
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:HIPK2-BRAF (FusionGDB2 ID:36401) |
Fusion Gene Summary for HIPK2-BRAF |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: HIPK2-BRAF | Fusion gene ID: 36401 | Hgene | Tgene | Gene symbol | HIPK2 | BRAF | Gene ID | 28996 | 673 |
| Gene name | homeodomain interacting protein kinase 2 | B-Raf proto-oncogene, serine/threonine kinase | |
| Synonyms | PRO0593 | B-RAF1|B-raf|BRAF1|NS7|RAFB1 | |
| Cytomap | 7q34 | 7q34 | |
| Type of gene | protein-coding | protein-coding | |
| Description | homeodomain-interacting protein kinase 2hHIPk2 | serine/threonine-protein kinase B-raf94 kDa B-raf proteinB-Raf proto-oncogene serine/threonine-protein kinase (p94)B-Raf serine/threonine-proteinmurine sarcoma viral (v-raf) oncogene homolog B1proto-oncogene B-Rafv-raf murine sarcoma viral oncogene | |
| Modification date | 20200313 | 20200329 | |
| UniProtAcc | . | P15056 | |
| Ensembl transtripts involved in fusion gene | ENST00000342645, ENST00000406875, ENST00000428878, | ENST00000288602, | |
| Fusion gene scores | * DoF score | 17 X 13 X 12=2652 | 48 X 58 X 16=44544 |
| # samples | 25 | 69 | |
| ** MAII score | log2(25/2652*10)=-3.4070807754505 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(69/44544*10)=-6.0124909441832 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: HIPK2 [Title/Abstract] AND BRAF [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | HIPK2(139297947)-BRAF(140449218), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | HIPK2 | GO:0006468 | protein phosphorylation | 19448668 |
| Hgene | HIPK2 | GO:0006978 | DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator | 14647468 |
| Hgene | HIPK2 | GO:0045766 | positive regulation of angiogenesis | 19046997 |
| Hgene | HIPK2 | GO:0060395 | SMAD protein signal transduction | 12874272 |
| Tgene | BRAF | GO:0000186 | activation of MAPKK activity | 29433126 |
| Tgene | BRAF | GO:0006468 | protein phosphorylation | 17563371 |
| Tgene | BRAF | GO:0010828 | positive regulation of glucose transmembrane transport | 23010278 |
| Tgene | BRAF | GO:0033138 | positive regulation of peptidyl-serine phosphorylation | 19667065 |
| Tgene | BRAF | GO:0043066 | negative regulation of apoptotic process | 19667065 |
| Tgene | BRAF | GO:0070374 | positive regulation of ERK1 and ERK2 cascade | 22065586 |
| Tgene | BRAF | GO:0071277 | cellular response to calcium ion | 18567582 |
| Tgene | BRAF | GO:0090150 | establishment of protein localization to membrane | 23010278 |
 Fusion gene breakpoints across HIPK2 (5'-gene) Fusion gene breakpoints across HIPK2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
