCCTGGGCCTCATCTGGACCCTGATCCTGCACTACTCCATCTCCATGCCCATGTGGGACGAGGAGGAGGATGAGGAGGCCAAGAAGCA GACCCCCAAGCAGAGGCTCCTGGGCTGGATCCAGAACAAGCTGCCGCAGCTGCCCATCACCAACTTCAGCCGGGACTGGCAGAGCGGCCG GGCCCTGGGCGCCCTGGTGGACAGCTGTGCCCCGGGCCTGTGTCCTGACTGGGACTCTTGGGACGCCAGCAAGCCCGTTACCAATGCGCG AGAGGCCATGCAGCAGGCGGATGACTGGCTGGGCATCCCCCAGGTGATCACCCCCGAGGAGATTGTGGACCCCAACGTGGACGAGCACTC TGTCATGACCTACCTGTCCCAGTTCCCCAAGGCCAAGCTGAAGCCAGGGGCTCCCTTGCGGCCCAAACTGAACCCGAAGAAAGCCCGTGC CTACGGGCCAGGCATCGTCCTGGATTCAGGTGATGGCGTCACCCACAATGTCCCCATCTATGAAGGCTATGCCCTGCCCCATGCCATCAT GCGCCTGGACTTGGCTGGCCGTGACCTCACGGACTACCTCATGAAGATCCTCACAGAGAGAGGCTATTCCTTTGTGACCACAGCTGAGAG AGAAATTGTGCGAGACATCAAGGAGAAGCTGTGCTATGTGGCCCTGGATTTTGAGAATGAGATGGCCACAGCAGCTTCCTCTTCCTCCCT GGAGAAGAGCTATGAGCTGCCAGATGGGCAGGTTATCACCATTGGCAATGAGCGCTTCCGCTGCCCTGAGACCCTCTTCCAGCCTTCCTT TATTGGCATGGAGTCCGCTGGAATTCATGAGACAACCTACAATTCCATCATGAAGTGTGACATTGACATCCGTAAGGACTTATATGCCAA CAATGTCCTCTCTGGGGGCACCACCATGTACCCTGGCATTGCTGACAGGATGCAGAAGGAGATCACAGCCCTGGCCCCCAGCACCATGAA GATCAAGATTATTGCTCCCCCAGAGCGGAAGTACTCAGTCTGGATCGGGGGCTCTATCCTGGCCTCTCTCTCCACCTTCCAGCAGATGTG GATCAGCAAGCCTGAGTATGATGAGGCAGGGCCCTCCATTGTCCACAGGAAGTGCTTCTAAAGTCAGAACAGGTTCTCCAAGGATCCCCT CGAGACTACTCTGTTACCAGTCATGAAACATTAAAACCTACAAGCCTTACTTCTCTGTGTGGGGCTCTTTTTTCCTGGGCTATGTCTCAT ACACAGTGCTAAGGACTTTTCACACATTACTTTTAATCCATGCAATAGTGCTGTAAGGTAGGTGCTATCATTATACCCATATTACAGATG >30627_30627_1_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000344736_ACTG2_chr2_74140612_ENST00000345517_length(amino acids)=527AA_BP=301 MQHRPPGFSRLKMSSSHSRAGQSAAGAAPGGGVDTRDAEMPATEKDLAEDAPWKKIQQNTFTRWCNEHLKCVSKRIANLQTDLSDGLRLI ALLEVLSQKKMHRKHNQRPTFRQMQLENVSVALEFLDRESIKLVSIDSKAIVDGNLKLILGLIWTLILHYSISMPMWDEEEDEEAKKQTP KQRLLGWIQNKLPQLPITNFSRDWQSGRALGALVDSCAPGLCPDWDSWDASKPVTNAREAMQQADDWLGIPQVITPEEIVDPNVDEHSVM TYLSQFPKAKLKPGAPLRPKLNPKKARAYGPGIVLDSGDGVTHNVPIYEGYALPHAIMRLDLAGRDLTDYLMKILTERGYSFVTTAEREI VRDIKEKLCYVALDFENEMATAASSSSLEKSYELPDGQVITIGNERFRCPETLFQPSFIGMESAGIHETTYNSIMKCDIDIRKDLYANNV -------------------------------------------------------------- >30627_30627_2_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000344736_ACTG2_chr2_74140612_ENST00000409731_length(transcript)=1881nt_BP=911nt GCCGAGGCTGCAGCACCGCCCCCCCGGCTTCTCGCGCCTCAAAATGAGTAGCTCCCACTCTCGGGCGGGCCAGAGCGCAGCAGGCGCGGC TCCGGGCGGCGGCGTCGACACGCGGGACGCCGAGATGCCGGCCACCGAGAAGGACCTGGCGGAGGACGCGCCGTGGAAGAAGATCCAGCA GAACACTTTCACGCGCTGGTGCAACGAGCACCTGAAGTGCGTGAGCAAGCGCATCGCCAACCTGCAGACGGACCTGAGCGACGGGCTGCG GCTTATCGCGCTGTTGGAGGTGCTCAGCCAGAAGAAGATGCACCGCAAGCACAACCAGCGGCCCACTTTCCGCCAAATGCAGCTTGAGAA CGTGTCGGTGGCGCTCGAGTTCCTGGACCGCGAGAGCATCAAACTGGTGTCCATCGACAGCAAGGCCATCGTGGACGGGAACCTGAAGCT GATCCTGGGCCTCATCTGGACCCTGATCCTGCACTACTCCATCTCCATGCCCATGTGGGACGAGGAGGAGGATGAGGAGGCCAAGAAGCA