
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:HLA-DPA1-SQSTM1 (FusionGDB2 ID:36669) |
Fusion Gene Summary for HLA-DPA1-SQSTM1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: HLA-DPA1-SQSTM1 | Fusion gene ID: 36669 | Hgene | Tgene | Gene symbol | HLA-DPA1 | SQSTM1 | Gene ID | 3113 | 8878 |
| Gene name | major histocompatibility complex, class II, DP alpha 1 | sequestosome 1 | |
| Synonyms | DP(W3)|DP(W4)|DPA1|HLA-DP1A|HLA-DPB1|HLADP|HLASB|PLT1 | A170|DMRV|FTDALS3|NADGP|OSIL|PDB3|ZIP3|p60|p62|p62B | |
| Cytomap | 6p21.32 | 5q35.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | HLA class II histocompatibility antigen, DP alpha 1 chainHLA DPA1HLA-DPA1*01:03:01:NewHLA-DPA1*02:01:01:NEWHLA-SB alpha chainMHC class II DP alpha chainMHC class II DP3-alphaMHC class II HLA-DPA1 antigen | sequestosome-1EBI3-associated protein of 60 kDaEBI3-associated protein p60EBIAPautophagy receptor p62oxidative stress induced likephosphotyrosine independent ligand for the Lck SH2 domain p62phosphotyrosine-independent ligand for the Lck SH2 domain | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | P20036 | Q13501 | |
| Ensembl transtripts involved in fusion gene | ENST00000428995, ENST00000374808, ENST00000383224, ENST00000415247, ENST00000419277, ENST00000422504, ENST00000443117, ENST00000454805, ENST00000463066, ENST00000474244, ENST00000475500, ENST00000480521, ENST00000484255, ENST00000488565, ENST00000494775, ENST00000502285, ENST00000515317, ENST00000549300, ENST00000550456, ENST00000550806, ENST00000551196, ENST00000551933, ENST00000552158, | ENST00000360718, ENST00000389805, ENST00000402874, ENST00000506690, ENST00000510187, ENST00000376929, | |
| Fusion gene scores | * DoF score | 7 X 8 X 3=168 | 33 X 21 X 17=11781 |
| # samples | 10 | 38 | |
| ** MAII score | log2(10/168*10)=-0.748461233004036 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(38/11781*10)=-4.95431877505661 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: HLA-DPA1 [Title/Abstract] AND SQSTM1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | HLA-DPA1(33032794)-SQSTM1(179250858), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | HLA-DPA1 | GO:0071346 | cellular response to interferon-gamma | 8568247 |
| Tgene | SQSTM1 | GO:0006914 | autophagy | 20452972 |
| Tgene | SQSTM1 | GO:0007032 | endosome organization | 27368102 |
| Tgene | SQSTM1 | GO:0031397 | negative regulation of protein ubiquitination | 20452972 |
| Tgene | SQSTM1 | GO:0061635 | regulation of protein complex stability | 25127057 |
| Tgene | SQSTM1 | GO:1905719 | protein localization to perinuclear region of cytoplasm | 27368102 |
 Fusion gene breakpoints across HLA-DPA1 (5'-gene) Fusion gene breakpoints across HLA-DPA1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene breakpoints across SQSTM1 (3'-gene) Fusion gene breakpoints across SQSTM1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
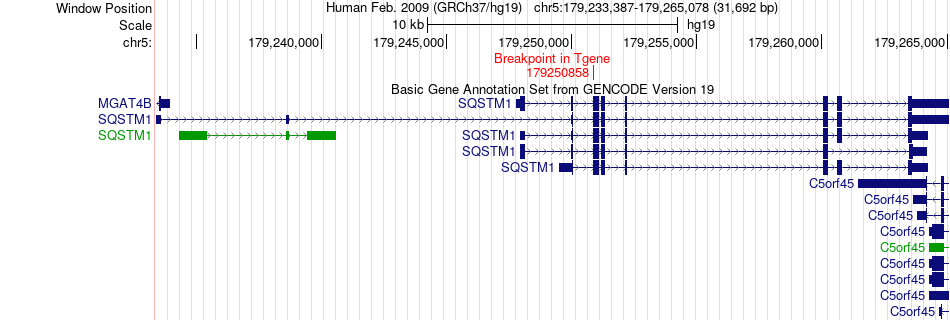 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | PAAD | TCGA-IB-7654-01A | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
Top |
Fusion Gene ORF analysis for HLA-DPA1-SQSTM1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000428995 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| 5CDS-intron | ENST00000428995 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| 5CDS-intron | ENST00000428995 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| 5CDS-intron | ENST00000428995 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| 5CDS-intron | ENST00000428995 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| In-frame | ENST00000428995 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000374808 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000383224 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000415247 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000419277 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000422504 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000443117 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000454805 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000463066 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000474244 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000475500 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000480521 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000484255 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000488565 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000494775 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000502285 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000515317 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000549300 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000550456 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000550806 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000551196 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000551933 