
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:GRB10-SCIN (FusionGDB2 ID:34626) |
Fusion Gene Summary for GRB10-SCIN |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: GRB10-SCIN | Fusion gene ID: 34626 | Hgene | Tgene | Gene symbol | GRB10 | SCIN | Gene ID | 2887 | 85477 |
| Gene name | growth factor receptor bound protein 10 | scinderin | |
| Synonyms | GRB-IR|Grb-10|IRBP|MEG1|RSS | - | |
| Cytomap | 7p12.1 | 7p21.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | growth factor receptor-bound protein 10GRB10 adapter proteinGRB10 adaptor proteininsulin receptor-binding protein Grb-IRmaternally expressed gene 1 | adseverin | |
| Modification date | 20200327 | 20200313 | |
| UniProtAcc | Q13322 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000335866, ENST00000357271, ENST00000398810, ENST00000398812, ENST00000401949, ENST00000402497, ENST00000402578, ENST00000403097, ENST00000406641, ENST00000407526, ENST00000439599, ENST00000483819, | ENST00000473722, ENST00000445618, ENST00000519209, ENST00000297029, | |
| Fusion gene scores | * DoF score | 13 X 10 X 8=1040 | 5 X 6 X 5=150 |
| # samples | 16 | 6 | |
| ** MAII score | log2(16/1040*10)=-2.70043971814109 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(6/150*10)=-1.32192809488736 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: GRB10 [Title/Abstract] AND SCIN [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | GRB10(50737419)-SCIN(12662426), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | GRB10-SCIN seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | GRB10 | GO:0030178 | negative regulation of Wnt signaling pathway | 17376403 |
| Hgene | GRB10 | GO:0030949 | positive regulation of vascular endothelial growth factor receptor signaling pathway | 15060076 |
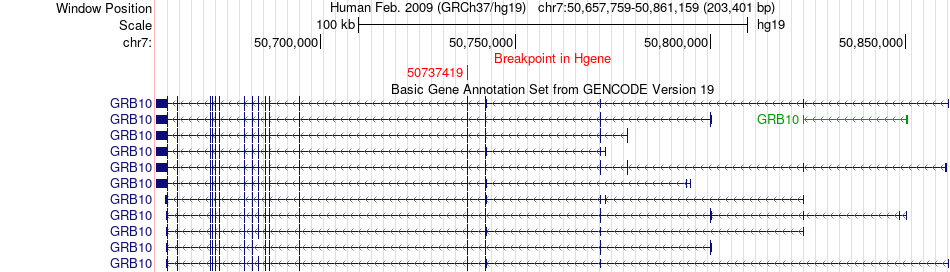
 Fusion gene breakpoints across GRB10 (5'-gene) Fusion gene breakpoints across GRB10 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene breakpoints across SCIN (3'-gene) Fusion gene breakpoints across SCIN (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
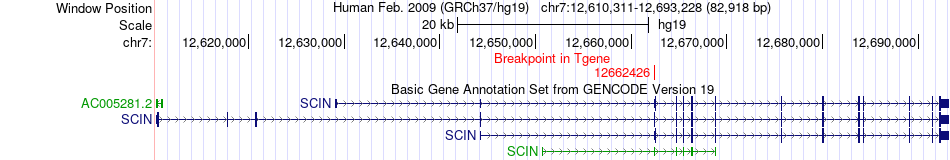 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | OV | TCGA-24-1557-01A | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
Top |
Fusion Gene ORF analysis for GRB10-SCIN |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000335866 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000357271 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000398810 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000398812 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000401949 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000402497 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000402578 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000403097 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000406641 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000407526 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-3UTR | ENST00000439599 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000335866 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000335866 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000357271 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000357271 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000398810 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000398810 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000398812 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000398812 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000401949 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000401949 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000402497 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000402497 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000402578 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000402578 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000403097 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000403097 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000406641 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000406641 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000407526 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000407526 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000439599 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| 