
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:HMCES-PCNP (FusionGDB2 ID:36776) |
Fusion Gene Summary for HMCES-PCNP |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: HMCES-PCNP | Fusion gene ID: 36776 | Hgene | Tgene | Gene symbol | HMCES | PCNP | Gene ID | 56941 | 57092 |
| Gene name | 5-hydroxymethylcytosine binding, ES cell specific | PEST proteolytic signal containing nuclear protein | |
| Synonyms | C3orf37|DC12|SRAPD1 | - | |
| Cytomap | 3q21.3 | 3q12.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | abasic site processing protein HMCES5-hydroxymethylcytosine (hmC) binding, ES cell-specificES cell-specific 5hmC-binding proteinSOS response associated peptidase domain containing 1SRAP domain-containing protein 1UPF0361 protein C3orf37embryonic ste | PEST proteolytic signal-containing nuclear proteinPEST-containing nuclear protein | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q96FZ2 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000383463, ENST00000389735, ENST00000417226, ENST00000502878, | ENST00000486406, ENST00000469941, ENST00000296024, ENST00000265260, | |
| Fusion gene scores | * DoF score | 4 X 5 X 2=40 | 10 X 13 X 3=390 |
| # samples | 5 | 14 | |
| ** MAII score | log2(5/40*10)=0.321928094887362 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(14/390*10)=-1.47804729680464 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: HMCES [Title/Abstract] AND PCNP [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | HMCES(128998758)-PCNP(101309059), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | HMCES | GO:0006974 | cellular response to DNA damage stimulus | 30554877 |
| Tgene | PCNP | GO:0016567 | protein ubiquitination | 14741369 |
| Tgene | PCNP | GO:0043161 | proteasome-mediated ubiquitin-dependent protein catabolic process | 14741369 |
 Fusion gene breakpoints across HMCES (5'-gene) Fusion gene breakpoints across HMCES (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
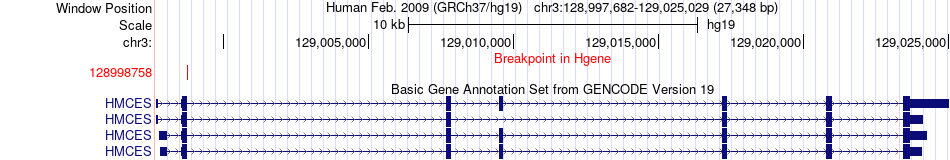 |
 Fusion gene breakpoints across PCNP (3'-gene) Fusion gene breakpoints across PCNP (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
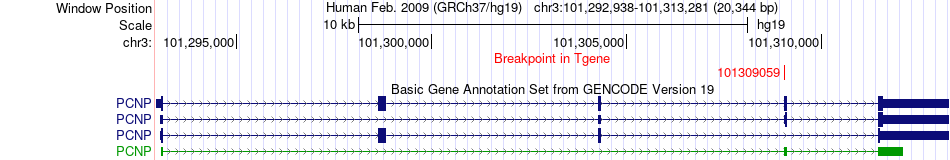 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-CG-5724-01A | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