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-3CDS | ENST00000552158 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000374808 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000374808 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000374808 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000374808 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000374808 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000383224 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000383224 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000383224 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000383224 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000383224 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000415247 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000415247 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000415247 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000415247 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000415247 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000419277 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000419277 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000419277 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000419277 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000419277 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000422504 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000422504 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000422504 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000422504 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000422504 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000443117 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000443117 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000443117 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000443117 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000443117 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000454805 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000454805 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000454805 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000454805 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000454805 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000463066 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000463066 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000463066 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000463066 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000463066 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000474244 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000474244 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000474244 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000474244 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000474244 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000475500 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000475500 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000475500 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000475500 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000475500 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000480521 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000480521 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000480521 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000480521 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000480521 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000484255 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000484255 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000484255 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000484255 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000484255 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000488565 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000488565 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000488565 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000488565 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000488565 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000494775 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000494775 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000494775 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000494775 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000494775 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000502285 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000502285 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000502285 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000502285 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000502285 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000515317 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000515317 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000515317 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000515317 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000515317 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000549300 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000549300 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000549300 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000549300 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000549300 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550456 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550456 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550456 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550456 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550456 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550806 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550806 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550806 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550806 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000550806 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551196 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551196 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551196 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551196 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551196 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551933 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551933 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551933 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551933 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000551933 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000552158 | ENST00000360718 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000552158 | ENST00000389805 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000552158 | ENST00000402874 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000552158 | ENST00000506690 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
| intron-intron | ENST00000552158 | ENST00000510187 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000428995 | HLA-DPA1 | chr6 | 33032794 | - | ENST00000376929 | SQSTM1 | chr5 | 179250858 | + | 3661 | 1157 | 1216 | 2178 | 320 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000428995 | ENST00000376929 | HLA-DPA1 | chr6 | 33032794 | - | SQSTM1 | chr5 | 179250858 | + | 0.021658076 | 0.97834194 |
Top |
Fusion Genomic Features for HLA-DPA1-SQSTM1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| HLA-DPA1 | chr6 | 33032793 | - | SQSTM1 | chr5 | 179250857 | + | 0.99916697 | 0.000833039 |
| HLA-DPA1 | chr6 | 33032793 | - | SQSTM1 | chr5 | 179250857 | + | 0.99916697 | 0.000833039 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
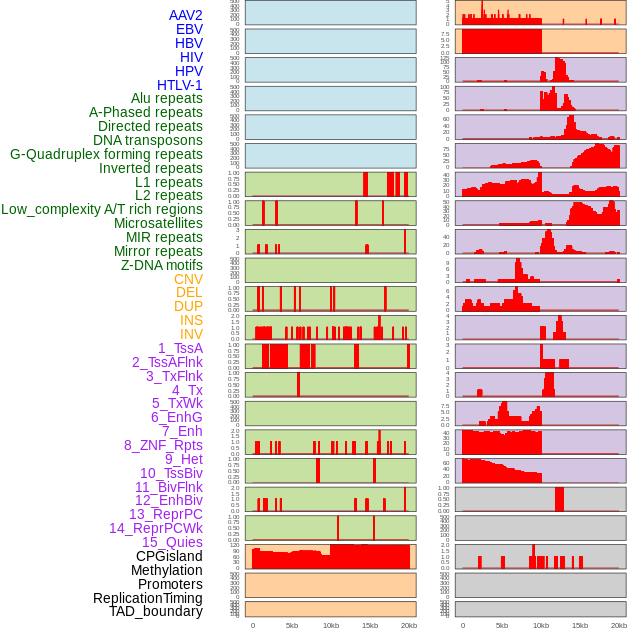 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
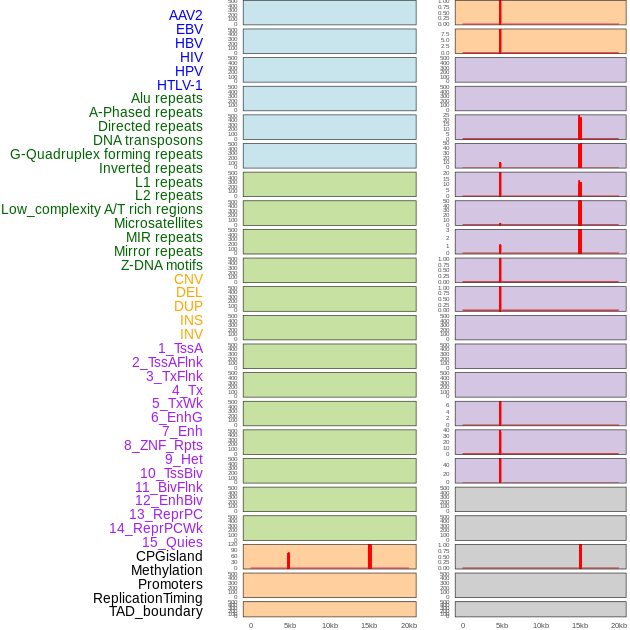 |
Top |
Fusion Protein Features for HLA-DPA1-SQSTM1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr6:33032794/chr5:179250858) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| HLA-DPA1 | SQSTM1 |
| FUNCTION: Binds peptides derived from antigens that access the endocytic route of antigen presenting cells (APC) and presents them on the cell surface for recognition by the CD4 T-cells. The peptide binding cleft accommodates peptides of 10-30 residues. The peptides presented by MHC class II molecules are generated mostly by degradation of proteins that access the endocytic route, where they are processed by lysosomal proteases and other hydrolases. Exogenous antigens that have been endocytosed by the APC are thus readily available for presentation via MHC II molecules, and for this reason this antigen presentation pathway is usually referred to as exogenous. As membrane proteins on their way to degradation in lysosomes as part of their normal turn-over are also contained in the endosomal/lysosomal compartments, exogenous antigens must compete with those derived from endogenous components. Autophagy is also a source of endogenous peptides, autophagosomes constitutively fuse with MHC class II loading compartments. In addition to APCs, other cells of the gastrointestinal tract, such as epithelial cells, express MHC class II molecules and CD74 and act as APCs, which is an unusual trait of the GI tract. To produce a MHC class II molecule that presents an antigen, three MHC class II molecules (heterodimers of an alpha and a beta chain) associate with a CD74 trimer in the ER to form a heterononamer. Soon after the entry of this complex into the endosomal/lysosomal system where antigen processing occurs, CD74 undergoes a sequential degradation by various proteases, including CTSS and CTSL, leaving a small fragment termed CLIP (class-II-associated invariant chain peptide). The removal of CLIP is facilitated by HLA-DM via direct binding to the alpha-beta-CLIP complex so that CLIP is released. HLA-DM