
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:HDAC7-CHUK (FusionGDB2 ID:35857) |
Fusion Gene Summary for HDAC7-CHUK |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: HDAC7-CHUK | Fusion gene ID: 35857 | Hgene | Tgene | Gene symbol | HDAC7 | CHUK | Gene ID | 51564 | 1147 |
| Gene name | histone deacetylase 7 | component of inhibitor of nuclear factor kappa B kinase complex | |
| Synonyms | HD7|HD7A|HDAC7A | IKBKA|IKK-alpha|IKK1|IKKA|NFKBIKA|TCF16 | |
| Cytomap | 12q13.11 | 10q24.31 | |
| Type of gene | protein-coding | protein-coding | |
| Description | histone deacetylase 7histone deacetylase 7A | inhibitor of nuclear factor kappa-B kinase subunit alphaI-kappa-B kinase 1I-kappa-B kinase-alphaIKK-a kinaseIkB kinase alpha subunitNuclear factor NFkappaB inhibitor kinase alphaTCF-16conserved helix-loop-helix ubiquitous kinasetranscription facto | |
| Modification date | 20200327 | 20200327 | |
| UniProtAcc | Q8WUI4 | O15111 | |
| Ensembl transtripts involved in fusion gene | ENST00000080059, ENST00000354334, ENST00000552960, ENST00000380610, ENST00000427332, ENST00000488927, | ENST00000590930, ENST00000370397, | |
| Fusion gene scores | * DoF score | 11 X 8 X 6=528 | 1 X 1 X 1=1 |
| # samples | 15 | 1 | |
| ** MAII score | log2(15/528*10)=-1.81557542886257 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(1/1*10)=3.32192809488736 | |
| Context | PubMed: HDAC7 [Title/Abstract] AND CHUK [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | HDAC7(48213550)-CHUK(101969547), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | HDAC7 | GO:0032703 | negative regulation of interleukin-2 production | 17360565 |
| Tgene | CHUK | GO:0006468 | protein phosphorylation | 20434986 |
| Tgene | CHUK | GO:0045944 | positive regulation of transcription by RNA polymerase II | 23091055 |
| Tgene | CHUK | GO:0071356 | cellular response to tumor necrosis factor | 23091055 |
 Fusion gene breakpoints across HDAC7 (5'-gene) Fusion gene breakpoints across HDAC7 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
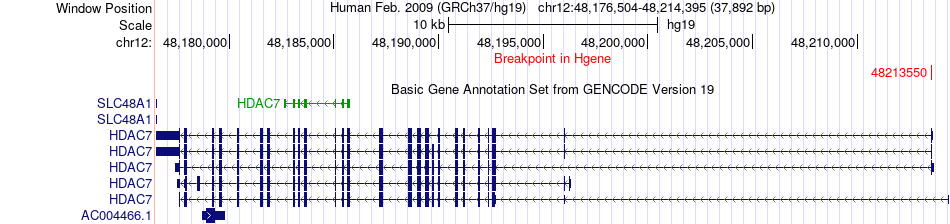 |
 Fusion gene breakpoints across CHUK (3'-gene) Fusion gene breakpoints across CHUK (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
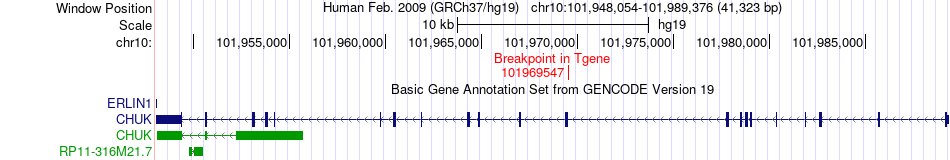 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChiTaRS5.0 | N/A | DA482203 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
Top |
Fusion Gene ORF analysis for HDAC7-CHUK |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000080059 | ENST00000590930 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| 5CDS-intron | ENST00000354334 | ENST00000590930 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| 5CDS-intron | ENST00000552960 | ENST00000590930 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| In-frame | ENST00000080059 | ENST00000370397 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| In-frame | ENST00000354334 | ENST00000370397 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| In-frame | ENST00000552960 | ENST00000370397 