Top |
Fusion Gene ORF analysis for HMCES-PCNP |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000383463 | ENST00000486406 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-3UTR | ENST00000389735 | ENST00000486406 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-3UTR | ENST00000417226 | ENST00000486406 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-3UTR | ENST00000502878 | ENST00000486406 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-5UTR | ENST00000383463 | ENST00000469941 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-5UTR | ENST00000389735 | ENST00000469941 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-5UTR | ENST00000417226 | ENST00000469941 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-5UTR | ENST00000502878 | ENST00000469941 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-intron | ENST00000383463 | ENST00000296024 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-intron | ENST00000389735 | ENST00000296024 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-intron | ENST00000417226 | ENST00000296024 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| 5CDS-intron | ENST00000502878 | ENST00000296024 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| In-frame | ENST00000383463 | ENST00000265260 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| In-frame | ENST00000389735 | ENST00000265260 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| In-frame | ENST00000417226 | ENST00000265260 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
| In-frame | ENST00000502878 | ENST00000265260 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000383463 | HMCES | chr3 | 128998758 | + | ENST00000265260 | PCNP | chr3 | 101309059 | + | 2139 | 272 | 89 | 454 | 121 |
| ENST00000417226 | HMCES | chr3 | 128998758 | + | ENST00000265260 | PCNP | chr3 | 101309059 | + | 2133 | 266 | 83 | 448 | 121 |
| ENST00000502878 | HMCES | chr3 | 128998758 | + | ENST00000265260 | PCNP | chr3 | 101309059 | + | 2321 | 454 | 113 | 490 | 125 |
| ENST00000389735 | HMCES | chr3 | 128998758 | + | ENST00000265260 | PCNP | chr3 | 101309059 | + | 2311 | 444 | 261 | 626 | 121 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000383463 | ENST00000265260 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + | 0.0879772 | 0.9120228 |
