
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:HNRNPA2B1-CALM1 (FusionGDB2 ID:37063) |
Fusion Gene Summary for HNRNPA2B1-CALM1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: HNRNPA2B1-CALM1 | Fusion gene ID: 37063 | Hgene | Tgene | Gene symbol | HNRNPA2B1 | CALM1 | Gene ID | 3181 | 801 |
| Gene name | heterogeneous nuclear ribonucleoprotein A2/B1 | calmodulin 1 | |
| Synonyms | HNRNPA2|HNRNPB1|HNRPA2|HNRPA2B1|HNRPB1|IBMPFD2|RNPA2|SNRPB1 | CALML2|CAM2|CAM3|CAMB|CAMC|CAMI|CAMIII|CPVT4|DD132|LQT14|PHKD|caM | |
| Cytomap | 7p15.2 | 14q32.11 | |
| Type of gene | protein-coding | protein-coding | |
| Description | heterogeneous nuclear ribonucleoproteins A2/B1HNRNPA2B1/MYC fusionepididymis secretory sperm binding proteinhnRNP A2 / hnRNP B1nuclear ribonucleoprotein particle A2 protein | calmodulin-1Calmodulin-2Calmodulin-3calmodulin 1 (phosphorylase kinase, delta)phosphorylase kinase subunit deltaphosphorylase kinase, delta subunitprepro-calmodulin 1 | |
| Modification date | 20200314 | 20200329 | |
| UniProtAcc | P22626 | P0DP23 | |
| Ensembl transtripts involved in fusion gene | ENST00000354667, ENST00000356674, ENST00000476233, | ENST00000544280, ENST00000553542, ENST00000555132, ENST00000356978, ENST00000447653, | |
| Fusion gene scores | * DoF score | 43 X 37 X 14=22274 | 13 X 14 X 3=546 |
| # samples | 50 | 15 | |
| ** MAII score | log2(50/22274*10)=-5.47728875772656 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(15/546*10)=-1.86393845042397 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: HNRNPA2B1 [Title/Abstract] AND CALM1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | HNRNPA2B1(26240192)-CALM1(90866409), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | HNRNPA2B1 | GO:0006397 | mRNA processing | 2557628 |
| Hgene | HNRNPA2B1 | GO:0006406 | mRNA export from nucleus | 10567417 |
| Hgene | HNRNPA2B1 | GO:0031053 | primary miRNA processing | 26321680 |
| Hgene | HNRNPA2B1 | GO:0050658 | RNA transport | 17004321 |
| Hgene | HNRNPA2B1 | GO:1990428 | miRNA transport | 24356509 |
| Tgene | CALM1 | GO:0010880 | regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum | 20226167 |
| Tgene | CALM1 | GO:0032516 | positive regulation of phosphoprotein phosphatase activity | 8631777 |
| Tgene | CALM1 | GO:0035307 | positive regulation of protein dephosphorylation | 8631777 |
| Tgene | CALM1 | GO:0051343 | positive regulation of cyclic-nucleotide phosphodiesterase activity | 8631777 |
| Tgene | CALM1 | GO:0051592 | response to calcium ion | 7607248 |
| Tgene | CALM1 | GO:0060314 | regulation of ryanodine-sensitive calcium-release channel activity | 22067155 |
| Tgene | CALM1 | GO:0060315 | negative regulation of ryanodine-sensitive calcium-release channel activity | 26164367 |
| Tgene | CALM1 | GO:0060316 | positive regulation of ryanodine-sensitive calcium-release channel activity | 20226167 |
 Fusion gene breakpoints across HNRNPA2B1 (5'-gene) Fusion gene breakpoints across HNRNPA2B1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
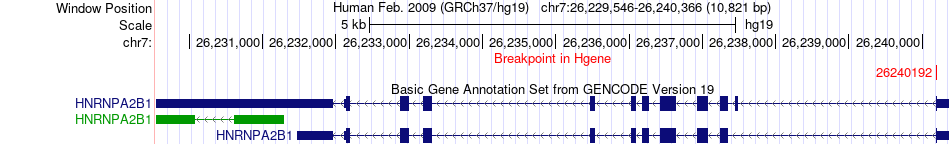 |