GACCCCCAAGCAGAGGCTCCTGGGCTGGATCCAGAACAAGCTGCCGCAGCTGCCCATCACCAACTTCAGCCGGGACTGGCAGAGCGGCCG GGCCCTGGGCGCCCTGGTGGACAGCTGTGCCCCGGGCCTGTGTCCTGACTGGGACTCTTGGGACGCCAGCAAGCCCGTTACCAATGCGCG AGAGGCCATGCAGCAGGCGGATGACTGGCTGGGCATCCCCCAGGTGATCACCCCCGAGGAGATTGTGGACCCCAACGTGGACGAGCACTC TGTCATGACCTACCTGTCCCAGTTCCCCAAGGCCAAGCTGAAGCCAGGGGCTCCCTTGCGGCCCAAACTGAACCCGAAGAAAGCCCGTGC CTACGGGCCAGGCATCGTCCTGGATTCAGGTGATGGCGTCACCCACAATGTCCCCATCTATGAAGGCTATGCCCTGCCCCATGCCATCAT GCGCCTGGACTTGGCTGGCCGTGACCTCACGGACTACCTCATGAAGATCCTCACAGAGAGAGGCTATTCCTTTGTGACCACAGCTGAGAG AGAAATTGTGCGAGACATCAAGGAGAAGCTGTGCTATGTGGCCCTGGATTTTGAGAATGAGATGGCCACAGCAGCTTCCTCTTCCTCCCT GGAGAAGAGCTATGAGCTGCCAGATGGGCAGGTTATCACCATTGGCAATGAGCGCTTCCGCTGCCCTGAGACCCTCTTCCAGCCTTCCTT TATTGGCATGGAGTCCGCTGGAATTCATGAGACAACCTACAATTCCATCATGAAGTGTGACATTGACATCCGTAAGGACTTATATGCCAA CAATGTCCTCTCTGGGGGCACCACCATGTACCCTGGCATTGCTGACAGGATGCAGAAGGAGATCACAGCCCTGGCCCCCAGCACCATGAA GATCAAGATTATTGCTCCCCCAGAGCGGAAGTACTCAGTCTGGATCGGGGGCTCTATCCTGGCCTCTCTCTCCACCTTCCAGCAGATGTG GATCAGCAAGCCTGAGTATGATGAGGCAGGGCCCTCCATTGTCCACAGGAAGTGCTTCTAAAGTCAGAACAGGTTCTCCAAGGATCCCCT CGAGACTACTCTGTTACCAGTCATGAAACATTAAAACCTACAAGCCTTACTTCTCTGTGTGGGGCTCTTTTTTCCTGGGCTATGTCTCAT ACACAGTGCTAAGGACTTTTCACACATTACTTTTAATCCATGCAATAGTGCTGTAAGGTAGGTGCTATCATTATACCCATATTACAGATG >30627_30627_2_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000344736_ACTG2_chr2_74140612_ENST00000409731_length(amino acids)=527AA_BP=301 MQHRPPGFSRLKMSSSHSRAGQSAAGAAPGGGVDTRDAEMPATEKDLAEDAPWKKIQQNTFTRWCNEHLKCVSKRIANLQTDLSDGLRLI ALLEVLSQKKMHRKHNQRPTFRQMQLENVSVALEFLDRESIKLVSIDSKAIVDGNLKLILGLIWTLILHYSISMPMWDEEEDEEAKKQTP KQRLLGWIQNKLPQLPITNFSRDWQSGRALGALVDSCAPGLCPDWDSWDASKPVTNAREAMQQADDWLGIPQVITPEEIVDPNVDEHSVM TYLSQFPKAKLKPGAPLRPKLNPKKARAYGPGIVLDSGDGVTHNVPIYEGYALPHAIMRLDLAGRDLTDYLMKILTERGYSFVTTAEREI VRDIKEKLCYVALDFENEMATAASSSSLEKSYELPDGQVITIGNERFRCPETLFQPSFIGMESAGIHETTYNSIMKCDIDIRKDLYANNV -------------------------------------------------------------- >30627_30627_3_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000360319_ACTG2_chr2_74140612_ENST00000345517_length(transcript)=1876nt_BP=906nt GGCTGCAGCACCGCCCCCCCGGCTTCTCGCGCCTCAAAATGAGTAGCTCCCACTCTCGGGCGGGCCAGAGCGCAGCAGGCGCGGCTCCGG GCGGCGGCGTCGACACGCGGGACGCCGAGATGCCGGCCACCGAGAAGGACCTGGCGGAGGACGCGCCGTGGAAGAAGATCCAGCAGAACA CTTTCACGCGCTGGTGCAACGAGCACCTGAAGTGCGTGAGCAAGCGCATCGCCAACCTGCAGACGGACCTGAGCGACGGGCTGCGGCTTA TCGCGCTGTTGGAGGTGCTCAGCCAGAAGAAGATGCACCGCAAGCACAACCAGCGGCCCACTTTCCGCCAAATGCAGCTTGAGAACGTGT CGGTGGCGCTCGAGTTCCTGGACCGCGAGAGCATCAAACTGGTGTCCATCGACAGCAAGGCCATCGTGGACGGGAACCTGAAGCTGATCC TGGGCCTCATCTGGACCCTGATCCTGCACTACTCCATCTCCATGCCCATGTGGGACGAGGAGGAGGATGAGGAGGCCAAGAAGCAGACCC CCAAGCAGAGGCTCCTGGGCTGGATCCAGAACAAGCTGCCGCAGCTGCCCATCACCAACTTCAGCCGGGACTGGCAGAGCGGCCGGGCCC TGGGCGCCCTGGTGGACAGCTGTGCCCCGGGCCTGTGTCCTGACTGGGACTCTTGGGACGCCAGCAAGCCCGTTACCAATGCGCGAGAGG