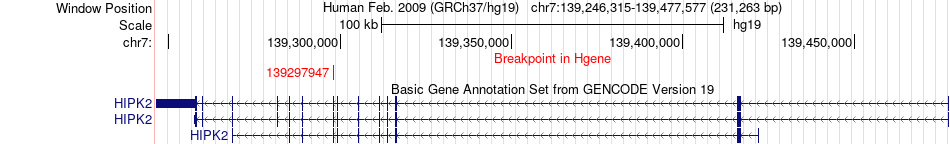 |
 Fusion gene breakpoints across BRAF (3'-gene) Fusion gene breakpoints across BRAF (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
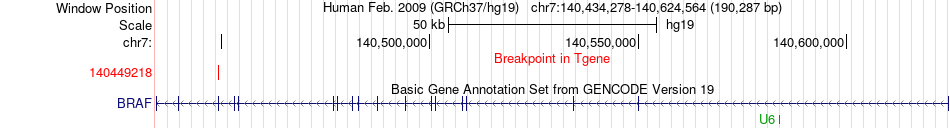 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-VQ-A92D | HIPK2 | chr7 | 139297947 | - | BRAF | chr7 | 140449218 | - |
Top |
Fusion Gene ORF analysis for HIPK2-BRAF |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000342645 | ENST00000288602 | HIPK2 | chr7 | 139297947 | - | BRAF | chr7 | 140449218 | - |
| In-frame | ENST00000406875 | ENST00000288602 | HIPK2 | chr7 | 139297947 | - | BRAF | chr7 | 140449218 | - |
| In-frame | ENST00000428878 | ENST00000288602 | HIPK2 | chr7 | 139297947 | - | BRAF | chr7 | 140449218 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000406875 | HIPK2 | chr7 | 139297947 | - | ENST00000288602 | BRAF | chr7 | 140449218 | - | 2766 | 2207 | 95 | 2647 | 850 |
| ENST00000428878 | HIPK2 | chr7 | 139297947 | - | ENST00000288602 | BRAF | chr7 | 140449218 | - | 2745 | 2186 | 155 | 2626 | 823 |
| ENST00000342645 | HIPK2 | chr7 | 139297947 | - | ENST00000288602 | BRAF | chr7 | 140449218 | - | 2671 | 2112 | 21 | 2552 | 843 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000406875 | ENST00000288602 | HIPK2 | chr7 | 139297947 | - | BRAF | chr7 | 140449218 | - | 0.004015429 | 0.99598455 |
| ENST00000428878 | ENST00000288602 | HIPK2 | chr7 | 139297947 | - | BRAF | chr7 | 140449218 | - | 0.005234031 | 0.994766 |
| ENST00000342645 | ENST00000288602 | HIPK2 | chr7 | 139297947 | - | BRAF | chr7 | 140449218 | - | 0.003625919 | 0.9963741 |
Top |
Fusion Genomic Features for HIPK2-BRAF |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
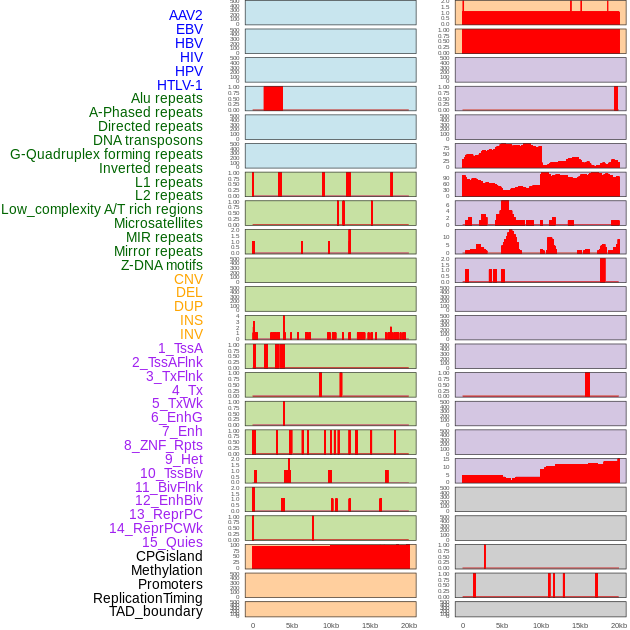 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for HIPK2-BRAF |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr7:139297947/chr7:140449218) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| . | BRAF |
| FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. | FUNCTION: Protein kinase involved in the transduction of mitogenic signals from the cell membrane to the nucleus (Probable). Phosphorylates MAP2K1, and thereby activates the MAP kinase signal transduction pathway (PubMed:21441910, PubMed:29433126). May play a role in the postsynaptic responses of hippocampal neurons (PubMed:1508179). {ECO:0000269|PubMed:1508179, ECO:0000269|PubMed:21441910, ECO:0000269|PubMed:29433126, ECO:0000305}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 199_527 | 704 | 1199.0 | Domain | Protein kinase |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 199_527 | 677 | 1172.0 | Domain | Protein kinase |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 205_213 | 704 | 1199.0 | Nucleotide binding | ATP |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 205_213 | 677 | 1172.0 | Nucleotide binding | ATP |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 97_230 | 704 | 1199.0 | Region | Transcriptional corepression |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 97_230 | 677 | 1172.0 | Region | Transcriptional corepression |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 1088_1094 | 704 | 1199.0 | Compositional bias | Note=Poly-Ala |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 1088_1094 | 677 | 1172.0 | Compositional bias | Note=Poly-Ala |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 802_805 | 704 | 1199.0 | Motif | Note=Nuclear localization signal 1 (NLS1) |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 832_835 | 704 | 1199.0 | Motif | Note=Nuclear localization signal 2 (NLS2) |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 802_805 | 677 | 1172.0 | Motif | Note=Nuclear localization signal 1 (NLS1) |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 832_835 | 677 | 1172.0 | Motif | Note=Nuclear localization signal 2 (NLS2) |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 873_980 | 704 | 1199.0 | Region | Required for localization to nuclear speckles |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 984_1198 | 704 | 1199.0 | Region | Note=Autoinhibitory domain (AID) |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 873_980 | 677 | 1172.0 | Region | Required for localization to nuclear speckles |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 984_1198 | 677 | 1172.0 | Region | Note=Autoinhibitory domain (AID) |
| Tgene | BRAF | chr7:139297947 | chr7:140449218 | ENST00000288602 | 14 | 18 | 122_129 | 620 | 767.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | BRAF | chr7:139297947 | chr7:140449218 | ENST00000288602 | 14 | 18 | 428_432 | 620 | 767.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | BRAF | chr7:139297947 | chr7:140449218 | ENST00000288602 | 14 | 18 | 6_11 | 620 | 767.0 | Compositional bias | Note=Poly-Gly | |
| Tgene | BRAF | chr7:139297947 | chr7:140449218 | ENST00000288602 | 14 | 18 | 155_227 | 620 | 767.0 | Domain | RBD | |
| Tgene | BRAF | chr7:139297947 | chr7:140449218 | ENST00000288602 | 14 | 18 | 457_717 | 620 | 767.0 | Domain | Protein kinase | |
| Tgene | BRAF | chr7:139297947 | chr7:140449218 | ENST00000288602 | 14 | 18 | 463_471 | 620 | 767.0 | Nucleotide binding | ATP | |
| Tgene | BRAF | chr7:139297947 | chr7:140449218 | ENST00000288602 | 14 | 18 | 234_280 | 620 | 767.0 | Zinc finger | Phorbol-ester/DAG-type |