5CDS-5UTR | ENST00000439599 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| Frame-shift | ENST00000335866 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| Frame-shift | ENST00000401949 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| Frame-shift | ENST00000402497 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| Frame-shift | ENST00000403097 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| Frame-shift | ENST00000406641 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| Frame-shift | ENST00000407526 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| In-frame | ENST00000357271 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| In-frame | ENST00000398810 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| In-frame | ENST00000398812 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| In-frame | ENST00000402578 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| In-frame | ENST00000439599 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| intron-3CDS | ENST00000483819 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| intron-3UTR | ENST00000483819 | ENST00000473722 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| intron-5UTR | ENST00000483819 | ENST00000445618 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
| intron-5UTR | ENST00000483819 | ENST00000519209 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000398810 | GRB10 | chr7 | 50737419 | - | ENST00000297029 | SCIN | chr7 | 12662426 | + | 2985 | 615 | 132 | 2096 | 654 |
| ENST00000439599 | GRB10 | chr7 | 50737419 | - | ENST00000297029 | SCIN | chr7 | 12662426 | + | 2856 | 486 | 0 | 1967 | 655 |
| ENST00000398812 | GRB10 | chr7 | 50737419 | - | ENST00000297029 | SCIN | chr7 | 12662426 | + | 2905 | 535 | 31 | 2016 | 661 |
| ENST00000402578 | GRB10 | chr7 | 50737419 | - | ENST00000297029 | SCIN | chr7 | 12662426 | + | 2965 | 595 | 244 | 2076 | 610 |
| ENST00000357271 | GRB10 | chr7 | 50737419 | - | ENST00000297029 | SCIN | chr7 | 12662426 | + | 2905 | 535 | 31 | 2016 | 661 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000398810 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + | 0.00123156 | 0.9987684 |
| ENST00000439599 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + | 0.000930468 | 0.9990695 |
| ENST00000398812 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + | 0.000536722 | 0.99946326 |
| ENST00000402578 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + | 0.001049041 | 0.99895096 |
| ENST00000357271 | ENST00000297029 | GRB10 | chr7 | 50737419 | - | SCIN | chr7 | 12662426 | + | 0.000536722 | 0.99946326 |
Top |
Fusion Genomic Features for GRB10-SCIN |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| GRB10 | chr7 | 50737418 | - | SCIN | chr7 | 12662425 | + | 0.17624581 | 0.82375413 |
| GRB10 | chr7 | 50737418 | - | SCIN | chr7 | 12662425 | + | 0.17624581 | 0.82375413 |
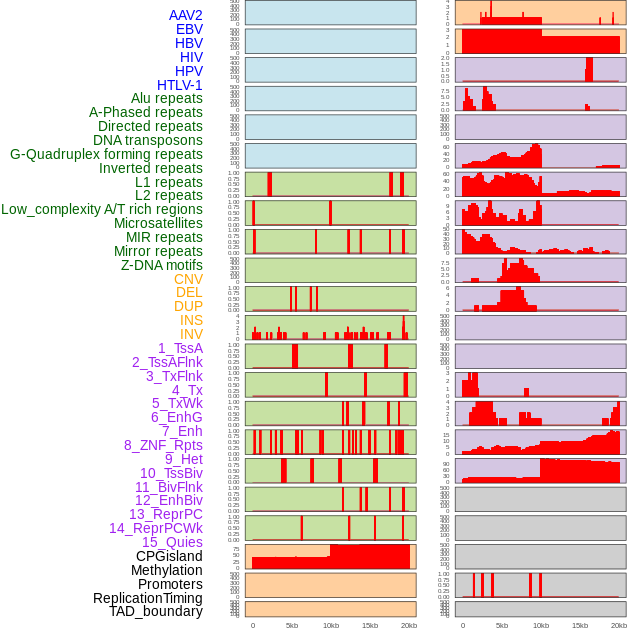
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
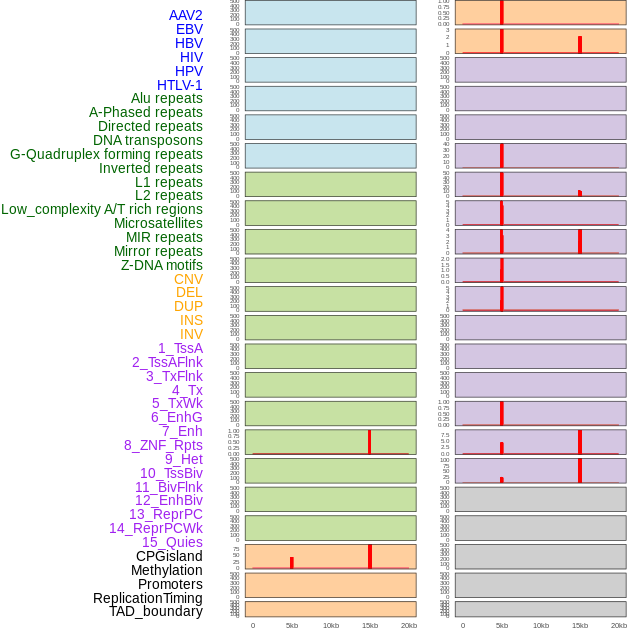
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for GRB10-SCIN |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr7:50737419/chr7:12662426) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| GRB10 | . |
| FUNCTION: Adapter protein which modulates coupling of a number of cell surface receptor kinases with specific signaling pathways. Binds to, and suppress signals from, activated receptors tyrosine kinases, including the insulin (INSR) and insulin-like growth factor (IGF1R) receptors. The inhibitory effect can be achieved by 2 mechanisms: interference with the signaling pathway and increased receptor degradation. Delays and reduces AKT1 phosphorylation in response to insulin stimulation. Blocks association between INSR and IRS1 and IRS2 and prevents insulin-stimulated IRS1 and IRS2 tyrosine phosphorylation. Recruits NEDD4 to IGF1R, leading to IGF1R ubiquitination, increased internalization and degradation by both the proteasomal and lysosomal pathways. May play a role in mediating insulin-stimulated ubiquitination of INSR, leading to proteasomal degradation. Negatively regulates Wnt signaling by interacting with LRP6 intracellular portion and interfering with the binding of AXIN1 to LRP6. Positive regulator of the KDR/VEGFR-2 signaling pathway. May inhibit NEDD4-mediated degradation of