 Fusion gene breakpoints across CALM1 (3'-gene) Fusion gene breakpoints across CALM1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
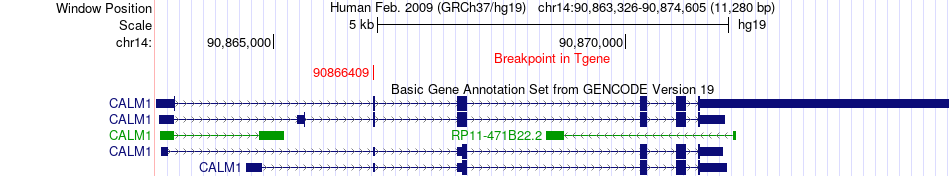 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | ERR315406 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
Top |
Fusion Gene ORF analysis for HNRNPA2B1-CALM1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000354667 | ENST00000544280 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| 5CDS-5UTR | ENST00000354667 | ENST00000553542 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| 5CDS-5UTR | ENST00000356674 | ENST00000544280 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| 5CDS-5UTR | ENST00000356674 | ENST00000553542 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| 5CDS-intron | ENST00000354667 | ENST00000555132 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| 5CDS-intron | ENST00000356674 | ENST00000555132 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| In-frame | ENST00000354667 | ENST00000356978 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| In-frame | ENST00000354667 | ENST00000447653 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| In-frame | ENST00000356674 | ENST00000356978 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| In-frame | ENST00000356674 | ENST00000447653 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| intron-3CDS | ENST00000476233 | ENST00000356978 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| intron-3CDS | ENST00000476233 | ENST00000447653 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| intron-5UTR | ENST00000476233 | ENST00000544280 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| intron-5UTR | ENST00000476233 | ENST00000553542 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
| intron-intron | ENST00000476233 | ENST00000555132 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000354667 | HNRNPA2B1 | chr7 | 26240192 | - | ENST00000356978 | CALM1 | chr14 | 90866409 | + | 4166 | 175 | 169 | 621 | 150 |
| ENST00000354667 | HNRNPA2B1 | chr7 | 26240192 | - | ENST00000447653 | CALM1 | chr14 | 90866409 | + | 977 | 175 | 169 | 621 | 150 |
| ENST00000356674 | HNRNPA2B1 | chr7 | 26240192 | - | ENST00000356978 | CALM1 | chr14 | 90866409 | + | 4166 | 175 | 169 | 621 | 150 |
| ENST00000356674 | HNRNPA2B1 | chr7 | 26240192 | - | ENST00000447653 | CALM1 | chr14 | 90866409 | + | 977 | 175 | 169 | 621 | 150 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000354667 | ENST00000356978 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + | 0.000774509 | 0.9992255 |
| ENST00000354667 | ENST00000447653 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + | 0.001439246 | 0.9985607 |
| ENST00000356674 | ENST00000356978 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + | 0.000774509 | 0.9992255 |
| ENST00000356674 | ENST00000447653 | HNRNPA2B1 | chr7 | 26240192 | - | CALM1 | chr14 | 90866409 | + | 0.001439246 | 0.9985607 |
Top |