Top |
Fusion Gene Sequence for HIPK2-BRAF |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >36401_36401_1_HIPK2-BRAF_HIPK2_chr7_139297947_ENST00000342645_BRAF_chr7_140449218_ENST00000288602_length(transcript)=2671nt_BP=2112nt CTTCAGCCTGTTTATGAGAGTATGGCCTCACATGTGCAAGTTTTCTCCCCTCACACCCTTCAATCAAGTGCCTTCTGTAGTGTGAAGAAA CTGAAAATAGAGCCGAGTTCCAACTGGGACATGACTGGGTACGGCTCCCACAGCAAAGTGTATAGCCAGAGCAAGAACATCCCCCTGTCG CAGCCAGCCACCACAACCGTCAGCACCTCCTTGCCGGTCCCAAACCCAAGCCTACCTTACGAGCAGACCATCGTCTTCCCAGGAAGCACC GGGCACATCGTGGTCACCTCAGCAAGCAGCACTTCTGTCACCGGGCAAGTCCTCGGCGGACCACACAACCTAATGCGTCGAAGCACTGTG AGCCTCCTTGATACCTACCAAAAATGTGGACTCAAGCGTAAGAGCGAGGAGATCGAGAACACAAGCAGCGTGCAGATCATCGAGGAGCAT CCACCCATGATTCAGAATAATGCAAGCGGGGCCACTGTCGCCACTGCCACCACGTCTACTGCCACCTCCAAAAACAGCGGCTCCAACAGC GAGGGCGACTATCAGCTGGTGCAGCATGAGGTGCTGTGCTCCATGACCAACACCTACGAGGTCTTAGAGTTCTTGGGCCGAGGGACGTTT GGGCAAGTGGTCAAGTGCTGGAAACGGGGCACCAATGAGATCGTAGCCATCAAGATCCTGAAGAACCACCCATCCTATGCCCGACAAGGT CAGATTGAAGTGAGCATCCTGGCCCGGTTGAGCACGGAGAGTGCCGATGACTATAACTTCGTCCGGGCCTACGAATGCTTCCAGCACAAG AACCACACGTGCTTGGTCTTCGAGATGTTGGAGCAGAACCTCTATGACTTTCTGAAGCAAAACAAGTTTAGCCCCTTGCCCCTCAAATAC ATTCGCCCAGTTCTCCAGCAGGTAGCCACAGCCCTGATGAAACTCAAAAGCCTAGGTCTTATCCACGCTGACCTCAAACCAGAAAACATC ATGCTGGTGGATCCATCTAGACAACCATACAGAGTCAAGGTCATCGACTTTGGTTCAGCCAGCCACGTCTCCAAGGCTGTGTGCTCCACC TACTTGCAGTCCAGATATTACAGGGCCCCTGAGATCATCCTTGGTTTACCATTTTGTGAGGCAATTGACATGTGGTCCCTGGGCTGTGTT ATTGCAGAATTGTTCCTGGGTTGGCCGTTATATCCAGGAGCTTCGGAGTATGATCAGATTCGGTATATTTCACAAACACAGGGTTTGCCT GCTGAATATTTATTAAGCGCCGGGACAAAGACAACTAGGTTTTTCAACCGTGACACGGACTCACCATATCCTTTGTGGAGACTGAAGACA CCAGATGACCATGAAGCAGAGACAGGGATTAAGTCAAAAGAAGCAAGAAAGTACATTTTCAACTGTTTAGATGATATGGCCCAGGTGAAC ATGACGACAGATTTGGAAGGGAGCGACATGTTGGTAGAAAAGGCTGACCGGCGGGAGTTCATTGACCTGTTGAAGAAGATGCTGACCATT GATGCTGACAAGAGAATCACTCCAATCGAAACCCTGAACCATCCCTTTGTCACCATGACACACTTACTCGATTTTCCCCACAGCACACAC GTCAAATCATGTTTCCAGAACATGGAGATCTGCAAGCGTCGGGTGAATATGTATGACACGGTGAACCAGAGCAAAACCCCTTTCATCACG CACGTGGCCCCCAGCACGTCCACCAACCTGACCATGACCTTTAACAACCAGCTGACCACTGTCCACAACCAGGCTCCCTCCTCTACCAGT GCCACTATTTCCTTAGCCAATCCCGAAGTCTCCATACTAAACTACCCATCTACACTCTACCAGCCCTCAGCGGCATCCATGGCTGCAGTG GCCCAGCGGAGCATGCCCCTGCAGACAGGAACAGCCCAGATTTGTGCCCGGCCTGACCCGTTCCAGCAAGCTCTCATCGTGTGTCCCCCC GGCTTCCAAGGCTTGCAGGCCTCTCCCTCTAAGCACGCTGGCTACTCGGTGCGAATGGAAAATGCAGTTCCCATCGTCACTCAAGCCCCA GGAGCTCAGCCTCTTCAGATCCAACCAGGTCTGCTTGCCCAGGCACCAGAAGTCATCAGAATGCAAGATAAAAATCCATACAGCTTTCAG TCAGATGTATATGCATTTGGAATTGTTCTGTATGAATTGATGACTGGACAGTTACCTTATTCAAACATCAACAACAGGGACCAGATAATT TTTATGGTGGGACGAGGATACCTGTCTCCAGATCTCAGTAAGGTACGGAGTAACTGTCCAAAAGCCATGAAGAGATTAATGGCAGAGTGC CTCAAAAAGAAAAGAGATGAGAGACCACTCTTTCCCCAAATTCTCGCCTCTATTGAGCTGCTGGCCCGCTCATTGCCAAAAATTCACCGC