KDR/VEGFR-2. {ECO:0000269|PubMed:12493740, ECO:0000269|PubMed:15060076, ECO:0000269|PubMed:16434550, ECO:0000269|PubMed:17376403}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 364_715 | 222 | 716.0 | Region | Note=Ca(2+)-dependent actin binding | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 112_119 | 0 | 469.0 | Region | Polyphosphoinositide binding | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 138_146 | 0 | 469.0 | Region | Polyphosphoinositide binding | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 1_363 | 0 | 469.0 | Region | Actin-severing | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 364_715 | 0 | 469.0 | Region | Note=Ca(2+)-dependent actin binding | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 112_119 | 0 | 469.0 | Region | Polyphosphoinositide binding | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 138_146 | 0 | 469.0 | Region | Polyphosphoinositide binding | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 1_363 | 0 | 469.0 | Region | Actin-severing | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 364_715 | 0 | 469.0 | Region | Note=Ca(2+)-dependent actin binding | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 265_307 | 222 | 716.0 | Repeat | Note=Gelsolin-like 3 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 398_451 | 222 | 716.0 | Repeat | Note=Gelsolin-like 4 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 523_564 | 222 | 716.0 | Repeat | Note=Gelsolin-like 5 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 626_668 | 222 | 716.0 | Repeat | Note=Gelsolin-like 6 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 148_188 | 0 | 469.0 | Repeat | Note=Gelsolin-like 2 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 265_307 | 0 | 469.0 | Repeat | Note=Gelsolin-like 3 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 27_76 | 0 | 469.0 | Repeat | Note=Gelsolin-like 1 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 398_451 | 0 | 469.0 | Repeat | Note=Gelsolin-like 4 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 523_564 | 0 | 469.0 | Repeat | Note=Gelsolin-like 5 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000445618 | 0 | 13 | 626_668 | 0 | 469.0 | Repeat | Note=Gelsolin-like 6 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 148_188 | 0 | 469.0 | Repeat | Note=Gelsolin-like 2 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 265_307 | 0 | 469.0 | Repeat | Note=Gelsolin-like 3 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 27_76 | 0 | 469.0 | Repeat | Note=Gelsolin-like 1 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 398_451 | 0 | 469.0 | Repeat | Note=Gelsolin-like 4 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 523_564 | 0 | 469.0 | Repeat | Note=Gelsolin-like 5 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000519209 | 1 | 14 | 626_668 | 0 | 469.0 | Repeat | Note=Gelsolin-like 6 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000335866 | - | 5 | 17 | 166_250 | 110 | 537.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000335866 | - | 5 | 17 | 290_399 | 110 | 537.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000335866 | - | 5 | 17 | 493_574 | 110 | 537.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000357271 | - | 4 | 15 | 166_250 | 168 | 549.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000357271 | - | 4 | 15 | 290_399 | 168 | 549.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000357271 | - | 4 | 15 | 493_574 | 168 | 549.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000398810 | - | 4 | 16 | 166_250 | 110 | 537.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000398810 | - | 4 | 16 | 290_399 | 110 | 537.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000398810 | - | 4 | 16 | 493_574 | 110 | 537.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000398812 | - | 4 | 16 | 166_250 | 168 | 595.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000398812 | - | 4 | 16 | 290_399 | 168 | 595.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000398812 | - | 4 | 16 | 493_574 | 168 | 595.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000401949 | - | 7 | 19 | 166_250 | 168 | 595.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000401949 | - | 7 | 19 | 290_399 | 168 | 595.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000401949 | - | 7 | 19 | 493_574 | 168 | 595.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000402497 | - | 4 | 16 | 166_250 | 110 | 537.