Fusion Genomic Features for HNRNPA2B1-CALM1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| HNRNPA2B1 | chr7 | 26240191 | - | CALM1 | chr14 | 90866408 | + | 9.63E-07 | 0.99999905 |
| HNRNPA2B1 | chr7 | 26240191 | - | CALM1 | chr14 | 90866408 | + | 9.63E-07 | 0.99999905 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
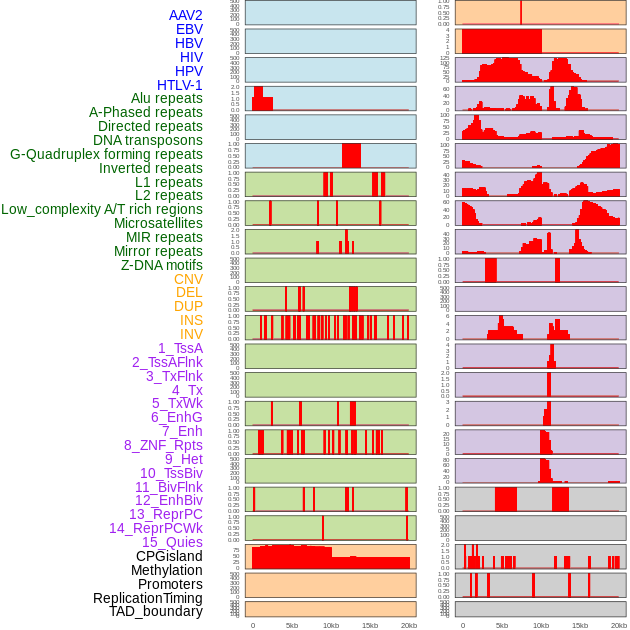 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
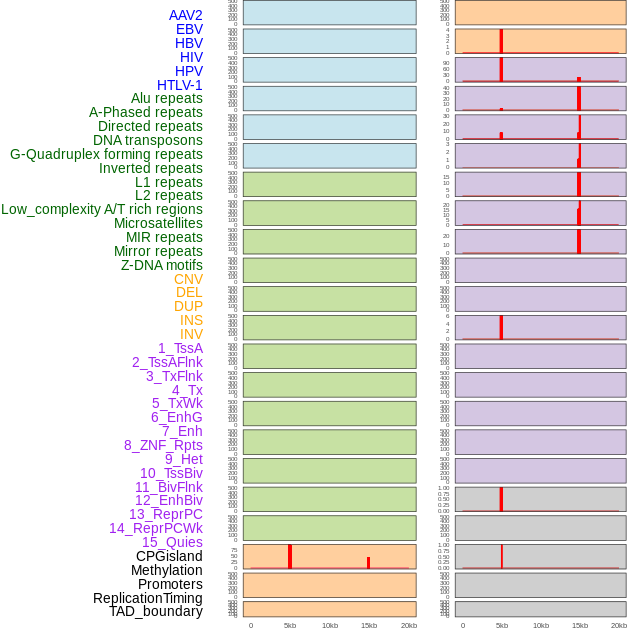 |
Top |
Fusion Protein Features for HNRNPA2B1-CALM1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr7:26240192/chr14:90866409) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| HNRNPA2B1 | CALM1 |
| FUNCTION: Heterogeneous nuclear ribonucleoprotein (hnRNP) that associates with nascent pre-mRNAs, packaging them into hnRNP particles. The hnRNP particle arrangement on nascent hnRNA is non-random and sequence-dependent and serves to condense and stabilize the transcripts and minimize tangling and knotting. Packaging plays a role in various processes such as transcription, pre-mRNA processing, RNA nuclear export, subcellular location, mRNA translation and stability of mature mRNAs (PubMed:19099192). Forms hnRNP particles with at least 20 other different hnRNP and heterogeneous nuclear RNA in the nucleus. Involved in transport of specific mRNAs to the cytoplasm in oligodendrocytes and neurons: acts by specifically recognizing and binding the A2RE (21 nucleotide hnRNP A2 response element) or the A2RE11 (derivative 11 nucleotide oligonucleotide) sequence motifs present on some mRNAs, and promotes their transport to the cytoplasm (PubMed:10567417). Specifically binds single-stranded telomeric DNA sequences, protecting telomeric DNA repeat against endonuclease digestion (By similarity). Also binds other RNA molecules, such as primary miRNA (pri-miRNAs): acts as a nuclear 'reader' of the N6-methyladenosine (m6A) mark by specifically recognizing and binding a subset of nuclear m6A-containing pri-miRNAs. Binding to m6A-containing pri-miRNAs promotes pri-miRNA processing by enhancing binding of DGCR8 to pri-miRNA transcripts (PubMed:26321680). Involved in miRNA sorting into exosomes following sumoylation, possibly by binding (m6A)-containing pre-miRNAs (PubMed:24356509). Acts as a regulator of efficiency of mRNA splicing, possibly by binding to m6A-containing pre-mRNAs (PubMed:26321680). Plays also a role in the activation of the innate immune response (PubMed:31320558). Mechanistically, senses the presence of viral DNA in the nucleus, homodimerizes and is demethylated by JMJD6 (PubMed:31320558). In turn, translocates to the cytoplasm where it activates the TBK1-IRF3 pathway, leading to interferon alpha/beta production (PubMed:31320558). {ECO:0000250|UniProtKB:A7VJC2, ECO:0000269|PubMed:10567417, ECO:0000269|PubMed:24356509, ECO:0000269|PubMed:26321680, ECO:0000303|PubMed:19099192}.; FUNCTION: (Microbial infection) Involved in the transport of HIV-1 genomic RNA out of the nucleus, to the microtubule organizing center (MTOC), and then from the MTOC to the cytoplasm: acts by specifically recognizing and binding the A2RE (21 nucleotide hnRNP A2 