AGTGCATCAGAACCCTCCTTGAATCGGGCTGGTTTCCAAACAGAGGATTTTAGTCTATATGCTTGTGCTTCTCCAAAAACACCCATCCAG GCAGGGGGATATGGTGCGTTTCCTGTCCACTGAAACAAATGAGTGAGAGAGTTCAGGAGAGTAGCAACAAAAGGAAAATAAATGAACATA >36401_36401_1_HIPK2-BRAF_HIPK2_chr7_139297947_ENST00000342645_BRAF_chr7_140449218_ENST00000288602_length(amino acids)=843AA_BP=691 MASHVQVFSPHTLQSSAFCSVKKLKIEPSSNWDMTGYGSHSKVYSQSKNIPLSQPATTTVSTSLPVPNPSLPYEQTIVFPGSTGHIVVTS ASSTSVTGQVLGGPHNLMRRSTVSLLDTYQKCGLKRKSEEIENTSSVQIIEEHPPMIQNNASGATVATATTSTATSKNSGSNSEGDYQLV QHEVLCSMTNTYEVLEFLGRGTFGQVVKCWKRGTNEIVAIKILKNHPSYARQGQIEVSILARLSTESADDYNFVRAYECFQHKNHTCLVF EMLEQNLYDFLKQNKFSPLPLKYIRPVLQQVATALMKLKSLGLIHADLKPENIMLVDPSRQPYRVKVIDFGSASHVSKAVCSTYLQSRYY RAPEIILGLPFCEAIDMWSLGCVIAELFLGWPLYPGASEYDQIRYISQTQGLPAEYLLSAGTKTTRFFNRDTDSPYPLWRLKTPDDHEAE TGIKSKEARKYIFNCLDDMAQVNMTTDLEGSDMLVEKADRREFIDLLKKMLTIDADKRITPIETLNHPFVTMTHLLDFPHSTHVKSCFQN MEICKRRVNMYDTVNQSKTPFITHVAPSTSTNLTMTFNNQLTTVHNQAPSSTSATISLANPEVSILNYPSTLYQPSAASMAAVAQRSMPL QTGTAQICARPDPFQQALIVCPPGFQGLQASPSKHAGYSVRMENAVPIVTQAPGAQPLQIQPGLLAQAPEVIRMQDKNPYSFQSDVYAFG IVLYELMTGQLPYSNINNRDQIIFMVGRGYLSPDLSKVRSNCPKAMKRLMAECLKKKRDERPLFPQILASIELLARSLPKIHRSASEPSL -------------------------------------------------------------- >36401_36401_2_HIPK2-BRAF_HIPK2_chr7_139297947_ENST00000406875_BRAF_chr7_140449218_ENST00000288602_length(transcript)=2766nt_BP=2207nt CTCAAGATGGCAGATTCCGACTGAGGCTGGGGGGGCCGAGCTCGCGCGCCGCTTTCCCGTCCCCGTTGCCATGAACCGCGGACACCCCGG CCCCGATGGCCCCCGTGTACGAAGGTATGGCCTCACATGTGCAAGTTTTCTCCCCTCACACCCTTCAATCAAGTGCCTTCTGTAGTGTGA AGAAACTGAAAATAGAGCCGAGTTCCAACTGGGACATGACTGGGTACGGCTCCCACAGCAAAGTGTATAGCCAGAGCAAGAACATCCCCC TGTCGCAGCCAGCCACCACAACCGTCAGCACCTCCTTGCCGGTCCCAAACCCAAGCCTACCTTACGAGCAGACCATCGTCTTCCCAGGAA GCACCGGGCACATCGTGGTCACCTCAGCAAGCAGCACTTCTGTCACCGGGCAAGTCCTCGGCGGACCACACAACCTAATGCGTCGAAGCA CTGTGAGCCTCCTTGATACCTACCAAAAATGTGGACTCAAGCGTAAGAGCGAGGAGATCGAGAACACAAGCAGCGTGCAGATCATCGAGG AGCATCCACCCATGATTCAGAATAATGCAAGCGGGGCCACTGTCGCCACTGCCACCACGTCTACTGCCACCTCCAAAAACAGCGGCTCCA ACAGCGAGGGCGACTATCAGCTGGTGCAGCATGAGGTGCTGTGCTCCATGACCAACACCTACGAGGTCTTAGAGTTCTTGGGCCGAGGGA CGTTTGGGCAAGTGGTCAAGTGCTGGAAACGGGGCACCAATGAGATCGTAGCCATCAAGATCCTGAAGAACCACCCATCCTATGCCCGAC AAGGTCAGATTGAAGTGAGCATCCTGGCCCGGTTGAGCACGGAGAGTGCCGATGACTATAACTTCGTCCGGGCCTACGAATGCTTCCAGC ACAAGAACCACACGTGCTTGGTCTTCGAGATGTTGGAGCAGAACCTCTATGACTTTCTGAAGCAAAACAAGTTTAGCCCCTTGCCCCTCA AATACATTCGCCCAGTTCTCCAGCAGGTAGCCACAGCCCTGATGAAACTCAAAAGCCTAGGTCTTATCCACGCTGACCTCAAACCAGAAA ACATCATGCTGGTGGATCCATCTAGACAACCATACAGAGTCAAGGTCATCGACTTTGGTTCAGCCAGCCACGTCTCCAAGGCTGTGTGCT CCACCTACTTGCAGTCCAGATATTACAGGGCCCCTGAGATCATCCTTGGTTTACCATTTTGTGAGGCAATTGACATGTGGTCCCTGGGCT