stabilizes MHC class II molecules until primary high affinity antigenic peptides are bound. The MHC II molecule bound to a peptide is then transported to the cell membrane surface. In B-cells, the interaction between HLA-DM and MHC class II molecules is regulated by HLA-DO. Primary dendritic cells (DCs) also to express HLA-DO. Lysosomal microenvironment has been implicated in the regulation of antigen loading into MHC II molecules, increased acidification produces increased proteolysis and efficient peptide loading. | FUNCTION: Autophagy receptor required for selective macroautophagy (aggrephagy). Functions as a bridge between polyubiquitinated cargo and autophagosomes. Interacts directly with both the cargo to become degraded and an autophagy modifier of the MAP1 LC3 family (PubMed:16286508, PubMed:20168092, PubMed:24128730, PubMed:28404643, PubMed:22622177). Along with WDFY3, involved in the formation and autophagic degradation of cytoplasmic ubiquitin-containing inclusions (p62 bodies, ALIS/aggresome-like induced structures). Along with WDFY3, required to recruit ubiquitinated proteins to PML bodies in the nucleus (PubMed:24128730, PubMed:20168092). May regulate the activation of NFKB1 by TNF-alpha, nerve growth factor (NGF) and interleukin-1. May play a role in titin/TTN downstream signaling in muscle cells. May regulate signaling cascades through ubiquitination. Adapter that mediates the interaction between TRAF6 and CYLD (By similarity). May be involved in cell differentiation, apoptosis, immune response and regulation of K(+) channels. Involved in endosome organization by retaining vesicles in the perinuclear cloud: following ubiquitination by RNF26, attracts specific vesicle-associated adapters, forming a molecular bridge that restrains cognate vesicles in the perinuclear region and organizes the endosomal pathway for efficient cargo transport (PubMed:27368102). Promotes relocalization of 'Lys-63'-linked ubiquitinated STING1 to autophagosomes (PubMed:29496741). Acts as an activator of the NFE2L2/NRF2 pathway via interaction with KEAP1: interaction inactivates the BCR(KEAP1) complex, promoting nuclear accumulation of NFE2L2/NRF2 and subsequent expression of cytoprotective genes (PubMed:20452972, PubMed:28380357, PubMed:33393215). {ECO:0000250|UniProtKB:O08623, ECO:0000250|UniProtKB:Q64337, ECO:0000269|PubMed:10356400, ECO:0000269|PubMed:10747026, ECO:0000269|PubMed:11244088, ECO:0000269|PubMed:12471037, ECO:0000269|PubMed:15340068, ECO:0000269|PubMed:15802564, ECO:0000269|PubMed:15911346, ECO:0000269|PubMed:15953362, ECO:0000269|PubMed:16079148, ECO:0000269|PubMed:16286508, ECO:0000269|PubMed:19931284, ECO:0000269|PubMed:20168092, ECO:0000269|PubMed:20452972, ECO:0000269|PubMed:22622177, ECO:0000269|PubMed:24128730, ECO:0000269|PubMed:27368102, ECO:0000269|PubMed:28380357, ECO:0000269|PubMed:28404643, ECO:0000269|PubMed:29496741, ECO:0000269|PubMed:33393215}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000428995 | - | 5 | 5 | 118_210 | 375 | 301.3333333333333 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000428995 | - | 5 | 5 | 116_209 | 375 | 301.3333333333333 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000428995 | - | 5 | 5 | 210_222 | 375 | 301.3333333333333 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000428995 | - | 5 | 5 | 29_115 | 375 | 301.3333333333333 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000428995 | - | 5 | 5 | 246_260 | 375 | 301.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000428995 | - | 5 | 5 | 29_222 | 375 | 301.3333333333333 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000428995 | - | 5 | 5 | 223_245 | 375 | 301.3333333333333 | Transmembrane | Helical |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 272_294 | 16 | 357.0 | Compositional bias | Note=Ser-rich | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 272_294 | 16 | 357.0 | Compositional bias | Note=Ser-rich | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 272_294 | 100 | 441.0 | Compositional bias | Note=Ser-rich | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 272_294 | 16 | 357.0 | Compositional bias | Note=Ser-rich | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 389_434 | 16 | 357.0 | Domain | UBA | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 389_434 | 16 | 357.0 | Domain | UBA | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 389_434 | 100 | 441.0 | Domain | UBA | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 389_434 | 16 | 357.0 | Domain | UBA | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 228_233 | 16 | 357.0 | Motif | Note=TRAF6-binding | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 336_341 | 16 | 357.0 | Motif | Note=LIR | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 228_233 | 16 | 357.0 | Motif | Note=TRAF6-binding | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 336_341 | 16 | 357.0 | Motif | Note=LIR | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 