response element) sequence motifs present on HIV-1 genomic RNA, and promotes its transport. {ECO:0000269|PubMed:15294897, ECO:0000269|PubMed:17004321}. | FUNCTION: Calmodulin mediates the control of a large number of enzymes, ion channels, aquaporins and other proteins through calcium-binding. Among the enzymes to be stimulated by the calmodulin-calcium complex are a number of protein kinases and phosphatases. Together with CCP110 and centrin, is involved in a genetic pathway that regulates the centrosome cycle and progression through cytokinesis (PubMed:16760425). Is a regulator of voltage-dependent L-type calcium channels (PubMed:31454269). Mediates calcium-dependent inactivation of CACNA1C (PubMed:26969752). Positively regulates calcium-activated potassium channel activity of KCNN2 (PubMed:27165696). Forms a potassium channel complex with KCNQ1 and regulates electrophysiological activity of the channel via calcium-binding (PubMed:25441029). Acts as a sensor to modulate the endomplasmic reticulum contacts with other organelles mediated by VMP1:ATP2A2 (PubMed:28890335). {ECO:0000269|PubMed:16760425, ECO:0000269|PubMed:23893133, ECO:0000269|PubMed:26969752, ECO:0000269|PubMed:27165696, ECO:0000269|PubMed:28890335, ECO:0000269|PubMed:31454269}.; FUNCTION: (Microbial infection) Required for Legionella pneumophila SidJ glutamylase activity. {ECO:0000269|PubMed:31330532}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000354667 | - | 1 | 12 | 202_353 | 2 | 1115.3333333333333 | Compositional bias | Note=Gly-rich |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000356674 | - | 1 | 11 | 202_353 | 2 | 439.3333333333333 | Compositional bias | Note=Gly-rich |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000354667 | - | 1 | 12 | 112_191 | 2 | 1115.3333333333333 | Domain | RRM 2 |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000354667 | - | 1 | 12 | 21_104 | 2 | 1115.3333333333333 | Domain | RRM 1 |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000356674 | - | 1 | 11 | 112_191 | 2 | 439.3333333333333 | Domain | RRM 2 |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000356674 | - | 1 | 11 | 21_104 | 2 | 439.3333333333333 | Domain | RRM 1 |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000354667 | - | 1 | 12 | 9_15 | 2 | 1115.3333333333333 | Motif | Nuclear localization signal |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000356674 | - | 1 | 11 | 9_15 | 2 | 439.3333333333333 | Motif | Nuclear localization signal |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000354667 | - | 1 | 12 | 193_353 | 2 | 1115.3333333333333 | Region | Low complexity (LC) region |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000354667 | - | 1 | 12 | 308_347 | 2 | 1115.3333333333333 | Region | Nuclear targeting sequence |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000356674 | - | 1 | 11 | 193_353 | 2 | 439.3333333333333 | Region | Low complexity (LC) region |
| Hgene | HNRNPA2B1 | chr7:26240192 | chr14:90866409 | ENST00000356674 | - | 1 | 11 | 308_347 | 2 | 439.3333333333333 | Region | Nuclear targeting sequence |