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| intron-3CDS | ENST00000380610 | ENST00000370397 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| intron-3CDS | ENST00000427332 | ENST00000370397 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| intron-3CDS | ENST00000488927 | ENST00000370397 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| intron-intron | ENST00000380610 | ENST00000590930 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| intron-intron | ENST00000427332 | ENST00000590930 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
| intron-intron | ENST00000488927 | ENST00000590930 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000080059 | HDAC7 | chr12 | 48213550 | - | ENST00000370397 | CHUK | chr10 | 101969547 | - | 2624 | 19 | 16 | 1323 | 435 |
| ENST00000354334 | HDAC7 | chr12 | 48213550 | - | ENST00000370397 | CHUK | chr10 | 101969547 | - | 2705 | 100 | 61 | 1404 | 447 |
| ENST00000552960 | HDAC7 | chr12 | 48213550 | - | ENST00000370397 | CHUK | chr10 | 101969547 | - | 2725 | 120 | 81 | 1424 | 447 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000080059 | ENST00000370397 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - | 0.000208736 | 0.9997913 |
| ENST00000354334 | ENST00000370397 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - | 0.000199119 | 0.99980086 |
| ENST00000552960 | ENST00000370397 | HDAC7 | chr12 | 48213550 | - | CHUK | chr10 | 101969547 | - | 0.000204617 | 0.9997954 |
Top |
Fusion Genomic Features for HDAC7-CHUK |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
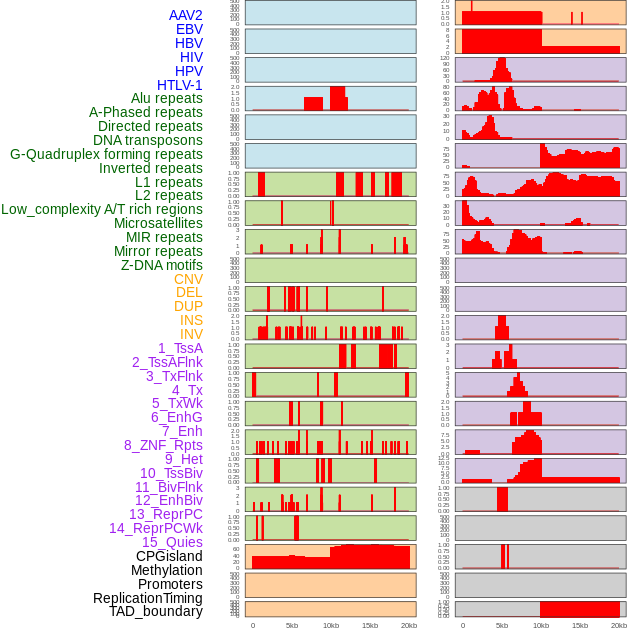 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for HDAC7-CHUK |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:48213550/chr10:101969547) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| HDAC7 | CHUK |
| FUNCTION: Responsible for the deacetylation of lysine residues on the N-terminal part of the core histones (H2A, H2B, H3 and H4). Histone deacetylation gives a tag for epigenetic repression and plays an important role in transcriptional regulation, cell cycle progression and developmental events. Histone deacetylases act via the formation of large multiprotein complexes. Involved in muscle maturation by repressing transcription of myocyte enhancer factors such as MEF2A, MEF2B and MEF2C. During muscle differentiation, it shuttles into the cytoplasm, allowing the expression of myocyte enhancer factors (By similarity). May be involved in Epstein-Barr virus (EBV) latency, possibly by repressing the viral BZLF1 gene. Positively regulates the transcriptional repressor activity of FOXP3 (PubMed:17360565). Serves as a corepressor of RARA, causing its deacetylation and inhibition of RARE DNA element binding (PubMed:28167758). In association with RARA, plays a role in the repression of microRNA-10a and thereby in the inflammatory response (PubMed:28167758). {ECO:0000250|UniProtKB:Q8C2B3, ECO:0000269|PubMed:12239305, ECO:0000269|PubMed:17360565, ECO:0000269|PubMed:28167758}. | FUNCTION: Serine kinase that plays an essential role in the NF-kappa-B signaling pathway which is activated by multiple stimuli such as inflammatory cytokines, bacterial or viral products, DNA damages or other cellular stresses. Acts as part of the canonical IKK complex in the conventional pathway of NF-kappa-B activation and phosphorylates inhibitors of NF-kappa-B on serine residues. These modifications allow polyubiquitination of the inhibitors and subsequent degradation by the proteasome. In turn, free NF-kappa-B is translocated into the nucleus and activates the transcription of hundreds of genes involved in immune response, growth control, or protection against apoptosis. Negatively regulates the pathway by phosphorylating the scaffold protein TAXBP1 and thus promoting the assembly of the A20/TNFAIP3 ubiquitin-editing complex (composed of A20/TNFAIP3, TAX1BP1, and the E3 ligases ITCH and RNF11). Therefore, CHUK plays a key role in the negative feedback of NF-kappa-B canonical signaling to limit inflammatory gene activation. As part of the non-canonical pathway of NF-kappa-B activation, the MAP3K14-activated CHUK/IKKA homodimer phosphorylates NFKB2/p100 associated with RelB, inducing its proteolytic processing to NFKB2/p52 and the formation of NF-kappa-B RelB-p52 complexes. In turn, these complexes regulate genes encoding molecules involved in B-cell survival and lymphoid organogenesis. Participates also in the negative feedback of the non-canonical NF-kappa-B signaling pathway by phosphorylating and destabilizing MAP3K14/NIK. Within the nucleus, phosphorylates CREBBP and consequently increases both its transcriptional and histone acetyltransferase activities. Modulates chromatin accessibility at NF-kappa-B-responsive promoters by phosphorylating histones H3 at 'Ser-10' that are subsequently acetylated at 'Lys-14' by CREBBP. Additionally, phosphorylates the CREBBP-interacting protein NCOA3. Also phosphorylates FOXO3 and may regulate this pro-apoptotic transcription factor (PubMed:12789342, PubMed:15084260, PubMed:17434128, PubMed:20434986, PubMed:20501937, PubMed:21765415). Phosphorylates RIPK1 at 'Ser-25' which represses its kinase activity and consequently prevents TNF-mediated RIPK1-dependent cell death (By similarity). {ECO:0000250|UniProtKB:Q60680, ECO:0000269|PubMed:12789342, ECO:0000269|PubMed:15084260, ECO:0000269|PubMed:17434128, ECO:0000269|PubMed:20434986, ECO:0000269|PubMed:20501937, ECO:0000269|PubMed:21765415}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | CHUK | chr12:48213550 | chr10:101969547 | ENST00000370397 | 8 | 21 | 455_476 | 311 | 746.0 | Region | Note=Leucine-zipper | |