| ENST00000417226 | ENST00000265260 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + | 0.13309391 | 0.8669061 |
| ENST00000502878 | ENST00000265260 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + | 0.20192197 | 0.79807806 |
| ENST00000389735 | ENST00000265260 | HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309059 | + | 0.08395294 | 0.91604704 |
Top |
Fusion Genomic Features for HMCES-PCNP |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309058 | + | 0.01544915 | 0.9845509 |
| HMCES | chr3 | 128998758 | + | PCNP | chr3 | 101309058 | + | 0.01544915 | 0.9845509 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
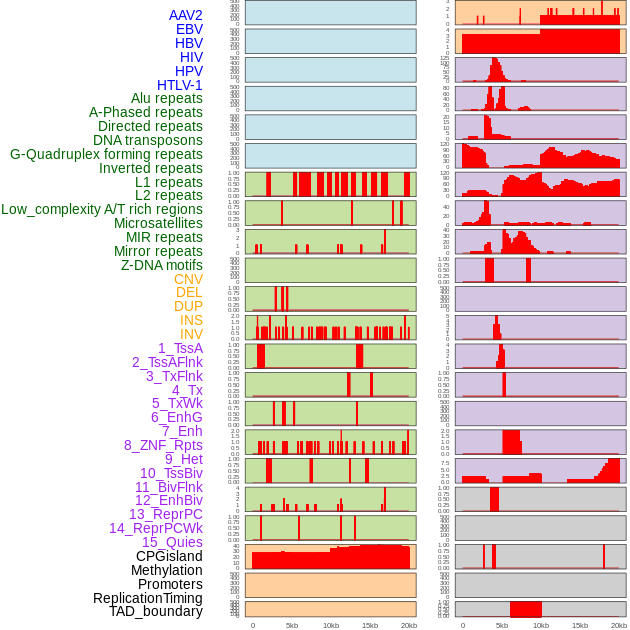 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
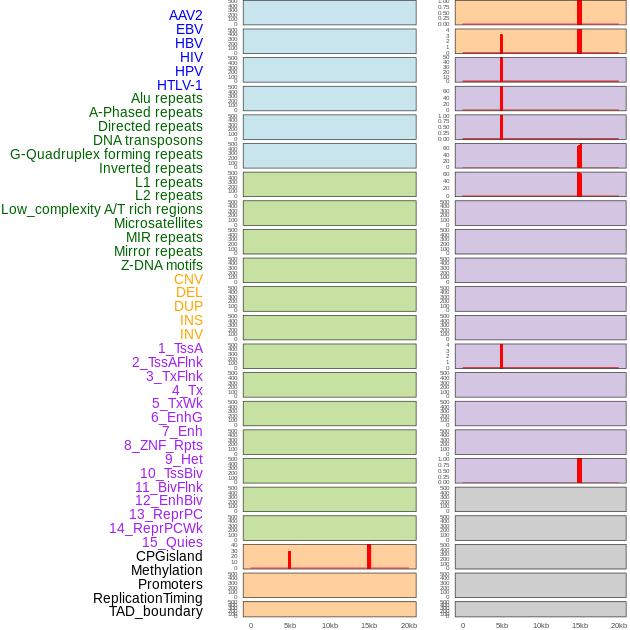 |
Top |
Fusion Protein Features for HMCES-PCNP |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr3:128998758/chr3:101309059) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| HMCES | . |
| FUNCTION: Sensor of abasic sites in single-stranded DNA (ssDNA) required to preserve genome integrity by promoting error-free repair of abasic sites (PubMed:30554877, PubMed:31235915, PubMed:31235913). Acts as an enzyme that recognizes and binds abasic sites in ssDNA at replication forks and chemically modifies the lesion by forming a covalent cross-link with DNA: forms a stable thiazolidine linkage between a ring-opened abasic site and the alpha-amino and sulfhydryl substituents of its N-terminal catalytic cysteine residue (PubMed:30554877, PubMed:31235913). The HMCES DNA-protein cross-link is then degraded by the proteasome (PubMed:30554877). Promotes error-free repair of abasic sites by acting as a 'suicide' enzyme that is degraded, thereby protecting abasic sites from translesion synthesis (TLS) polymerases and endonucleases that are error-prone and would generate mutations and double-strand breaks (PubMed:30554877). Has preference for ssDNA, but can also accommodate double-stranded DNA with 3' or 5' overhang (dsDNA), and dsDNA-ssDNA 3' junction (PubMed:31235915, PubMed:31806351). Also involved in class switch recombination (CSR) in B-cells independently of the formation of a DNA-protein cross-link: acts by binding and protecting ssDNA overhangs to promote DNA double-strand break