Top |
Fusion Gene Sequence for HNRNPA2B1-CALM1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >37063_37063_1_HNRNPA2B1-CALM1_HNRNPA2B1_chr7_26240192_ENST00000354667_CALM1_chr14_90866409_ENST00000356978_length(transcript)=4166nt_BP=175nt AGTAGCAGCAGCGCCGGGTCCCGTGCGGAGGTGCTCCTCGCAGAGTTGTTTCTCGAGCAGCGGCAGTTCTCACTACAGCGCCAGGACGAG TCCGGTTCGTGTTCGTCCGCGGAGATCTCTCTCATCTCGCTCGGCTGCGGGAAATCGGGCTGAAGCGACTGAGTCCGCGATGGAGGCTGA TCAGCTGACCGAAGAACAGATTGCTGAATTCAAGGAAGCCTTCTCCCTATTTGATAAAGATGGCGATGGCACCATCACAACAAAGGAACT TGGAACTGTCATGAGGTCACTGGGTCAGAACCCAACAGAAGCTGAATTGCAGGATATGATCAATGAAGTGGATGCTGATGGTAATGGCAC CATTGACTTCCCCGAATTTTTGACTATGATGGCTAGAAAAATGAAAGATACAGATAGTGAAGAAGAAATCCGTGAGGCATTCCGAGTCTT TGACAAGGATGGCAATGGTTATATCAGTGCAGCAGAACTACGTCACGTCATGACAAACTTAGGAGAAAAACTAACAGATGAAGAAGTAGA TGAAATGATCAGAGAAGCAGATATTGATGGAGACGGACAAGTCAACTATGAAGAATTCGTACAGATGATGACTGCAAAATGAAGACCTAC TTTCAACTCCTTTTTCCCCCCTCTAGAAGAATCAAATTGAATCTTTTACTTACCTCTTGCAAAAAAAAGAAAAAAGAAAAAAGTTCATTT ATTCATTCTGTTTCTATATAGCAAAACTGAATGTCAAAAGTACCTTCTGTCCACACACACAAAATCTGCATGTATTGGTTGGTGGTCCTG TCCCCTAAAGATCAAGCTACACATCAGTTTTACAATATAAATACTTGTACTACCTTAATGATAAGGACTCCTTAAAGTTCCATTTGCTAA TGATTAATACACTGTTTGGGCTGGCCAGTTTTTCATGCATGCAGCTTGACGATTGAGCACAGTCAGGCCTTTGTATTAAAAATGAAAAAT GAAAAAACAAATTCAAAACCTATTCAAATGGGTTCTAGTTCAATTTGTTTAGTATAAATTGTCATAGCTGGTTTACTGAAAACAAACACA TTTAAAATTGGTTTACCTCAGGATGACGTGCAGAAAAATGGGTGAAGGATAAACCGTTGAGACGTGGCCCCACTGGTAGGATGGTCCTCT TGTACTTCGTGTGCTCCGACCCATGGTGACGATGACACACCCTGGTGGCATGCCCGTGTATGTTGGTTTAGCGTTGTCTGCATTGTTCTA GAGTGAAACAGGTGTCAGGCTGTCACTGTTCACACAAATTTTTAATAAGAAACATTTACCAAGGGAGCATCTTTGGACTCTCTGTTTTTA AAACCTTCTGAACCATGACTTGGAGCCGGCAGAGTAGGCTGTGGCTGTGGACTTCAGCACAACCATCAACATTGCTGTTCAAAGAAATTA CAGTTTACGTCCATTCCAAGTTGTAAATGCTAGTCTTTTTTTTTTTTTTTCCAATAAAAAGACCATTAACTTAAAGTGGTGTTAAATGCT TTGTAAAGCTGAGATCTAAATGGGGACAAGGCAGGTGGAGGGGAGGCCAGTGTACATGTAAATGCCCACAGCCCAGCATTGGGTTTCCCT CCCAAGGCCCCAGCACCAACCTCTGAGCCCAAGACCTTGCCTGAAAACAAGCAGATACCGATTGCTTCATCCTATTTATGGACATGTAGG TCTAGTTGCATTTTCACTGGGGGGAGGGGGGAAGGTGAATTATGGTAACTTTTAATGATCTATTCAGGCAGTAGAGCTCTTAAGGAAAAA AAAAAACCCACTTTCTCTCAAGCATGTATTTAGGGGTTGTTCTCAATTGTGCTGCTGATTACCTGTCTTATGTAACTACTTGAGACCATC TGCAAGAGACATGATTTAGTGTGTCTGTAATTCAATCTTCGCTGTGTGTGGTAGAAGCAGTAGTCACTTTTGTAAGCCAGTCTCTTCATG CCTAAAAGACACTACCAGTCACCTTTGATTCGCGACTTTTAATTTATGATTATACTTAGCCTCCTCCTCCTTTTTTTTTTTTTCCCAAGT TGACTTGACTTTGCTTTTTTCCCCCCAAGTAGAACTAATGCTAGCTTCCAGCTTGAAAGTAAAACTCCAGTGTGGAGTGAATTTTGTGTC TAATTATAAACCTGTAACCAAAACTCAGACATCTGGTACTGGTCTTTGCATTGAGATTGGTCCCTGTAAAACCCCCTTTAAAAGCATATT GCATTTAGTACAGAGCTCTTTTTTGAAATGAAGGCTGGAGATGTGCATTTTTCACGGTGTTAACTGGTTGTATCTTATTAGCAAGGAGAT TGGGGTTTTGAGTGTTTGCGTGGGTGGTTTCAATTTGCCAGGGAACAGTGGCAGGCTGCTAGCAAGGCAGTGAGAAGCTCTTGGCAGCCA AATGGGTGCATTCAGGGCTGATTTATAGAGACCCTTGGCTTCTCCTTCTCCTACTCCCTGTCTTTCTGGCATTTTGTAGCTTGTTAGATT TTCTGCCAGAGGGGTGGGTCAGAGCAGTGGAGGGGAGACATCGCCCATGTGCTTCTGCTACTGGTCCTTGGGCTGGGTGGTTGGTAGAGG AGATGTTGACACTATGAGCTAAGGGTTGGCTTTTGTAATTACCTGAATCTGAAAGGAATGCCTAAGGTTACCTTGGGGTTTCTCTTCTGG TGAGATAGGGTTCCTGGTTTGAGTAAGTTAATGTCCTGGATATTTCTTGTGGCAGGGGGTGGTCAAAGAGCCTGATTGCTGACCCAGTCT