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000402497 | - | 4 | 16 | 290_399 | 110 | 537.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000402497 | - | 4 | 16 | 493_574 | 110 | 537.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000402578 | - | 4 | 16 | 166_250 | 110 | 537.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000402578 | - | 4 | 16 | 290_399 | 110 | 537.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000402578 | - | 4 | 16 | 493_574 | 110 | 537.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000403097 | - | 6 | 18 | 166_250 | 162 | 589.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000403097 | - | 6 | 18 | 290_399 | 162 | 589.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000403097 | - | 6 | 18 | 493_574 | 162 | 589.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000406641 | - | 5 | 17 | 166_250 | 110 | 537.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000406641 | - | 5 | 17 | 290_399 | 110 | 537.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000406641 | - | 5 | 17 | 493_574 | 110 | 537.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000407526 | - | 4 | 16 | 166_250 | 110 | 537.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000407526 | - | 4 | 16 | 290_399 | 110 | 537.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000407526 | - | 4 | 16 | 493_574 | 110 | 537.0 | Domain | SH2 |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000439599 | - | 4 | 16 | 166_250 | 162 | 589.0 | Domain | Ras-associating |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000439599 | - | 4 | 16 | 290_399 | 162 | 589.0 | Domain | PH |
| Hgene | GRB10 | chr7:50737419 | chr7:12662426 | ENST00000439599 | - | 4 | 16 | 493_574 | 162 | 589.0 | Domain | SH2 |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 112_119 | 222 | 716.0 | Region | Polyphosphoinositide binding | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 138_146 | 222 | 716.0 | Region | Polyphosphoinositide binding | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 1_363 | 222 | 716.0 | Region | Actin-severing | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 148_188 | 222 | 716.0 | Repeat | Note=Gelsolin-like 2 | |
| Tgene | SCIN | chr7:50737419 | chr7:12662426 | ENST00000297029 | 3 | 16 | 27_76 | 222 | 716.0 | Repeat | Note=Gelsolin-like 1 |
Top |
Fusion Gene Sequence for GRB10-SCIN |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >34626_34626_1_GRB10-SCIN_GRB10_chr7_50737419_ENST00000357271_SCIN_chr7_12662426_ENST00000297029_length(transcript)=2905nt_BP=535nt AAATGTAATTTGAAGAAGGCAGAAGGAACCCATGGCTTTAGCCGGCTGCCCAGATTCCTTTTTGCACCATCCGTACTACCAGGACAAGGT GGAGCAGACACCTCGCAGTCAACAAGACCCGGCAGGACCAGGACTCCCCGCACAGTCTGACCGACTTGCGAATCACCAGGAGGATGATGT GGACCTGGAAGCCCTGGTGAACGATATGAATGCATCCCTGGAGAGCCTGTACTCGGCCTGCAGCATGCAGTCAGACACGGTGCCCCTCCT GCAGAATGGCCAGCATGCCCGCAGCCAGCCTCGGGCTTCAGGCCCTCCTCGGTCCATCCAGCCACAGGTGTCCCCGAGGCAGAGGGTGCA GCGCTCCCAGCCTGTGCACATCCTCGCTGTCAGGCGCCTTCAGGAGGAAGACCAGCAGTTTAGAACCTCATCTCTGCCGGCCATCCCCAA TCCTTTTCCTGAACTCTGTGGCCCTGGGAGCCCCCCTGTGCTCACGCCGGGTTCTTTACCTCCGAGCCAGGCCGCCGCAAAGCAGGTCTT AGGGGAAAAGCCAGAGCTTCCAGATGGAGGTGATGATGATGACATTATAGCAGACATAAGTAACAGGAAAATGGCTAAACTATACATGGT TTCAGATGCAAGTGGCTCCATGAGAGTGACTGTGGTGGCAGAAGAAAACCCCTTCTCAATGGCAATGCTGCTGTCTGAAGAATGCTTTAT TTTGGACCACGGGGCTGCCAAACAAATTTTCGTATGGAAAGGTAAAGATGCTAATCCCCAAGAGAGGAAGGCTGCAATGAAGACAGCTGA AGAATTTCTACAGCAAATGAATTATTCCAAGAATACCCAAATTCAAGTTCTTCCAGAAGGAGGTGAAACACCAATCTTCAAACAGTTTTT TAAGGACTGGAGAGATAAAGATCAGAGTGATGGCTTCGGGAAAGTTTATGTCACAGAGAAAGTGGCTCAAATAAAACAAATTCCCTTTGA TGCCTCAAAATTACACAGTTCTCCGCAGATGGCAGCCCAGCACAATATGGTGGATGATGGTTCTGGCAAAGTGGAGATTTGGCGTGTAGA AAACAATGGTAGGATCCAAGTTGACCAAAACTCATATGGTGAATTCTATGGTGGTGACTGCTACATCATACTCTACACCTATCCCAGAGG ACAGATTATCTACACGTGGCAAGGAGCAAATGCCACACGAGATGAGCTGACAACATCTGCGTTCCTGACTGTTCAGTTGGATCGGTCCCT TGGAGGACAGGCTGTGCAGATCCGAGTCTCCCAAGGCAAAGAGCCTGTTCACCTACTGAGTTTGTTCAAAGACAAACCGCTCATTATTTA CAAGAATGGAACATCAAAGAAAGGAGGTCAGGCACCTGCTCCCCCTACACGCCTCTTTCAAGTCCGGAGAAACCTGGCATCTATCACCAG AATTGTGGAGGTTGATGTTGATGCAAATTCACTGAATTCTAACGATGTTTTTGTCCTGAAACTGCCACAAAATAGTGGCTACATCTGGGT AGGAAAAGGTGCTAGCCAGGAGGAGGAGAAAGGAGCAGAGTATGTAGCAAGTGTCCTAAAGTGCAAAACCTTAAGGATCCAAGAAGGCGA GGAGCCAGAGGAGTTCTGGAATTCCCTTGGAGGGAAAAAAGACTACCAGACCTCACCACTACTGGAAACCCAGGCTGAAGACCATCCACC TCGGCTTTACGGCTGCTCTAACAAAACTGGAAGATTTGTTATTGAAGAGATTCCAGGAGAGTTCACCCAGGATGATTTAGCTGAAGATGA