repair through the microhomology-mediated alternative-end-joining (Alt-EJ) pathway (By similarity). Acts as a protease: mediates autocatalytic processing of its N-terminal methionine in order to expose the catalytic cysteine (By similarity). {ECO:0000250|UniProtKB:Q8R1M0, ECO:0000269|PubMed:30554877, ECO:0000269|PubMed:31235913, ECO:0000269|PubMed:31235915, ECO:0000269|PubMed:31806351}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HMCES | chr3:128998758 | chr3:101309059 | ENST00000383463 | + | 2 | 7 | 332_338 | 61 | 355.0 | Motif | PIP-box |
| Hgene | HMCES | chr3:128998758 | chr3:101309059 | ENST00000389735 | + | 2 | 7 | 332_338 | 61 | 355.0 | Motif | PIP-box |
| Hgene | HMCES | chr3:128998758 | chr3:101309059 | ENST00000502878 | + | 2 | 7 | 332_338 | 61 | 355.0 | Motif | PIP-box |
Top |
Fusion Gene Sequence for HMCES-PCNP |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >36776_36776_1_HMCES-PCNP_HMCES_chr3_128998758_ENST00000383463_PCNP_chr3_101309059_ENST00000265260_length(transcript)=2139nt_BP=272nt TGGCCAAAGCGGAGCGGAGCGGGGGTGAGGAGAGTCGAGGGAGGTGACGCGCGCTGCCGGGGCGAGGTTGCGAGGGGCGGTGTTGAAGAA TGTGTGGGCGAACATCCTGTCACTTACCTAGAGATGTTCTCACGAGAGCTTGCGCCTACCAGGATCGGCGGGGCCAGCAGCGGCTCCCGG AGTGGAGGGACCCTGATAAGTACTGCCCCTCTTACAACAAGAGTCCTCAATCCAACAGCCCAGTGCTTCTGTCTCGACTGCACTTTGAGA AGAGTGAACCAGAGGAAATGCCTCCAGAAGCAAAGATGAGGATGAAGAATATTGGAAGGGATACACCAACATCAGCTGGACCAAACTCCT TCAATAAAGGAAAGCATGGGTTTTCTGATAACCAGAAGCTGTGGGAGCGAAATATAAAATCTCATCTTGGAAATGTCCATGACCAAGACA ATTAAATGATGTTTTGAAATTGGGGTGTGGGGTGGGTGTAAAGTTAAAAGGAACAGTTTCCTTTTTTAAAGAATGGTATAAGACTATCTT TGGAGCCGCTTTTTTTTTCTTTTTCATTTTTTTAAAAGATTGAGTGGTACACTAATAAATGAGAGTTTGAAATTAGAGGTAATTTATGTT TTATATACAGATTTCAAGACATTTGCTAATTTTGTAGTTTCATGTGATTAGTTTCCAAAGGTTACAGATAATAAAGAAATCAGAAATGGT ACCTTTTTAAGAATTGCATATTTTTTTAGACACAACTATTAGTACATTAAGAGGGAAGCAAAGTTACTGTCTATTTAAAACTGCAAGCAG TTAACTCTCTTAACTCCCTTATTACCTAAACTTGTCTGGCTCCCAGGAACAGCCTTATAGAGAGAGGGAGTATTGTATTGGGAAGAAAAT GTTACTGAACTATTGACTGAAAGTAAATTTAGATAAAATACAGCTTTTTTCCTTATGGGCATTTGTTTTGTTTCAAGTCATCATAAACTA GGTATTGCATTGCTATCCGTGGATACAGACGCTTAGCTCTTAAAAGATTTTTTTTTTATGTAAACTGTTGAATATTTGAAATAGTCCACT TCACCTTAATGGGTCTTGTCTATCTTCATTAGTCTTCAAAGAAAAACCATTTGCTACCAAAGTAAATCAGTATTTTGAATGTGCTTCTCT TGTTTTTTGTTTATTAGCTAGTTCCTGTAAGCATTTCCACCAGAACTTGAGGCAAATCGTAAGGAAGCTGTTTCTTTTAAAACACAAACC