
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:GLG1-FA2H (FusionGDB2 ID:33246) |
Fusion Gene Summary for GLG1-FA2H |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: GLG1-FA2H | Fusion gene ID: 33246 | Hgene | Tgene | Gene symbol | GLG1 | FA2H | Gene ID | 2734 | 79152 |
| Gene name | golgi glycoprotein 1 | fatty acid 2-hydroxylase | |
| Synonyms | CFR-1|ESL-1|MG-160|MG160 | FAAH|FAH1|FAXDC1|SCS7|SPG35 | |
| Cytomap | 16q23.1 | 16q23.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | Golgi apparatus protein 1E-selectin ligand 1cysteine-rich fibroblast growth factor receptorgolgi sialoglycoprotein MG-160 | fatty acid 2-hydroxylasefatty acid alpha-hydroxylasefatty acid hydroxylase domain containing 1spastic paraplegia 35 (autosomal recessive) | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q92896 | Q7L5A8 | |
| Ensembl transtripts involved in fusion gene | ENST00000205061, ENST00000422840, ENST00000447066, | ENST00000219368, ENST00000544337, | |
| Fusion gene scores | * DoF score | 23 X 21 X 12=5796 | 3 X 3 X 2=18 |
| # samples | 29 | 3 | |
| ** MAII score | log2(29/5796*10)=-4.320932789542 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(3/18*10)=0.736965594166206 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: GLG1 [Title/Abstract] AND FA2H [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | FA2H(74773921)-GLG1(74506309), # samples:2 GLG1(74508444)-FA2H(74748167), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | FA2H | GO:0046513 | ceramide biosynthetic process | 15337768 |
 Fusion gene breakpoints across GLG1 (5'-gene) Fusion gene breakpoints across GLG1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
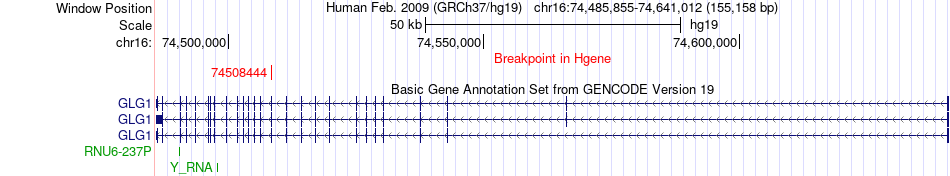 |
 Fusion gene breakpoints across FA2H (3'-gene) Fusion gene breakpoints across FA2H (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
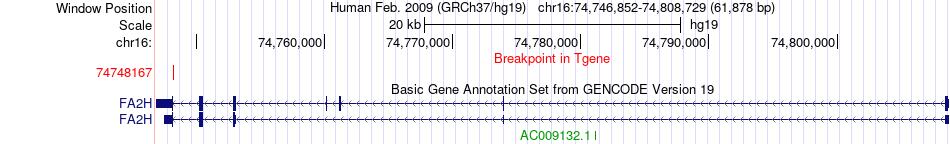 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | PAAD | TCGA-RL-AAAS-01A | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - |
Top |
Fusion Gene ORF analysis for GLG1-FA2H |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000205061 | ENST00000219368 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - |
| In-frame | ENST00000205061 | ENST00000544337 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - |
| In-frame | ENST00000422840 | ENST00000219368 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - |
| In-frame | ENST00000422840 | ENST00000544337 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - |
| In-frame | ENST00000447066 | ENST00000219368 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - |
| In-frame | ENST00000447066 | ENST00000544337 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000205061 | GLG1 | chr16 | 74508444 | - | ENST00000219368 | FA2H | chr16 | 74748167 | - | 3387 | 2072 | 20 | 2140 | 706 |
| ENST00000205061 | GLG1 | chr16 | 74508444 | - | ENST00000544337 | FA2H | chr16 | 74748167 | - | 2744 | 2072 | 20 | 2140 | 706 |
| ENST00000422840 | GLG1 | chr16 | 74508444 | - | ENST00000219368 | FA2H | chr16 | 74748167 | - | 3367 | 2052 | 0 | 2120 | 706 |
| ENST00000422840 | GLG1 | chr16 | 74508444 | - | ENST00000544337 | FA2H | chr16 | 74748167 | - | 2724 | 2052 | 0 | 2120 | 706 |
| ENST00000447066 | GLG1 | chr16 | 74508444 | - | ENST00000219368 | FA2H | chr16 | 74748167 | - | 3352 | 2037 | 18 | 2105 | 695 |
| ENST00000447066 | GLG1 | chr16 | 74508444 | - | ENST00000544337 | FA2H | chr16 | 74748167 | - | 2709 | 2037 | 18 | 2105 | 695 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000205061 | ENST00000219368 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - | 0.002034318 | 0.9979657 |
| ENST00000205061 | ENST00000544337 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - | 0.003681607 | 0.99631834 |
| ENST00000422840 | ENST00000219368 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - | 0.001915154 | 0.9980849 |
| ENST00000422840 | ENST00000544337 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - | 0.003430223 | 0.99656975 |
| ENST00000447066 | ENST00000219368 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - | 0.002968915 | 0.9970311 |
| ENST00000447066 | ENST00000544337 | GLG1 | chr16 | 74508444 | - | FA2H | chr16 | 74748167 | - | 0.005305613 | 0.9946944 |
Top |
Fusion Genomic Features for GLG1-FA2H |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
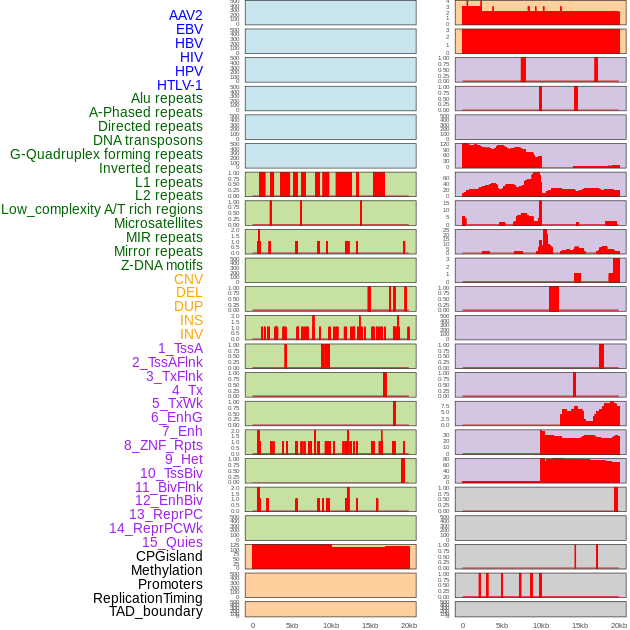 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for GLG1-FA2H |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr16:74773921/chr16:74506309) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| GLG1 | FA2H |