TGTCATGTTACTAGATGCTTGGGAACAGATATTTATTTGGATTGGCAAAGATGCTAATGAAGTTGAGAAAAAAGAATCTCTGAAGTCTGC CAAAATGTACCTTGAGACAGACCCTTCTGGAAGAGACAAGAGGACACCAATTGTCATCATAAAACAGGGCCATGAGCCACCCACATTCAC AGGCTGGTTCCTGGGCTGGGATTCCAGCAAGTGGTAAATTGGTATTTGTAAAAAGCAAACAAACATTACAAGGCAGTTATCTCATTGCTG TTTTGGGAGAGGAACGGGAAAAGCTTTTTGCTTATTTGTCTTTTGAAAATTAAGGCTGGGCGCGGTGGCTCACACCTGTAATCCCAGCAC TTTGAGAGGATGAGGTAGGCGGATCACTGGGGTCAGGATTTCGAGACCAGCCTGGCCAACATGGCGAAACCTCGCCTCTACTAAAAATAC AAAAAAATTAGCTGCGCGTGGTGGTGCACGCCTGTAGTCCCTGCTACTTGGAAGGCTGAGACAGGAAAATTGCTTGAACCCAGGAGGCTG AGGTTGCAGTGAGCCAGGATTGCGCCACCACACTCCAGCCTGGGCAACAGAGACTCTGTCTCAAAAAAAAAAAAAAAAAAAATTAACCTC AACCAAACGCCTATTTTTTAATGCTTAGTTTTGGCTTGAAATTCTTCTTCACCTGGAGTTTTCTTACGTTAATACATTAAAATAACCTGA AGGAAACTTTCGTTATGGGCCAATATTAGCTCTTATGGAAACTGATTTATATCTTTTTTATGACCTTTCAAAAGTAAAATACTATGCATA ATCTAGAAAGATTTGTCAGATAGGAAATTTTATAATGCATTAGCCATTAGTCAGAGTTGTTTTTTAACATGCCAGAGAAAAAGTTGTAAT GCCTTGGAAGTTATTCTCTTTTCTATAGATTGTGTTCAAAAGATGTGTATACTTGCAATAAGGTTTATGTTAAGGTGGCTTTAACAATTG GTTGCTATTTTCCTTCTTTAGTCAATATGACTTTGATGTCATTAACTGCTAAGTGTTATTTTCTAAGAGGAATTTTATCTTTTTTTTAAC >34626_34626_1_GRB10-SCIN_GRB10_chr7_50737419_ENST00000357271_SCIN_chr7_12662426_ENST00000297029_length(amino acids)=661AA_BP=167 MALAGCPDSFLHHPYYQDKVEQTPRSQQDPAGPGLPAQSDRLANHQEDDVDLEALVNDMNASLESLYSACSMQSDTVPLLQNGQHARSQP RASGPPRSIQPQVSPRQRVQRSQPVHILAVRRLQEEDQQFRTSSLPAIPNPFPELCGPGSPPVLTPGSLPPSQAAAKQVLGEKPELPDGG DDDDIIADISNRKMAKLYMVSDASGSMRVTVVAEENPFSMAMLLSEECFILDHGAAKQIFVWKGKDANPQERKAAMKTAEEFLQQMNYSK NTQIQVLPEGGETPIFKQFFKDWRDKDQSDGFGKVYVTEKVAQIKQIPFDASKLHSSPQMAAQHNMVDDGSGKVEIWRVENNGRIQVDQN SYGEFYGGDCYIILYTYPRGQIIYTWQGANATRDELTTSAFLTVQLDRSLGGQAVQIRVSQGKEPVHLLSLFKDKPLIIYKNGTSKKGGQ APAPPTRLFQVRRNLASITRIVEVDVDANSLNSNDVFVLKLPQNSGYIWVGKGASQEEEKGAEYVASVLKCKTLRIQEGEEPEEFWNSLG GKKDYQTSPLLETQAEDHPPRLYGCSNKTGRFVIEEIPGEFTQDDLAEDDVMLLDAWEQIFIWIGKDANEVEKKESLKSAKMYLETDPSG -------------------------------------------------------------- >34626_34626_2_GRB10-SCIN_GRB10_chr7_50737419_ENST00000398810_SCIN_chr7_12662426_ENST00000297029_length(transcript)=2985nt_BP=615nt CGCAACTTTGCCTCCCAGGGAACAAACATCCTCCTTCTAAGTGGTAGATGTGGGTGAGCTGACCCTGCTGGAGTCTGTCCCCTGGGCTAC CCTCTGCTTCCCCCCATTGTGAGTGGTCCGTGAAGCACAGCGTTGACCAGACCTAAACCTGTTTGCTCCCAGGACAAGGTGGAGCAGACA CCTCGCAGTCAACAAGACCCGGCAGGACCAGGACTCCCCGCACAGTCTGACCGACTTGCGAATCACCAGGAGGATGATGTGGACCTGGAA GCCCTGGTGAACGATATGAATGCATCCCTGGAGAGCCTGTACTCGGCCTGCAGCATGCAGTCAGACACGGTGCCCCTCCTGCAGAATGGC CAGCATGCCCGCAGCCAGCCTCGGGCTTCAGGCCCTCCTCGGTCCATCCAGCCACAGGTGTCCCCGAGGCAGAGGGTGCAGCGCTCCCAG CCTGTGCACATCCTCGCTGTCAGGCGCCTTCAGGAGGAAGACCAGCAGTTTAGAACCTCATCTCTGCCGGCCATCCCCAATCCTTTTCCT GAACTCTGTGGCCCTGGGAGCCCCCCTGTGCTCACGCCGGGTTCTTTACCTCCGAGCCAGGCCGCCGCAAAGCAGGTCTTAGGGGAAAAG CCAGAGCTTCCAGATGGAGGTGATGATGATGACATTATAGCAGACATAAGTAACAGGAAAATGGCTAAACTATACATGGTTTCAGATGCA AGTGGCTCCATGAGAGTGACTGTGGTGGCAGAAGAAAACCCCTTCTCAATGGCAATGCTGCTGTCTGAAGAATGCTTTATTTTGGACCAC GGGGCTGCCAAACAAATTTTCGTATGGAAAGGTAAAGATGCTAATCCCCAAGAGAGGAAGGCTGCAATGAAGACAGCTGAAGAATTTCTA CAGCAAATGAATTATTCCAAGAATACCCAAATTCAAGTTCTTCCAGAAGGAGGTGAAACACCAATCTTCAAACAGTTTTTTAAGGACTGG AGAGATAAAGATCAGAGTGATGGCTTCGGGAAAGTTTATGTCACAGAGAAAGTGGCTCAAATAAAACAAATTCCCTTTGATGCCTCAAAA TTACACAGTTCTCCGCAGATGGCAGCCCAGCACAATATGGTGGATGATGGTTCTGGCAAAGTGGAGATTTGGCGTGTAGAAAACAATGGT AGGATCCAAGTTGACCAAAACTCATATGGTGAATTCTATGGTGGTGACTGCTACATCATACTCTACACCTATCCCAGAGGACAGATTATC TACACGTGGCAAGGAGCAAATGCCACACGAGATGAGCTGACAACATCTGCGTTCCTGACTGTTCAGTTGGATCGGTCCCTTGGAGGACAG GCTGTGCAGATCCGAGTCTCCCAAGGCAAAGAGCCTGTTCACCTACTGAGTTTGTTCAAAGACAAACCGCTCATTATTTACAAGAATGGA ACATCAAAGAAAGGAGGTCAGGCACCTGCTCCCCCTACACGCCTCTTTCAAGTCCGGAGAAACCTGGCATCTATCACCAGAATTGTGGAG GTTGATGTTGATGCAAATTCACTGAATTCTAACGATGTTTTTGTCCTGAAACTGCCACAAAATAGTGGCTACATCTGGGTAGGAAAAGGT GCTAGCCAGGAGGAGGAGAAAGGAGCAGAGTATGTAGCAAGTGTCCTAAAGTGCAAAACCTTAAGGATCCAAGAAGGCGAGGAGCCAGAG GAGTTCTGGAATTCCCTTGGAGGGAAAAAAGACTACCAGACCTCACCACTACTGGAAACCCAGGCTGAAGACCATCCACCTCGGCTTTAC GGCTGCTCTAACAAAACTGGAAGATTTGTTATTGAAGAGATTCCAGGAGAGTTCACCCAGGATGATTTAGCTGAAGATGATGTCATGTTA CTAGATGCTTGGGAACAGATATTTATTTGGATTGGCAAAGATGCTAATGAAGTTGAGAAAAAAGAATCTCTGAAGTCTGCCAAAATGTAC CTTGAGACAGACCCTTCTGGAAGAGACAAGAGGACACCAATTGTCATCATAAAACAGGGCCATGAGCCACCCACATTCACAGGCTGGTTC CTGGGCTGGGATTCCAGCAAGTGGTAAATTGGTATTTGTAAAAAGCAAACAAACATTACAAGGCAGTTATCTCATTGCTGTTTTGGGAGA GGAACGGGAAAAGCTTTTTGCTTATTTGTCTTTTGAAAATTAAGGCTGGGCGCGGTGGCTCACACCTGTAATCCCAGCACTTTGAGAGGA