228_233 | 100 | 441.0 | Motif | Note=TRAF6-binding | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 336_341 | 100 | 441.0 | Motif | Note=LIR | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 228_233 | 16 | 357.0 | Motif | Note=TRAF6-binding | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 336_341 | 16 | 357.0 | Motif | Note=LIR | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 170_220 | 16 | 357.0 | Region | Note=LIM protein-binding (LB) | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 321_342 | 16 | 357.0 | Region | MAP1LC3B-binding | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 170_220 | 16 | 357.0 | Region | Note=LIM protein-binding (LB) | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 321_342 | 16 | 357.0 | Region | MAP1LC3B-binding | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 170_220 | 100 | 441.0 | Region | Note=LIM protein-binding (LB) | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 321_342 | 100 | 441.0 | Region | MAP1LC3B-binding | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 170_220 | 16 | 357.0 | Region | Note=LIM protein-binding (LB) | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 321_342 | 16 | 357.0 | Region | MAP1LC3B-binding | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 122_167 | 16 | 357.0 | Zinc finger | ZZ-type | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 122_167 | 16 | 357.0 | Zinc finger | ZZ-type | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 122_167 | 100 | 441.0 | Zinc finger | ZZ-type | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 122_167 | 16 | 357.0 | Zinc finger | ZZ-type |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000374808 | - | 1 | 6 | 118_210 | 0 | 499.6666666666667 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000383224 | - | 1 | 6 | 118_210 | 0 | 499.6666666666667 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000415247 | - | 1 | 5 | 118_210 | 0 | 443.6666666666667 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000419277 | - | 1 | 6 | 118_210 | 0 | 499.6666666666667 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000422504 | - | 1 | 6 | 118_210 | 0 | 499.6666666666667 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000443117 | - | 1 | 6 | 118_210 | 0 | 499.6666666666667 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000454805 | - | 1 | 6 | 118_210 | 0 | 499.6666666666667 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000515317 | - | 1 | 6 | 118_210 | 0 | 499.6666666666667 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000549300 | - | 1 | 5 | 118_210 | 0 | 301.3333333333333 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550456 | - | 1 | 5 | 118_210 | 0 | 301.3333333333333 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550806 | - | 1 | 5 | 118_210 | 0 | 301.3333333333333 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551196 | - | 1 | 5 | 118_210 | 0 | 301.3333333333333 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551933 | - | 1 | 5 | 118_210 | 0 | 301.3333333333333 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000552158 | - | 1 | 5 | 118_210 | 0 | 301.3333333333333 | Domain | Note=Ig-like C1-type |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000374808 | - | 1 | 6 | 116_209 | 0 | 499.6666666666667 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000374808 | - | 1 | 6 | 210_222 | 0 | 499.6666666666667 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000374808 | - | 1 | 6 | 29_115 | 0 | 499.6666666666667 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000383224 | - | 1 | 6 | 116_209 | 0 | 499.6666666666667 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000383224 | - | 1 | 6 | 210_222 | 0 | 499.6666666666667 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000383224 | - | 1 | 6 | 29_115 | 0 | 499.6666666666667 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000415247 | - | 1 | 5 | 116_209 | 0 | 443.6666666666667 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000415247 | - | 1 | 5 | 210_222 | 0 | 443.6666666666667 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000415247 | - | 1 | 5 | 29_115 | 0 | 443.6666666666667 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000419277 | - | 1 | 6 | 116_209 | 0 | 499.6666666666667 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000419277 | - | 1 | 6 | 210_222 | 0 | 499.6666666666667 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000419277 | - | 1 | 6 | 29_115 | 0 | 499.6666666666667 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000422504 | - | 1 | 6 | 116_209 | 0 | 499.6666666666667 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000422504 | - | 1 | 6 | 210_222 | 0 | 499.6666666666667 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000422504 | - | 1 | 6 | 29_115 | 0 | 499.6666666666667 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000443117 | - | 1 | 6 | 116_209 | 0 | 499.6666666666667 