| FUNCTION: Binds fibroblast growth factor and E-selectin (cell-adhesion lectin on endothelial cells mediating the binding of neutrophils). {ECO:0000269|PubMed:8985126}. | FUNCTION: Catalyzes the hydroxylation of free fatty acids at the C-2 position to produce 2-hydroxy fatty acids, which are building blocks of sphingolipids and glycosphingolipids common in neural tissue and epidermis (PubMed:15337768, PubMed:15863841, PubMed:17355976, PubMed:22517924). FA2H is stereospecific for the production of (R)-2-hydroxy fatty acids (PubMed:22517924). Plays an essential role in the synthesis of galactosphingolipids of the myelin sheath (By similarity). Responsible for the synthesis of sphingolipids and glycosphingolipids involved in the formation of epidermal lamellar bodies critical for skin permeability barrier (PubMed:17355976). Participates in the synthesis of glycosphingolipids and a fraction of type II wax diesters in sebaceous gland, specifically regulating hair follicle homeostasis (By similarity). Involved in the synthesis of sphingolipids of plasma membrane rafts, controlling lipid raft mobility and trafficking of raft-associated proteins (By similarity). {ECO:0000250|UniProtKB:Q5MPP0, ECO:0000269|PubMed:15337768, ECO:0000269|PubMed:15863841, ECO:0000269|PubMed:17355976, ECO:0000269|PubMed:22517924}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 54_57 | 684 | 1204.0 | Compositional bias | Note=Poly-Gly |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 71_86 | 684 | 1204.0 | Compositional bias | Note=Poly-Gln |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 54_57 | 684 | 1180.0 | Compositional bias | Note=Poly-Gly |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 71_86 | 684 | 1180.0 | Compositional bias | Note=Poly-Gln |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 54_57 | 673 | 1193.0 | Compositional bias | Note=Poly-Gly |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 71_86 | 673 | 1193.0 | Compositional bias | Note=Poly-Gln |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 116_149 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 1 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 150_212 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 2 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 215_278 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 3 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 286_346 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 4 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 347_413 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 5 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 414_473 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 6 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 475_537 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 7 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 538_604 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 8 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 609_668 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 9 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 116_149 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 1 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 150_212 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 2 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 215_278 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 3 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 286_346 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 4 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 347_413 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 5 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 414_473 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 6 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 475_537 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 7 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 538_604 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 8 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 609_668 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 9 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 116_149 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 1 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 150_212 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 2 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 215_278 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 3 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 286_346 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 4 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 347_413 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 5 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 414_473 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 6 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 475_537 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 7 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 