ACCACCAAAAATTTAAATGTACATATTGCTTAAGTATTTGGCTGTTTTTATTTTTTAAAAGGTATAAACACCAAAAAAAAAATTAACATT GTATGAAGATGGAAAATAAGAAGATGCACTTTCTGTAACTTTGTCTAAGGATTTAAATTACTAACTTATGAACTCCAATTTGAATTGAAC TTAACTATCGGCTTTCTTACTGGTAAAATTATATGGTTTATTTTAAATGCGTACATATTGACCAATGGCCTCTGAAAAAGCACATTTTAG ATACTGAAATTGAAGGAAAGAAAATGCATCTTCAAACATTTTTTGGAATCTCACCACATATACTTTGTTAGATTTGTGTATTGTAGGGTG TTTGTTTTGTATTTTTGTATTGTATATGAACTTTTTTTAAATGTGACAGTTAAACACATCTTTAAAAGCATAGTCACAGACAAAAGCATA CAGTATAAAAATTTCCTTGAAAACTCCTACAATATTATATTTGGAGGCAGCTTCAGACTGTTTTATTGGTGGTAGCTGCTTGCTGAGGTC TTTTAGTTGGTAATAACTCCAGAGAAGCAGCCTGTGTATATTCCTAACACTTTGTTCACTAGCATTTAAGTTTAGAATAAGCCCAAGTAA GACAATGGAAATGTATATAGAACTCTTAGTTCTTACATGATTTAATTATATCGATACATGAATTTAACTTACTTTAATGTAGGCAAACTA TCAATTTTTTGTCCATTTTCCTGTTTGTTAAAATAACATACCTCTCCTACGTATTATTTTCTTGACCCAAATGAAATATTAACCTAAGGT >36776_36776_1_HMCES-PCNP_HMCES_chr3_128998758_ENST00000383463_PCNP_chr3_101309059_ENST00000265260_length(amino acids)=121AA_BP=61 MCGRTSCHLPRDVLTRACAYQDRRGQQRLPEWRDPDKYCPSYNKSPQSNSPVLLSRLHFEKSEPEEMPPEAKMRMKNIGRDTPTSAGPNS -------------------------------------------------------------- >36776_36776_2_HMCES-PCNP_HMCES_chr3_128998758_ENST00000389735_PCNP_chr3_101309059_ENST00000265260_length(transcript)=2311nt_BP=444nt GCGAGCTGACAGAGGGGAGCAGAGGCCCGAGCGGACGCGAGGCGACGCGGAGAGGGCGGCCTGGGCCGAAGTGAGGCGACGCGGGGCGGC GGAGTCTGAGGGCCGGAGCGGAGCGGAGGCGACGCGGGGAGGGGCGAGGGATCGCGGCCGGTGGCTGGGGGCCACCTGCTCCCCGAGCAC CGGCCCCCGCTCCGGGGAGCAGAGTCCGGAGCGGGATCCGCGGCCCACAGGTTCGCAGGTTGCGAGGGGCGGTGTTGAAGAATGTGTGGG CGAACATCCTGTCACTTACCTAGAGATGTTCTCACGAGAGCTTGCGCCTACCAGGATCGGCGGGGCCAGCAGCGGCTCCCGGAGTGGAGG GACCCTGATAAGTACTGCCCCTCTTACAACAAGAGTCCTCAATCCAACAGCCCAGTGCTTCTGTCTCGACTGCACTTTGAGAAGAGTGAA CCAGAGGAAATGCCTCCAGAAGCAAAGATGAGGATGAAGAATATTGGAAGGGATACACCAACATCAGCTGGACCAAACTCCTTCAATAAA GGAAAGCATGGGTTTTCTGATAACCAGAAGCTGTGGGAGCGAAATATAAAATCTCATCTTGGAAATGTCCATGACCAAGACAATTAAATG ATGTTTTGAAATTGGGGTGTGGGGTGGGTGTAAAGTTAAAAGGAACAGTTTCCTTTTTTAAAGAATGGTATAAGACTATCTTTGGAGCCG CTTTTTTTTTCTTTTTCATTTTTTTAAAAGATTGAGTGGTACACTAATAAATGAGAGTTTGAAATTAGAGGTAATTTATGTTTTATATAC AGATTTCAAGACATTTGCTAATTTTGTAGTTTCATGTGATTAGTTTCCAAAGGTTACAGATAATAAAGAAATCAGAAATGGTACCTTTTT AAGAATTGCATATTTTTTTAGACACAACTATTAGTACATTAAGAGGGAAGCAAAGTTACTGTCTATTTAAAACTGCAAGCAGTTAACTCT CTTAACTCCCTTATTACCTAAACTTGTCTGGCTCCCAGGAACAGCCTTATAGAGAGAGGGAGTATTGTATTGGGAAGAAAATGTTACTGA