| Tgene | CHUK | chr12:48213550 | chr10:101969547 | ENST00000370397 | 8 | 21 | 738_743 | 311 | 746.0 | Region | Note=NEMO-binding |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000080059 | - | 1 | 26 | 197_203 | 6 | 992.0 | Compositional bias | Note=Poly-Ser |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000080059 | - | 1 | 26 | 368_373 | 6 | 992.0 | Compositional bias | Note=Poly-Pro |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000354334 | - | 1 | 25 | 197_203 | 6 | 955.0 | Compositional bias | Note=Poly-Ser |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000354334 | - | 1 | 25 | 368_373 | 6 | 955.0 | Compositional bias | Note=Poly-Pro |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000427332 | - | 1 | 26 | 197_203 | 0 | 953.0 | Compositional bias | Note=Poly-Ser |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000427332 | - | 1 | 26 | 368_373 | 0 | 953.0 | Compositional bias | Note=Poly-Pro |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000552960 | - | 1 | 25 | 197_203 | 6 | 975.0 | Compositional bias | Note=Poly-Ser |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000552960 | - | 1 | 25 | 368_373 | 6 | 975.0 | Compositional bias | Note=Poly-Pro |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000080059 | - | 1 | 26 | 918_952 | 6 | 992.0 | Motif | Nuclear export signal |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000354334 | - | 1 | 25 | 918_952 | 6 | 955.0 | Motif | Nuclear export signal |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000427332 | - | 1 | 26 | 918_952 | 0 | 953.0 | Motif | Nuclear export signal |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000552960 | - | 1 | 25 | 918_952 | 6 | 975.0 | Motif | Nuclear export signal |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000080059 | - | 1 | 26 | 1_268 | 6 | 992.0 | Region | Transcription repression 1 |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000080059 | - | 1 | 26 | 218_546 | 6 | 992.0 | Region | Transcription repression 2 |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000080059 | - | 1 | 26 | 518_865 | 6 | 992.0 | Region | Note=Histone deacetylase |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000354334 | - | 1 | 25 | 1_268 | 6 | 955.0 | Region | Transcription repression 1 |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000354334 | - | 1 | 25 | 218_546 | 6 | 955.0 | Region | Transcription repression 2 |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000354334 | - | 1 | 25 | 518_865 | 6 | 955.0 | Region | Note=Histone deacetylase |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000427332 | - | 1 | 26 | 1_268 | 0 | 953.0 | Region | Transcription repression 1 |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000427332 | - | 1 | 26 | 218_546 | 0 | 953.0 | Region | Transcription repression 2 |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000427332 | - | 1 | 26 | 518_865 | 0 | 953.0 | Region | Note=Histone deacetylase |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000552960 | - | 1 | 25 | 1_268 | 6 | 975.0 | Region | Transcription repression 1 |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000552960 | - | 1 | 25 | 218_546 | 6 | 975.0 | Region | Transcription repression 2 |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000552960 | - | 1 | 25 | 518_865 | 6 | 975.0 | Region | Note=Histone deacetylase |