538_604 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 8 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 609_668 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 9 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 1041_1101 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 16 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 670_728 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 10 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 729_788 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 11 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 796_856 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 12 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 858_911 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 13 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 912_979 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 14 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 980_1035 | 684 | 1204.0 | Repeat | Note=Cys-rich GLG1 15 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 1041_1101 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 16 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 670_728 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 10 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 729_788 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 11 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 796_856 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 12 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 858_911 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 13 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 912_979 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 14 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 980_1035 | 684 | 1180.0 | Repeat | Note=Cys-rich GLG1 15 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 1041_1101 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 16 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 670_728 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 10 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 729_788 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 11 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 796_856 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 12 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 858_911 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 13 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 912_979 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 14 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 980_1035 | 673 | 1193.0 | Repeat | Note=Cys-rich GLG1 15 |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 1167_1179 | 684 | 1204.0 | Topological domain | Cytoplasmic |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 30_1145 | 684 | 1204.0 | Topological domain | Extracellular |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 1167_1179 | 684 | 1180.0 | Topological domain | Cytoplasmic |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 30_1145 | 684 | 1180.0 | Topological domain | Extracellular |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 1167_1179 | 673 | 1193.0 | Topological domain | Cytoplasmic |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 30_1145 | 673 | 1193.0 | Topological domain | Extracellular |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000205061 | - | 13 | 27 | 1146_1166 | 684 | 1204.0 | Transmembrane | Helical |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000422840 | - | 13 | 26 | 1146_1166 | 684 | 1180.0 | Transmembrane | Helical |
| Hgene | GLG1 | chr16:74508444 | chr16:74748167 | ENST00000447066 | - | 12 | 26 | 1146_1166 | 673 | 1193.0 | Transmembrane | Helical |
| Tgene | FA2H | chr16:74508444 | chr16:74748167 | ENST00000219368 | 5 | 7 | 219_361 | 346 | 373.0 | Domain | Fatty acid hydroxylase | |
| Tgene | FA2H | chr16:74508444 | chr16:74748167 | ENST00000219368 | 5 | 7 | 8_86 | 346 | 373.0 | Domain | Cytochrome b5 heme-binding | |
| Tgene | FA2H | chr16:74508444 | chr16:74748167 | ENST00000219368 | 5 | 7 | 168_188 | 346 | 373.0 | Transmembrane | Helical | |
| Tgene | FA2H | chr16:74508444 | chr16:74748167 | ENST00000219368 | 5 | 7 | 213_233 | 346 | 373.0 | Transmembrane | Helical | |
| Tgene | FA2H | chr16:74508444 | chr16:74748167 | ENST00000219368 | 5 | 7 | 268_288 | 346 | 373.0 | Transmembrane | Helical | |
| Tgene | FA2H | chr16:74508444 | chr16:74748167 | ENST00000219368 | 5 | 7 | 290_310 | 346 | 373.0 | Transmembrane | Helical |
Top |