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000443117 | - | 1 | 6 | 210_222 | 0 | 499.6666666666667 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000443117 | - | 1 | 6 | 29_115 | 0 | 499.6666666666667 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000454805 | - | 1 | 6 | 116_209 | 0 | 499.6666666666667 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000454805 | - | 1 | 6 | 210_222 | 0 | 499.6666666666667 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000454805 | - | 1 | 6 | 29_115 | 0 | 499.6666666666667 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000515317 | - | 1 | 6 | 116_209 | 0 | 499.6666666666667 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000515317 | - | 1 | 6 | 210_222 | 0 | 499.6666666666667 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000515317 | - | 1 | 6 | 29_115 | 0 | 499.6666666666667 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000549300 | - | 1 | 5 | 116_209 | 0 | 301.3333333333333 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000549300 | - | 1 | 5 | 210_222 | 0 | 301.3333333333333 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000549300 | - | 1 | 5 | 29_115 | 0 | 301.3333333333333 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550456 | - | 1 | 5 | 116_209 | 0 | 301.3333333333333 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550456 | - | 1 | 5 | 210_222 | 0 | 301.3333333333333 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550456 | - | 1 | 5 | 29_115 | 0 | 301.3333333333333 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550806 | - | 1 | 5 | 116_209 | 0 | 301.3333333333333 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550806 | - | 1 | 5 | 210_222 | 0 | 301.3333333333333 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550806 | - | 1 | 5 | 29_115 | 0 | 301.3333333333333 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551196 | - | 1 | 5 | 116_209 | 0 | 301.3333333333333 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551196 | - | 1 | 5 | 210_222 | 0 | 301.3333333333333 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551196 | - | 1 | 5 | 29_115 | 0 | 301.3333333333333 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551933 | - | 1 | 5 | 116_209 | 0 | 301.3333333333333 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551933 | - | 1 | 5 | 210_222 | 0 | 301.3333333333333 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551933 | - | 1 | 5 | 29_115 | 0 | 301.3333333333333 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000552158 | - | 1 | 5 | 116_209 | 0 | 301.3333333333333 | Region | Note=Alpha-2 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000552158 | - | 1 | 5 | 210_222 | 0 | 301.3333333333333 | Region | Note=Connecting peptide |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000552158 | - | 1 | 5 | 29_115 | 0 | 301.3333333333333 | Region | Note=Alpha-1 |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000374808 | - | 1 | 6 | 246_260 | 0 | 499.6666666666667 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000374808 | - | 1 | 6 | 29_222 | 0 | 499.6666666666667 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000383224 | - | 1 | 6 | 246_260 | 0 | 499.6666666666667 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000383224 | - | 1 | 6 | 29_222 | 0 | 499.6666666666667 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000415247 | - | 1 | 5 | 246_260 | 0 | 443.6666666666667 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000415247 | - | 1 | 5 | 29_222 | 0 | 443.6666666666667 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000419277 | - | 1 | 6 | 246_260 | 0 | 499.6666666666667 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000419277 | - | 1 | 6 | 29_222 | 0 | 499.6666666666667 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000422504 | - | 1 | 6 | 246_260 | 0 | 499.6666666666667 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000422504 | - | 1 | 6 | 29_222 | 0 | 499.6666666666667 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000443117 | - | 1 | 6 | 246_260 | 0 | 499.6666666666667 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000443117 | - | 1 | 6 | 29_222 | 0 | 499.6666666666667 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000454805 | - | 1 | 6 | 246_260 | 0 | 499.6666666666667 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000454805 | - | 1 | 6 | 29_222 | 0 | 499.6666666666667 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000515317 | - | 1 | 6 | 246_260 | 0 | 499.6666666666667 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000515317 | - | 1 | 6 | 29_222 | 0 | 499.6666666666667 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000549300 | - | 1 | 5 | 246_260 | 0 | 301.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000549300 | - | 1 | 5 | 29_222 | 0 | 301.3333333333333 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550456 | - | 1 | 5 | 246_260 | 0 | 301.