| Tgene | CHUK | chr12:48213550 | chr10:101969547 | ENST00000370397 | 8 | 21 | 15_302 | 311 | 746.0 | Domain | Protein kinase | |
| Tgene | CHUK | chr12:48213550 | chr10:101969547 | ENST00000370397 | 8 | 21 | 21_29 | 311 | 746.0 | Nucleotide binding | ATP |
Top |
Fusion Gene Sequence for HDAC7-CHUK |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >35857_35857_1_HDAC7-CHUK_HDAC7_chr12_48213550_ENST00000080059_CHUK_chr10_101969547_ENST00000370397_length(transcript)=2624nt_BP=19nt ATGCACAGCCCCGGCGCTGATAGTACACATCCTAAATATGACTTCTGCAAAGATAATTTCTTTTCTGTTACCACCTGATGAAAGTCTTCA TTCACTACAGTCTCGTATTGAGCGTGAAACTGGAATAAATACTGGTTCTCAAGAACTTCTTTCAGAGACAGGAATTTCTCTGGATCCTCG GAAACCAGCCTCTCAATGTGTTCTAGATGGAGTTAGAGGCTGTGATAGCTATATGGTTTATTTGTTTGATAAAAGTAAAACTGTATATGA AGGGCCATTTGCTTCCAGAAGTTTATCTGATTGTGTAAATTATATTGTACAGGACAGCAAAATACAGCTTCCAATTATACAGCTGCGTAA AGTGTGGGCTGAAGCAGTGCACTATGTGTCTGGACTAAAAGAAGACTATAGCAGGCTCTTTCAGGGACAAAGGGCAGCAATGTTAAGTCT TCTTAGATATAATGCTAACTTAACAAAAATGAAGAACACTTTGATCTCAGCATCACAACAACTGAAAGCTAAATTGGAGTTTTTTCACAA AAGCATTCAGCTTGACTTGGAGAGATACAGCGAGCAGATGACGTATGGGATATCTTCAGAAAAAATGCTAAAAGCATGGAAAGAAATGGA AGAAAAGGCCATCCACTATGCTGAGGTTGGTGTCATTGGATACCTGGAGGATCAGATTATGTCTTTGCATGCTGAAATCATGGAGCTACA GAAGAGCCCCTATGGAAGACGTCAGGGAGACTTGATGGAATCTCTGGAACAGCGTGCCATTGATCTATATAAGCAGTTAAAACACAGACC TTCAGATCACTCCTACAGTGACAGCACAGAGATGGTGAAAATCATTGTGCACACTGTGCAGAGTCAGGACCGTGTGCTCAAGGAGCTGTT TGGTCATTTGAGCAAGTTGTTGGGCTGTAAGCAGAAGATTATTGATCTACTCCCTAAGGTGGAAGTGGCCCTCAGTAATATCAAAGAAGC TGACAATACTGTCATGTTCATGCAGGGAAAAAGGCAGAAAGAAATATGGCATCTCCTTAAAATTGCCTGTACACAGAGTTCTGCCCGGTC CCTTGTAGGATCCAGTCTAGAAGGTGCAGTAACCCCTCAGACATCAGCATGGCTGCCCCCGACTTCAGCAGAACATGATCATTCTCTGTC ATGTGTGGTAACTCCTCAAGATGGGGAGACTTCAGCACAAATGATAGAAGAAAATTTGAACTGCCTTGGCCATTTAAGCACTATTATTCA TGAGGCAAATGAGGAACAGGGCAATAGTATGATGAATCTTGATTGGAGTTGGTTAACAGAATGAGTTGTCACTTGTTCACTGTCCCCAAA CCTATGGAAGTTGTTGCTATACATGTTGGAAATGTGTTTTTCCCCCATGAAACCATTCTTCAGACATCAGTCAATGGAAGAAATGGCTAT GAACAGAAACTACATTTCTACTATGATCAGAAGAACATGATTTTACAAGTATAACAGTTTTGAGTAATTCAAGCCTCTAAACAGACAGGA ATTTAGAAAAAGTCAATGTACTTGTTTGAATATTTGTTTTAATACCACAGCTATTTAGAAGCATCATCACGACACATTTGCCTTCAGTCT TGGTAAAACATTACTTATTTAACTGATTAAAAATACCTTCTATGTATTAGTGTCAACTTTTAACTTTTGGGCGTAAGACCAAATGTAGTT TTGTATACAGAGAAGAAAACCTCAAGTAATAGGCATTTTAAGTAAAAGTCTACCTGTGTTTTTTTCTAAAAAGGCTGCTCACAAGTTCTA TTTCTTGAAGAATAAATTCTACCTCCTTGTGTTGCACTGAACAGGTTCTCTTCCTGGCATCATAAGGAGTTGGTGTAATCATTTTAAATT CCACTGAAAATTTAACAGTATCCCCTTCTCATCGAAGGGATTGTGTATCTGTGCTTCTAATATTAGTTGGCTTTCATAAATCATGTTGTT GTGTGTATATGTATTTAAGATGTACATTTAATAATATCAAAGAGAAGATGCCTGTTAATTTATAATGTATTTGAAAATTACATGTTTTTT CATTTGTAAAAATGAGTCATTTGTTTAAACAATCTTTCATGTCTTGTCATACAAATTTATAAAGGTCTGCACTCCTTTATCTGTAATTGT AATTCCAAAATCCAAAAAGCTCTGAAAACAAGGTTTCCATAAGCTTGGTGACAAAATTCATTTGCTTGCAATCTAATCTGAACTGACCTT GAATCTTTTTATCCCATTTAGTGTGAATATTCCTTTATTTTGCTGCTTGATGATGAGAGGGAGGGCTGCTGCCACAGACTGTGGTGAGGG CTGGTTAATGTAGTATGGTATATGCACAAAACTACTTTTCTAAAATCTAAAATTTCATAATTCTGAAACAACTTGCCCCAAGGGTTTCAG AGAAAGGACTGTGGACCTCTATCATCTGCTAAGTAATTTAGAAGATATTATTTGTCTTAAAAAATGTGAAATGCTTTTATATTCTAATAG