Fusion Gene Sequence for GLG1-FA2H |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >33246_33246_1_GLG1-FA2H_GLG1_chr16_74508444_ENST00000205061_FA2H_chr16_74748167_ENST00000219368_length(transcript)=3387nt_BP=2072nt GAGCTCGCCGCGGACTCAAGATGGCGGCGTGTGGACGTGTACGGAGGATGTTCCGCTTGTCGGCGGCGCTGCATCTGCTGCTGCTATTCG CGGCCGGGGCCGAGAAACTCCCCGGCCAGGGCGTCCACAGCCAGGGCCAGGGTCCCGGGGCCAACTTTGTGTCCTTCGTAGGGCAGGCCG GAGGCGGCGGCCCGGCGGGTCAGCAGCTGCCCCAGCTGCCTCAGTCATCGCAGCTTCAGCAGCAACAGCAGCAGCAGCAACAGCAACAGC AGCCTCAGCCGCCGCAGCCGCCTTTCCCGGCGGGTGGGCCTCCGGCCCGGCGGGGAGGAGCGGGGGCTGGTGGGGGCTGGAAGCTGGCGG AGGAAGAGTCCTGCAGGGAGGACGTGACCCGCGTGTGCCCTAAGCACACCTGGAGCAACAACCTGGCGGTGCTCGAGTGCCTGCAGGATG TGAGGGAGCCTGAAAATGAAATTTCTTCAGACTGCAATCATTTGTTGTGGAATTATAAGCTGAACCTAACTACAGATCCCAAATTTGAAT CTGTGGCCAGAGAGGTTTGCAAATCTACTATAACAGAGATTAAAGAATGTGCTGATGAACCGGTTGGAAAAGGTTACATGGTTTCCTGCT TGGTGGATCACCGAGGCAACATCACTGAGTATCAGTGTCACCAGTACATTACCAAGATGACGGCCATCATTTTTAGTGATTACCGTTTAA TCTGTGGCTTCATGGATGACTGCAAAAATGACATCAACATTCTGAAATGTGGCAGTATTCGGCTTGGAGAAAAGGATGCACATTCACAAG GTGAGGTGGTATCATGCTTGGAGAAAGGCCTGGTGAAAGAAGCAGAAGAAAGAGAACCCAAGATTCAAGTTTCTGAACTCTGCAAGAAAG CCATTCTCCGGGTGGCTGAGCTGTCATCGGATGACTTTCACTTAGACCGGCATTTATATTTTGCTTGCCGAGATGATCGGGAGCGTTTTT GTGAAAATACACAAGCTGGTGAGGGCAGAGTGTATAAGTGCCTCTTTAACCATAAATTTGAAGAATCCATGAGTGAAAAGTGTCGAGAAG CACTTACAACCCGCCAAAAGCTGATTGCCCAGGATTATAAAGTCAGTTATTCATTGGCCAAATCCTGTAAAAGTGACTTGAAGAAATACC GGTGCAATGTGGAAAACCTTCCGCGATCGCGTGAAGCCAGGCTCTCCTACTTGTTAATGTGCCTGGAGTCAGCTGTACACAGAGGGCGAC AAGTCAGCAGTGAGTGCCAGGGGGAGATGCTGGATTACCGACGCATGTTGATGGAAGACTTTTCTCTGAGCCCTGAGATCATCCTAAGCT GTCGGGGGGAGATTGAACACCATTGTTCCGGATTACATCGAAAAGGGCGGACCCTACACTGTCTGATGAAAGTAGTTCGAGGGGAGAAGG GGAACCTTGGAATGAACTGCCAGCAGGCGCTTCAAACACTGATTCAGGAGACTGACCCTGGTGCAGATTACCGCATTGATCGAGCTTTGA ATGAAGCTTGTGAATCTGTAATCCAGACAGCCTGCAAACATATAAGATCTGGAGACCCAATGATCTTGTCGTGCCTGATGGAACATTTAT ACACAGAGAAGATGGTAGAAGACTGTGAACACCGTCTCTTAGAGCTGCAGTATTTCATCTCCCGGGATTGGAAGCTGGACCCTGTCCTGT ACCGCAAGTGCCAGGGAGACGCTTCTCGTCTTTGCCACACCCACGGTTGGAATGAGACCAGTGAATTTATGCCTCAGGGAGCTGTGTTCT CTTGTTTATACAGACACGCCTACCGCACTGAGGAACAGGGAAGGAGGCTCTCACGGGAGTGCCGAGCTGAAGTCCAAAGGATCCTACACC AGCGTGCCATGGATGTCAAGCTGGATCCTGCCCTCCAGGATAAGTGCCTGATTGATCTGGGAAAATGGTGCAGTGAGAAAACAGAGACTG GACAGGAGCTGGAGTGCCTTCAGGACCATCTGGATGACTTGGTGGTGGAGTGTAGAGATATAGTTGGCAACCTCACTGAGTTAGAATCAG AGGATTTGGTATCAGCACTAAATTGTGGGATTACTGTTTCCACACCCTCACTCCAGAGAAACCCCACCTGAAGACGCAGTGACAACTCCC ACCCCCTCCGTCCTGCCCTCAGCCCGGCCCTGGCCCCTTCCCGACCCCCACCCGCCATTCAGACCCCATTAAGAAGGTTGGCTTGGCCAG GCAGGATGGGCTGTGTCCGGCCCTGCAGCCTAGTGGAAGGTGCTGAGGGGGCCCTGAGGCAGGACCGCCCTCCTGACCCCTGGTAGGAGG GTCACATCCACTTGGTGGCCAGGTGGCCCTTGGTGACCCACTTCTTCCTGGAGCGTCCCTGCCTAGAGCTCAGCCCACAGGACTGCTTCA GGCCGTGGCCACAGGTAGCAGCCGCAAGGGGAAATGAAGAAAACTGAGCCCTCGTGGCCACCTGTGTCACCCTTGTGCCTTAGCCTCATG GGCTGCCTAGGAGCTGCCTGCACGGCACAGCTCGCTTTCACAGTCAGAAGTGGGTCTGTGGGATCTGTGGTCCCTGTCCTCCCTGCTGTC CCTTCTGGGGAGGCTTTGGTGGCTCTGAGGTGGACAAAGAGCTCTCGCAAGAAGAGACAGCGTGATGCCTCCCACAGTCCACCCCAGACC CTGGGGCAGCCCCTCTGGCCCTGCCAGCTGCCTGCGTCGTTGGGCCCAGGGTGGCTGGCAGGAGTCCCAGCTGCTTGCTTTAGGACCTGG CAGCTTTTCTTGCCGTCCCTCCCCTGCCTCCAGAATCACAGCCCTTCTCCCCAAGGGAGGCTGAGGAGGCTTCTCCACCAGTGGCAGCCC CACCCCGTCCCTGGCCATTCTTGGCCTCCACCCCGCTCAGGCCCCTACTCGGGCGCTCCCAGAAGGAGCCACCTCTCAGTGCCTCACCTC CCCCTGCCTCCCAGCCTCCGCAGATGAGGTTCCTGCCCCTTCCTCCTCGTAACCAAAACCCTCACTGCTCCCAGGACGGTCTTATTTATA AACCAGATACATGTTCTTAGTCTGGTCCCAGACCAAGGAGCTGGTCAGACGGCCCTTTCTAATCCTACATGTTGAGCTTATGTAAAAAAT GTTGTTTCCTCCTGTTTTTGGTTCCTTTCTTACCCACAAACCATTACTACTTGAAACTTAAAAAACTCGCCAAGTGTAAAGGCTAAAGAG AAGCAGTTTGACGGACCTTGTGATTTGTACTGTTTGCTGCGGAGCTATTTAAAGATTTTGGAATAAATATACAAAACTACGGTTGTGAAA >33246_33246_1_GLG1-FA2H_GLG1_chr16_74508444_ENST00000205061_FA2H_chr16_74748167_ENST00000219368_length(amino acids)=706AA_BP=684 MAACGRVRRMFRLSAALHLLLLFAAGAEKLPGQGVHSQGQGPGANFVSFVGQAGGGGPAGQQLPQLPQSSQLQQQQQQQQQQQQPQPPQP PFPAGGPPARRGGAGAGGGWKLAEEESCREDVTRVCPKHTWSNNLAVLECLQDVREPENEISSDCNHLLWNYKLNLTTDPKFESVAREVC KSTITEIKECADEPVGKGYMVSCLVDHRGNITEYQCHQYITKMTAIIFSDYRLICGFMDDCKNDINILKCGSIRLGEKDAHSQGEVVSCL EKGLVKEAEEREPKIQVSELCKKAILRVAELSSDDFHLDRHLYFACRDDRERFCENTQAGEGRVYKCLFNHKFEESMSEKCREALTTRQK LIAQDYKVSYSLAKSCKSDLKKYRCNVENLPRSREARLSYLLMCLESAVHRGRQVSSECQGEMLDYRRMLMEDFSLSPEIILSCRGEIEH HCSGLHRKGRTLHCLMKVVRGEKGNLGMNCQQALQTLIQETDPGADYRIDRALNEACESVIQTACKHIRSGDPMILSCLMEHLYTEKMVE DCEHRLLELQYFISRDWKLDPVLYRKCQGDASRLCHTHGWNETSEFMPQGAVFSCLYRHAYRTEEQGRRLSRECRAEVQRILHQRAMDVK -------------------------------------------------------------- >33246_33246_2_GLG1-FA2H_GLG1_chr16_74508444_ENST00000205061_FA2H_chr16_74748167_ENST00000544337_length(transcript)=2744nt_BP=2072nt GAGCTCGCCGCGGACTCAAGATGGCGGCGTGTGGACGTGTACGGAGGATGTTCCGCTTGTCGGCGGCGCTGCATCTGCTGCTGCTATTCG CGGCCGGGGCCGAGAAACTCCCCGGCCAGGGCGTCCACAGCCAGGGCCAGGGTCCCGGGGCCAACTTTGTGTCCTTCGTAGGGCAGGCCG GAGGCGGCGGCCCGGCGGGTCAGCAGCTGCCCCAGCTGCCTCAGTCATCGCAGCTTCAGCAGCAACAGCAGCAGCAGCAACAGCAACAGC AGCCTCAGCCGCCGCAGCCGCCTTTCCCGGCGGGTGGGCCTCCGGCCCGGCGGGGAGGAGCGGGGGCTGGTGGGGGCTGGAAGCTGGCGG AGGAAGAGTCCTGCAGGGAGGACGTGACCCGCGTGTGCCCTAAGCACACCTGGAGCAACAACCTGGCGGTGCTCGAGTGCCTGCAGGATG TGAGGGAGCCTGAAAATGAAATTTCTTCAGACTGCAATCATTTGTTGTGGAATTATAAGCTGAACCTAACTACAGATCCCAAATTTGAAT