
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:HNRNPAB-SAMM50 (FusionGDB2 ID:37121) |
Fusion Gene Summary for HNRNPAB-SAMM50 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: HNRNPAB-SAMM50 | Fusion gene ID: 37121 | Hgene | Tgene | Gene symbol | HNRNPAB | SAMM50 | Gene ID | 3182 | 25813 |
| Gene name | heterogeneous nuclear ribonucleoprotein A/B | SAMM50 sorting and assembly machinery component | |
| Synonyms | ABBP1|HNRPAB | CGI-51|OMP85|SAM50|TOB55|TRG-3|YNL026W | |
| Cytomap | 5q35.3 | 22q13.31 | |
| Type of gene | protein-coding | protein-coding | |
| Description | heterogeneous nuclear ribonucleoprotein A/BABBP-1APOBEC1-binding protein 1apobec-1 binding protein 1apolipoprotein B mRNA editing enzyme, catalytic polypeptide 1-binding protein 1hnRNP A/BhnRNP type A/B protein | sorting and assembly machinery component 50 homologsorting and assembly machinery 50kDatransformation-related gene 3 protein | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | Q99729 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000355836, ENST00000358344, ENST00000504898, ENST00000506259, ENST00000506339, ENST00000514633, ENST00000515193, | ENST00000396202, ENST00000493161, ENST00000350028, | |
| Fusion gene scores | * DoF score | 4 X 4 X 3=48 | 11 X 12 X 5=660 |
| # samples | 4 | 13 | |
| ** MAII score | log2(4/48*10)=-0.263034405833794 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(13/660*10)=-2.34395440121736 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: HNRNPAB [Title/Abstract] AND SAMM50 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | HNRNPAB(177634226)-SAMM50(44371935), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Tgene | SAMM50 | GO:0045040 | protein import into mitochondrial outer membrane | 15644312 |
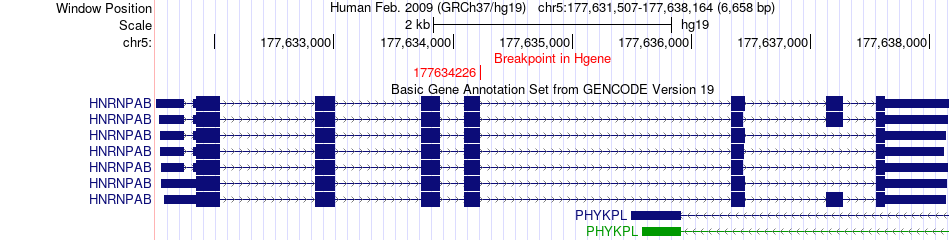
 Fusion gene breakpoints across HNRNPAB (5'-gene) Fusion gene breakpoints across HNRNPAB (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene breakpoints across SAMM50 (3'-gene) Fusion gene breakpoints across SAMM50 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
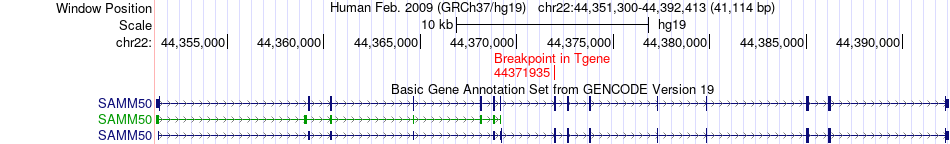 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SKCM | TCGA-BF-A5EQ-01A | HNRNPAB | chr5 | 177634226 | - | SAMM50 | chr22 | 44371935 | + |
| ChimerDB4 | SKCM | TCGA-BF-A5EQ-01A | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
Top |
Fusion Gene ORF analysis for HNRNPAB-SAMM50 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000355836 | ENST00000396202 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000355836 | ENST00000493161 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000358344 | ENST00000396202 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000358344 | ENST00000493161 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000504898 | ENST00000396202 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000504898 | ENST00000493161 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000506259 | ENST00000396202 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000506259 | ENST00000493161 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000506339 | ENST00000396202 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000506339 | ENST00000493161 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000514633 | ENST00000396202 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000514633 | ENST00000493161 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000515193 | ENST00000396202 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| 5CDS-intron | ENST00000515193 | ENST00000493161 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| In-frame | ENST00000355836 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| In-frame | ENST00000358344 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| In-frame | ENST00000504898 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| In-frame | ENST00000506259 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| In-frame | ENST00000506339 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| In-frame | ENST00000514633 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
| In-frame | ENST00000515193 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000358344 | HNRNPAB | chr5 | 177634226 | + | ENST00000350028 | SAMM50 | chr22 | 44371935 | + | 1838 | 926 | 131 | 1687 | 518 |
| ENST00000506339 | HNRNPAB | chr5 | 177634226 | + | ENST00000350028 | SAMM50 | chr22 | 44371935 | + | 1812 | 900 | 105 | 1661 | 518 |
| ENST00000355836 | HNRNPAB | chr5 | 177634226 | + | ENST00000350028 | SAMM50 | chr22 | 44371935 | + | 1806 | 894 | 99 | 1655 | 518 |
| ENST00000514633 | HNRNPAB | chr5 | 177634226 | + | ENST00000350028 | SAMM50 | chr22 | 44371935 | + | 1804 | 892 | 97 | 1653 | 518 |
| ENST00000515193 | HNRNPAB | chr5 | 177634226 | + | ENST00000350028 | SAMM50 | chr22 | 44371935 | + | 1801 | 889 | 94 | 1650 | 518 |
| ENST00000506259 | HNRNPAB | chr5 | 177634226 | + | ENST00000350028 | SAMM50 | chr22 | 44371935 | + | 1871 | 959 | 248 | 1720 | 490 |
| ENST00000504898 | HNRNPAB | chr5 | 177634226 | + | ENST00000350028 | SAMM50 | chr22 | 44371935 | + | 1848 | 936 | 225 | 1697 | 490 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000358344 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + | 0.002335408 | 0.9976646 |
| ENST00000506339 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + | 0.002468026 | 0.9975319 |
| ENST00000355836 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + | 0.00246795 | 0.997532 |
| ENST00000514633 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + | 0.002482579 | 0.99751747 |
| ENST00000515193 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + | 0.002392422 | 0.9976076 |
| ENST00000506259 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + | 0.002538175 | 0.9974618 |
| ENST00000504898 | ENST00000350028 | HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371935 | + | 0.002577911 | 0.9974221 |