CCATGCAGCAGGCGGATGACTGGCTGGGCATCCCCCAGGTGATCACCCCCGAGGAGATTGTGGACCCCAACGTGGACGAGCACTCTGTCA TGACCTACCTGTCCCAGTTCCCCAAGGCCAAGCTGAAGCCAGGGGCTCCCTTGCGGCCCAAACTGAACCCGAAGAAAGCCCGTGCCTACG GGCCAGGCATCGTCCTGGATTCAGGTGATGGCGTCACCCACAATGTCCCCATCTATGAAGGCTATGCCCTGCCCCATGCCATCATGCGCC TGGACTTGGCTGGCCGTGACCTCACGGACTACCTCATGAAGATCCTCACAGAGAGAGGCTATTCCTTTGTGACCACAGCTGAGAGAGAAA TTGTGCGAGACATCAAGGAGAAGCTGTGCTATGTGGCCCTGGATTTTGAGAATGAGATGGCCACAGCAGCTTCCTCTTCCTCCCTGGAGA AGAGCTATGAGCTGCCAGATGGGCAGGTTATCACCATTGGCAATGAGCGCTTCCGCTGCCCTGAGACCCTCTTCCAGCCTTCCTTTATTG GCATGGAGTCCGCTGGAATTCATGAGACAACCTACAATTCCATCATGAAGTGTGACATTGACATCCGTAAGGACTTATATGCCAACAATG TCCTCTCTGGGGGCACCACCATGTACCCTGGCATTGCTGACAGGATGCAGAAGGAGATCACAGCCCTGGCCCCCAGCACCATGAAGATCA AGATTATTGCTCCCCCAGAGCGGAAGTACTCAGTCTGGATCGGGGGCTCTATCCTGGCCTCTCTCTCCACCTTCCAGCAGATGTGGATCA GCAAGCCTGAGTATGATGAGGCAGGGCCCTCCATTGTCCACAGGAAGTGCTTCTAAAGTCAGAACAGGTTCTCCAAGGATCCCCTCGAGA CTACTCTGTTACCAGTCATGAAACATTAAAACCTACAAGCCTTACTTCTCTGTGTGGGGCTCTTTTTTCCTGGGCTATGTCTCATACACA GTGCTAAGGACTTTTCACACATTACTTTTAATCCATGCAATAGTGCTGTAAGGTAGGTGCTATCATTATACCCATATTACAGATGAGGAA >30627_30627_3_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000360319_ACTG2_chr2_74140612_ENST00000345517_length(amino acids)=527AA_BP=301 LQHRPPGFSRLKMSSSHSRAGQSAAGAAPGGGVDTRDAEMPATEKDLAEDAPWKKIQQNTFTRWCNEHLKCVSKRIANLQTDLSDGLRLI ALLEVLSQKKMHRKHNQRPTFRQMQLENVSVALEFLDRESIKLVSIDSKAIVDGNLKLILGLIWTLILHYSISMPMWDEEEDEEAKKQTP KQRLLGWIQNKLPQLPITNFSRDWQSGRALGALVDSCAPGLCPDWDSWDASKPVTNAREAMQQADDWLGIPQVITPEEIVDPNVDEHSVM TYLSQFPKAKLKPGAPLRPKLNPKKARAYGPGIVLDSGDGVTHNVPIYEGYALPHAIMRLDLAGRDLTDYLMKILTERGYSFVTTAEREI VRDIKEKLCYVALDFENEMATAASSSSLEKSYELPDGQVITIGNERFRCPETLFQPSFIGMESAGIHETTYNSIMKCDIDIRKDLYANNV -------------------------------------------------------------- >30627_30627_4_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000360319_ACTG2_chr2_74140612_ENST00000409731_length(transcript)=1876nt_BP=906nt GGCTGCAGCACCGCCCCCCCGGCTTCTCGCGCCTCAAAATGAGTAGCTCCCACTCTCGGGCGGGCCAGAGCGCAGCAGGCGCGGCTCCGG GCGGCGGCGTCGACACGCGGGACGCCGAGATGCCGGCCACCGAGAAGGACCTGGCGGAGGACGCGCCGTGGAAGAAGATCCAGCAGAACA CTTTCACGCGCTGGTGCAACGAGCACCTGAAGTGCGTGAGCAAGCGCATCGCCAACCTGCAGACGGACCTGAGCGACGGGCTGCGGCTTA TCGCGCTGTTGGAGGTGCTCAGCCAGAAGAAGATGCACCGCAAGCACAACCAGCGGCCCACTTTCCGCCAAATGCAGCTTGAGAACGTGT CGGTGGCGCTCGAGTTCCTGGACCGCGAGAGCATCAAACTGGTGTCCATCGACAGCAAGGCCATCGTGGACGGGAACCTGAAGCTGATCC TGGGCCTCATCTGGACCCTGATCCTGCACTACTCCATCTCCATGCCCATGTGGGACGAGGAGGAGGATGAGGAGGCCAAGAAGCAGACCC CCAAGCAGAGGCTCCTGGGCTGGATCCAGAACAAGCTGCCGCAGCTGCCCATCACCAACTTCAGCCGGGACTGGCAGAGCGGCCGGGCCC TGGGCGCCCTGGTGGACAGCTGTGCCCCGGGCCTGTGTCCTGACTGGGACTCTTGGGACGCCAGCAAGCCCGTTACCAATGCGCGAGAGG CCATGCAGCAGGCGGATGACTGGCTGGGCATCCCCCAGGTGATCACCCCCGAGGAGATTGTGGACCCCAACGTGGACGAGCACTCTGTCA