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550456 | - | 1 | 5 | 29_222 | 0 | 301.3333333333333 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550806 | - | 1 | 5 | 246_260 | 0 | 301.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550806 | - | 1 | 5 | 29_222 | 0 | 301.3333333333333 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551196 | - | 1 | 5 | 246_260 | 0 | 301.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551196 | - | 1 | 5 | 29_222 | 0 | 301.3333333333333 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551933 | - | 1 | 5 | 246_260 | 0 | 301.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551933 | - | 1 | 5 | 29_222 | 0 | 301.3333333333333 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000552158 | - | 1 | 5 | 246_260 | 0 | 301.3333333333333 | Topological domain | Cytoplasmic |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000552158 | - | 1 | 5 | 29_222 | 0 | 301.3333333333333 | Topological domain | Extracellular |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000374808 | - | 1 | 6 | 223_245 | 0 | 499.6666666666667 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000383224 | - | 1 | 6 | 223_245 | 0 | 499.6666666666667 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000415247 | - | 1 | 5 | 223_245 | 0 | 443.6666666666667 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000419277 | - | 1 | 6 | 223_245 | 0 | 499.6666666666667 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000422504 | - | 1 | 6 | 223_245 | 0 | 499.6666666666667 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000443117 | - | 1 | 6 | 223_245 | 0 | 499.6666666666667 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000454805 | - | 1 | 6 | 223_245 | 0 | 499.6666666666667 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000515317 | - | 1 | 6 | 223_245 | 0 | 499.6666666666667 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000549300 | - | 1 | 5 | 223_245 | 0 | 301.3333333333333 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550456 | - | 1 | 5 | 223_245 | 0 | 301.3333333333333 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000550806 | - | 1 | 5 | 223_245 | 0 | 301.3333333333333 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551196 | - | 1 | 5 | 223_245 | 0 | 301.3333333333333 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000551933 | - | 1 | 5 | 223_245 | 0 | 301.3333333333333 | Transmembrane | Helical |
| Hgene | HLA-DPA1 | chr6:33032794 | chr5:179250858 | ENST00000552158 | - | 1 | 5 | 223_245 | 0 | 301.3333333333333 | Transmembrane | Helical |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 3_102 | 16 | 357.0 | Domain | PB1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 3_102 | 16 | 357.0 | Domain | PB1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 3_102 | 100 | 441.0 | Domain | PB1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 3_102 | 16 | 357.0 | Domain | PB1 |
Top |
Fusion Gene Sequence for HLA-DPA1-SQSTM1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >36669_36669_1_HLA-DPA1-SQSTM1_HLA-DPA1_chr6_33032794_ENST00000428995_SQSTM1_chr5_179250858_ENST00000376929_length(transcript)=3661nt_BP=1157nt GCTCACAGTCATCAATTATAGACCCCACAACATGCGCCCTGAAGACAGAATGTTCCATATCAGAGCTGTGATCTTGAGAGCCCTCTCCTT GGCTTTCCTGCTGAGTCTCCGAGGAGCTGGGGCCATCAAGGCGGACCATGTGTCAACTTATGCCGCGTTTGTACAGACGCATAGACCAAC AGGGGAGTTTATGTTTGAATTTGATGAAGATGAGATGTTCTATGTGGATCTGGACAAGAAGGAGACCGTCTGGCATCTGGAGGAGTTTGG CCAAGCCTTTTCCTTTGAGGCTCAGGGCGGGCTGGCTAACATTGCTATATTGAACAACAACTTGAATACCTTGATCCAGCGTTCCAACCA CACTCAGGCCACCAACGATCCCCCTGAGGTGACCGTGTTTCCCAAGGAGCCTGTGGAGCTGGGCCAGCCCAACACCCTCATCTGCCACAT TGACAAGTTCTTCCCACCAGTGCTCAACGTCACGTGGCTGTGCAACGGGGAGCTGGTCACTGAGGGTGTCGCTGAGAGCCTCTTCCTGCC CAGAACAGATTACAGCTTCCACAAGTTCCATTACCTGACCTTTGTGCCCTCAGCAGAGGACTTCTATGACTGCAGGGTGGAGCACTGGGG CTTGGACCAGCCGCTCCTCAAGCACTGGGAGGCCCAAGAGCCAATCCAGATGCCTGAGACAACGGAGACTGTGCTCTGTGCCCTGGGCCT GGTGCTGGGCCTAGTCGGCATCATCGTGGGCACCGTCCTCATCATAAAGTCTCTGCGTTCTGGCCATGACCCCCGGGCCCAGGGGACCCT GTGAAATACTGTAAAGGTGACAAAATATCTGAACAGAAGAGGACTTAGGAGAGATCTGAACTCCAGCTGCCCTACAAACTCCATCTCAGC TTTTCTTCTCACTTCATGTGAAAACTACTCCAGTGGCTGACTGAATTGCTGACCCTTCAAGCTCTGTCCTTATCCATTACCTCAAAGCAG TCATTCCTTAGTAAAGTTTCCAACAAATAGAAATTAATGACACTTTGGTAGCACTAATATGGAGATTATCCTTTCATTGAGCCTTTTATC CTCTGTTCTCCTTTGAAGAACCCCTCACTGTCACCTTCCCGAGAATACCCTAAGACCAATAAATACTTCAGTATTTCAGAAAAAAGAGTG CCGGCGGGACCACCGCCCACCGTGTGCTCAGGAGGCGCCCCGCAACATGGTGCACCCCAATGTGATCTGCGATGGCTGCAATGGGCCTGT GGTAGGAACCCGCTACAAGTGCAGCGTCTGCCCAGACTACGACTTGTGTAGCGTCTGCGAGGGAAAGGGCTTGCACCGGGGGCACACCAA GCTCGCATTCCCCAGCCCCTTCGGGCACCTGTCTGAGGGCTTCTCGCACAGCCGCTGGCTCCGGAAGGTGAAACACGGACACTTCGGGTG GCCAGGATGGGAAATGGGTCCACCAGGAAACTGGAGCCCACGTCCTCCTCGTGCAGGGGAGGCCCGCCCTGGCCCCACGGCAGAATCAGC TTCTGGTCCATCGGAGGATCCGAGTGTGAATTTCCTGAAGAACGTTGGGGAGAGTGTGGCAGCTGCCCTTAGCCCTCTGGGCATTGAAGT TGATATCGATGTGGAGCACGGAGGGAAAAGAAGCCGCCTGACCCCCGTCTCTCCAGAGAGTTCCAGCACAGAGGAGAAGAGCAGCTCACA GCCAAGCAGCTGCTGCTCTGACCCCAGCAAGCCGGGTGGGAATGTTGAGGGCGCCACGCAGTCTCTGGCGGAGCAGATGAGGAAGATCGC CTTGGAGTCCGAGGGGCGCCCTGAGGAACAGATGGAGTCGGATAACTGTTCAGGAGGAGATGATGACTGGACCCATCTGTCTTCAAAAGA