Top |
Fusion Genomic Features for HNRNPAB-SAMM50 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371934 | + | 0.000179608 | 0.9998204 |
| HNRNPAB | chr5 | 177634226 | + | SAMM50 | chr22 | 44371934 | + | 0.000179608 | 0.9998204 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for HNRNPAB-SAMM50 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr5:177634226/chr22:44371935) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| HNRNPAB | . |
| FUNCTION: Binds single-stranded RNA. Has a high affinity for G-rich and U-rich regions of hnRNA. Also binds to APOB mRNA transcripts around the RNA editing site. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000355836 | + | 5 | 7 | 69_154 | 223 | 286.0 | Domain | RRM 1 |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000358344 | + | 5 | 8 | 69_154 | 223 | 333.0 | Domain | RRM 1 |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000504898 | + | 4 | 7 | 69_154 | 223 | 333.0 | Domain | RRM 1 |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000506259 | + | 4 | 6 | 69_154 | 223 | 286.0 | Domain | RRM 1 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000355836 | + | 5 | 7 | 241_324 | 223 | 286.0 | Compositional bias | Note=Gly-rich |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000358344 | + | 5 | 8 | 241_324 | 223 | 333.0 | Compositional bias | Note=Gly-rich |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000504898 | + | 4 | 7 | 241_324 | 223 | 333.0 | Compositional bias | Note=Gly-rich |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000506259 | + | 4 | 6 | 241_324 | 223 | 286.0 | Compositional bias | Note=Gly-rich |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000355836 | + | 5 | 7 | 153_233 | 223 | 286.0 | Domain | RRM 2 |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000358344 | + | 5 | 8 | 153_233 | 223 | 333.0 | Domain | RRM 2 |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000504898 | + | 4 | 7 | 153_233 | 223 | 333.0 | Domain | RRM 2 |
| Hgene | HNRNPAB | chr5:177634226 | chr22:44371935 | ENST00000506259 | + | 4 | 6 | 153_233 | 223 | 286.0 | Domain | RRM 2 |
| Tgene | SAMM50 | chr5:177634226 | chr22:44371935 | ENST00000350028 | 6 | 15 | 45_125 | 216 | 470.0 | Domain | POTRA |
Top |
Fusion Gene Sequence for HNRNPAB-SAMM50 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >37121_37121_1_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000355836_SAMM50_chr22_44371935_ENST00000350028_length(transcript)=1806nt_BP=894nt CGGCACCGCGCGGGACGGAGCTTGGCTGTTGGTCGGTGGGTTCCCGTGCGGCGGCGGCCAAGGAGGAGGAGACACAGTTGGAGCAGCTCC GTGGGCTGACTGGGGCGAGGCCTCAGCAGCGCGAGCTTGAGTGCGGCCGAGCCTGCGGCGCCTTCCCCTGCGGGTGGGGACGAGCGGGCC CCGCGGCGTCATCGGCGGCGAGGAGCCGCCGCGCCTCGGCCTAGCATGTCGGAAGCGGGCGAGGAGCAGCCCATGGAGACGACGGGCGCC ACCGAGAACGGACATGAGGCCGTCCCCGAAGGCGAGTCGCCGGCCGGGGCTGGCACGGGCGCCGCGGCGGGGGCTGGAGGCGCGACCGCG GCGCCCCCGAGCGGGAATCAGAACGGCGCCGAGGGCGACCAGATCAACGCCAGCAAGAACGAGGAGGACGCGGGAAAAATGTTCGTTGGT