CTGTGGCCAGAGAGGTTTGCAAATCTACTATAACAGAGATTAAAGAATGTGCTGATGAACCGGTTGGAAAAGGTTACATGGTTTCCTGCT TGGTGGATCACCGAGGCAACATCACTGAGTATCAGTGTCACCAGTACATTACCAAGATGACGGCCATCATTTTTAGTGATTACCGTTTAA TCTGTGGCTTCATGGATGACTGCAAAAATGACATCAACATTCTGAAATGTGGCAGTATTCGGCTTGGAGAAAAGGATGCACATTCACAAG GTGAGGTGGTATCATGCTTGGAGAAAGGCCTGGTGAAAGAAGCAGAAGAAAGAGAACCCAAGATTCAAGTTTCTGAACTCTGCAAGAAAG CCATTCTCCGGGTGGCTGAGCTGTCATCGGATGACTTTCACTTAGACCGGCATTTATATTTTGCTTGCCGAGATGATCGGGAGCGTTTTT GTGAAAATACACAAGCTGGTGAGGGCAGAGTGTATAAGTGCCTCTTTAACCATAAATTTGAAGAATCCATGAGTGAAAAGTGTCGAGAAG CACTTACAACCCGCCAAAAGCTGATTGCCCAGGATTATAAAGTCAGTTATTCATTGGCCAAATCCTGTAAAAGTGACTTGAAGAAATACC GGTGCAATGTGGAAAACCTTCCGCGATCGCGTGAAGCCAGGCTCTCCTACTTGTTAATGTGCCTGGAGTCAGCTGTACACAGAGGGCGAC AAGTCAGCAGTGAGTGCCAGGGGGAGATGCTGGATTACCGACGCATGTTGATGGAAGACTTTTCTCTGAGCCCTGAGATCATCCTAAGCT GTCGGGGGGAGATTGAACACCATTGTTCCGGATTACATCGAAAAGGGCGGACCCTACACTGTCTGATGAAAGTAGTTCGAGGGGAGAAGG GGAACCTTGGAATGAACTGCCAGCAGGCGCTTCAAACACTGATTCAGGAGACTGACCCTGGTGCAGATTACCGCATTGATCGAGCTTTGA ATGAAGCTTGTGAATCTGTAATCCAGACAGCCTGCAAACATATAAGATCTGGAGACCCAATGATCTTGTCGTGCCTGATGGAACATTTAT ACACAGAGAAGATGGTAGAAGACTGTGAACACCGTCTCTTAGAGCTGCAGTATTTCATCTCCCGGGATTGGAAGCTGGACCCTGTCCTGT ACCGCAAGTGCCAGGGAGACGCTTCTCGTCTTTGCCACACCCACGGTTGGAATGAGACCAGTGAATTTATGCCTCAGGGAGCTGTGTTCT CTTGTTTATACAGACACGCCTACCGCACTGAGGAACAGGGAAGGAGGCTCTCACGGGAGTGCCGAGCTGAAGTCCAAAGGATCCTACACC AGCGTGCCATGGATGTCAAGCTGGATCCTGCCCTCCAGGATAAGTGCCTGATTGATCTGGGAAAATGGTGCAGTGAGAAAACAGAGACTG GACAGGAGCTGGAGTGCCTTCAGGACCATCTGGATGACTTGGTGGTGGAGTGTAGAGATATAGTTGGCAACCTCACTGAGTTAGAATCAG AGGATTTGGTATCAGCACTAAATTGTGGGATTACTGTTTCCACACCCTCACTCCAGAGAAACCCCACCTGAAGACGCAGTGACAACTCCC ACCCCCTCCGTCCTGCCCTCAGCCCGGCCCTGGCCCCTTCCCGACCCCCACCCGCCATTCAGACCCCATTAAGAAGGTTGGCTTGGCCAG GCAGGATGGGCTGTGTCCGGCCCTGCAGCCTAGTGGAAGGTGCTGAGGGGGCCCTGAGGCAGGACCGCCCTCCTGACCCCTGGTAGGAGG GTCACATCCACTTGGTGGCCAGGTGGCCCTTGGTGACCCACTTCTTCCTGGAGCGTCCCTGCCTAGAGCTCAGCCCACAGGACTGCTTCA GGCCGTGGCCACAGGTAGCAGCCGCAAGGGGAAATGAAGAAAACTGAGCCCTCGTGGCCACCTGTGTCACCCTTGTGCCTTAGCCTCATG GGCTGCCTAGGAGCTGCCTGCACGGCACAGCTCGCTTTCACAGTCAGAAGTGGGTCTGTGGGATCTGTGGTCCCTGTCCTCCCTGCTGTC CCTTCTGGGGAGGCTTTGGTGGCTCTGAGGTGGACAAAGAGCTCTCGCAAGAAGAGACAGCGTGATGCCTCCCACAGTCCACCCCAGACC >33246_33246_2_GLG1-FA2H_GLG1_chr16_74508444_ENST00000205061_FA2H_chr16_74748167_ENST00000544337_length(amino acids)=706AA_BP=684 MAACGRVRRMFRLSAALHLLLLFAAGAEKLPGQGVHSQGQGPGANFVSFVGQAGGGGPAGQQLPQLPQSSQLQQQQQQQQQQQQPQPPQP PFPAGGPPARRGGAGAGGGWKLAEEESCREDVTRVCPKHTWSNNLAVLECLQDVREPENEISSDCNHLLWNYKLNLTTDPKFESVAREVC KSTITEIKECADEPVGKGYMVSCLVDHRGNITEYQCHQYITKMTAIIFSDYRLICGFMDDCKNDINILKCGSIRLGEKDAHSQGEVVSCL EKGLVKEAEEREPKIQVSELCKKAILRVAELSSDDFHLDRHLYFACRDDRERFCENTQAGEGRVYKCLFNHKFEESMSEKCREALTTRQK LIAQDYKVSYSLAKSCKSDLKKYRCNVENLPRSREARLSYLLMCLESAVHRGRQVSSECQGEMLDYRRMLMEDFSLSPEIILSCRGEIEH HCSGLHRKGRTLHCLMKVVRGEKGNLGMNCQQALQTLIQETDPGADYRIDRALNEACESVIQTACKHIRSGDPMILSCLMEHLYTEKMVE DCEHRLLELQYFISRDWKLDPVLYRKCQGDASRLCHTHGWNETSEFMPQGAVFSCLYRHAYRTEEQGRRLSRECRAEVQRILHQRAMDVK -------------------------------------------------------------- >33246_33246_3_GLG1-FA2H_GLG1_chr16_74508444_ENST00000422840_FA2H_chr16_74748167_ENST00000219368_length(transcript)=3367nt_BP=2052nt ATGGCGGCGTGTGGACGTGTACGGAGGATGTTCCGCTTGTCGGCGGCGCTGCATCTGCTGCTGCTATTCGCGGCCGGGGCCGAGAAACTC CCCGGCCAGGGCGTCCACAGCCAGGGCCAGGGTCCCGGGGCCAACTTTGTGTCCTTCGTAGGGCAGGCCGGAGGCGGCGGCCCGGCGGGT CAGCAGCTGCCCCAGCTGCCTCAGTCATCGCAGCTTCAGCAGCAACAGCAGCAGCAGCAACAGCAACAGCAGCCTCAGCCGCCGCAGCCG CCTTTCCCGGCGGGTGGGCCTCCGGCCCGGCGGGGAGGAGCGGGGGCTGGTGGGGGCTGGAAGCTGGCGGAGGAAGAGTCCTGCAGGGAG GACGTGACCCGCGTGTGCCCTAAGCACACCTGGAGCAACAACCTGGCGGTGCTCGAGTGCCTGCAGGATGTGAGGGAGCCTGAAAATGAA ATTTCTTCAGACTGCAATCATTTGTTGTGGAATTATAAGCTGAACCTAACTACAGATCCCAAATTTGAATCTGTGGCCAGAGAGGTTTGC AAATCTACTATAACAGAGATTAAAGAATGTGCTGATGAACCGGTTGGAAAAGGTTACATGGTTTCCTGCTTGGTGGATCACCGAGGCAAC ATCACTGAGTATCAGTGTCACCAGTACATTACCAAGATGACGGCCATCATTTTTAGTGATTACCGTTTAATCTGTGGCTTCATGGATGAC TGCAAAAATGACATCAACATTCTGAAATGTGGCAGTATTCGGCTTGGAGAAAAGGATGCACATTCACAAGGTGAGGTGGTATCATGCTTG GAGAAAGGCCTGGTGAAAGAAGCAGAAGAAAGAGAACCCAAGATTCAAGTTTCTGAACTCTGCAAGAAAGCCATTCTCCGGGTGGCTGAG CTGTCATCGGATGACTTTCACTTAGACCGGCATTTATATTTTGCTTGCCGAGATGATCGGGAGCGTTTTTGTGAAAATACACAAGCTGGT GAGGGCAGAGTGTATAAGTGCCTCTTTAACCATAAATTTGAAGAATCCATGAGTGAAAAGTGTCGAGAAGCACTTACAACCCGCCAAAAG CTGATTGCCCAGGATTATAAAGTCAGTTATTCATTGGCCAAATCCTGTAAAAGTGACTTGAAGAAATACCGGTGCAATGTGGAAAACCTT CCGCGATCGCGTGAAGCCAGGCTCTCCTACTTGTTAATGTGCCTGGAGTCAGCTGTACACAGAGGGCGACAAGTCAGCAGTGAGTGCCAG GGGGAGATGCTGGATTACCGACGCATGTTGATGGAAGACTTTTCTCTGAGCCCTGAGATCATCCTAAGCTGTCGGGGGGAGATTGAACAC CATTGTTCCGGATTACATCGAAAAGGGCGGACCCTACACTGTCTGATGAAAGTAGTTCGAGGGGAGAAGGGGAACCTTGGAATGAACTGC CAGCAGGCGCTTCAAACACTGATTCAGGAGACTGACCCTGGTGCAGATTACCGCATTGATCGAGCTTTGAATGAAGCTTGTGAATCTGTA ATCCAGACAGCCTGCAAACATATAAGATCTGGAGACCCAATGATCTTGTCGTGCCTGATGGAACATTTATACACAGAGAAGATGGTAGAA