TTTTTCACTTTGTGTATTAAATGGTTTTTAAATTACTTTCTTGATCTCTATTCATTATAAAAATCAGATTATAATAAAACAGTTGAATAT >35857_35857_1_HDAC7-CHUK_HDAC7_chr12_48213550_ENST00000080059_CHUK_chr10_101969547_ENST00000370397_length(amino acids)=435AA_BP=1 MIVHILNMTSAKIISFLLPPDESLHSLQSRIERETGINTGSQELLSETGISLDPRKPASQCVLDGVRGCDSYMVYLFDKSKTVYEGPFAS RSLSDCVNYIVQDSKIQLPIIQLRKVWAEAVHYVSGLKEDYSRLFQGQRAAMLSLLRYNANLTKMKNTLISASQQLKAKLEFFHKSIQLD LERYSEQMTYGISSEKMLKAWKEMEEKAIHYAEVGVIGYLEDQIMSLHAEIMELQKSPYGRRQGDLMESLEQRAIDLYKQLKHRPSDHSY SDSTEMVKIIVHTVQSQDRVLKELFGHLSKLLGCKQKIIDLLPKVEVALSNIKEADNTVMFMQGKRQKEIWHLLKIACTQSSARSLVGSS -------------------------------------------------------------- >35857_35857_2_HDAC7-CHUK_HDAC7_chr12_48213550_ENST00000354334_CHUK_chr10_101969547_ENST00000370397_length(transcript)=2705nt_BP=100nt GGCGACTGGGGGGATGTGAGGCCGGCGCCCCAGCCCCCCGCCCCGCCATGAGCCCCCCGCTCTGAGGGCCCCGGCCCCTGGATGCACAGC CCCGGCGCTGATAGTACACATCCTAAATATGACTTCTGCAAAGATAATTTCTTTTCTGTTACCACCTGATGAAAGTCTTCATTCACTACA GTCTCGTATTGAGCGTGAAACTGGAATAAATACTGGTTCTCAAGAACTTCTTTCAGAGACAGGAATTTCTCTGGATCCTCGGAAACCAGC CTCTCAATGTGTTCTAGATGGAGTTAGAGGCTGTGATAGCTATATGGTTTATTTGTTTGATAAAAGTAAAACTGTATATGAAGGGCCATT TGCTTCCAGAAGTTTATCTGATTGTGTAAATTATATTGTACAGGACAGCAAAATACAGCTTCCAATTATACAGCTGCGTAAAGTGTGGGC TGAAGCAGTGCACTATGTGTCTGGACTAAAAGAAGACTATAGCAGGCTCTTTCAGGGACAAAGGGCAGCAATGTTAAGTCTTCTTAGATA TAATGCTAACTTAACAAAAATGAAGAACACTTTGATCTCAGCATCACAACAACTGAAAGCTAAATTGGAGTTTTTTCACAAAAGCATTCA GCTTGACTTGGAGAGATACAGCGAGCAGATGACGTATGGGATATCTTCAGAAAAAATGCTAAAAGCATGGAAAGAAATGGAAGAAAAGGC CATCCACTATGCTGAGGTTGGTGTCATTGGATACCTGGAGGATCAGATTATGTCTTTGCATGCTGAAATCATGGAGCTACAGAAGAGCCC CTATGGAAGACGTCAGGGAGACTTGATGGAATCTCTGGAACAGCGTGCCATTGATCTATATAAGCAGTTAAAACACAGACCTTCAGATCA CTCCTACAGTGACAGCACAGAGATGGTGAAAATCATTGTGCACACTGTGCAGAGTCAGGACCGTGTGCTCAAGGAGCTGTTTGGTCATTT GAGCAAGTTGTTGGGCTGTAAGCAGAAGATTATTGATCTACTCCCTAAGGTGGAAGTGGCCCTCAGTAATATCAAAGAAGCTGACAATAC TGTCATGTTCATGCAGGGAAAAAGGCAGAAAGAAATATGGCATCTCCTTAAAATTGCCTGTACACAGAGTTCTGCCCGGTCCCTTGTAGG ATCCAGTCTAGAAGGTGCAGTAACCCCTCAGACATCAGCATGGCTGCCCCCGACTTCAGCAGAACATGATCATTCTCTGTCATGTGTGGT AACTCCTCAAGATGGGGAGACTTCAGCACAAATGATAGAAGAAAATTTGAACTGCCTTGGCCATTTAAGCACTATTATTCATGAGGCAAA TGAGGAACAGGGCAATAGTATGATGAATCTTGATTGGAGTTGGTTAACAGAATGAGTTGTCACTTGTTCACTGTCCCCAAACCTATGGAA GTTGTTGCTATACATGTTGGAAATGTGTTTTTCCCCCATGAAACCATTCTTCAGACATCAGTCAATGGAAGAAATGGCTATGAACAGAAA CTACATTTCTACTATGATCAGAAGAACATGATTTTACAAGTATAACAGTTTTGAGTAATTCAAGCCTCTAAACAGACAGGAATTTAGAAA AAGTCAATGTACTTGTTTGAATATTTGTTTTAATACCACAGCTATTTAGAAGCATCATCACGACACATTTGCCTTCAGTCTTGGTAAAAC ATTACTTATTTAACTGATTAAAAATACCTTCTATGTATTAGTGTCAACTTTTAACTTTTGGGCGTAAGACCAAATGTAGTTTTGTATACA GAGAAGAAAACCTCAAGTAATAGGCATTTTAAGTAAAAGTCTACCTGTGTTTTTTTCTAAAAAGGCTGCTCACAAGTTCTATTTCTTGAA GAATAAATTCTACCTCCTTGTGTTGCACTGAACAGGTTCTCTTCCTGGCATCATAAGGAGTTGGTGTAATCATTTTAAATTCCACTGAAA ATTTAACAGTATCCCCTTCTCATCGAAGGGATTGTGTATCTGTGCTTCTAATATTAGTTGGCTTTCATAAATCATGTTGTTGTGTGTATA TGTATTTAAGATGTACATTTAATAATATCAAAGAGAAGATGCCTGTTAATTTATAATGTATTTGAAAATTACATGTTTTTTCATTTGTAA AAATGAGTCATTTGTTTAAACAATCTTTCATGTCTTGTCATACAAATTTATAAAGGTCTGCACTCCTTTATCTGTAATTGTAATTCCAAA