ACTATTGACTGAAAGTAAATTTAGATAAAATACAGCTTTTTTCCTTATGGGCATTTGTTTTGTTTCAAGTCATCATAAACTAGGTATTGC ATTGCTATCCGTGGATACAGACGCTTAGCTCTTAAAAGATTTTTTTTTTATGTAAACTGTTGAATATTTGAAATAGTCCACTTCACCTTA ATGGGTCTTGTCTATCTTCATTAGTCTTCAAAGAAAAACCATTTGCTACCAAAGTAAATCAGTATTTTGAATGTGCTTCTCTTGTTTTTT GTTTATTAGCTAGTTCCTGTAAGCATTTCCACCAGAACTTGAGGCAAATCGTAAGGAAGCTGTTTCTTTTAAAACACAAACCACCACCAA AAATTTAAATGTACATATTGCTTAAGTATTTGGCTGTTTTTATTTTTTAAAAGGTATAAACACCAAAAAAAAAATTAACATTGTATGAAG ATGGAAAATAAGAAGATGCACTTTCTGTAACTTTGTCTAAGGATTTAAATTACTAACTTATGAACTCCAATTTGAATTGAACTTAACTAT CGGCTTTCTTACTGGTAAAATTATATGGTTTATTTTAAATGCGTACATATTGACCAATGGCCTCTGAAAAAGCACATTTTAGATACTGAA ATTGAAGGAAAGAAAATGCATCTTCAAACATTTTTTGGAATCTCACCACATATACTTTGTTAGATTTGTGTATTGTAGGGTGTTTGTTTT GTATTTTTGTATTGTATATGAACTTTTTTTAAATGTGACAGTTAAACACATCTTTAAAAGCATAGTCACAGACAAAAGCATACAGTATAA AAATTTCCTTGAAAACTCCTACAATATTATATTTGGAGGCAGCTTCAGACTGTTTTATTGGTGGTAGCTGCTTGCTGAGGTCTTTTAGTT GGTAATAACTCCAGAGAAGCAGCCTGTGTATATTCCTAACACTTTGTTCACTAGCATTTAAGTTTAGAATAAGCCCAAGTAAGACAATGG AAATGTATATAGAACTCTTAGTTCTTACATGATTTAATTATATCGATACATGAATTTAACTTACTTTAATGTAGGCAAACTATCAATTTT TTGTCCATTTTCCTGTTTGTTAAAATAACATACCTCTCCTACGTATTATTTTCTTGACCCAAATGAAATATTAACCTAAGGTCAAGCTGG >36776_36776_2_HMCES-PCNP_HMCES_chr3_128998758_ENST00000389735_PCNP_chr3_101309059_ENST00000265260_length(amino acids)=121AA_BP=61 MCGRTSCHLPRDVLTRACAYQDRRGQQRLPEWRDPDKYCPSYNKSPQSNSPVLLSRLHFEKSEPEEMPPEAKMRMKNIGRDTPTSAGPNS -------------------------------------------------------------- >36776_36776_3_HMCES-PCNP_HMCES_chr3_128998758_ENST00000417226_PCNP_chr3_101309059_ENST00000265260_length(transcript)=2133nt_BP=266nt AAGCGGAGCGGAGCGGGGGTGAGGAGAGTCGAGGGAGGTGACGCGCGCTGCCGGGGCGAGGTTGCGAGGGGCGGTGTTGAAGAATGTGTG GGCGAACATCCTGTCACTTACCTAGAGATGTTCTCACGAGAGCTTGCGCCTACCAGGATCGGCGGGGCCAGCAGCGGCTCCCGGAGTGGA GGGACCCTGATAAGTACTGCCCCTCTTACAACAAGAGTCCTCAATCCAACAGCCCAGTGCTTCTGTCTCGACTGCACTTTGAGAAGAGTG AACCAGAGGAAATGCCTCCAGAAGCAAAGATGAGGATGAAGAATATTGGAAGGGATACACCAACATCAGCTGGACCAAACTCCTTCAATA AAGGAAAGCATGGGTTTTCTGATAACCAGAAGCTGTGGGAGCGAAATATAAAATCTCATCTTGGAAATGTCCATGACCAAGACAATTAAA TGATGTTTTGAAATTGGGGTGTGGGGTGGGTGTAAAGTTAAAAGGAACAGTTTCCTTTTTTAAAGAATGGTATAAGACTATCTTTGGAGC CGCTTTTTTTTTCTTTTTCATTTTTTTAAAAGATTGAGTGGTACACTAATAAATGAGAGTTTGAAATTAGAGGTAATTTATGTTTTATAT