AGTGGACCCGTCTACAGGTGAACTCCAGTCCCTACAGATGCCAGAATCCGAAGGGCCAAGCTCTCTGGACCCCTCCCAGGAGGGACCCAC AGGGCTGAAGGAAGCTGCCTTGTACCCACATCTCCCGCCAGAGGCTGACCCGCGGCTGATTGAGTCCCTCTCCCAGATGCTGTCCATGGG CTTCTCTGATGAAGGCGGCTGGCTCACCAGGCTCCTGCAGACCAAGAACTATGACATCGGAGCGGCTCTGGACACCATCCAGTATTCAAA GCATCCCCCGCCGTTGTGACCACTTTTGCCCACCTCTTCTGCGTGCCCCTCTTCTGTCTCATAGTTGTGTTAAGCTTGCGTAGAATTGCA GGTCTCTGTACGGGCCAGTTTCTCTGCCTTCTTCCAGGATCAGGGGTTAGGGTGCAAGAAGCCATTTAGGGCAGCAAAACAAGTGACATG AAGGGAGGGTCCCTGTGTGTGTGTGTGCTGATGTTTCCTGGGTGCCCTGGCTCCTTGCAGCAGGGCTGGGCCTGCGAGACCCAAGGCTCA CTGCAGCGCGCTCCTGACCCCTCCCTGCAGGGGCTACGTTAGCAGCCCAGCACATAGCTTGCCTAATGGCTTTCACTTTCTCTTTTGTTT TAAATGACTCATAGGTCCCTGACATTTAGTTGATTATTTTCTGCTACAGACCTGGTACACTCTGATTTTAGATAAAGTAAGCCTAGGTGT TGTCAGCAGGCAGGCTGGGGAGGCCAGTGTTGTGGGCTTCCTGCTGGGACTGAGAAGGCTCACGAAGGGCATCCGCAATGTTGGTTTCAC TGAGAGCTGCCTCCTGGTCTCTTCACCACTGTAGTTCTCTCATTTCCAAACCATCAGCTGCTTTTAAAATAAGATCTCTTTGTAGCCATC CTGTTAAATTTGTAAACAATCTAATTAAATGGCATCAGCACTTTAACCAATGACGTTTGCATAGAGAGAAATGATTGACAGTAAGTTTAT TGTTAATGGTTCTTACAGAGTATCTTTAAAAGTGCCTTAGGGGAACCCTGTCCCTCCTAACAAGTGTATCTCGATTAATAACCTGCCAGT CCCAGATCACACATCATCATCGAAGTCTTCCCCAGTTATAAAGAGGTCACATAGTCGTGTGGGTCGAGGATTCTGTGCCTCCAGGACCAG GGGCCCACCCTCTGCCCAGGGAGTCCTTGCGTCCCATGAGGTCTTCCCGCAAGGCCTCTCAGACCCAGATGTGACGGGGTGTGTGGCCCG AGGAAGCTGGACAGCGGCAGTGGGCCTGCTGAGGCCTTCTCTTGAGGCCTGTGCTCTGGGGGTCCCTTGCTTAGCCTGTGCTGGACCAGC TGGCCTGGGGTCCCTCTGAAGAGACCTTGGCTGCTCACTGTCCACATGTGAACTTTTTCTAGGTGGCAGGACAAATTGCGCCCATTTAGA GGATGTGGCTGTAACCTGCTGGATGGGACTCCATAGCTCCTTCCCAGGACCCCTCAGCTCCCCGGCACTGCAGTCTGCAGAGTTCTCCTG GAGGCAGGGGCTGCTGCCTTGTTTCACCTTCCATGTCAGGCCAGCCTGTCCCTGAAAGAGAAGATGGCCATGCCCTCCATGTGTAAGAAC AATGCCAGGGCCCAGGAGGACCGCCTGCCCTGCCTGGGCCTTGGCTGGGCCTCTGGTTCTGACACTTTCTGCTGGAAGCTGTCAGGCTGG >36669_36669_1_HLA-DPA1-SQSTM1_HLA-DPA1_chr6_33032794_ENST00000428995_SQSTM1_chr5_179250858_ENST00000376929_length(amino acids)=320AA_BP= MVHPNVICDGCNGPVVGTRYKCSVCPDYDLCSVCEGKGLHRGHTKLAFPSPFGHLSEGFSHSRWLRKVKHGHFGWPGWEMGPPGNWSPRP PRAGEARPGPTAESASGPSEDPSVNFLKNVGESVAAALSPLGIEVDIDVEHGGKRSRLTPVSPESSSTEEKSSSQPSSCCSDPSKPGGNV EGATQSLAEQMRKIALESEGRPEEQMESDNCSGGDDDWTHLSSKEVDPSTGELQSLQMPESEGPSSLDPSQEGPTGLKEAALYPHLPPEA -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for HLA-DPA1-SQSTM1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 122_224 | 16.333333333333332 | 357.0 | GABRR3 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 122_224 | 16.333333333333332 | 357.0 | GABRR3 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 122_224 | 100.33333333333333 | 441.0 | GABRR3 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 122_224 | 16.333333333333332 | 357.0 | GABRR3 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 347_352 | 16.333333333333332 | 357.0 | KEAP1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 347_352 | 16.333333333333332 | 357.0 | KEAP1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 347_352 | 100.33333333333333 | 441.0 | KEAP1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 347_352 | 16.333333333333332 | 357.0 | KEAP1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 269_440 | 16.333333333333332 | 357.0 | NTRK1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 269_440 | 16.333333333333332 | 357.0 | NTRK1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 269_440 | 100.33333333333333 | 441.0 | NTRK1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 269_440 | 16.333333333333332 | 357.0 | NTRK1 | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 50_80 | 16.333333333333332 | 357.0 | PAWR | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 50_80 | 16.333333333333332 | 357.0 | PAWR | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 50_80 | 16.333333333333332 | 357.0 | PAWR | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 43_107 | 16.333333333333332 | 357.0 | PRKCZ and dimerization | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 43_107 | 16.333333333333332 | 357.0 | PRKCZ and dimerization | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 43_107 | 16.333333333333332 | 357.0 | PRKCZ and dimerization |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000360718 | 0 | 7 | 2_50 | 16.333333333333332 | 357.0 | LCK | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000376929 | 2 | 9 | 2_50 | 16.333333333333332 | 357.0 | LCK | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 2_50 | 100.33333333333333 | 441.0 | LCK | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000402874 | 1 | 8 | 2_50 | 16.333333333333332 | 357.0 | LCK | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 50_80 | 100.33333333333333 | 441.0 | PAWR | |
| Tgene | SQSTM1 | chr6:33032794 | chr5:179250858 | ENST00000389805 | 1 | 8 | 43_107 | 100.33333333333333 | 441.0 | PRKCZ and dimerization |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for HLA-DPA1-SQSTM1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for HLA-DPA1-SQSTM1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