TGAGGTAGGCGGATCACTGGGGTCAGGATTTCGAGACCAGCCTGGCCAACATGGCGAAACCTCGCCTCTACTAAAAATACAAAAAAATTA GCTGCGCGTGGTGGTGCACGCCTGTAGTCCCTGCTACTTGGAAGGCTGAGACAGGAAAATTGCTTGAACCCAGGAGGCTGAGGTTGCAGT GAGCCAGGATTGCGCCACCACACTCCAGCCTGGGCAACAGAGACTCTGTCTCAAAAAAAAAAAAAAAAAAAATTAACCTCAACCAAACGC CTATTTTTTAATGCTTAGTTTTGGCTTGAAATTCTTCTTCACCTGGAGTTTTCTTACGTTAATACATTAAAATAACCTGAAGGAAACTTT CGTTATGGGCCAATATTAGCTCTTATGGAAACTGATTTATATCTTTTTTATGACCTTTCAAAAGTAAAATACTATGCATAATCTAGAAAG ATTTGTCAGATAGGAAATTTTATAATGCATTAGCCATTAGTCAGAGTTGTTTTTTAACATGCCAGAGAAAAAGTTGTAATGCCTTGGAAG TTATTCTCTTTTCTATAGATTGTGTTCAAAAGATGTGTATACTTGCAATAAGGTTTATGTTAAGGTGGCTTTAACAATTGGTTGCTATTT TCCTTCTTTAGTCAATATGACTTTGATGTCATTAACTGCTAAGTGTTATTTTCTAAGAGGAATTTTATCTTTTTTTTAACCTTATTAAAA >34626_34626_2_GRB10-SCIN_GRB10_chr7_50737419_ENST00000398810_SCIN_chr7_12662426_ENST00000297029_length(amino acids)=654AA_BP=160 MTRPKPVCSQDKVEQTPRSQQDPAGPGLPAQSDRLANHQEDDVDLEALVNDMNASLESLYSACSMQSDTVPLLQNGQHARSQPRASGPPR SIQPQVSPRQRVQRSQPVHILAVRRLQEEDQQFRTSSLPAIPNPFPELCGPGSPPVLTPGSLPPSQAAAKQVLGEKPELPDGGDDDDIIA DISNRKMAKLYMVSDASGSMRVTVVAEENPFSMAMLLSEECFILDHGAAKQIFVWKGKDANPQERKAAMKTAEEFLQQMNYSKNTQIQVL PEGGETPIFKQFFKDWRDKDQSDGFGKVYVTEKVAQIKQIPFDASKLHSSPQMAAQHNMVDDGSGKVEIWRVENNGRIQVDQNSYGEFYG GDCYIILYTYPRGQIIYTWQGANATRDELTTSAFLTVQLDRSLGGQAVQIRVSQGKEPVHLLSLFKDKPLIIYKNGTSKKGGQAPAPPTR LFQVRRNLASITRIVEVDVDANSLNSNDVFVLKLPQNSGYIWVGKGASQEEEKGAEYVASVLKCKTLRIQEGEEPEEFWNSLGGKKDYQT SPLLETQAEDHPPRLYGCSNKTGRFVIEEIPGEFTQDDLAEDDVMLLDAWEQIFIWIGKDANEVEKKESLKSAKMYLETDPSGRDKRTPI -------------------------------------------------------------- >34626_34626_3_GRB10-SCIN_GRB10_chr7_50737419_ENST00000398812_SCIN_chr7_12662426_ENST00000297029_length(transcript)=2905nt_BP=535nt AAATGTAATTTGAAGAAGGCAGAAGGAACCCATGGCTTTAGCCGGCTGCCCAGATTCCTTTTTGCACCATCCGTACTACCAGGACAAGGT GGAGCAGACACCTCGCAGTCAACAAGACCCGGCAGGACCAGGACTCCCCGCACAGTCTGACCGACTTGCGAATCACCAGGAGGATGATGT GGACCTGGAAGCCCTGGTGAACGATATGAATGCATCCCTGGAGAGCCTGTACTCGGCCTGCAGCATGCAGTCAGACACGGTGCCCCTCCT GCAGAATGGCCAGCATGCCCGCAGCCAGCCTCGGGCTTCAGGCCCTCCTCGGTCCATCCAGCCACAGGTGTCCCCGAGGCAGAGGGTGCA GCGCTCCCAGCCTGTGCACATCCTCGCTGTCAGGCGCCTTCAGGAGGAAGACCAGCAGTTTAGAACCTCATCTCTGCCGGCCATCCCCAA TCCTTTTCCTGAACTCTGTGGCCCTGGGAGCCCCCCTGTGCTCACGCCGGGTTCTTTACCTCCGAGCCAGGCCGCCGCAAAGCAGGTCTT AGGGGAAAAGCCAGAGCTTCCAGATGGAGGTGATGATGATGACATTATAGCAGACATAAGTAACAGGAAAATGGCTAAACTATACATGGT TTCAGATGCAAGTGGCTCCATGAGAGTGACTGTGGTGGCAGAAGAAAACCCCTTCTCAATGGCAATGCTGCTGTCTGAAGAATGCTTTAT TTTGGACCACGGGGCTGCCAAACAAATTTTCGTATGGAAAGGTAAAGATGCTAATCCCCAAGAGAGGAAGGCTGCAATGAAGACAGCTGA AGAATTTCTACAGCAAATGAATTATTCCAAGAATACCCAAATTCAAGTTCTTCCAGAAGGAGGTGAAACACCAATCTTCAAACAGTTTTT TAAGGACTGGAGAGATAAAGATCAGAGTGATGGCTTCGGGAAAGTTTATGTCACAGAGAAAGTGGCTCAAATAAAACAAATTCCCTTTGA TGCCTCAAAATTACACAGTTCTCCGCAGATGGCAGCCCAGCACAATATGGTGGATGATGGTTCTGGCAAAGTGGAGATTTGGCGTGTAGA AAACAATGGTAGGATCCAAGTTGACCAAAACTCATATGGTGAATTCTATGGTGGTGACTGCTACATCATACTCTACACCTATCCCAGAGG ACAGATTATCTACACGTGGCAAGGAGCAAATGCCACACGAGATGAGCTGACAACATCTGCGTTCCTGACTGTTCAGTTGGATCGGTCCCT TGGAGGACAGGCTGTGCAGATCCGAGTCTCCCAAGGCAAAGAGCCTGTTCACCTACTGAGTTTGTTCAAAGACAAACCGCTCATTATTTA CAAGAATGGAACATCAAAGAAAGGAGGTCAGGCACCTGCTCCCCCTACACGCCTCTTTCAAGTCCGGAGAAACCTGGCATCTATCACCAG AATTGTGGAGGTTGATGTTGATGCAAATTCACTGAATTCTAACGATGTTTTTGTCCTGAAACTGCCACAAAATAGTGGCTACATCTGGGT AGGAAAAGGTGCTAGCCAGGAGGAGGAGAAAGGAGCAGAGTATGTAGCAAGTGTCCTAAAGTGCAAAACCTTAAGGATCCAAGAAGGCGA GGAGCCAGAGGAGTTCTGGAATTCCCTTGGAGGGAAAAAAGACTACCAGACCTCACCACTACTGGAAACCCAGGCTGAAGACCATCCACC TCGGCTTTACGGCTGCTCTAACAAAACTGGAAGATTTGTTATTGAAGAGATTCCAGGAGAGTTCACCCAGGATGATTTAGCTGAAGATGA TGTCATGTTACTAGATGCTTGGGAACAGATATTTATTTGGATTGGCAAAGATGCTAATGAAGTTGAGAAAAAAGAATCTCTGAAGTCTGC CAAAATGTACCTTGAGACAGACCCTTCTGGAAGAGACAAGAGGACACCAATTGTCATCATAAAACAGGGCCATGAGCCACCCACATTCAC AGGCTGGTTCCTGGGCTGGGATTCCAGCAAGTGGTAAATTGGTATTTGTAAAAAGCAAACAAACATTACAAGGCAGTTATCTCATTGCTG TTTTGGGAGAGGAACGGGAAAAGCTTTTTGCTTATTTGTCTTTTGAAAATTAAGGCTGGGCGCGGTGGCTCACACCTGTAATCCCAGCAC TTTGAGAGGATGAGGTAGGCGGATCACTGGGGTCAGGATTTCGAGACCAGCCTGGCCAACATGGCGAAACCTCGCCTCTACTAAAAATAC AAAAAAATTAGCTGCGCGTGGTGGTGCACGCCTGTAGTCCCTGCTACTTGGAAGGCTGAGACAGGAAAATTGCTTGAACCCAGGAGGCTG AGGTTGCAGTGAGCCAGGATTGCGCCACCACACTCCAGCCTGGGCAACAGAGACTCTGTCTCAAAAAAAAAAAAAAAAAAAATTAACCTC AACCAAACGCCTATTTTTTAATGCTTAGTTTTGGCTTGAAATTCTTCTTCACCTGGAGTTTTCTTACGTTAATACATTAAAATAACCTGA AGGAAACTTTCGTTATGGGCCAATATTAGCTCTTATGGAAACTGATTTATATCTTTTTTATGACCTTTCAAAAGTAAAATACTATGCATA ATCTAGAAAGATTTGTCAGATAGGAAATTTTATAATGCATTAGCCATTAGTCAGAGTTGTTTTTTAACATGCCAGAGAAAAAGTTGTAAT