GACTGTGAACACCGTCTCTTAGAGCTGCAGTATTTCATCTCCCGGGATTGGAAGCTGGACCCTGTCCTGTACCGCAAGTGCCAGGGAGAC GCTTCTCGTCTTTGCCACACCCACGGTTGGAATGAGACCAGTGAATTTATGCCTCAGGGAGCTGTGTTCTCTTGTTTATACAGACACGCC TACCGCACTGAGGAACAGGGAAGGAGGCTCTCACGGGAGTGCCGAGCTGAAGTCCAAAGGATCCTACACCAGCGTGCCATGGATGTCAAG CTGGATCCTGCCCTCCAGGATAAGTGCCTGATTGATCTGGGAAAATGGTGCAGTGAGAAAACAGAGACTGGACAGGAGCTGGAGTGCCTT CAGGACCATCTGGATGACTTGGTGGTGGAGTGTAGAGATATAGTTGGCAACCTCACTGAGTTAGAATCAGAGGATTTGGTATCAGCACTA AATTGTGGGATTACTGTTTCCACACCCTCACTCCAGAGAAACCCCACCTGAAGACGCAGTGACAACTCCCACCCCCTCCGTCCTGCCCTC AGCCCGGCCCTGGCCCCTTCCCGACCCCCACCCGCCATTCAGACCCCATTAAGAAGGTTGGCTTGGCCAGGCAGGATGGGCTGTGTCCGG CCCTGCAGCCTAGTGGAAGGTGCTGAGGGGGCCCTGAGGCAGGACCGCCCTCCTGACCCCTGGTAGGAGGGTCACATCCACTTGGTGGCC AGGTGGCCCTTGGTGACCCACTTCTTCCTGGAGCGTCCCTGCCTAGAGCTCAGCCCACAGGACTGCTTCAGGCCGTGGCCACAGGTAGCA GCCGCAAGGGGAAATGAAGAAAACTGAGCCCTCGTGGCCACCTGTGTCACCCTTGTGCCTTAGCCTCATGGGCTGCCTAGGAGCTGCCTG CACGGCACAGCTCGCTTTCACAGTCAGAAGTGGGTCTGTGGGATCTGTGGTCCCTGTCCTCCCTGCTGTCCCTTCTGGGGAGGCTTTGGT GGCTCTGAGGTGGACAAAGAGCTCTCGCAAGAAGAGACAGCGTGATGCCTCCCACAGTCCACCCCAGACCCTGGGGCAGCCCCTCTGGCC CTGCCAGCTGCCTGCGTCGTTGGGCCCAGGGTGGCTGGCAGGAGTCCCAGCTGCTTGCTTTAGGACCTGGCAGCTTTTCTTGCCGTCCCT CCCCTGCCTCCAGAATCACAGCCCTTCTCCCCAAGGGAGGCTGAGGAGGCTTCTCCACCAGTGGCAGCCCCACCCCGTCCCTGGCCATTC TTGGCCTCCACCCCGCTCAGGCCCCTACTCGGGCGCTCCCAGAAGGAGCCACCTCTCAGTGCCTCACCTCCCCCTGCCTCCCAGCCTCCG CAGATGAGGTTCCTGCCCCTTCCTCCTCGTAACCAAAACCCTCACTGCTCCCAGGACGGTCTTATTTATAAACCAGATACATGTTCTTAG TCTGGTCCCAGACCAAGGAGCTGGTCAGACGGCCCTTTCTAATCCTACATGTTGAGCTTATGTAAAAAATGTTGTTTCCTCCTGTTTTTG GTTCCTTTCTTACCCACAAACCATTACTACTTGAAACTTAAAAAACTCGCCAAGTGTAAAGGCTAAAGAGAAGCAGTTTGACGGACCTTG TGATTTGTACTGTTTGCTGCGGAGCTATTTAAAGATTTTGGAATAAATATACAAAACTACGGTTGTGAAATAAAAACTTAAATTGTATAT >33246_33246_3_GLG1-FA2H_GLG1_chr16_74508444_ENST00000422840_FA2H_chr16_74748167_ENST00000219368_length(amino acids)=706AA_BP=684 MAACGRVRRMFRLSAALHLLLLFAAGAEKLPGQGVHSQGQGPGANFVSFVGQAGGGGPAGQQLPQLPQSSQLQQQQQQQQQQQQPQPPQP PFPAGGPPARRGGAGAGGGWKLAEEESCREDVTRVCPKHTWSNNLAVLECLQDVREPENEISSDCNHLLWNYKLNLTTDPKFESVAREVC KSTITEIKECADEPVGKGYMVSCLVDHRGNITEYQCHQYITKMTAIIFSDYRLICGFMDDCKNDINILKCGSIRLGEKDAHSQGEVVSCL EKGLVKEAEEREPKIQVSELCKKAILRVAELSSDDFHLDRHLYFACRDDRERFCENTQAGEGRVYKCLFNHKFEESMSEKCREALTTRQK LIAQDYKVSYSLAKSCKSDLKKYRCNVENLPRSREARLSYLLMCLESAVHRGRQVSSECQGEMLDYRRMLMEDFSLSPEIILSCRGEIEH HCSGLHRKGRTLHCLMKVVRGEKGNLGMNCQQALQTLIQETDPGADYRIDRALNEACESVIQTACKHIRSGDPMILSCLMEHLYTEKMVE DCEHRLLELQYFISRDWKLDPVLYRKCQGDASRLCHTHGWNETSEFMPQGAVFSCLYRHAYRTEEQGRRLSRECRAEVQRILHQRAMDVK -------------------------------------------------------------- >33246_33246_4_GLG1-FA2H_GLG1_chr16_74508444_ENST00000422840_FA2H_chr16_74748167_ENST00000544337_length(transcript)=2724nt_BP=2052nt ATGGCGGCGTGTGGACGTGTACGGAGGATGTTCCGCTTGTCGGCGGCGCTGCATCTGCTGCTGCTATTCGCGGCCGGGGCCGAGAAACTC CCCGGCCAGGGCGTCCACAGCCAGGGCCAGGGTCCCGGGGCCAACTTTGTGTCCTTCGTAGGGCAGGCCGGAGGCGGCGGCCCGGCGGGT CAGCAGCTGCCCCAGCTGCCTCAGTCATCGCAGCTTCAGCAGCAACAGCAGCAGCAGCAACAGCAACAGCAGCCTCAGCCGCCGCAGCCG CCTTTCCCGGCGGGTGGGCCTCCGGCCCGGCGGGGAGGAGCGGGGGCTGGTGGGGGCTGGAAGCTGGCGGAGGAAGAGTCCTGCAGGGAG GACGTGACCCGCGTGTGCCCTAAGCACACCTGGAGCAACAACCTGGCGGTGCTCGAGTGCCTGCAGGATGTGAGGGAGCCTGAAAATGAA ATTTCTTCAGACTGCAATCATTTGTTGTGGAATTATAAGCTGAACCTAACTACAGATCCCAAATTTGAATCTGTGGCCAGAGAGGTTTGC AAATCTACTATAACAGAGATTAAAGAATGTGCTGATGAACCGGTTGGAAAAGGTTACATGGTTTCCTGCTTGGTGGATCACCGAGGCAAC ATCACTGAGTATCAGTGTCACCAGTACATTACCAAGATGACGGCCATCATTTTTAGTGATTACCGTTTAATCTGTGGCTTCATGGATGAC TGCAAAAATGACATCAACATTCTGAAATGTGGCAGTATTCGGCTTGGAGAAAAGGATGCACATTCACAAGGTGAGGTGGTATCATGCTTG GAGAAAGGCCTGGTGAAAGAAGCAGAAGAAAGAGAACCCAAGATTCAAGTTTCTGAACTCTGCAAGAAAGCCATTCTCCGGGTGGCTGAG CTGTCATCGGATGACTTTCACTTAGACCGGCATTTATATTTTGCTTGCCGAGATGATCGGGAGCGTTTTTGTGAAAATACACAAGCTGGT GAGGGCAGAGTGTATAAGTGCCTCTTTAACCATAAATTTGAAGAATCCATGAGTGAAAAGTGTCGAGAAGCACTTACAACCCGCCAAAAG CTGATTGCCCAGGATTATAAAGTCAGTTATTCATTGGCCAAATCCTGTAAAAGTGACTTGAAGAAATACCGGTGCAATGTGGAAAACCTT CCGCGATCGCGTGAAGCCAGGCTCTCCTACTTGTTAATGTGCCTGGAGTCAGCTGTACACAGAGGGCGACAAGTCAGCAGTGAGTGCCAG GGGGAGATGCTGGATTACCGACGCATGTTGATGGAAGACTTTTCTCTGAGCCCTGAGATCATCCTAAGCTGTCGGGGGGAGATTGAACAC CATTGTTCCGGATTACATCGAAAAGGGCGGACCCTACACTGTCTGATGAAAGTAGTTCGAGGGGAGAAGGGGAACCTTGGAATGAACTGC CAGCAGGCGCTTCAAACACTGATTCAGGAGACTGACCCTGGTGCAGATTACCGCATTGATCGAGCTTTGAATGAAGCTTGTGAATCTGTA ATCCAGACAGCCTGCAAACATATAAGATCTGGAGACCCAATGATCTTGTCGTGCCTGATGGAACATTTATACACAGAGAAGATGGTAGAA GACTGTGAACACCGTCTCTTAGAGCTGCAGTATTTCATCTCCCGGGATTGGAAGCTGGACCCTGTCCTGTACCGCAAGTGCCAGGGAGAC GCTTCTCGTCTTTGCCACACCCACGGTTGGAATGAGACCAGTGAATTTATGCCTCAGGGAGCTGTGTTCTCTTGTTTATACAGACACGCC TACCGCACTGAGGAACAGGGAAGGAGGCTCTCACGGGAGTGCCGAGCTGAAGTCCAAAGGATCCTACACCAGCGTGCCATGGATGTCAAG CTGGATCCTGCCCTCCAGGATAAGTGCCTGATTGATCTGGGAAAATGGTGCAGTGAGAAAACAGAGACTGGACAGGAGCTGGAGTGCCTT CAGGACCATCTGGATGACTTGGTGGTGGAGTGTAGAGATATAGTTGGCAACCTCACTGAGTTAGAATCAGAGGATTTGGTATCAGCACTA AATTGTGGGATTACTGTTTCCACACCCTCACTCCAGAGAAACCCCACCTGAAGACGCAGTGACAACTCCCACCCCCTCCGTCCTGCCCTC