GTGTTATTGCAGAATTGTTCCTGGGTTGGCCGTTATATCCAGGAGCTTCGGAGTATGATCAGATTCGGTATATTTCACAAACACAGGGTT TGCCTGCTGAATATTTATTAAGCGCCGGGACAAAGACAACTAGGTTTTTCAACCGTGACACGGACTCACCATATCCTTTGTGGAGACTGA AGACACCAGATGACCATGAAGCAGAGACAGGGATTAAGTCAAAAGAAGCAAGAAAGTACATTTTCAACTGTTTAGATGATATGGCCCAGG TGAACATGACGACAGATTTGGAAGGGAGCGACATGTTGGTAGAAAAGGCTGACCGGCGGGAGTTCATTGACCTGTTGAAGAAGATGCTGA CCATTGATGCTGACAAGAGAATCACTCCAATCGAAACCCTGAACCATCCCTTTGTCACCATGACACACTTACTCGATTTTCCCCACAGCA CACACGTCAAATCATGTTTCCAGAACATGGAGATCTGCAAGCGTCGGGTGAATATGTATGACACGGTGAACCAGAGCAAAACCCCTTTCA TCACGCACGTGGCCCCCAGCACGTCCACCAACCTGACCATGACCTTTAACAACCAGCTGACCACTGTCCACAACCAGGCTCCCTCCTCTA CCAGTGCCACTATTTCCTTAGCCAATCCCGAAGTCTCCATACTAAACTACCCATCTACACTCTACCAGCCCTCAGCGGCATCCATGGCTG CAGTGGCCCAGCGGAGCATGCCCCTGCAGACAGGAACAGCCCAGATTTGTGCCCGGCCTGACCCGTTCCAGCAAGCTCTCATCGTGTGTC CCCCCGGCTTCCAAGGCTTGCAGGCCTCTCCCTCTAAGCACGCTGGCTACTCGGTGCGAATGGAAAATGCAGTTCCCATCGTCACTCAAG CCCCAGGAGCTCAGCCTCTTCAGATCCAACCAGGTCTGCTTGCCCAGGCACCAGAAGTCATCAGAATGCAAGATAAAAATCCATACAGCT TTCAGTCAGATGTATATGCATTTGGAATTGTTCTGTATGAATTGATGACTGGACAGTTACCTTATTCAAACATCAACAACAGGGACCAGA TAATTTTTATGGTGGGACGAGGATACCTGTCTCCAGATCTCAGTAAGGTACGGAGTAACTGTCCAAAAGCCATGAAGAGATTAATGGCAG AGTGCCTCAAAAAGAAAAGAGATGAGAGACCACTCTTTCCCCAAATTCTCGCCTCTATTGAGCTGCTGGCCCGCTCATTGCCAAAAATTC ACCGCAGTGCATCAGAACCCTCCTTGAATCGGGCTGGTTTCCAAACAGAGGATTTTAGTCTATATGCTTGTGCTTCTCCAAAAACACCCA TCCAGGCAGGGGGATATGGTGCGTTTCCTGTCCACTGAAACAAATGAGTGAGAGAGTTCAGGAGAGTAGCAACAAAAGGAAAATAAATGA >36401_36401_2_HIPK2-BRAF_HIPK2_chr7_139297947_ENST00000406875_BRAF_chr7_140449218_ENST00000288602_length(amino acids)=850AA_BP=698 MAPVYEGMASHVQVFSPHTLQSSAFCSVKKLKIEPSSNWDMTGYGSHSKVYSQSKNIPLSQPATTTVSTSLPVPNPSLPYEQTIVFPGST GHIVVTSASSTSVTGQVLGGPHNLMRRSTVSLLDTYQKCGLKRKSEEIENTSSVQIIEEHPPMIQNNASGATVATATTSTATSKNSGSNS EGDYQLVQHEVLCSMTNTYEVLEFLGRGTFGQVVKCWKRGTNEIVAIKILKNHPSYARQGQIEVSILARLSTESADDYNFVRAYECFQHK NHTCLVFEMLEQNLYDFLKQNKFSPLPLKYIRPVLQQVATALMKLKSLGLIHADLKPENIMLVDPSRQPYRVKVIDFGSASHVSKAVCST YLQSRYYRAPEIILGLPFCEAIDMWSLGCVIAELFLGWPLYPGASEYDQIRYISQTQGLPAEYLLSAGTKTTRFFNRDTDSPYPLWRLKT PDDHEAETGIKSKEARKYIFNCLDDMAQVNMTTDLEGSDMLVEKADRREFIDLLKKMLTIDADKRITPIETLNHPFVTMTHLLDFPHSTH VKSCFQNMEICKRRVNMYDTVNQSKTPFITHVAPSTSTNLTMTFNNQLTTVHNQAPSSTSATISLANPEVSILNYPSTLYQPSAASMAAV AQRSMPLQTGTAQICARPDPFQQALIVCPPGFQGLQASPSKHAGYSVRMENAVPIVTQAPGAQPLQIQPGLLAQAPEVIRMQDKNPYSFQ SDVYAFGIVLYELMTGQLPYSNINNRDQIIFMVGRGYLSPDLSKVRSNCPKAMKRLMAECLKKKRDERPLFPQILASIELLARSLPKIHR -------------------------------------------------------------- >36401_36401_3_HIPK2-BRAF_HIPK2_chr7_139297947_ENST00000428878_BRAF_chr7_140449218_ENST00000288602_length(transcript)=2745nt_BP=2186nt CACGGGAGGCGGTGATGCGGGCGCGGGCGGCCTCGGCTGCGCCGAGAGCGGAGACACAGGCTCAAGATGGCAGATTCCGACTGAGGCTGG