GCCTTGGAAGTTATTCTCTTTTCTATAGATTGTGTTCAAAAGATGTGTATACTTGCAATAAGGTTTATGTTAAGGTGGCTTTAACAATTG GTTGCTATTTTCCTTCTTTAGTCAATATGACTTTGATGTCATTAACTGCTAAGTGTTATTTTCTAAGAGGAATTTTATCTTTTTTTTAAC >34626_34626_3_GRB10-SCIN_GRB10_chr7_50737419_ENST00000398812_SCIN_chr7_12662426_ENST00000297029_length(amino acids)=661AA_BP=167 MALAGCPDSFLHHPYYQDKVEQTPRSQQDPAGPGLPAQSDRLANHQEDDVDLEALVNDMNASLESLYSACSMQSDTVPLLQNGQHARSQP RASGPPRSIQPQVSPRQRVQRSQPVHILAVRRLQEEDQQFRTSSLPAIPNPFPELCGPGSPPVLTPGSLPPSQAAAKQVLGEKPELPDGG DDDDIIADISNRKMAKLYMVSDASGSMRVTVVAEENPFSMAMLLSEECFILDHGAAKQIFVWKGKDANPQERKAAMKTAEEFLQQMNYSK NTQIQVLPEGGETPIFKQFFKDWRDKDQSDGFGKVYVTEKVAQIKQIPFDASKLHSSPQMAAQHNMVDDGSGKVEIWRVENNGRIQVDQN SYGEFYGGDCYIILYTYPRGQIIYTWQGANATRDELTTSAFLTVQLDRSLGGQAVQIRVSQGKEPVHLLSLFKDKPLIIYKNGTSKKGGQ APAPPTRLFQVRRNLASITRIVEVDVDANSLNSNDVFVLKLPQNSGYIWVGKGASQEEEKGAEYVASVLKCKTLRIQEGEEPEEFWNSLG GKKDYQTSPLLETQAEDHPPRLYGCSNKTGRFVIEEIPGEFTQDDLAEDDVMLLDAWEQIFIWIGKDANEVEKKESLKSAKMYLETDPSG -------------------------------------------------------------- >34626_34626_4_GRB10-SCIN_GRB10_chr7_50737419_ENST00000402578_SCIN_chr7_12662426_ENST00000297029_length(transcript)=2965nt_BP=595nt ACCGACTCTGTGAATTGGGTACCTTCATGATACCCATTTTATAAATGAGAAAACTGAGGTTTTTACTATATGCTGGTCATCATGCTGAGG ACTTTACTTGGATTTTCTCTCCTGCCAATGCTACGCGGTGGGGACCTTCATGTTCTCCCCTTTGCAGATGAGATTATCAAAGCAGAAAAA GACCATCTCTCCTACTGAGTAACAGAGGAAACCAGGGCTTGCAGATATTGAGGATGATGTGGACCTGGAAGCCCTGGTGAACGATATGAA TGCATCCCTGGAGAGCCTGTACTCGGCCTGCAGCATGCAGTCAGACACGGTGCCCCTCCTGCAGAATGGCCAGCATGCCCGCAGCCAGCC TCGGGCTTCAGGCCCTCCTCGGTCCATCCAGCCACAGGTGTCCCCGAGGCAGAGGGTGCAGCGCTCCCAGCCTGTGCACATCCTCGCTGT CAGGCGCCTTCAGGAGGAAGACCAGCAGTTTAGAACCTCATCTCTGCCGGCCATCCCCAATCCTTTTCCTGAACTCTGTGGCCCTGGGAG CCCCCCTGTGCTCACGCCGGGTTCTTTACCTCCGAGCCAGGCCGCCGCAAAGCAGGTCTTAGGGGAAAAGCCAGAGCTTCCAGATGGAGG TGATGATGATGACATTATAGCAGACATAAGTAACAGGAAAATGGCTAAACTATACATGGTTTCAGATGCAAGTGGCTCCATGAGAGTGAC TGTGGTGGCAGAAGAAAACCCCTTCTCAATGGCAATGCTGCTGTCTGAAGAATGCTTTATTTTGGACCACGGGGCTGCCAAACAAATTTT CGTATGGAAAGGTAAAGATGCTAATCCCCAAGAGAGGAAGGCTGCAATGAAGACAGCTGAAGAATTTCTACAGCAAATGAATTATTCCAA GAATACCCAAATTCAAGTTCTTCCAGAAGGAGGTGAAACACCAATCTTCAAACAGTTTTTTAAGGACTGGAGAGATAAAGATCAGAGTGA TGGCTTCGGGAAAGTTTATGTCACAGAGAAAGTGGCTCAAATAAAACAAATTCCCTTTGATGCCTCAAAATTACACAGTTCTCCGCAGAT GGCAGCCCAGCACAATATGGTGGATGATGGTTCTGGCAAAGTGGAGATTTGGCGTGTAGAAAACAATGGTAGGATCCAAGTTGACCAAAA CTCATATGGTGAATTCTATGGTGGTGACTGCTACATCATACTCTACACCTATCCCAGAGGACAGATTATCTACACGTGGCAAGGAGCAAA TGCCACACGAGATGAGCTGACAACATCTGCGTTCCTGACTGTTCAGTTGGATCGGTCCCTTGGAGGACAGGCTGTGCAGATCCGAGTCTC CCAAGGCAAAGAGCCTGTTCACCTACTGAGTTTGTTCAAAGACAAACCGCTCATTATTTACAAGAATGGAACATCAAAGAAAGGAGGTCA GGCACCTGCTCCCCCTACACGCCTCTTTCAAGTCCGGAGAAACCTGGCATCTATCACCAGAATTGTGGAGGTTGATGTTGATGCAAATTC ACTGAATTCTAACGATGTTTTTGTCCTGAAACTGCCACAAAATAGTGGCTACATCTGGGTAGGAAAAGGTGCTAGCCAGGAGGAGGAGAA AGGAGCAGAGTATGTAGCAAGTGTCCTAAAGTGCAAAACCTTAAGGATCCAAGAAGGCGAGGAGCCAGAGGAGTTCTGGAATTCCCTTGG AGGGAAAAAAGACTACCAGACCTCACCACTACTGGAAACCCAGGCTGAAGACCATCCACCTCGGCTTTACGGCTGCTCTAACAAAACTGG AAGATTTGTTATTGAAGAGATTCCAGGAGAGTTCACCCAGGATGATTTAGCTGAAGATGATGTCATGTTACTAGATGCTTGGGAACAGAT ATTTATTTGGATTGGCAAAGATGCTAATGAAGTTGAGAAAAAAGAATCTCTGAAGTCTGCCAAAATGTACCTTGAGACAGACCCTTCTGG AAGAGACAAGAGGACACCAATTGTCATCATAAAACAGGGCCATGAGCCACCCACATTCACAGGCTGGTTCCTGGGCTGGGATTCCAGCAA GTGGTAAATTGGTATTTGTAAAAAGCAAACAAACATTACAAGGCAGTTATCTCATTGCTGTTTTGGGAGAGGAACGGGAAAAGCTTTTTG CTTATTTGTCTTTTGAAAATTAAGGCTGGGCGCGGTGGCTCACACCTGTAATCCCAGCACTTTGAGAGGATGAGGTAGGCGGATCACTGG GGTCAGGATTTCGAGACCAGCCTGGCCAACATGGCGAAACCTCGCCTCTACTAAAAATACAAAAAAATTAGCTGCGCGTGGTGGTGCACG CCTGTAGTCCCTGCTACTTGGAAGGCTGAGACAGGAAAATTGCTTGAACCCAGGAGGCTGAGGTTGCAGTGAGCCAGGATTGCGCCACCA CACTCCAGCCTGGGCAACAGAGACTCTGTCTCAAAAAAAAAAAAAAAAAAAATTAACCTCAACCAAACGCCTATTTTTTAATGCTTAGTT TTGGCTTGAAATTCTTCTTCACCTGGAGTTTTCTTACGTTAATACATTAAAATAACCTGAAGGAAACTTTCGTTATGGGCCAATATTAGC TCTTATGGAAACTGATTTATATCTTTTTTATGACCTTTCAAAAGTAAAATACTATGCATAATCTAGAAAGATTTGTCAGATAGGAAATTT TATAATGCATTAGCCATTAGTCAGAGTTGTTTTTTAACATGCCAGAGAAAAAGTTGTAATGCCTTGGAAGTTATTCTCTTTTCTATAGAT TGTGTTCAAAAGATGTGTATACTTGCAATAAGGTTTATGTTAAGGTGGCTTTAACAATTGGTTGCTATTTTCCTTCTTTAGTCAATATGA >34626_34626_4_GRB10-SCIN_GRB10_chr7_50737419_ENST00000402578_SCIN_chr7_12662426_ENST00000297029_length(amino acids)=610AA_BP=116 MEALVNDMNASLESLYSACSMQSDTVPLLQNGQHARSQPRASGPPRSIQPQVSPRQRVQRSQPVHILAVRRLQEEDQQFRTSSLPAIPNP FPELCGPGSPPVLTPGSLPPSQAAAKQVLGEKPELPDGGDDDDIIADISNRKMAKLYMVSDASGSMRVTVVAEENPFSMAMLLSEECFIL