TGACCTACCTGTCCCAGTTCCCCAAGGCCAAGCTGAAGCCAGGGGCTCCCTTGCGGCCCAAACTGAACCCGAAGAAAGCCCGTGCCTACG GGCCAGGCATCGTCCTGGATTCAGGTGATGGCGTCACCCACAATGTCCCCATCTATGAAGGCTATGCCCTGCCCCATGCCATCATGCGCC TGGACTTGGCTGGCCGTGACCTCACGGACTACCTCATGAAGATCCTCACAGAGAGAGGCTATTCCTTTGTGACCACAGCTGAGAGAGAAA TTGTGCGAGACATCAAGGAGAAGCTGTGCTATGTGGCCCTGGATTTTGAGAATGAGATGGCCACAGCAGCTTCCTCTTCCTCCCTGGAGA AGAGCTATGAGCTGCCAGATGGGCAGGTTATCACCATTGGCAATGAGCGCTTCCGCTGCCCTGAGACCCTCTTCCAGCCTTCCTTTATTG GCATGGAGTCCGCTGGAATTCATGAGACAACCTACAATTCCATCATGAAGTGTGACATTGACATCCGTAAGGACTTATATGCCAACAATG TCCTCTCTGGGGGCACCACCATGTACCCTGGCATTGCTGACAGGATGCAGAAGGAGATCACAGCCCTGGCCCCCAGCACCATGAAGATCA AGATTATTGCTCCCCCAGAGCGGAAGTACTCAGTCTGGATCGGGGGCTCTATCCTGGCCTCTCTCTCCACCTTCCAGCAGATGTGGATCA GCAAGCCTGAGTATGATGAGGCAGGGCCCTCCATTGTCCACAGGAAGTGCTTCTAAAGTCAGAACAGGTTCTCCAAGGATCCCCTCGAGA CTACTCTGTTACCAGTCATGAAACATTAAAACCTACAAGCCTTACTTCTCTGTGTGGGGCTCTTTTTTCCTGGGCTATGTCTCATACACA GTGCTAAGGACTTTTCACACATTACTTTTAATCCATGCAATAGTGCTGTAAGGTAGGTGCTATCATTATACCCATATTACAGATGAGGAA >30627_30627_4_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000360319_ACTG2_chr2_74140612_ENST00000409731_length(amino acids)=527AA_BP=301 LQHRPPGFSRLKMSSSHSRAGQSAAGAAPGGGVDTRDAEMPATEKDLAEDAPWKKIQQNTFTRWCNEHLKCVSKRIANLQTDLSDGLRLI ALLEVLSQKKMHRKHNQRPTFRQMQLENVSVALEFLDRESIKLVSIDSKAIVDGNLKLILGLIWTLILHYSISMPMWDEEEDEEAKKQTP KQRLLGWIQNKLPQLPITNFSRDWQSGRALGALVDSCAPGLCPDWDSWDASKPVTNAREAMQQADDWLGIPQVITPEEIVDPNVDEHSVM TYLSQFPKAKLKPGAPLRPKLNPKKARAYGPGIVLDSGDGVTHNVPIYEGYALPHAIMRLDLAGRDLTDYLMKILTERGYSFVTTAEREI VRDIKEKLCYVALDFENEMATAASSSSLEKSYELPDGQVITIGNERFRCPETLFQPSFIGMESAGIHETTYNSIMKCDIDIRKDLYANNV -------------------------------------------------------------- >30627_30627_5_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000369850_ACTG2_chr2_74140612_ENST00000345517_length(transcript)=2075nt_BP=1105nt GCGCGTCGCGCGCAGCGGACGCCGACAGAATCCCCGAGGCGCCTGGCGCGGGCGCGGGCGCGAAGGCGATCCGGGCGCCACCCCGCGGTC ATCGGTCACCGGTCGCTCTCAGGAACAGCAGCGCAACCTCTGCTCCCTGCCTCGCCTCCCGCGCGCCTAGGTGCCTGCGACTTTAATTAA AGGGCCGTCCCCTCGCCGAGGCTGCAGCACCGCCCCCCCGGCTTCTCGCGCCTCAAAATGAGTAGCTCCCACTCTCGGGCGGGCCAGAGC GCAGCAGGCGCGGCTCCGGGCGGCGGCGTCGACACGCGGGACGCCGAGATGCCGGCCACCGAGAAGGACCTGGCGGAGGACGCGCCGTGG AAGAAGATCCAGCAGAACACTTTCACGCGCTGGTGCAACGAGCACCTGAAGTGCGTGAGCAAGCGCATCGCCAACCTGCAGACGGACCTG AGCGACGGGCTGCGGCTTATCGCGCTGTTGGAGGTGCTCAGCCAGAAGAAGATGCACCGCAAGCACAACCAGCGGCCCACTTTCCGCCAA ATGCAGCTTGAGAACGTGTCGGTGGCGCTCGAGTTCCTGGACCGCGAGAGCATCAAACTGGTGTCCATCGACAGCAAGGCCATCGTGGAC GGGAACCTGAAGCTGATCCTGGGCCTCATCTGGACCCTGATCCTGCACTACTCCATCTCCATGCCCATGTGGGACGAGGAGGAGGATGAG GAGGCCAAGAAGCAGACCCCCAAGCAGAGGCTCCTGGGCTGGATCCAGAACAAGCTGCCGCAGCTGCCCATCACCAACTTCAGCCGGGAC TGGCAGAGCGGCCGGGCCCTGGGCGCCCTGGTGGACAGCTGTGCCCCGGGCCTGTGTCCTGACTGGGACTCTTGGGACGCCAGCAAGCCC GTTACCAATGCGCGAGAGGCCATGCAGCAGGCGGATGACTGGCTGGGCATCCCCCAGGTGATCACCCCCGAGGAGATTGTGGACCCCAAC