ACAGATTTCAAGACATTTGCTAATTTTGTAGTTTCATGTGATTAGTTTCCAAAGGTTACAGATAATAAAGAAATCAGAAATGGTACCTTT TTAAGAATTGCATATTTTTTTAGACACAACTATTAGTACATTAAGAGGGAAGCAAAGTTACTGTCTATTTAAAACTGCAAGCAGTTAACT CTCTTAACTCCCTTATTACCTAAACTTGTCTGGCTCCCAGGAACAGCCTTATAGAGAGAGGGAGTATTGTATTGGGAAGAAAATGTTACT GAACTATTGACTGAAAGTAAATTTAGATAAAATACAGCTTTTTTCCTTATGGGCATTTGTTTTGTTTCAAGTCATCATAAACTAGGTATT GCATTGCTATCCGTGGATACAGACGCTTAGCTCTTAAAAGATTTTTTTTTTATGTAAACTGTTGAATATTTGAAATAGTCCACTTCACCT TAATGGGTCTTGTCTATCTTCATTAGTCTTCAAAGAAAAACCATTTGCTACCAAAGTAAATCAGTATTTTGAATGTGCTTCTCTTGTTTT TTGTTTATTAGCTAGTTCCTGTAAGCATTTCCACCAGAACTTGAGGCAAATCGTAAGGAAGCTGTTTCTTTTAAAACACAAACCACCACC AAAAATTTAAATGTACATATTGCTTAAGTATTTGGCTGTTTTTATTTTTTAAAAGGTATAAACACCAAAAAAAAAATTAACATTGTATGA AGATGGAAAATAAGAAGATGCACTTTCTGTAACTTTGTCTAAGGATTTAAATTACTAACTTATGAACTCCAATTTGAATTGAACTTAACT ATCGGCTTTCTTACTGGTAAAATTATATGGTTTATTTTAAATGCGTACATATTGACCAATGGCCTCTGAAAAAGCACATTTTAGATACTG AAATTGAAGGAAAGAAAATGCATCTTCAAACATTTTTTGGAATCTCACCACATATACTTTGTTAGATTTGTGTATTGTAGGGTGTTTGTT TTGTATTTTTGTATTGTATATGAACTTTTTTTAAATGTGACAGTTAAACACATCTTTAAAAGCATAGTCACAGACAAAAGCATACAGTAT AAAAATTTCCTTGAAAACTCCTACAATATTATATTTGGAGGCAGCTTCAGACTGTTTTATTGGTGGTAGCTGCTTGCTGAGGTCTTTTAG TTGGTAATAACTCCAGAGAAGCAGCCTGTGTATATTCCTAACACTTTGTTCACTAGCATTTAAGTTTAGAATAAGCCCAAGTAAGACAAT GGAAATGTATATAGAACTCTTAGTTCTTACATGATTTAATTATATCGATACATGAATTTAACTTACTTTAATGTAGGCAAACTATCAATT TTTTGTCCATTTTCCTGTTTGTTAAAATAACATACCTCTCCTACGTATTATTTTCTTGACCCAAATGAAATATTAACCTAAGGTCAAGCT >36776_36776_3_HMCES-PCNP_HMCES_chr3_128998758_ENST00000417226_PCNP_chr3_101309059_ENST00000265260_length(amino acids)=121AA_BP=61 MCGRTSCHLPRDVLTRACAYQDRRGQQRLPEWRDPDKYCPSYNKSPQSNSPVLLSRLHFEKSEPEEMPPEAKMRMKNIGRDTPTSAGPNS -------------------------------------------------------------- >36776_36776_4_HMCES-PCNP_HMCES_chr3_128998758_ENST00000502878_PCNP_chr3_101309059_ENST00000265260_length(transcript)=2321nt_BP=454nt GAGGGGAAAGGAAGCGACGCGAGCTGACAGAGGGGAGCAGAGGCCCGAGCGGACGCGAGGCGACGCGGAGAGGGCGGCCTGGGCCGAAGT GAGGCGACGCGGGGCGGCGGAGTCTGAGGGCCGGAGCGGAGCGGAGGCGACGCGGGGAGGGGCGAGGGATCGCGGCCGGTGGCTGGGGGC CACCTGCTCCCCGAGCACCGGCCCCCGCTCCGGGGAGCAGAGTCCGGAGCGGGATCCGCGGCCCACAGGTTGCGAGGGGCGGTGTTGAAG AATGTGTGGGCGAACATCCTGTCACTTACCTAGAGATGTTCTCACGAGAGCTTGCGCCTACCAGGATCGGCGGGGCCAGCAGCGGCTCCC GGAGTGGAGGGACCCTGATAAGTACTGCCCCTCTTACAACAAGAGTCCTCAATCCAACAGCCCAGTGCTTCTGTCTCGACTGCACTTTGA