ATCCAAAAAGCTCTGAAAACAAGGTTTCCATAAGCTTGGTGACAAAATTCATTTGCTTGCAATCTAATCTGAACTGACCTTGAATCTTTT TATCCCATTTAGTGTGAATATTCCTTTATTTTGCTGCTTGATGATGAGAGGGAGGGCTGCTGCCACAGACTGTGGTGAGGGCTGGTTAAT GTAGTATGGTATATGCACAAAACTACTTTTCTAAAATCTAAAATTTCATAATTCTGAAACAACTTGCCCCAAGGGTTTCAGAGAAAGGAC TGTGGACCTCTATCATCTGCTAAGTAATTTAGAAGATATTATTTGTCTTAAAAAATGTGAAATGCTTTTATATTCTAATAGTTTTTCACT TTGTGTATTAAATGGTTTTTAAATTACTTTCTTGATCTCTATTCATTATAAAAATCAGATTATAATAAAACAGTTGAATATGGCTTAGGA >35857_35857_2_HDAC7-CHUK_HDAC7_chr12_48213550_ENST00000354334_CHUK_chr10_101969547_ENST00000370397_length(amino acids)=447AA_BP=13 MRAPAPGCTAPALIVHILNMTSAKIISFLLPPDESLHSLQSRIERETGINTGSQELLSETGISLDPRKPASQCVLDGVRGCDSYMVYLFD KSKTVYEGPFASRSLSDCVNYIVQDSKIQLPIIQLRKVWAEAVHYVSGLKEDYSRLFQGQRAAMLSLLRYNANLTKMKNTLISASQQLKA KLEFFHKSIQLDLERYSEQMTYGISSEKMLKAWKEMEEKAIHYAEVGVIGYLEDQIMSLHAEIMELQKSPYGRRQGDLMESLEQRAIDLY KQLKHRPSDHSYSDSTEMVKIIVHTVQSQDRVLKELFGHLSKLLGCKQKIIDLLPKVEVALSNIKEADNTVMFMQGKRQKEIWHLLKIAC -------------------------------------------------------------- >35857_35857_3_HDAC7-CHUK_HDAC7_chr12_48213550_ENST00000552960_CHUK_chr10_101969547_ENST00000370397_length(transcript)=2725nt_BP=120nt ACAGTCGCTCTGCAGCCTCCGGCGACTGGGGGGATGTGAGGCCGGCGCCCCAGCCCCCCGCCCCGCCATGAGCCCCCCGCTCTGAGGGCC CCGGCCCCTGGATGCACAGCCCCGGCGCTGATAGTACACATCCTAAATATGACTTCTGCAAAGATAATTTCTTTTCTGTTACCACCTGAT GAAAGTCTTCATTCACTACAGTCTCGTATTGAGCGTGAAACTGGAATAAATACTGGTTCTCAAGAACTTCTTTCAGAGACAGGAATTTCT CTGGATCCTCGGAAACCAGCCTCTCAATGTGTTCTAGATGGAGTTAGAGGCTGTGATAGCTATATGGTTTATTTGTTTGATAAAAGTAAA ACTGTATATGAAGGGCCATTTGCTTCCAGAAGTTTATCTGATTGTGTAAATTATATTGTACAGGACAGCAAAATACAGCTTCCAATTATA CAGCTGCGTAAAGTGTGGGCTGAAGCAGTGCACTATGTGTCTGGACTAAAAGAAGACTATAGCAGGCTCTTTCAGGGACAAAGGGCAGCA ATGTTAAGTCTTCTTAGATATAATGCTAACTTAACAAAAATGAAGAACACTTTGATCTCAGCATCACAACAACTGAAAGCTAAATTGGAG TTTTTTCACAAAAGCATTCAGCTTGACTTGGAGAGATACAGCGAGCAGATGACGTATGGGATATCTTCAGAAAAAATGCTAAAAGCATGG AAAGAAATGGAAGAAAAGGCCATCCACTATGCTGAGGTTGGTGTCATTGGATACCTGGAGGATCAGATTATGTCTTTGCATGCTGAAATC ATGGAGCTACAGAAGAGCCCCTATGGAAGACGTCAGGGAGACTTGATGGAATCTCTGGAACAGCGTGCCATTGATCTATATAAGCAGTTA AAACACAGACCTTCAGATCACTCCTACAGTGACAGCACAGAGATGGTGAAAATCATTGTGCACACTGTGCAGAGTCAGGACCGTGTGCTC AAGGAGCTGTTTGGTCATTTGAGCAAGTTGTTGGGCTGTAAGCAGAAGATTATTGATCTACTCCCTAAGGTGGAAGTGGCCCTCAGTAAT ATCAAAGAAGCTGACAATACTGTCATGTTCATGCAGGGAAAAAGGCAGAAAGAAATATGGCATCTCCTTAAAATTGCCTGTACACAGAGT TCTGCCCGGTCCCTTGTAGGATCCAGTCTAGAAGGTGCAGTAACCCCTCAGACATCAGCATGGCTGCCCCCGACTTCAGCAGAACATGAT CATTCTCTGTCATGTGTGGTAACTCCTCAAGATGGGGAGACTTCAGCACAAATGATAGAAGAAAATTTGAACTGCCTTGGCCATTTAAGC ACTATTATTCATGAGGCAAATGAGGAACAGGGCAATAGTATGATGAATCTTGATTGGAGTTGGTTAACAGAATGAGTTGTCACTTGTTCA CTGTCCCCAAACCTATGGAAGTTGTTGCTATACATGTTGGAAATGTGTTTTTCCCCCATGAAACCATTCTTCAGACATCAGTCAATGGAA GAAATGGCTATGAACAGAAACTACATTTCTACTATGATCAGAAGAACATGATTTTACAAGTATAACAGTTTTGAGTAATTCAAGCCTCTA AACAGACAGGAATTTAGAAAAAGTCAATGTACTTGTTTGAATATTTGTTTTAATACCACAGCTATTTAGAAGCATCATCACGACACATTT GCCTTCAGTCTTGGTAAAACATTACTTATTTAACTGATTAAAAATACCTTCTATGTATTAGTGTCAACTTTTAACTTTTGGGCGTAAGAC CAAATGTAGTTTTGTATACAGAGAAGAAAACCTCAAGTAATAGGCATTTTAAGTAAAAGTCTACCTGTGTTTTTTTCTAAAAAGGCTGCT