AGCCCGGCCCTGGCCCCTTCCCGACCCCCACCCGCCATTCAGACCCCATTAAGAAGGTTGGCTTGGCCAGGCAGGATGGGCTGTGTCCGG CCCTGCAGCCTAGTGGAAGGTGCTGAGGGGGCCCTGAGGCAGGACCGCCCTCCTGACCCCTGGTAGGAGGGTCACATCCACTTGGTGGCC AGGTGGCCCTTGGTGACCCACTTCTTCCTGGAGCGTCCCTGCCTAGAGCTCAGCCCACAGGACTGCTTCAGGCCGTGGCCACAGGTAGCA GCCGCAAGGGGAAATGAAGAAAACTGAGCCCTCGTGGCCACCTGTGTCACCCTTGTGCCTTAGCCTCATGGGCTGCCTAGGAGCTGCCTG CACGGCACAGCTCGCTTTCACAGTCAGAAGTGGGTCTGTGGGATCTGTGGTCCCTGTCCTCCCTGCTGTCCCTTCTGGGGAGGCTTTGGT GGCTCTGAGGTGGACAAAGAGCTCTCGCAAGAAGAGACAGCGTGATGCCTCCCACAGTCCACCCCAGACCCTGGGGCAGCCCCTCTGGCC >33246_33246_4_GLG1-FA2H_GLG1_chr16_74508444_ENST00000422840_FA2H_chr16_74748167_ENST00000544337_length(amino acids)=706AA_BP=684 MAACGRVRRMFRLSAALHLLLLFAAGAEKLPGQGVHSQGQGPGANFVSFVGQAGGGGPAGQQLPQLPQSSQLQQQQQQQQQQQQPQPPQP PFPAGGPPARRGGAGAGGGWKLAEEESCREDVTRVCPKHTWSNNLAVLECLQDVREPENEISSDCNHLLWNYKLNLTTDPKFESVAREVC KSTITEIKECADEPVGKGYMVSCLVDHRGNITEYQCHQYITKMTAIIFSDYRLICGFMDDCKNDINILKCGSIRLGEKDAHSQGEVVSCL EKGLVKEAEEREPKIQVSELCKKAILRVAELSSDDFHLDRHLYFACRDDRERFCENTQAGEGRVYKCLFNHKFEESMSEKCREALTTRQK LIAQDYKVSYSLAKSCKSDLKKYRCNVENLPRSREARLSYLLMCLESAVHRGRQVSSECQGEMLDYRRMLMEDFSLSPEIILSCRGEIEH HCSGLHRKGRTLHCLMKVVRGEKGNLGMNCQQALQTLIQETDPGADYRIDRALNEACESVIQTACKHIRSGDPMILSCLMEHLYTEKMVE DCEHRLLELQYFISRDWKLDPVLYRKCQGDASRLCHTHGWNETSEFMPQGAVFSCLYRHAYRTEEQGRRLSRECRAEVQRILHQRAMDVK -------------------------------------------------------------- >33246_33246_5_GLG1-FA2H_GLG1_chr16_74508444_ENST00000447066_FA2H_chr16_74748167_ENST00000219368_length(transcript)=3352nt_BP=2037nt GCTCGCCGCGGACTCAAGATGGCGGCGTGTGGACGTGTACGGAGGATGTTCCGCTTGTCGGCGGCGCTGCATCTGCTGCTGCTATTCGCG GCCGGGGCCGAGAAACTCCCCGGCCAGGGCGTCCACAGCCAGGGCCAGGGTCCCGGGGCCAACTTTGTGTCCTTCGTAGGGCAGGCCGGA GGCGGCGGCCCGGCGGGTCAGCAGCTGCCCCAGCTGCCTCAGTCATCGCAGCTTCAGCAGCAACAGCAGCAGCAGCAACAGCAACAGCAG CCTCAGCCGCCGCAGCCGCCTTTCCCGGCGGGTGGGCCTCCGGCCCGGCGGGGAGGAGCGGGGGCTGGTGGGGGCTGGAAGCTGGCGGAG GAAGAGTCCTGCAGGGAGGACGTGACCCGCGTGTGCCCTAAGCACACCTGGAGCAACAACCTGGCGGTGCTCGAGTGCCTGCAGGATGTG AGGGAGTTGTTGTGGAATTATAAGCTGAACCTAACTACAGATCCCAAATTTGAATCTGTGGCCAGAGAGGTTTGCAAATCTACTATAACA GAGATTAAAGAATGTGCTGATGAACCGGTTGGAAAAGGTTACATGGTTTCCTGCTTGGTGGATCACCGAGGCAACATCACTGAGTATCAG TGTCACCAGTACATTACCAAGATGACGGCCATCATTTTTAGTGATTACCGTTTAATCTGTGGCTTCATGGATGACTGCAAAAATGACATC AACATTCTGAAATGTGGCAGTATTCGGCTTGGAGAAAAGGATGCACATTCACAAGGTGAGGTGGTATCATGCTTGGAGAAAGGCCTGGTG AAAGAAGCAGAAGAAAGAGAACCCAAGATTCAAGTTTCTGAACTCTGCAAGAAAGCCATTCTCCGGGTGGCTGAGCTGTCATCGGATGAC TTTCACTTAGACCGGCATTTATATTTTGCTTGCCGAGATGATCGGGAGCGTTTTTGTGAAAATACACAAGCTGGTGAGGGCAGAGTGTAT AAGTGCCTCTTTAACCATAAATTTGAAGAATCCATGAGTGAAAAGTGTCGAGAAGCACTTACAACCCGCCAAAAGCTGATTGCCCAGGAT TATAAAGTCAGTTATTCATTGGCCAAATCCTGTAAAAGTGACTTGAAGAAATACCGGTGCAATGTGGAAAACCTTCCGCGATCGCGTGAA GCCAGGCTCTCCTACTTGTTAATGTGCCTGGAGTCAGCTGTACACAGAGGGCGACAAGTCAGCAGTGAGTGCCAGGGGGAGATGCTGGAT TACCGACGCATGTTGATGGAAGACTTTTCTCTGAGCCCTGAGATCATCCTAAGCTGTCGGGGGGAGATTGAACACCATTGTTCCGGATTA CATCGAAAAGGGCGGACCCTACACTGTCTGATGAAAGTAGTTCGAGGGGAGAAGGGGAACCTTGGAATGAACTGCCAGCAGGCGCTTCAA ACACTGATTCAGGAGACTGACCCTGGTGCAGATTACCGCATTGATCGAGCTTTGAATGAAGCTTGTGAATCTGTAATCCAGACAGCCTGC AAACATATAAGATCTGGAGACCCAATGATCTTGTCGTGCCTGATGGAACATTTATACACAGAGAAGATGGTAGAAGACTGTGAACACCGT CTCTTAGAGCTGCAGTATTTCATCTCCCGGGATTGGAAGCTGGACCCTGTCCTGTACCGCAAGTGCCAGGGAGACGCTTCTCGTCTTTGC CACACCCACGGTTGGAATGAGACCAGTGAATTTATGCCTCAGGGAGCTGTGTTCTCTTGTTTATACAGACACGCCTACCGCACTGAGGAA CAGGGAAGGAGGCTCTCACGGGAGTGCCGAGCTGAAGTCCAAAGGATCCTACACCAGCGTGCCATGGATGTCAAGCTGGATCCTGCCCTC CAGGATAAGTGCCTGATTGATCTGGGAAAATGGTGCAGTGAGAAAACAGAGACTGGACAGGAGCTGGAGTGCCTTCAGGACCATCTGGAT GACTTGGTGGTGGAGTGTAGAGATATAGTTGGCAACCTCACTGAGTTAGAATCAGAGGATTTGGTATCAGCACTAAATTGTGGGATTACT GTTTCCACACCCTCACTCCAGAGAAACCCCACCTGAAGACGCAGTGACAACTCCCACCCCCTCCGTCCTGCCCTCAGCCCGGCCCTGGCC CCTTCCCGACCCCCACCCGCCATTCAGACCCCATTAAGAAGGTTGGCTTGGCCAGGCAGGATGGGCTGTGTCCGGCCCTGCAGCCTAGTG GAAGGTGCTGAGGGGGCCCTGAGGCAGGACCGCCCTCCTGACCCCTGGTAGGAGGGTCACATCCACTTGGTGGCCAGGTGGCCCTTGGTG ACCCACTTCTTCCTGGAGCGTCCCTGCCTAGAGCTCAGCCCACAGGACTGCTTCAGGCCGTGGCCACAGGTAGCAGCCGCAAGGGGAAAT GAAGAAAACTGAGCCCTCGTGGCCACCTGTGTCACCCTTGTGCCTTAGCCTCATGGGCTGCCTAGGAGCTGCCTGCACGGCACAGCTCGC TTTCACAGTCAGAAGTGGGTCTGTGGGATCTGTGGTCCCTGTCCTCCCTGCTGTCCCTTCTGGGGAGGCTTTGGTGGCTCTGAGGTGGAC AAAGAGCTCTCGCAAGAAGAGACAGCGTGATGCCTCCCACAGTCCACCCCAGACCCTGGGGCAGCCCCTCTGGCCCTGCCAGCTGCCTGC GTCGTTGGGCCCAGGGTGGCTGGCAGGAGTCCCAGCTGCTTGCTTTAGGACCTGGCAGCTTTTCTTGCCGTCCCTCCCCTGCCTCCAGAA TCACAGCCCTTCTCCCCAAGGGAGGCTGAGGAGGCTTCTCCACCAGTGGCAGCCCCACCCCGTCCCTGGCCATTCTTGGCCTCCACCCCG CTCAGGCCCCTACTCGGGCGCTCCCAGAAGGAGCCACCTCTCAGTGCCTCACCTCCCCCTGCCTCCCAGCCTCCGCAGATGAGGTTCCTG CCCCTTCCTCCTCGTAACCAAAACCCTCACTGCTCCCAGGACGGTCTTATTTATAAACCAGATACATGTTCTTAGTCTGGTCCCAGACCA AGGAGCTGGTCAGACGGCCCTTTCTAATCCTACATGTTGAGCTTATGTAAAAAATGTTGTTTCCTCCTGTTTTTGGTTCCTTTCTTACCC ACAAACCATTACTACTTGAAACTTAAAAAACTCGCCAAGTGTAAAGGCTAAAGAGAAGCAGTTTGACGGACCTTGTGATTTGTACTGTTT