GTGGACGAGCACTCTGTCATGACCTACCTGTCCCAGTTCCCCAAGGCCAAGCTGAAGCCAGGGGCTCCCTTGCGGCCCAAACTGAACCCG AAGAAAGCCCGTGCCTACGGGCCAGGCATCGTCCTGGATTCAGGTGATGGCGTCACCCACAATGTCCCCATCTATGAAGGCTATGCCCTG CCCCATGCCATCATGCGCCTGGACTTGGCTGGCCGTGACCTCACGGACTACCTCATGAAGATCCTCACAGAGAGAGGCTATTCCTTTGTG ACCACAGCTGAGAGAGAAATTGTGCGAGACATCAAGGAGAAGCTGTGCTATGTGGCCCTGGATTTTGAGAATGAGATGGCCACAGCAGCT TCCTCTTCCTCCCTGGAGAAGAGCTATGAGCTGCCAGATGGGCAGGTTATCACCATTGGCAATGAGCGCTTCCGCTGCCCTGAGACCCTC TTCCAGCCTTCCTTTATTGGCATGGAGTCCGCTGGAATTCATGAGACAACCTACAATTCCATCATGAAGTGTGACATTGACATCCGTAAG GACTTATATGCCAACAATGTCCTCTCTGGGGGCACCACCATGTACCCTGGCATTGCTGACAGGATGCAGAAGGAGATCACAGCCCTGGCC CCCAGCACCATGAAGATCAAGATTATTGCTCCCCCAGAGCGGAAGTACTCAGTCTGGATCGGGGGCTCTATCCTGGCCTCTCTCTCCACC TTCCAGCAGATGTGGATCAGCAAGCCTGAGTATGATGAGGCAGGGCCCTCCATTGTCCACAGGAAGTGCTTCTAAAGTCAGAACAGGTTC TCCAAGGATCCCCTCGAGACTACTCTGTTACCAGTCATGAAACATTAAAACCTACAAGCCTTACTTCTCTGTGTGGGGCTCTTTTTTCCT GGGCTATGTCTCATACACAGTGCTAAGGACTTTTCACACATTACTTTTAATCCATGCAATAGTGCTGTAAGGTAGGTGCTATCATTATAC CCATATTACAGATGAGGAAATTGAGGCTCAGAGAAGTCAAGGACTTGCGAAGATCACACAGATTCCAGATTAAAATTCAAGTATCTGACT >30627_30627_5_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000369850_ACTG2_chr2_74140612_ENST00000345517_length(amino acids)=527AA_BP=301 MQHRPPGFSRLKMSSSHSRAGQSAAGAAPGGGVDTRDAEMPATEKDLAEDAPWKKIQQNTFTRWCNEHLKCVSKRIANLQTDLSDGLRLI ALLEVLSQKKMHRKHNQRPTFRQMQLENVSVALEFLDRESIKLVSIDSKAIVDGNLKLILGLIWTLILHYSISMPMWDEEEDEEAKKQTP KQRLLGWIQNKLPQLPITNFSRDWQSGRALGALVDSCAPGLCPDWDSWDASKPVTNAREAMQQADDWLGIPQVITPEEIVDPNVDEHSVM TYLSQFPKAKLKPGAPLRPKLNPKKARAYGPGIVLDSGDGVTHNVPIYEGYALPHAIMRLDLAGRDLTDYLMKILTERGYSFVTTAEREI VRDIKEKLCYVALDFENEMATAASSSSLEKSYELPDGQVITIGNERFRCPETLFQPSFIGMESAGIHETTYNSIMKCDIDIRKDLYANNV -------------------------------------------------------------- >30627_30627_6_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000369850_ACTG2_chr2_74140612_ENST00000409731_length(transcript)=2075nt_BP=1105nt GCGCGTCGCGCGCAGCGGACGCCGACAGAATCCCCGAGGCGCCTGGCGCGGGCGCGGGCGCGAAGGCGATCCGGGCGCCACCCCGCGGTC ATCGGTCACCGGTCGCTCTCAGGAACAGCAGCGCAACCTCTGCTCCCTGCCTCGCCTCCCGCGCGCCTAGGTGCCTGCGACTTTAATTAA AGGGCCGTCCCCTCGCCGAGGCTGCAGCACCGCCCCCCCGGCTTCTCGCGCCTCAAAATGAGTAGCTCCCACTCTCGGGCGGGCCAGAGC GCAGCAGGCGCGGCTCCGGGCGGCGGCGTCGACACGCGGGACGCCGAGATGCCGGCCACCGAGAAGGACCTGGCGGAGGACGCGCCGTGG AAGAAGATCCAGCAGAACACTTTCACGCGCTGGTGCAACGAGCACCTGAAGTGCGTGAGCAAGCGCATCGCCAACCTGCAGACGGACCTG AGCGACGGGCTGCGGCTTATCGCGCTGTTGGAGGTGCTCAGCCAGAAGAAGATGCACCGCAAGCACAACCAGCGGCCCACTTTCCGCCAA ATGCAGCTTGAGAACGTGTCGGTGGCGCTCGAGTTCCTGGACCGCGAGAGCATCAAACTGGTGTCCATCGACAGCAAGGCCATCGTGGAC GGGAACCTGAAGCTGATCCTGGGCCTCATCTGGACCCTGATCCTGCACTACTCCATCTCCATGCCCATGTGGGACGAGGAGGAGGATGAG GAGGCCAAGAAGCAGACCCCCAAGCAGAGGCTCCTGGGCTGGATCCAGAACAAGCTGCCGCAGCTGCCCATCACCAACTTCAGCCGGGAC