GGCCTGAGCTGGGATACTAGCAAAAAAGATTTAAAAGACTATTTTACTAAATTTGGAGAGGTCGTTGACTGTACAATAAAAATGGATCCC AACACTGGACGGTCAAGAGGGTTTGGGTTTATCCTGTTCAAAGATGCAGCCAGTGTGGAGAAGGTCCTAGACCAGAAGGAGCACAGGCTG GATGGCCGTGTCATTGACCCTAAAAAGGCCATGGCTATGAAGAAGGACCCGGTGAAGAAAATCTTCGTTGGGGGTCTGAATCCTGAAGCC ACTGAGGAAAAGATCAGGGAGTACTTTGGCGAGTTTGGGGAGATTGAGGCCATTGAATTGCCAATGGATCCAAAGTTGAACAAAAGACGA GGTTTTGTGTTTATCACCTTTAAAGAAGAAGAACCCGTGAAGAAGGTTCTGGAGAAAAAGTTCCATACTGTCAGTGGAAGCAAGTTTCCC ATATGGAAGACCAGCCACACTGTCAAGTGGGAAGGCGTATGGCGAGAACTGGGCTGCCTCTCAAGGACGGCGTCATTTGCTGTTCGAAAA GAAAGCGGACATTCACTGAAATCATCTCTTTCGCACGCCATGGTCATCGATTCTCGGAATTCTTCCATCTTACCAAGGAGAGGTGCTTTG CTGAAAGTTAACCAGGAACTGGCAGGCTACACTGGCGGGGATGTGAGCTTCATCAAAGAAGATTTTGAACTTCAGTTGAACAAGCAACTC ATATTTGATTCAGTTTTTTCAGCGTCTTTCTGGGGCGGAATGTTGGTACCCATTGGTGATAAGCCGTCAAGCATTGCTGATAGGTTTTAC CTTGGGGGACCCACAAGCATCCGCGGATTCAGCATGCACAGCATCGGGCCACAGAGCGAAGGAGACTACCTAGGTGGAGAAGCGTACTGG GCCGGCGGCCTGCACCTCTACACCCCATTACCTTTCCGGCCAGGCCAGGGTGGCTTTGGAGAACTTTTCCGAACACACTTCTTTCTCAAC GCAGGAAACCTCTGCAACCTCAACTATGGGGAGGGCCCCAAAGCTCATATTCGTAAGCTGGCTGAGTGCATCCGCTGGTCGTACGGGGCC GGGATTGTCCTCAGGCTTGGCAACATCGCTCGGTTGGAACTTAATTACTGCGTCCCCATGGGAGTACAGACAGGCGACAGGATATGTGAT GGCGTCCAGTTTGGAGCTGGGATAAGGTTCCTGTAGCCGACACCCCTACAGGAGAAGCTCTGGGACTGGGGCAGCAGCAAGGCGCCCATG CCACACACCGTCTCTCGAGGAAACGCGGTTCAGCGATTCTTTGACTGCGGACCCTGTGGGAAACCCCGTCAATAAATGTTAAAGACACAC >37121_37121_1_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000355836_SAMM50_chr22_44371935_ENST00000350028_length(amino acids)=518AA_BP=265 MGRGLSSASLSAAEPAAPSPAGGDERAPRRHRRRGAAAPRPSMSEAGEEQPMETTGATENGHEAVPEGESPAGAGTGAAAGAGGATAAPP SGNQNGAEGDQINASKNEEDAGKMFVGGLSWDTSKKDLKDYFTKFGEVVDCTIKMDPNTGRSRGFGFILFKDAASVEKVLDQKEHRLDGR VIDPKKAMAMKKDPVKKIFVGGLNPEATEEKIREYFGEFGEIEAIELPMDPKLNKRRGFVFITFKEEEPVKKVLEKKFHTVSGSKFPIWK TSHTVKWEGVWRELGCLSRTASFAVRKESGHSLKSSLSHAMVIDSRNSSILPRRGALLKVNQELAGYTGGDVSFIKEDFELQLNKQLIFD SVFSASFWGGMLVPIGDKPSSIADRFYLGGPTSIRGFSMHSIGPQSEGDYLGGEAYWAGGLHLYTPLPFRPGQGGFGELFRTHFFLNAGN -------------------------------------------------------------- >37121_37121_2_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000358344_SAMM50_chr22_44371935_ENST00000350028_length(transcript)=1838nt_BP=926nt AAAGGGCGCCACGAGTCGGCATTGTCAGGCGGCGGCACCGCGCGGGACGGAGCTTGGCTGTTGGTCGGTGGGTTCCCGTGCGGCGGCGGC CAAGGAGGAGGAGACACAGTTGGAGCAGCTCCGTGGGCTGACTGGGGCGAGGCCTCAGCAGCGCGAGCTTGAGTGCGGCCGAGCCTGCGG CGCCTTCCCCTGCGGGTGGGGACGAGCGGGCCCCGCGGCGTCATCGGCGGCGAGGAGCCGCCGCGCCTCGGCCTAGCATGTCGGAAGCGG GCGAGGAGCAGCCCATGGAGACGACGGGCGCCACCGAGAACGGACATGAGGCCGTCCCCGAAGGCGAGTCGCCGGCCGGGGCTGGCACGG GCGCCGCGGCGGGGGCTGGAGGCGCGACCGCGGCGCCCCCGAGCGGGAATCAGAACGGCGCCGAGGGCGACCAGATCAACGCCAGCAAGA ACGAGGAGGACGCGGGAAAAATGTTCGTTGGTGGCCTGAGCTGGGATACTAGCAAAAAAGATTTAAAAGACTATTTTACTAAATTTGGAG AGGTCGTTGACTGTACAATAAAAATGGATCCCAACACTGGACGGTCAAGAGGGTTTGGGTTTATCCTGTTCAAAGATGCAGCCAGTGTGG AGAAGGTCCTAGACCAGAAGGAGCACAGGCTGGATGGCCGTGTCATTGACCCTAAAAAGGCCATGGCTATGAAGAAGGACCCGGTGAAGA AAATCTTCGTTGGGGGTCTGAATCCTGAAGCCACTGAGGAAAAGATCAGGGAGTACTTTGGCGAGTTTGGGGAGATTGAGGCCATTGAAT