GGGGGCCGAGCTCGCGCGCCGCTTTCCCGTCCCCGTTGCCATGAACCGCGGACACCCCGGCCCCGATGGCCCCCGTGTACGAAGGTATGG CCTCACATGTGCAAGTTTTCTCCCCTCACACCCTTCAATCAAGTGCCTTCTGTAGTGTGAAGAAACTGAAAATAGAGCCGAGTTCCAACT GGGACATGACTGGGTACGGCTCCCACAGCAAAGTGTATAGCCAGAGCAAGAACATCCCCCTGTCGCAGCCAGCCACCACAACCGTCAGCA CCTCCTTGCCGGTCCCAAACCCAAGCCTACCTTACGAGCAGACCATCGTCTTCCCAGGAAGCACCGGGCACATCGTGGTCACCTCAGCAA GCAGCACTTCTGTCACCGGGCAAGTCCTCGGCGGACCACACAACCTAATGCGTCGAAGCACTGTGAGCCTCCTTGATACCTACCAAAAAT GTGGACTCAAGCGTAAGAGCGAGGAGATCGAGAACACAAGCAGCGTGCAGATCATCGAGGAGCATCCACCCATGATTCAGAATAATGCAA GCGGGGCCACTGTCGCCACTGCCACCACGTCTACTGCCACCTCCAAAAACAGCGGCTCCAACAGCGAGGGCGACTATCAGCTGGTGCAGC ATGAGGTGCTGTGCTCCATGACCAACACCTACGAGGTCTTAGAGTTCTTGGGCCGAGGGACGTTTGGGCAAGTGGTCAAGTGCTGGAAAC GGGGCACCAATGAGATCGTAGCCATCAAGATCCTGAAGAACCACCCATCCTATGCCCGACAAGGTCAGATTGAAGTGAGCATCCTGGCCC GGTTGAGCACGGAGAGTGCCGATGACTATAACTTCGTCCGGGCCTACGAATGCTTCCAGCACAAGAACCACACGTGCTTGGTCTTCGAGA TGTTGGAGCAGAACCTCTATGACTTTCTGAAGCAAAACAAGTTTAGCCCCTTGCCCCTCAAATACATTCGCCCAGTTCTCCAGCAGGTAG CCACAGCCCTGATGAAACTCAAAAGCCTAGGTCTTATCCACGCTGACCTCAAACCAGAAAACATCATGCTGGTGGATCCATCTAGACAAC CATACAGAGTCAAGGTCATCGACTTTGGTTCAGCCAGCCACGTCTCCAAGGCTGTGTGCTCCACCTACTTGCAGTCCAGATATTACAGGG CCCCTGAGATCATCCTTGGTTTACCATTTTGTGAGGCAATTGACATGTGGTCCCTGGGCTGTGTTATTGCAGAATTGTTCCTGGGTTGGC CGTTATATCCAGGAGCTTCGGAGTATGATCAGATTCGGTATATTTCACAAACACAGGGTTTGCCTGCTGAATATTTATTAAGCGCCGGGA CAAAGACAACTAGGTTTTTCAACCGTGACACGGACTCACCATATCCTTTGTGGAGACTGAAGACACCAGATGACCATGAAGCAGAGACAG GGATTAAGTCAAAAGAAGCAAGAAAGTACATTTTCAACTGTTTAGATGATATGGCCCAGGTGAACATGACGACAGATTTGGAAGGGAGCG ACATGTTGGTAGAAAAGGCTGACCGGCGGGAGTTCATTGACCTGTTGAAGAAGATGCTGACCATTGATGCTGACAAGAGAATCACTCCAA TCGAAACCCTGAACCATCCCTTTGTCACCATGACACACTTACTCGATTTTCCCCACAGCACACACGTCAAATCATGTTTCCAGAACATGG AGATCTGCAAGCGTCGGGTGAATATGTATGACACGGTGAACCAGAGCAAAACCCCTTTCATCACGCACGTGGCCCCCAGCACGTCCACCA ACCTGACCATGACCTTTAACAACCAGCTGACCACTGTCCACAACCAGCCCTCAGCGGCATCCATGGCTGCAGTGGCCCAGCGGAGCATGC CCCTGCAGACAGGAACAGCCCAGATTTGTGCCCGGCCTGACCCGTTCCAGCAAGCTCTCATCGTGTGTCCCCCCGGCTTCCAAGGCTTGC AGGCCTCTCCCTCTAAGCACGCTGGCTACTCGGTGCGAATGGAAAATGCAGTTCCCATCGTCACTCAAGCCCCAGGAGCTCAGCCTCTTC AGATCCAACCAGGTCTGCTTGCCCAGGCACCAGAAGTCATCAGAATGCAAGATAAAAATCCATACAGCTTTCAGTCAGATGTATATGCAT TTGGAATTGTTCTGTATGAATTGATGACTGGACAGTTACCTTATTCAAACATCAACAACAGGGACCAGATAATTTTTATGGTGGGACGAG GATACCTGTCTCCAGATCTCAGTAAGGTACGGAGTAACTGTCCAAAAGCCATGAAGAGATTAATGGCAGAGTGCCTCAAAAAGAAAAGAG ATGAGAGACCACTCTTTCCCCAAATTCTCGCCTCTATTGAGCTGCTGGCCCGCTCATTGCCAAAAATTCACCGCAGTGCATCAGAACCCT CCTTGAATCGGGCTGGTTTCCAAACAGAGGATTTTAGTCTATATGCTTGTGCTTCTCCAAAAACACCCATCCAGGCAGGGGGATATGGTG CGTTTCCTGTCCACTGAAACAAATGAGTGAGAGAGTTCAGGAGAGTAGCAACAAAAGGAAAATAAATGAACATATGTTTGCTTATATGTT >36401_36401_3_HIPK2-BRAF_HIPK2_chr7_139297947_ENST00000428878_BRAF_chr7_140449218_ENST00000288602_length(amino acids)=823AA_BP=671 MAPVYEGMASHVQVFSPHTLQSSAFCSVKKLKIEPSSNWDMTGYGSHSKVYSQSKNIPLSQPATTTVSTSLPVPNPSLPYEQTIVFPGST GHIVVTSASSTSVTGQVLGGPHNLMRRSTVSLLDTYQKCGLKRKSEEIENTSSVQIIEEHPPMIQNNASGATVATATTSTATSKNSGSNS EGDYQLVQHEVLCSMTNTYEVLEFLGRGTFGQVVKCWKRGTNEIVAIKILKNHPSYARQGQIEVSILARLSTESADDYNFVRAYECFQHK NHTCLVFEMLEQNLYDFLKQNKFSPLPLKYIRPVLQQVATALMKLKSLGLIHADLKPENIMLVDPSRQPYRVKVIDFGSASHVSKAVCST YLQSRYYRAPEIILGLPFCEAIDMWSLGCVIAELFLGWPLYPGASEYDQIRYISQTQGLPAEYLLSAGTKTTRFFNRDTDSPYPLWRLKT PDDHEAETGIKSKEARKYIFNCLDDMAQVNMTTDLEGSDMLVEKADRREFIDLLKKMLTIDADKRITPIETLNHPFVTMTHLLDFPHSTH VKSCFQNMEICKRRVNMYDTVNQSKTPFITHVAPSTSTNLTMTFNNQLTTVHNQPSAASMAAVAQRSMPLQTGTAQICARPDPFQQALIV CPPGFQGLQASPSKHAGYSVRMENAVPIVTQAPGAQPLQIQPGLLAQAPEVIRMQDKNPYSFQSDVYAFGIVLYELMTGQLPYSNINNRD QIIFMVGRGYLSPDLSKVRSNCPKAMKRLMAECLKKKRDERPLFPQILASIELLARSLPKIHRSASEPSLNRAGFQTEDFSLYACASPKT -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for HIPK2-BRAF |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 189_520 | 704.0 | 1199.0 | DAXX |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 189_520 | 677.0 | 1172.0 | DAXX |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 935_1049 | 704.0 | 1199.0 | AXIN1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 935_1049 | 677.0 | 1172.0 | AXIN1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 774_876 | 704.0 | 1199.0 | CTBP1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 774_876 | 677.0 | 1172.0 | CTBP1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 787_897 | 704.0 | 1199.0 | HMGA1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 787_897 | 677.0 | 1172.0 | HMGA1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 752_897 | 704.0 | 1199.0 | POU4F1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 752_897 | 677.0 | 1172.0 | POU4F1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 539_844 | 704.0 | 1199.0 | SKI and SMAD1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 539_844 | 677.0 | 1172.0 | SKI and SMAD1 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 846_941 | 704.0 | 1199.0 | TP53 and TP73 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 846_941 | 677.0 | 1172.0 | TP53 and TP73 |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000406875 | - | 9 | 15 | 873_907 | 704.0 | 1199.0 | UBE2I |
| Hgene | HIPK2 | chr7:139297947 | chr7:140449218 | ENST00000428878 | - | 9 | 15 | 873_907 | 677.0 | 1172.0 | UBE2I |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for HIPK2-BRAF |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for HIPK2-BRAF |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