TGGCAGAGCGGCCGGGCCCTGGGCGCCCTGGTGGACAGCTGTGCCCCGGGCCTGTGTCCTGACTGGGACTCTTGGGACGCCAGCAAGCCC GTTACCAATGCGCGAGAGGCCATGCAGCAGGCGGATGACTGGCTGGGCATCCCCCAGGTGATCACCCCCGAGGAGATTGTGGACCCCAAC GTGGACGAGCACTCTGTCATGACCTACCTGTCCCAGTTCCCCAAGGCCAAGCTGAAGCCAGGGGCTCCCTTGCGGCCCAAACTGAACCCG AAGAAAGCCCGTGCCTACGGGCCAGGCATCGTCCTGGATTCAGGTGATGGCGTCACCCACAATGTCCCCATCTATGAAGGCTATGCCCTG CCCCATGCCATCATGCGCCTGGACTTGGCTGGCCGTGACCTCACGGACTACCTCATGAAGATCCTCACAGAGAGAGGCTATTCCTTTGTG ACCACAGCTGAGAGAGAAATTGTGCGAGACATCAAGGAGAAGCTGTGCTATGTGGCCCTGGATTTTGAGAATGAGATGGCCACAGCAGCT TCCTCTTCCTCCCTGGAGAAGAGCTATGAGCTGCCAGATGGGCAGGTTATCACCATTGGCAATGAGCGCTTCCGCTGCCCTGAGACCCTC TTCCAGCCTTCCTTTATTGGCATGGAGTCCGCTGGAATTCATGAGACAACCTACAATTCCATCATGAAGTGTGACATTGACATCCGTAAG GACTTATATGCCAACAATGTCCTCTCTGGGGGCACCACCATGTACCCTGGCATTGCTGACAGGATGCAGAAGGAGATCACAGCCCTGGCC CCCAGCACCATGAAGATCAAGATTATTGCTCCCCCAGAGCGGAAGTACTCAGTCTGGATCGGGGGCTCTATCCTGGCCTCTCTCTCCACC TTCCAGCAGATGTGGATCAGCAAGCCTGAGTATGATGAGGCAGGGCCCTCCATTGTCCACAGGAAGTGCTTCTAAAGTCAGAACAGGTTC TCCAAGGATCCCCTCGAGACTACTCTGTTACCAGTCATGAAACATTAAAACCTACAAGCCTTACTTCTCTGTGTGGGGCTCTTTTTTCCT GGGCTATGTCTCATACACAGTGCTAAGGACTTTTCACACATTACTTTTAATCCATGCAATAGTGCTGTAAGGTAGGTGCTATCATTATAC CCATATTACAGATGAGGAAATTGAGGCTCAGAGAAGTCAAGGACTTGCGAAGATCACACAGATTCCAGATTAAAATTCAAGTATCTGACT >30627_30627_6_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000369850_ACTG2_chr2_74140612_ENST00000409731_length(amino acids)=527AA_BP=301 MQHRPPGFSRLKMSSSHSRAGQSAAGAAPGGGVDTRDAEMPATEKDLAEDAPWKKIQQNTFTRWCNEHLKCVSKRIANLQTDLSDGLRLI ALLEVLSQKKMHRKHNQRPTFRQMQLENVSVALEFLDRESIKLVSIDSKAIVDGNLKLILGLIWTLILHYSISMPMWDEEEDEEAKKQTP KQRLLGWIQNKLPQLPITNFSRDWQSGRALGALVDSCAPGLCPDWDSWDASKPVTNAREAMQQADDWLGIPQVITPEEIVDPNVDEHSVM TYLSQFPKAKLKPGAPLRPKLNPKKARAYGPGIVLDSGDGVTHNVPIYEGYALPHAIMRLDLAGRDLTDYLMKILTERGYSFVTTAEREI VRDIKEKLCYVALDFENEMATAASSSSLEKSYELPDGQVITIGNERFRCPETLFQPSFIGMESAGIHETTYNSIMKCDIDIRKDLYANNV -------------------------------------------------------------- >30627_30627_7_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000422373_ACTG2_chr2_74140612_ENST00000345517_length(transcript)=2087nt_BP=1117nt ATTCGCGTGGAGGCGCGTCGCGCGCAGCGGACGCCGACAGAATCCCCGAGGCGCCTGGCGCGGGCGCGGGCGCGAAGGCGATCCGGGCGC CACCCCGCGGTCATCGGTCACCGGTCGCTCTCAGGAACAGCAGCGCAACCTCTGCTCCCTGCCTCGCCTCCCGCGCGCCTAGGTGCCTGC GACTTTAATTAAAGGGCCGTCCCCTCGCCGAGGCTGCAGCACCGCCCCCCCGGCTTCTCGCGCCTCAAAATGAGTAGCTCCCACTCTCGG GCGGGCCAGAGCGCAGCAGGCGCGGCTCCGGGCGGCGGCGTCGACACGCGGGACGCCGAGATGCCGGCCACCGAGAAGGACCTGGCGGAG GACGCGCCGTGGAAGAAGATCCAGCAGAACACTTTCACGCGCTGGTGCAACGAGCACCTGAAGTGCGTGAGCAAGCGCATCGCCAACCTG CAGACGGACCTGAGCGACGGGCTGCGGCTTATCGCGCTGTTGGAGGTGCTCAGCCAGAAGAAGATGCACCGCAAGCACAACCAGCGGCCC ACTTTCCGCCAAATGCAGCTTGAGAACGTGTCGGTGGCGCTCGAGTTCCTGGACCGCGAGAGCATCAAACTGGTGTCCATCGACAGCAAG GCCATCGTGGACGGGAACCTGAAGCTGATCCTGGGCCTCATCTGGACCCTGATCCTGCACTACTCCATCTCCATGCCCATGTGGGACGAG