CAGGCCTGTGGTCGATGACCTCTCGGTAGTTTCAAAGGGGGCTGGAGGGGGATATTTGACTTGTTTTTTCGAAATGTAGCCTTCTAACCC TCAAGTCTTTAGAAGCTGGGTGGACTCTTAGTGGTCCTGCAGCGTATCCTAAAAGACTACCTTTGAAACAGGATTCTTGTATGGCCAGGA TCCTGTCTGGGAACCAGAAACCCTACACCCTCCCCCTCCAGGGAATGCTGAGTTCCAGTTTTGAGCAGAGGTGAGGCAGAATCCACTGTA GCCTTCCGCCCTGGTATTTGGGGGGATGACCAGCCCAGGCGTTGGGTGTTAGTCTGCATGAGTTTGTGAGAGGAAATAGCTGGGTGTCCT GGCAGTGCCCTTGAAGTTGGTTAGGACCTTCCTGTAAACTCTTGCCCCTACTTCTAACTACTCTATAAATATATACATATATTTATATAT AAAGTGATTAGTTGAACTGGCATCCTGCTTTAGCCTGAGACTTGCCATAAGAAACTGCTGAGTACTTGGCAAACCCTTTCATAGTTTTGT TCTCCATCTGTTTGGGGTAGGTGTTGAGCGAGGCAAATGGATCTCGATATTTCAGATGGGCTTTTGATGCACTGTTGCCAAGGAAGGCTT TTTCTGATTTTTTGACAAATGAATTTTTGCACACTTTCATTGGTGTCTTTCGGCAACTTACACACATTGAAAATGAGCTATTGTACATAT TTTTATATTCTCTTTATAAATGCATGTCTGATTGTACTTGTAACAATATTGTAATGAACGGCTGTGCAGTAGGCCCAGCGCTGCTGTGTC TCGTCAGAGGAATAGCTTACCACGAACCCCTCAGCATACTGGGAATCTCTTCCTGAACAACGAATGTAAATTTGGTCAAGTCTACTCTTC CGTTCATTCAATTATTTTAAGCATTTGAATTATTTATTGTATATCCTAAATATATTTCTCCTTTGGCAGTGACTAGATTTCCACTAATGT GTCTTAATCTATCCCTCCAGCTGGCAGTTACTGTTTTTTTAATCCCCTGAAGTTGTCCTGTAGGAGACAGAAATTCTTTGCTGTCTGTAT CCCTTGGAGTAAGAAGGTAGTGGCATGGGTGGAGTGTGTGTTCTTTCTCCAAATCTATTATGATGTTTATTAAACACTTCTGTAGCAAAG ATGGTGGTAGTTCTTTTGTTACTGAAGTTGCCCTTCACCATGGCTATTTGAAAAGGAGATGTACTTGGACGTTTCTGTAAATCTTGAGAT AAACTGTTTGGAGATTTAACCACCTCTCTGATGGGGGACCAACTCTATGGAAATTGTAAATACGTTTTATTTATAAACCTGGCACTGTAT >37063_37063_1_HNRNPA2B1-CALM1_HNRNPA2B1_chr7_26240192_ENST00000354667_CALM1_chr14_90866409_ENST00000356978_length(amino acids)=150AA_BP=2 MEADQLTEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLTMMARKMKDTDSEEEIREA -------------------------------------------------------------- >37063_37063_2_HNRNPA2B1-CALM1_HNRNPA2B1_chr7_26240192_ENST00000354667_CALM1_chr14_90866409_ENST00000447653_length(transcript)=977nt_BP=175nt AGTAGCAGCAGCGCCGGGTCCCGTGCGGAGGTGCTCCTCGCAGAGTTGTTTCTCGAGCAGCGGCAGTTCTCACTACAGCGCCAGGACGAG TCCGGTTCGTGTTCGTCCGCGGAGATCTCTCTCATCTCGCTCGGCTGCGGGAAATCGGGCTGAAGCGACTGAGTCCGCGATGGAGGCTGA TCAGCTGACCGAAGAACAGATTGCTGAATTCAAGGAAGCCTTCTCCCTATTTGATAAAGATGGCGATGGCACCATCACAACAAAGGAACT TGGAACTGTCATGAGGTCACTGGGTCAGAACCCAACAGAAGCTGAATTGCAGGATATGATCAATGAAGTGGATGCTGATGGTAATGGCAC CATTGACTTCCCCGAATTTTTGACTATGATGGCTAGAAAAATGAAAGATACAGATAGTGAAGAAGAAATCCGTGAGGCATTCCGAGTCTT TGACAAGGATGGCAATGGTTATATCAGTGCAGCAGAACTACGTCACGTCATGACAAACTTAGGAGAAAAACTAACAGATGAAGAAGTAGA TGAAATGATCAGAGAAGCAGATATTGATGGAGACGGACAAGTCAACTATGAAGAATTCGTACAGATGATGACTGCAAAATGAAGACCTAC TTTCAACTCCTTTTTCCCCCCTCTAGAAGAATCAAATTGAATCTTTTACTTACCTCTTGCAAAAAAAAGAAAAAAGAAAAAAGTTCATTT ATTCATTCTGTTTCTATATAGCAAAACTGAATGTCAAAAGTACCTTCTGTCCACACACACAAAATCTGCATGTATTGGTTGGTGGTCCTG TCCCCTAAAGATCAAGCTACACATCAGTTTTACAATATAAATACTTGTACTACCTTAATGATAAGGACTCCTTAAAGTTCCATTTGCTAA >37063_37063_2_HNRNPA2B1-CALM1_HNRNPA2B1_chr7_26240192_ENST00000354667_CALM1_chr14_90866409_ENST00000447653_length(amino acids)=150AA_BP=2 MEADQLTEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLTMMARKMKDTDSEEEIREA -------------------------------------------------------------- >37063_37063_3_HNRNPA2B1-CALM1_HNRNPA2B1_chr7_26240192_ENST00000356674_CALM1_chr14_90866409_ENST00000356978_length(transcript)=4166nt_BP=175nt AGTAGCAGCAGCGCCGGGTCCCGTGCGGAGGTGCTCCTCGCAGAGTTGTTTCTCGAGCAGCGGCAGTTCTCACTACAGCGCCAGGACGAG TCCGGTTCGTGTTCGTCCGCGGAGATCTCTCTCATCTCGCTCGGCTGCGGGAAATCGGGCTGAAGCGACTGAGTCCGCGATGGAGGCTGA TCAGCTGACCGAAGAACAGATTGCTGAATTCAAGGAAGCCTTCTCCCTATTTGATAAAGATGGCGATGGCACCATCACAACAAAGGAACT TGGAACTGTCATGAGGTCACTGGGTCAGAACCCAACAGAAGCTGAATTGCAGGATATGATCAATGAAGTGGATGCTGATGGTAATGGCAC CATTGACTTCCCCGAATTTTTGACTATGATGGCTAGAAAAATGAAAGATACAGATAGTGAAGAAGAAATCCGTGAGGCATTCCGAGTCTT TGACAAGGATGGCAATGGTTATATCAGTGCAGCAGAACTACGTCACGTCATGACAAACTTAGGAGAAAAACTAACAGATGAAGAAGTAGA TGAAATGATCAGAGAAGCAGATATTGATGGAGACGGACAAGTCAACTATGAAGAATTCGTACAGATGATGACTGCAAAATGAAGACCTAC TTTCAACTCCTTTTTCCCCCCTCTAGAAGAATCAAATTGAATCTTTTACTTACCTCTTGCAAAAAAAAGAAAAAAGAAAAAAGTTCATTT ATTCATTCTGTTTCTATATAGCAAAACTGAATGTCAAAAGTACCTTCTGTCCACACACACAAAATCTGCATGTATTGGTTGGTGGTCCTG TCCCCTAAAGATCAAGCTACACATCAGTTTTACAATATAAATACTTGTACTACCTTAATGATAAGGACTCCTTAAAGTTCCATTTGCTAA TGATTAATACACTGTTTGGGCTGGCCAGTTTTTCATGCATGCAGCTTGACGATTGAGCACAGTCAGGCCTTTGTATTAAAAATGAAAAAT GAAAAAACAAATTCAAAACCTATTCAAATGGGTTCTAGTTCAATTTGTTTAGTATAAATTGTCATAGCTGGTTTACTGAAAACAAACACA TTTAAAATTGGTTTACCTCAGGATGACGTGCAGAAAAATGGGTGAAGGATAAACCGTTGAGACGTGGCCCCACTGGTAGGATGGTCCTCT TGTACTTCGTGTGCTCCGACCCATGGTGACGATGACACACCCTGGTGGCATGCCCGTGTATGTTGGTTTAGCGTTGTCTGCATTGTTCTA GAGTGAAACAGGTGTCAGGCTGTCACTGTTCACACAAATTTTTAATAAGAAACATTTACCAAGGGAGCATCTTTGGACTCTCTGTTTTTA AAACCTTCTGAACCATGACTTGGAGCCGGCAGAGTAGGCTGTGGCTGTGGACTTCAGCACAACCATCAACATTGCTGTTCAAAGAAATTA CAGTTTACGTCCATTCCAAGTTGTAAATGCTAGTCTTTTTTTTTTTTTTTCCAATAAAAAGACCATTAACTTAAAGTGGTGTTAAATGCT TTGTAAAGCTGAGATCTAAATGGGGACAAGGCAGGTGGAGGGGAGGCCAGTGTACATGTAAATGCCCACAGCCCAGCATTGGGTTTCCCT CCCAAGGCCCCAGCACCAACCTCTGAGCCCAAGACCTTGCCTGAAAACAAGCAGATACCGATTGCTTCATCCTATTTATGGACATGTAGG TCTAGTTGCATTTTCACTGGGGGGAGGGGGGAAGGTGAATTATGGTAACTTTTAATGATCTATTCAGGCAGTAGAGCTCTTAAGGAAAAA AAAAAACCCACTTTCTCTCAAGCATGTATTTAGGGGTTGTTCTCAATTGTGCTGCTGATTACCTGTCTTATGTAACTACTTGAGACCATC TGCAAGAGACATGATTTAGTGTGTCTGTAATTCAATCTTCGCTGTGTGTGGTAGAAGCAGTAGTCACTTTTGTAAGCCAGTCTCTTCATG CCTAAAAGACACTACCAGTCACCTTTGATTCGCGACTTTTAATTTATGATTATACTTAGCCTCCTCCTCCTTTTTTTTTTTTTCCCAAGT TGACTTGACTTTGCTTTTTTCCCCCCAAGTAGAACTAATGCTAGCTTCCAGCTTGAAAGTAAAACTCCAGTGTGGAGTGAATTTTGTGTC TAATTATAAACCTGTAACCAAAACTCAGACATCTGGTACTGGTCTTTGCATTGAGATTGGTCCCTGTAAAACCCCCTTTAAAAGCATATT GCATTTAGTACAGAGCTCTTTTTTGAAATGAAGGCTGGAGATGTGCATTTTTCACGGTGTTAACTGGTTGTATCTTATTAGCAAGGAGAT TGGGGTTTTGAGTGTTTGCGTGGGTGGTTTCAATTTGCCAGGGAACAGTGGCAGGCTGCTAGCAAGGCAGTGAGAAGCTCTTGGCAGCCA AATGGGTGCATTCAGGGCTGATTTATAGAGACCCTTGGCTTCTCCTTCTCCTACTCCCTGTCTTTCTGGCATTTTGTAGCTTGTTAGATT TTCTGCCAGAGGGGTGGGTCAGAGCAGTGGAGGGGAGACATCGCCCATGTGCTTCTGCTACTGGTCCTTGGGCTGGGTGGTTGGTAGAGG AGATGTTGACACTATGAGCTAAGGGTTGGCTTTTGTAATTACCTGAATCTGAAAGGAATGCCTAAGGTTACCTTGGGGTTTCTCTTCTGG TGAGATAGGGTTCCTGGTTTGAGTAAGTTAATGTCCTGGATATTTCTTGTGGCAGGGGGTGGTCAAAGAGCCTGATTGCTGACCCAGTCT CAGGCCTGTGGTCGATGACCTCTCGGTAGTTTCAAAGGGGGCTGGAGGGGGATATTTGACTTGTTTTTTCGAAATGTAGCCTTCTAACCC