CACAAGTTCTATTTCTTGAAGAATAAATTCTACCTCCTTGTGTTGCACTGAACAGGTTCTCTTCCTGGCATCATAAGGAGTTGGTGTAAT CATTTTAAATTCCACTGAAAATTTAACAGTATCCCCTTCTCATCGAAGGGATTGTGTATCTGTGCTTCTAATATTAGTTGGCTTTCATAA ATCATGTTGTTGTGTGTATATGTATTTAAGATGTACATTTAATAATATCAAAGAGAAGATGCCTGTTAATTTATAATGTATTTGAAAATT ACATGTTTTTTCATTTGTAAAAATGAGTCATTTGTTTAAACAATCTTTCATGTCTTGTCATACAAATTTATAAAGGTCTGCACTCCTTTA TCTGTAATTGTAATTCCAAAATCCAAAAAGCTCTGAAAACAAGGTTTCCATAAGCTTGGTGACAAAATTCATTTGCTTGCAATCTAATCT GAACTGACCTTGAATCTTTTTATCCCATTTAGTGTGAATATTCCTTTATTTTGCTGCTTGATGATGAGAGGGAGGGCTGCTGCCACAGAC TGTGGTGAGGGCTGGTTAATGTAGTATGGTATATGCACAAAACTACTTTTCTAAAATCTAAAATTTCATAATTCTGAAACAACTTGCCCC AAGGGTTTCAGAGAAAGGACTGTGGACCTCTATCATCTGCTAAGTAATTTAGAAGATATTATTTGTCTTAAAAAATGTGAAATGCTTTTA TATTCTAATAGTTTTTCACTTTGTGTATTAAATGGTTTTTAAATTACTTTCTTGATCTCTATTCATTATAAAAATCAGATTATAATAAAA >35857_35857_3_HDAC7-CHUK_HDAC7_chr12_48213550_ENST00000552960_CHUK_chr10_101969547_ENST00000370397_length(amino acids)=447AA_BP=13 MRAPAPGCTAPALIVHILNMTSAKIISFLLPPDESLHSLQSRIERETGINTGSQELLSETGISLDPRKPASQCVLDGVRGCDSYMVYLFD KSKTVYEGPFASRSLSDCVNYIVQDSKIQLPIIQLRKVWAEAVHYVSGLKEDYSRLFQGQRAAMLSLLRYNANLTKMKNTLISASQQLKA KLEFFHKSIQLDLERYSEQMTYGISSEKMLKAWKEMEEKAIHYAEVGVIGYLEDQIMSLHAEIMELQKSPYGRRQGDLMESLEQRAIDLY KQLKHRPSDHSYSDSTEMVKIIVHTVQSQDRVLKELFGHLSKLLGCKQKIIDLLPKVEVALSNIKEADNTVMFMQGKRQKEIWHLLKIAC -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for HDAC7-CHUK |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000080059 | - | 1 | 26 | 49_149 | 6.333333333333333 | 992.0 | MEF2A |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000354334 | - | 1 | 25 | 49_149 | 6.333333333333333 | 955.0 | MEF2A |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000427332 | - | 1 | 26 | 49_149 | 0 | 953.0 | MEF2A |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000552960 | - | 1 | 25 | 49_149 | 6.333333333333333 | 975.0 | MEF2A |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000080059 | - | 1 | 26 | 1_98 | 6.333333333333333 | 992.0 | MEF2C |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000354334 | - | 1 | 25 | 1_98 | 6.333333333333333 | 955.0 | MEF2C |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000427332 | - | 1 | 26 | 1_98 | 0 | 953.0 | MEF2C |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000552960 | - | 1 | 25 | 1_98 | 6.333333333333333 | 975.0 | MEF2C |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000080059 | - | 1 | 26 | 877_952 | 6.333333333333333 | 992.0 | SIN3A |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000354334 | - | 1 | 25 | 877_952 | 6.333333333333333 | 955.0 | SIN3A |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000427332 | - | 1 | 26 | 877_952 | 0 | 953.0 | SIN3A |
| Hgene | HDAC7 | chr12:48213550 | chr10:101969547 | ENST00000552960 | - | 1 | 25 | 877_952 | 6.333333333333333 | 975.0 | SIN3A |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for HDAC7-CHUK |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for HDAC7-CHUK |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