GAAGAGTGAACCAGAGGAAATGCCTCCAGAAGCAAAGATGAGGATGAAGAATATTGGAAGGGATACACCAACATCAGCTGGACCAAACTC CTTCAATAAAGGAAAGCATGGGTTTTCTGATAACCAGAAGCTGTGGGAGCGAAATATAAAATCTCATCTTGGAAATGTCCATGACCAAGA CAATTAAATGATGTTTTGAAATTGGGGTGTGGGGTGGGTGTAAAGTTAAAAGGAACAGTTTCCTTTTTTAAAGAATGGTATAAGACTATC TTTGGAGCCGCTTTTTTTTTCTTTTTCATTTTTTTAAAAGATTGAGTGGTACACTAATAAATGAGAGTTTGAAATTAGAGGTAATTTATG TTTTATATACAGATTTCAAGACATTTGCTAATTTTGTAGTTTCATGTGATTAGTTTCCAAAGGTTACAGATAATAAAGAAATCAGAAATG GTACCTTTTTAAGAATTGCATATTTTTTTAGACACAACTATTAGTACATTAAGAGGGAAGCAAAGTTACTGTCTATTTAAAACTGCAAGC AGTTAACTCTCTTAACTCCCTTATTACCTAAACTTGTCTGGCTCCCAGGAACAGCCTTATAGAGAGAGGGAGTATTGTATTGGGAAGAAA ATGTTACTGAACTATTGACTGAAAGTAAATTTAGATAAAATACAGCTTTTTTCCTTATGGGCATTTGTTTTGTTTCAAGTCATCATAAAC TAGGTATTGCATTGCTATCCGTGGATACAGACGCTTAGCTCTTAAAAGATTTTTTTTTTATGTAAACTGTTGAATATTTGAAATAGTCCA CTTCACCTTAATGGGTCTTGTCTATCTTCATTAGTCTTCAAAGAAAAACCATTTGCTACCAAAGTAAATCAGTATTTTGAATGTGCTTCT CTTGTTTTTTGTTTATTAGCTAGTTCCTGTAAGCATTTCCACCAGAACTTGAGGCAAATCGTAAGGAAGCTGTTTCTTTTAAAACACAAA CCACCACCAAAAATTTAAATGTACATATTGCTTAAGTATTTGGCTGTTTTTATTTTTTAAAAGGTATAAACACCAAAAAAAAAATTAACA TTGTATGAAGATGGAAAATAAGAAGATGCACTTTCTGTAACTTTGTCTAAGGATTTAAATTACTAACTTATGAACTCCAATTTGAATTGA ACTTAACTATCGGCTTTCTTACTGGTAAAATTATATGGTTTATTTTAAATGCGTACATATTGACCAATGGCCTCTGAAAAAGCACATTTT AGATACTGAAATTGAAGGAAAGAAAATGCATCTTCAAACATTTTTTGGAATCTCACCACATATACTTTGTTAGATTTGTGTATTGTAGGG TGTTTGTTTTGTATTTTTGTATTGTATATGAACTTTTTTTAAATGTGACAGTTAAACACATCTTTAAAAGCATAGTCACAGACAAAAGCA TACAGTATAAAAATTTCCTTGAAAACTCCTACAATATTATATTTGGAGGCAGCTTCAGACTGTTTTATTGGTGGTAGCTGCTTGCTGAGG TCTTTTAGTTGGTAATAACTCCAGAGAAGCAGCCTGTGTATATTCCTAACACTTTGTTCACTAGCATTTAAGTTTAGAATAAGCCCAAGT AAGACAATGGAAATGTATATAGAACTCTTAGTTCTTACATGATTTAATTATATCGATACATGAATTTAACTTACTTTAATGTAGGCAAAC TATCAATTTTTTGTCCATTTTCCTGTTTGTTAAAATAACATACCTCTCCTACGTATTATTTTCTTGACCCAAATGAAATATTAACCTAAG >36776_36776_4_HMCES-PCNP_HMCES_chr3_128998758_ENST00000502878_PCNP_chr3_101309059_ENST00000265260_length(amino acids)=125AA_BP= MRAGAERRRRGEGRGIAAGGWGPPAPRAPAPAPGSRVRSGIRGPQVARGGVEECVGEHPVTYLEMFSRELAPTRIGGASSGSRSGGTLIS -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for HMCES-PCNP |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for HMCES-PCNP |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for HMCES-PCNP |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