TCAAGTCTTTAGAAGCTGGGTGGACTCTTAGTGGTCCTGCAGCGTATCCTAAAAGACTACCTTTGAAACAGGATTCTTGTATGGCCAGGA TCCTGTCTGGGAACCAGAAACCCTACACCCTCCCCCTCCAGGGAATGCTGAGTTCCAGTTTTGAGCAGAGGTGAGGCAGAATCCACTGTA GCCTTCCGCCCTGGTATTTGGGGGGATGACCAGCCCAGGCGTTGGGTGTTAGTCTGCATGAGTTTGTGAGAGGAAATAGCTGGGTGTCCT GGCAGTGCCCTTGAAGTTGGTTAGGACCTTCCTGTAAACTCTTGCCCCTACTTCTAACTACTCTATAAATATATACATATATTTATATAT AAAGTGATTAGTTGAACTGGCATCCTGCTTTAGCCTGAGACTTGCCATAAGAAACTGCTGAGTACTTGGCAAACCCTTTCATAGTTTTGT TCTCCATCTGTTTGGGGTAGGTGTTGAGCGAGGCAAATGGATCTCGATATTTCAGATGGGCTTTTGATGCACTGTTGCCAAGGAAGGCTT TTTCTGATTTTTTGACAAATGAATTTTTGCACACTTTCATTGGTGTCTTTCGGCAACTTACACACATTGAAAATGAGCTATTGTACATAT TTTTATATTCTCTTTATAAATGCATGTCTGATTGTACTTGTAACAATATTGTAATGAACGGCTGTGCAGTAGGCCCAGCGCTGCTGTGTC TCGTCAGAGGAATAGCTTACCACGAACCCCTCAGCATACTGGGAATCTCTTCCTGAACAACGAATGTAAATTTGGTCAAGTCTACTCTTC CGTTCATTCAATTATTTTAAGCATTTGAATTATTTATTGTATATCCTAAATATATTTCTCCTTTGGCAGTGACTAGATTTCCACTAATGT GTCTTAATCTATCCCTCCAGCTGGCAGTTACTGTTTTTTTAATCCCCTGAAGTTGTCCTGTAGGAGACAGAAATTCTTTGCTGTCTGTAT CCCTTGGAGTAAGAAGGTAGTGGCATGGGTGGAGTGTGTGTTCTTTCTCCAAATCTATTATGATGTTTATTAAACACTTCTGTAGCAAAG ATGGTGGTAGTTCTTTTGTTACTGAAGTTGCCCTTCACCATGGCTATTTGAAAAGGAGATGTACTTGGACGTTTCTGTAAATCTTGAGAT AAACTGTTTGGAGATTTAACCACCTCTCTGATGGGGGACCAACTCTATGGAAATTGTAAATACGTTTTATTTATAAACCTGGCACTGTAT >37063_37063_3_HNRNPA2B1-CALM1_HNRNPA2B1_chr7_26240192_ENST00000356674_CALM1_chr14_90866409_ENST00000356978_length(amino acids)=150AA_BP=2 MEADQLTEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLTMMARKMKDTDSEEEIREA -------------------------------------------------------------- >37063_37063_4_HNRNPA2B1-CALM1_HNRNPA2B1_chr7_26240192_ENST00000356674_CALM1_chr14_90866409_ENST00000447653_length(transcript)=977nt_BP=175nt AGTAGCAGCAGCGCCGGGTCCCGTGCGGAGGTGCTCCTCGCAGAGTTGTTTCTCGAGCAGCGGCAGTTCTCACTACAGCGCCAGGACGAG TCCGGTTCGTGTTCGTCCGCGGAGATCTCTCTCATCTCGCTCGGCTGCGGGAAATCGGGCTGAAGCGACTGAGTCCGCGATGGAGGCTGA TCAGCTGACCGAAGAACAGATTGCTGAATTCAAGGAAGCCTTCTCCCTATTTGATAAAGATGGCGATGGCACCATCACAACAAAGGAACT TGGAACTGTCATGAGGTCACTGGGTCAGAACCCAACAGAAGCTGAATTGCAGGATATGATCAATGAAGTGGATGCTGATGGTAATGGCAC CATTGACTTCCCCGAATTTTTGACTATGATGGCTAGAAAAATGAAAGATACAGATAGTGAAGAAGAAATCCGTGAGGCATTCCGAGTCTT TGACAAGGATGGCAATGGTTATATCAGTGCAGCAGAACTACGTCACGTCATGACAAACTTAGGAGAAAAACTAACAGATGAAGAAGTAGA TGAAATGATCAGAGAAGCAGATATTGATGGAGACGGACAAGTCAACTATGAAGAATTCGTACAGATGATGACTGCAAAATGAAGACCTAC TTTCAACTCCTTTTTCCCCCCTCTAGAAGAATCAAATTGAATCTTTTACTTACCTCTTGCAAAAAAAAGAAAAAAGAAAAAAGTTCATTT ATTCATTCTGTTTCTATATAGCAAAACTGAATGTCAAAAGTACCTTCTGTCCACACACACAAAATCTGCATGTATTGGTTGGTGGTCCTG TCCCCTAAAGATCAAGCTACACATCAGTTTTACAATATAAATACTTGTACTACCTTAATGATAAGGACTCCTTAAAGTTCCATTTGCTAA >37063_37063_4_HNRNPA2B1-CALM1_HNRNPA2B1_chr7_26240192_ENST00000356674_CALM1_chr14_90866409_ENST00000447653_length(amino acids)=150AA_BP=2 MEADQLTEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFLTMMARKMKDTDSEEEIREA -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for HNRNPA2B1-CALM1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for HNRNPA2B1-CALM1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for HNRNPA2B1-CALM1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