GCTGCGGAGCTATTTAAAGATTTTGGAATAAATATACAAAACTACGGTTGTGAAATAAAAACTTAAATTGTATATTTTGAAAAATAAAAC >33246_33246_5_GLG1-FA2H_GLG1_chr16_74508444_ENST00000447066_FA2H_chr16_74748167_ENST00000219368_length(amino acids)=695AA_BP=673 MAACGRVRRMFRLSAALHLLLLFAAGAEKLPGQGVHSQGQGPGANFVSFVGQAGGGGPAGQQLPQLPQSSQLQQQQQQQQQQQQPQPPQP PFPAGGPPARRGGAGAGGGWKLAEEESCREDVTRVCPKHTWSNNLAVLECLQDVRELLWNYKLNLTTDPKFESVAREVCKSTITEIKECA DEPVGKGYMVSCLVDHRGNITEYQCHQYITKMTAIIFSDYRLICGFMDDCKNDINILKCGSIRLGEKDAHSQGEVVSCLEKGLVKEAEER EPKIQVSELCKKAILRVAELSSDDFHLDRHLYFACRDDRERFCENTQAGEGRVYKCLFNHKFEESMSEKCREALTTRQKLIAQDYKVSYS LAKSCKSDLKKYRCNVENLPRSREARLSYLLMCLESAVHRGRQVSSECQGEMLDYRRMLMEDFSLSPEIILSCRGEIEHHCSGLHRKGRT LHCLMKVVRGEKGNLGMNCQQALQTLIQETDPGADYRIDRALNEACESVIQTACKHIRSGDPMILSCLMEHLYTEKMVEDCEHRLLELQY FISRDWKLDPVLYRKCQGDASRLCHTHGWNETSEFMPQGAVFSCLYRHAYRTEEQGRRLSRECRAEVQRILHQRAMDVKLDPALQDKCLI -------------------------------------------------------------- >33246_33246_6_GLG1-FA2H_GLG1_chr16_74508444_ENST00000447066_FA2H_chr16_74748167_ENST00000544337_length(transcript)=2709nt_BP=2037nt GCTCGCCGCGGACTCAAGATGGCGGCGTGTGGACGTGTACGGAGGATGTTCCGCTTGTCGGCGGCGCTGCATCTGCTGCTGCTATTCGCG GCCGGGGCCGAGAAACTCCCCGGCCAGGGCGTCCACAGCCAGGGCCAGGGTCCCGGGGCCAACTTTGTGTCCTTCGTAGGGCAGGCCGGA GGCGGCGGCCCGGCGGGTCAGCAGCTGCCCCAGCTGCCTCAGTCATCGCAGCTTCAGCAGCAACAGCAGCAGCAGCAACAGCAACAGCAG CCTCAGCCGCCGCAGCCGCCTTTCCCGGCGGGTGGGCCTCCGGCCCGGCGGGGAGGAGCGGGGGCTGGTGGGGGCTGGAAGCTGGCGGAG GAAGAGTCCTGCAGGGAGGACGTGACCCGCGTGTGCCCTAAGCACACCTGGAGCAACAACCTGGCGGTGCTCGAGTGCCTGCAGGATGTG AGGGAGTTGTTGTGGAATTATAAGCTGAACCTAACTACAGATCCCAAATTTGAATCTGTGGCCAGAGAGGTTTGCAAATCTACTATAACA GAGATTAAAGAATGTGCTGATGAACCGGTTGGAAAAGGTTACATGGTTTCCTGCTTGGTGGATCACCGAGGCAACATCACTGAGTATCAG TGTCACCAGTACATTACCAAGATGACGGCCATCATTTTTAGTGATTACCGTTTAATCTGTGGCTTCATGGATGACTGCAAAAATGACATC AACATTCTGAAATGTGGCAGTATTCGGCTTGGAGAAAAGGATGCACATTCACAAGGTGAGGTGGTATCATGCTTGGAGAAAGGCCTGGTG AAAGAAGCAGAAGAAAGAGAACCCAAGATTCAAGTTTCTGAACTCTGCAAGAAAGCCATTCTCCGGGTGGCTGAGCTGTCATCGGATGAC TTTCACTTAGACCGGCATTTATATTTTGCTTGCCGAGATGATCGGGAGCGTTTTTGTGAAAATACACAAGCTGGTGAGGGCAGAGTGTAT AAGTGCCTCTTTAACCATAAATTTGAAGAATCCATGAGTGAAAAGTGTCGAGAAGCACTTACAACCCGCCAAAAGCTGATTGCCCAGGAT TATAAAGTCAGTTATTCATTGGCCAAATCCTGTAAAAGTGACTTGAAGAAATACCGGTGCAATGTGGAAAACCTTCCGCGATCGCGTGAA GCCAGGCTCTCCTACTTGTTAATGTGCCTGGAGTCAGCTGTACACAGAGGGCGACAAGTCAGCAGTGAGTGCCAGGGGGAGATGCTGGAT TACCGACGCATGTTGATGGAAGACTTTTCTCTGAGCCCTGAGATCATCCTAAGCTGTCGGGGGGAGATTGAACACCATTGTTCCGGATTA CATCGAAAAGGGCGGACCCTACACTGTCTGATGAAAGTAGTTCGAGGGGAGAAGGGGAACCTTGGAATGAACTGCCAGCAGGCGCTTCAA ACACTGATTCAGGAGACTGACCCTGGTGCAGATTACCGCATTGATCGAGCTTTGAATGAAGCTTGTGAATCTGTAATCCAGACAGCCTGC AAACATATAAGATCTGGAGACCCAATGATCTTGTCGTGCCTGATGGAACATTTATACACAGAGAAGATGGTAGAAGACTGTGAACACCGT CTCTTAGAGCTGCAGTATTTCATCTCCCGGGATTGGAAGCTGGACCCTGTCCTGTACCGCAAGTGCCAGGGAGACGCTTCTCGTCTTTGC CACACCCACGGTTGGAATGAGACCAGTGAATTTATGCCTCAGGGAGCTGTGTTCTCTTGTTTATACAGACACGCCTACCGCACTGAGGAA CAGGGAAGGAGGCTCTCACGGGAGTGCCGAGCTGAAGTCCAAAGGATCCTACACCAGCGTGCCATGGATGTCAAGCTGGATCCTGCCCTC CAGGATAAGTGCCTGATTGATCTGGGAAAATGGTGCAGTGAGAAAACAGAGACTGGACAGGAGCTGGAGTGCCTTCAGGACCATCTGGAT GACTTGGTGGTGGAGTGTAGAGATATAGTTGGCAACCTCACTGAGTTAGAATCAGAGGATTTGGTATCAGCACTAAATTGTGGGATTACT GTTTCCACACCCTCACTCCAGAGAAACCCCACCTGAAGACGCAGTGACAACTCCCACCCCCTCCGTCCTGCCCTCAGCCCGGCCCTGGCC CCTTCCCGACCCCCACCCGCCATTCAGACCCCATTAAGAAGGTTGGCTTGGCCAGGCAGGATGGGCTGTGTCCGGCCCTGCAGCCTAGTG GAAGGTGCTGAGGGGGCCCTGAGGCAGGACCGCCCTCCTGACCCCTGGTAGGAGGGTCACATCCACTTGGTGGCCAGGTGGCCCTTGGTG ACCCACTTCTTCCTGGAGCGTCCCTGCCTAGAGCTCAGCCCACAGGACTGCTTCAGGCCGTGGCCACAGGTAGCAGCCGCAAGGGGAAAT GAAGAAAACTGAGCCCTCGTGGCCACCTGTGTCACCCTTGTGCCTTAGCCTCATGGGCTGCCTAGGAGCTGCCTGCACGGCACAGCTCGC TTTCACAGTCAGAAGTGGGTCTGTGGGATCTGTGGTCCCTGTCCTCCCTGCTGTCCCTTCTGGGGAGGCTTTGGTGGCTCTGAGGTGGAC AAAGAGCTCTCGCAAGAAGAGACAGCGTGATGCCTCCCACAGTCCACCCCAGACCCTGGGGCAGCCCCTCTGGCCCTGCCAGCTGCCTGC >33246_33246_6_GLG1-FA2H_GLG1_chr16_74508444_ENST00000447066_FA2H_chr16_74748167_ENST00000544337_length(amino acids)=695AA_BP=673 MAACGRVRRMFRLSAALHLLLLFAAGAEKLPGQGVHSQGQGPGANFVSFVGQAGGGGPAGQQLPQLPQSSQLQQQQQQQQQQQQPQPPQP PFPAGGPPARRGGAGAGGGWKLAEEESCREDVTRVCPKHTWSNNLAVLECLQDVRELLWNYKLNLTTDPKFESVAREVCKSTITEIKECA DEPVGKGYMVSCLVDHRGNITEYQCHQYITKMTAIIFSDYRLICGFMDDCKNDINILKCGSIRLGEKDAHSQGEVVSCLEKGLVKEAEER EPKIQVSELCKKAILRVAELSSDDFHLDRHLYFACRDDRERFCENTQAGEGRVYKCLFNHKFEESMSEKCREALTTRQKLIAQDYKVSYS LAKSCKSDLKKYRCNVENLPRSREARLSYLLMCLESAVHRGRQVSSECQGEMLDYRRMLMEDFSLSPEIILSCRGEIEHHCSGLHRKGRT LHCLMKVVRGEKGNLGMNCQQALQTLIQETDPGADYRIDRALNEACESVIQTACKHIRSGDPMILSCLMEHLYTEKMVEDCEHRLLELQY FISRDWKLDPVLYRKCQGDASRLCHTHGWNETSEFMPQGAVFSCLYRHAYRTEEQGRRLSRECRAEVQRILHQRAMDVKLDPALQDKCLI -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for GLG1-FA2H |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for GLG1-FA2H |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for GLG1-FA2H |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