GAGGAGGATGAGGAGGCCAAGAAGCAGACCCCCAAGCAGAGGCTCCTGGGCTGGATCCAGAACAAGCTGCCGCAGCTGCCCATCACCAAC TTCAGCCGGGACTGGCAGAGCGGCCGGGCCCTGGGCGCCCTGGTGGACAGCTGTGCCCCGGGCCTGTGTCCTGACTGGGACTCTTGGGAC GCCAGCAAGCCCGTTACCAATGCGCGAGAGGCCATGCAGCAGGCGGATGACTGGCTGGGCATCCCCCAGGTGATCACCCCCGAGGAGATT GTGGACCCCAACGTGGACGAGCACTCTGTCATGACCTACCTGTCCCAGTTCCCCAAGGCCAAGCTGAAGCCAGGGGCTCCCTTGCGGCCC AAACTGAACCCGAAGAAAGCCCGTGCCTACGGGCCAGGCATCGTCCTGGATTCAGGTGATGGCGTCACCCACAATGTCCCCATCTATGAA GGCTATGCCCTGCCCCATGCCATCATGCGCCTGGACTTGGCTGGCCGTGACCTCACGGACTACCTCATGAAGATCCTCACAGAGAGAGGC TATTCCTTTGTGACCACAGCTGAGAGAGAAATTGTGCGAGACATCAAGGAGAAGCTGTGCTATGTGGCCCTGGATTTTGAGAATGAGATG GCCACAGCAGCTTCCTCTTCCTCCCTGGAGAAGAGCTATGAGCTGCCAGATGGGCAGGTTATCACCATTGGCAATGAGCGCTTCCGCTGC CCTGAGACCCTCTTCCAGCCTTCCTTTATTGGCATGGAGTCCGCTGGAATTCATGAGACAACCTACAATTCCATCATGAAGTGTGACATT GACATCCGTAAGGACTTATATGCCAACAATGTCCTCTCTGGGGGCACCACCATGTACCCTGGCATTGCTGACAGGATGCAGAAGGAGATC ACAGCCCTGGCCCCCAGCACCATGAAGATCAAGATTATTGCTCCCCCAGAGCGGAAGTACTCAGTCTGGATCGGGGGCTCTATCCTGGCC TCTCTCTCCACCTTCCAGCAGATGTGGATCAGCAAGCCTGAGTATGATGAGGCAGGGCCCTCCATTGTCCACAGGAAGTGCTTCTAAAGT CAGAACAGGTTCTCCAAGGATCCCCTCGAGACTACTCTGTTACCAGTCATGAAACATTAAAACCTACAAGCCTTACTTCTCTGTGTGGGG CTCTTTTTTCCTGGGCTATGTCTCATACACAGTGCTAAGGACTTTTCACACATTACTTTTAATCCATGCAATAGTGCTGTAAGGTAGGTG CTATCATTATACCCATATTACAGATGAGGAAATTGAGGCTCAGAGAAGTCAAGGACTTGCGAAGATCACACAGATTCCAGATTAAAATTC >30627_30627_7_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000422373_ACTG2_chr2_74140612_ENST00000345517_length(amino acids)=527AA_BP=301 MQHRPPGFSRLKMSSSHSRAGQSAAGAAPGGGVDTRDAEMPATEKDLAEDAPWKKIQQNTFTRWCNEHLKCVSKRIANLQTDLSDGLRLI ALLEVLSQKKMHRKHNQRPTFRQMQLENVSVALEFLDRESIKLVSIDSKAIVDGNLKLILGLIWTLILHYSISMPMWDEEEDEEAKKQTP KQRLLGWIQNKLPQLPITNFSRDWQSGRALGALVDSCAPGLCPDWDSWDASKPVTNAREAMQQADDWLGIPQVITPEEIVDPNVDEHSVM TYLSQFPKAKLKPGAPLRPKLNPKKARAYGPGIVLDSGDGVTHNVPIYEGYALPHAIMRLDLAGRDLTDYLMKILTERGYSFVTTAEREI VRDIKEKLCYVALDFENEMATAASSSSLEKSYELPDGQVITIGNERFRCPETLFQPSFIGMESAGIHETTYNSIMKCDIDIRKDLYANNV -------------------------------------------------------------- >30627_30627_8_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000422373_ACTG2_chr2_74140612_ENST00000409731_length(transcript)=2087nt_BP=1117nt ATTCGCGTGGAGGCGCGTCGCGCGCAGCGGACGCCGACAGAATCCCCGAGGCGCCTGGCGCGGGCGCGGGCGCGAAGGCGATCCGGGCGC CACCCCGCGGTCATCGGTCACCGGTCGCTCTCAGGAACAGCAGCGCAACCTCTGCTCCCTGCCTCGCCTCCCGCGCGCCTAGGTGCCTGC GACTTTAATTAAAGGGCCGTCCCCTCGCCGAGGCTGCAGCACCGCCCCCCCGGCTTCTCGCGCCTCAAAATGAGTAGCTCCCACTCTCGG GCGGGCCAGAGCGCAGCAGGCGCGGCTCCGGGCGGCGGCGTCGACACGCGGGACGCCGAGATGCCGGCCACCGAGAAGGACCTGGCGGAG GACGCGCCGTGGAAGAAGATCCAGCAGAACACTTTCACGCGCTGGTGCAACGAGCACCTGAAGTGCGTGAGCAAGCGCATCGCCAACCTG CAGACGGACCTGAGCGACGGGCTGCGGCTTATCGCGCTGTTGGAGGTGCTCAGCCAGAAGAAGATGCACCGCAAGCACAACCAGCGGCCC