DHGAAKQIFVWKGKDANPQERKAAMKTAEEFLQQMNYSKNTQIQVLPEGGETPIFKQFFKDWRDKDQSDGFGKVYVTEKVAQIKQIPFDA SKLHSSPQMAAQHNMVDDGSGKVEIWRVENNGRIQVDQNSYGEFYGGDCYIILYTYPRGQIIYTWQGANATRDELTTSAFLTVQLDRSLG GQAVQIRVSQGKEPVHLLSLFKDKPLIIYKNGTSKKGGQAPAPPTRLFQVRRNLASITRIVEVDVDANSLNSNDVFVLKLPQNSGYIWVG KGASQEEEKGAEYVASVLKCKTLRIQEGEEPEEFWNSLGGKKDYQTSPLLETQAEDHPPRLYGCSNKTGRFVIEEIPGEFTQDDLAEDDV -------------------------------------------------------------- >34626_34626_5_GRB10-SCIN_GRB10_chr7_50737419_ENST00000439599_SCIN_chr7_12662426_ENST00000297029_length(transcript)=2856nt_BP=486nt ATGCAAGCTGCCGGCCCTCTGTTCCGTAGTAAGGACAAGGTGGAGCAGACACCTCGCAGTCAACAAGACCCGGCAGGACCAGGACTCCCC GCACAGTCTGACCGACTTGCGAATCACCAGGAGGATGATGTGGACCTGGAAGCCCTGGTGAACGATATGAATGCATCCCTGGAGAGCCTG TACTCGGCCTGCAGCATGCAGTCAGACACGGTGCCCCTCCTGCAGAATGGCCAGCATGCCCGCAGCCAGCCTCGGGCTTCAGGCCCTCCT CGGTCCATCCAGCCACAGGTGTCCCCGAGGCAGAGGGTGCAGCGCTCCCAGCCTGTGCACATCCTCGCTGTCAGGCGCCTTCAGGAGGAA GACCAGCAGTTTAGAACCTCATCTCTGCCGGCCATCCCCAATCCTTTTCCTGAACTCTGTGGCCCTGGGAGCCCCCCTGTGCTCACGCCG GGTTCTTTACCTCCGAGCCAGGCCGCCGCAAAGCAGGTCTTAGGGGAAAAGCCAGAGCTTCCAGATGGAGGTGATGATGATGACATTATA GCAGACATAAGTAACAGGAAAATGGCTAAACTATACATGGTTTCAGATGCAAGTGGCTCCATGAGAGTGACTGTGGTGGCAGAAGAAAAC CCCTTCTCAATGGCAATGCTGCTGTCTGAAGAATGCTTTATTTTGGACCACGGGGCTGCCAAACAAATTTTCGTATGGAAAGGTAAAGAT GCTAATCCCCAAGAGAGGAAGGCTGCAATGAAGACAGCTGAAGAATTTCTACAGCAAATGAATTATTCCAAGAATACCCAAATTCAAGTT CTTCCAGAAGGAGGTGAAACACCAATCTTCAAACAGTTTTTTAAGGACTGGAGAGATAAAGATCAGAGTGATGGCTTCGGGAAAGTTTAT GTCACAGAGAAAGTGGCTCAAATAAAACAAATTCCCTTTGATGCCTCAAAATTACACAGTTCTCCGCAGATGGCAGCCCAGCACAATATG GTGGATGATGGTTCTGGCAAAGTGGAGATTTGGCGTGTAGAAAACAATGGTAGGATCCAAGTTGACCAAAACTCATATGGTGAATTCTAT GGTGGTGACTGCTACATCATACTCTACACCTATCCCAGAGGACAGATTATCTACACGTGGCAAGGAGCAAATGCCACACGAGATGAGCTG ACAACATCTGCGTTCCTGACTGTTCAGTTGGATCGGTCCCTTGGAGGACAGGCTGTGCAGATCCGAGTCTCCCAAGGCAAAGAGCCTGTT CACCTACTGAGTTTGTTCAAAGACAAACCGCTCATTATTTACAAGAATGGAACATCAAAGAAAGGAGGTCAGGCACCTGCTCCCCCTACA CGCCTCTTTCAAGTCCGGAGAAACCTGGCATCTATCACCAGAATTGTGGAGGTTGATGTTGATGCAAATTCACTGAATTCTAACGATGTT TTTGTCCTGAAACTGCCACAAAATAGTGGCTACATCTGGGTAGGAAAAGGTGCTAGCCAGGAGGAGGAGAAAGGAGCAGAGTATGTAGCA AGTGTCCTAAAGTGCAAAACCTTAAGGATCCAAGAAGGCGAGGAGCCAGAGGAGTTCTGGAATTCCCTTGGAGGGAAAAAAGACTACCAG ACCTCACCACTACTGGAAACCCAGGCTGAAGACCATCCACCTCGGCTTTACGGCTGCTCTAACAAAACTGGAAGATTTGTTATTGAAGAG ATTCCAGGAGAGTTCACCCAGGATGATTTAGCTGAAGATGATGTCATGTTACTAGATGCTTGGGAACAGATATTTATTTGGATTGGCAAA GATGCTAATGAAGTTGAGAAAAAAGAATCTCTGAAGTCTGCCAAAATGTACCTTGAGACAGACCCTTCTGGAAGAGACAAGAGGACACCA ATTGTCATCATAAAACAGGGCCATGAGCCACCCACATTCACAGGCTGGTTCCTGGGCTGGGATTCCAGCAAGTGGTAAATTGGTATTTGT AAAAAGCAAACAAACATTACAAGGCAGTTATCTCATTGCTGTTTTGGGAGAGGAACGGGAAAAGCTTTTTGCTTATTTGTCTTTTGAAAA TTAAGGCTGGGCGCGGTGGCTCACACCTGTAATCCCAGCACTTTGAGAGGATGAGGTAGGCGGATCACTGGGGTCAGGATTTCGAGACCA GCCTGGCCAACATGGCGAAACCTCGCCTCTACTAAAAATACAAAAAAATTAGCTGCGCGTGGTGGTGCACGCCTGTAGTCCCTGCTACTT GGAAGGCTGAGACAGGAAAATTGCTTGAACCCAGGAGGCTGAGGTTGCAGTGAGCCAGGATTGCGCCACCACACTCCAGCCTGGGCAACA GAGACTCTGTCTCAAAAAAAAAAAAAAAAAAAATTAACCTCAACCAAACGCCTATTTTTTAATGCTTAGTTTTGGCTTGAAATTCTTCTT CACCTGGAGTTTTCTTACGTTAATACATTAAAATAACCTGAAGGAAACTTTCGTTATGGGCCAATATTAGCTCTTATGGAAACTGATTTA TATCTTTTTTATGACCTTTCAAAAGTAAAATACTATGCATAATCTAGAAAGATTTGTCAGATAGGAAATTTTATAATGCATTAGCCATTA GTCAGAGTTGTTTTTTAACATGCCAGAGAAAAAGTTGTAATGCCTTGGAAGTTATTCTCTTTTCTATAGATTGTGTTCAAAAGATGTGTA TACTTGCAATAAGGTTTATGTTAAGGTGGCTTTAACAATTGGTTGCTATTTTCCTTCTTTAGTCAATATGACTTTGATGTCATTAACTGC >34626_34626_5_GRB10-SCIN_GRB10_chr7_50737419_ENST00000439599_SCIN_chr7_12662426_ENST00000297029_length(amino acids)=655AA_BP=161 MQAAGPLFRSKDKVEQTPRSQQDPAGPGLPAQSDRLANHQEDDVDLEALVNDMNASLESLYSACSMQSDTVPLLQNGQHARSQPRASGPP RSIQPQVSPRQRVQRSQPVHILAVRRLQEEDQQFRTSSLPAIPNPFPELCGPGSPPVLTPGSLPPSQAAAKQVLGEKPELPDGGDDDDII ADISNRKMAKLYMVSDASGSMRVTVVAEENPFSMAMLLSEECFILDHGAAKQIFVWKGKDANPQERKAAMKTAEEFLQQMNYSKNTQIQV LPEGGETPIFKQFFKDWRDKDQSDGFGKVYVTEKVAQIKQIPFDASKLHSSPQMAAQHNMVDDGSGKVEIWRVENNGRIQVDQNSYGEFY GGDCYIILYTYPRGQIIYTWQGANATRDELTTSAFLTVQLDRSLGGQAVQIRVSQGKEPVHLLSLFKDKPLIIYKNGTSKKGGQAPAPPT RLFQVRRNLASITRIVEVDVDANSLNSNDVFVLKLPQNSGYIWVGKGASQEEEKGAEYVASVLKCKTLRIQEGEEPEEFWNSLGGKKDYQ TSPLLETQAEDHPPRLYGCSNKTGRFVIEEIPGEFTQDDLAEDDVMLLDAWEQIFIWIGKDANEVEKKESLKSAKMYLETDPSGRDKRTP -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for GRB10-SCIN |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for GRB10-SCIN |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for GRB10-SCIN |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