ACTTTCCGCCAAATGCAGCTTGAGAACGTGTCGGTGGCGCTCGAGTTCCTGGACCGCGAGAGCATCAAACTGGTGTCCATCGACAGCAAG GCCATCGTGGACGGGAACCTGAAGCTGATCCTGGGCCTCATCTGGACCCTGATCCTGCACTACTCCATCTCCATGCCCATGTGGGACGAG GAGGAGGATGAGGAGGCCAAGAAGCAGACCCCCAAGCAGAGGCTCCTGGGCTGGATCCAGAACAAGCTGCCGCAGCTGCCCATCACCAAC TTCAGCCGGGACTGGCAGAGCGGCCGGGCCCTGGGCGCCCTGGTGGACAGCTGTGCCCCGGGCCTGTGTCCTGACTGGGACTCTTGGGAC GCCAGCAAGCCCGTTACCAATGCGCGAGAGGCCATGCAGCAGGCGGATGACTGGCTGGGCATCCCCCAGGTGATCACCCCCGAGGAGATT GTGGACCCCAACGTGGACGAGCACTCTGTCATGACCTACCTGTCCCAGTTCCCCAAGGCCAAGCTGAAGCCAGGGGCTCCCTTGCGGCCC AAACTGAACCCGAAGAAAGCCCGTGCCTACGGGCCAGGCATCGTCCTGGATTCAGGTGATGGCGTCACCCACAATGTCCCCATCTATGAA GGCTATGCCCTGCCCCATGCCATCATGCGCCTGGACTTGGCTGGCCGTGACCTCACGGACTACCTCATGAAGATCCTCACAGAGAGAGGC TATTCCTTTGTGACCACAGCTGAGAGAGAAATTGTGCGAGACATCAAGGAGAAGCTGTGCTATGTGGCCCTGGATTTTGAGAATGAGATG GCCACAGCAGCTTCCTCTTCCTCCCTGGAGAAGAGCTATGAGCTGCCAGATGGGCAGGTTATCACCATTGGCAATGAGCGCTTCCGCTGC CCTGAGACCCTCTTCCAGCCTTCCTTTATTGGCATGGAGTCCGCTGGAATTCATGAGACAACCTACAATTCCATCATGAAGTGTGACATT GACATCCGTAAGGACTTATATGCCAACAATGTCCTCTCTGGGGGCACCACCATGTACCCTGGCATTGCTGACAGGATGCAGAAGGAGATC ACAGCCCTGGCCCCCAGCACCATGAAGATCAAGATTATTGCTCCCCCAGAGCGGAAGTACTCAGTCTGGATCGGGGGCTCTATCCTGGCC TCTCTCTCCACCTTCCAGCAGATGTGGATCAGCAAGCCTGAGTATGATGAGGCAGGGCCCTCCATTGTCCACAGGAAGTGCTTCTAAAGT CAGAACAGGTTCTCCAAGGATCCCCTCGAGACTACTCTGTTACCAGTCATGAAACATTAAAACCTACAAGCCTTACTTCTCTGTGTGGGG CTCTTTTTTCCTGGGCTATGTCTCATACACAGTGCTAAGGACTTTTCACACATTACTTTTAATCCATGCAATAGTGCTGTAAGGTAGGTG CTATCATTATACCCATATTACAGATGAGGAAATTGAGGCTCAGAGAAGTCAAGGACTTGCGAAGATCACACAGATTCCAGATTAAAATTC >30627_30627_8_FLNA-ACTG2_FLNA_chrX_153595765_ENST00000422373_ACTG2_chr2_74140612_ENST00000409731_length(amino acids)=527AA_BP=301 MQHRPPGFSRLKMSSSHSRAGQSAAGAAPGGGVDTRDAEMPATEKDLAEDAPWKKIQQNTFTRWCNEHLKCVSKRIANLQTDLSDGLRLI ALLEVLSQKKMHRKHNQRPTFRQMQLENVSVALEFLDRESIKLVSIDSKAIVDGNLKLILGLIWTLILHYSISMPMWDEEEDEEAKKQTP KQRLLGWIQNKLPQLPITNFSRDWQSGRALGALVDSCAPGLCPDWDSWDASKPVTNAREAMQQADDWLGIPQVITPEEIVDPNVDEHSVM TYLSQFPKAKLKPGAPLRPKLNPKKARAYGPGIVLDSGDGVTHNVPIYEGYALPHAIMRLDLAGRDLTDYLMKILTERGYSFVTTAEREI VRDIKEKLCYVALDFENEMATAASSSSLEKSYELPDGQVITIGNERFRCPETLFQPSFIGMESAGIHETTYNSIMKCDIDIRKDLYANNV -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for FLNA-ACTG2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000360319 | - | 4 | 46 | 1490_1607 | 289.3333333333333 | 2640.0 | furin |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000369850 | - | 5 | 48 | 1490_1607 | 289.3333333333333 | 2648.0 | furin |
| Hgene | FLNA | chrX:153595765 | chr2:74140612 | ENST00000422373 | - | 5 | 47 | 1490_1607 | 289.3333333333333 | 2640.0 | furin |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for FLNA-ACTG2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for FLNA-ACTG2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