TGCCAATGGATCCAAAGTTGAACAAAAGACGAGGTTTTGTGTTTATCACCTTTAAAGAAGAAGAACCCGTGAAGAAGGTTCTGGAGAAAA AGTTCCATACTGTCAGTGGAAGCAAGTTTCCCATATGGAAGACCAGCCACACTGTCAAGTGGGAAGGCGTATGGCGAGAACTGGGCTGCC TCTCAAGGACGGCGTCATTTGCTGTTCGAAAAGAAAGCGGACATTCACTGAAATCATCTCTTTCGCACGCCATGGTCATCGATTCTCGGA ATTCTTCCATCTTACCAAGGAGAGGTGCTTTGCTGAAAGTTAACCAGGAACTGGCAGGCTACACTGGCGGGGATGTGAGCTTCATCAAAG AAGATTTTGAACTTCAGTTGAACAAGCAACTCATATTTGATTCAGTTTTTTCAGCGTCTTTCTGGGGCGGAATGTTGGTACCCATTGGTG ATAAGCCGTCAAGCATTGCTGATAGGTTTTACCTTGGGGGACCCACAAGCATCCGCGGATTCAGCATGCACAGCATCGGGCCACAGAGCG AAGGAGACTACCTAGGTGGAGAAGCGTACTGGGCCGGCGGCCTGCACCTCTACACCCCATTACCTTTCCGGCCAGGCCAGGGTGGCTTTG GAGAACTTTTCCGAACACACTTCTTTCTCAACGCAGGAAACCTCTGCAACCTCAACTATGGGGAGGGCCCCAAAGCTCATATTCGTAAGC TGGCTGAGTGCATCCGCTGGTCGTACGGGGCCGGGATTGTCCTCAGGCTTGGCAACATCGCTCGGTTGGAACTTAATTACTGCGTCCCCA TGGGAGTACAGACAGGCGACAGGATATGTGATGGCGTCCAGTTTGGAGCTGGGATAAGGTTCCTGTAGCCGACACCCCTACAGGAGAAGC TCTGGGACTGGGGCAGCAGCAAGGCGCCCATGCCACACACCGTCTCTCGAGGAAACGCGGTTCAGCGATTCTTTGACTGCGGACCCTGTG >37121_37121_2_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000358344_SAMM50_chr22_44371935_ENST00000350028_length(amino acids)=518AA_BP=265 MGRGLSSASLSAAEPAAPSPAGGDERAPRRHRRRGAAAPRPSMSEAGEEQPMETTGATENGHEAVPEGESPAGAGTGAAAGAGGATAAPP SGNQNGAEGDQINASKNEEDAGKMFVGGLSWDTSKKDLKDYFTKFGEVVDCTIKMDPNTGRSRGFGFILFKDAASVEKVLDQKEHRLDGR VIDPKKAMAMKKDPVKKIFVGGLNPEATEEKIREYFGEFGEIEAIELPMDPKLNKRRGFVFITFKEEEPVKKVLEKKFHTVSGSKFPIWK TSHTVKWEGVWRELGCLSRTASFAVRKESGHSLKSSLSHAMVIDSRNSSILPRRGALLKVNQELAGYTGGDVSFIKEDFELQLNKQLIFD SVFSASFWGGMLVPIGDKPSSIADRFYLGGPTSIRGFSMHSIGPQSEGDYLGGEAYWAGGLHLYTPLPFRPGQGGFGELFRTHFFLNAGN -------------------------------------------------------------- >37121_37121_3_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000504898_SAMM50_chr22_44371935_ENST00000350028_length(transcript)=1848nt_BP=936nt TCGGTGGGTTCCCGTGCGGCGGCGGCCAAGGAGGAGGAGACACAGTTGGAGCAGCTCCGTGGGCTGACTGGGGCGAGGCCTCAGCAGCGC GAGCTTGAGTGCGGCCGAGCCTGCGGCGCCTTCCCCTGCGGGTGGGGACGAGCGGGCCCCGCGGCGTCATCGGCGGCGAGGTGAGCGGCG GCCGGCGGGCGGGAGCTCTGGCGGGAGCCGCGCCGGCCTGACGCGCTGTCGCCGCTGGTTGCAGGAGCCGCCGCGCCTCGGCCTAGCATG TCGGAAGCGGGCGAGGAGCAGCCCATGGAGACGACGGGCGCCACCGAGAACGGACATGAGGCCGTCCCCGAAGGCGAGTCGCCGGCCGGG GCTGGCACGGGCGCCGCGGCGGGGGCTGGAGGCGCGACCGCGGCGCCCCCGAGCGGGAATCAGAACGGCGCCGAGGGCGACCAGATCAAC GCCAGCAAGAACGAGGAGGACGCGGGAAAAATGTTCGTTGGTGGCCTGAGCTGGGATACTAGCAAAAAAGATTTAAAAGACTATTTTACT AAATTTGGAGAGGTCGTTGACTGTACAATAAAAATGGATCCCAACACTGGACGGTCAAGAGGGTTTGGGTTTATCCTGTTCAAAGATGCA GCCAGTGTGGAGAAGGTCCTAGACCAGAAGGAGCACAGGCTGGATGGCCGTGTCATTGACCCTAAAAAGGCCATGGCTATGAAGAAGGAC CCGGTGAAGAAAATCTTCGTTGGGGGTCTGAATCCTGAAGCCACTGAGGAAAAGATCAGGGAGTACTTTGGCGAGTTTGGGGAGATTGAG GCCATTGAATTGCCAATGGATCCAAAGTTGAACAAAAGACGAGGTTTTGTGTTTATCACCTTTAAAGAAGAAGAACCCGTGAAGAAGGTT CTGGAGAAAAAGTTCCATACTGTCAGTGGAAGCAAGTTTCCCATATGGAAGACCAGCCACACTGTCAAGTGGGAAGGCGTATGGCGAGAA CTGGGCTGCCTCTCAAGGACGGCGTCATTTGCTGTTCGAAAAGAAAGCGGACATTCACTGAAATCATCTCTTTCGCACGCCATGGTCATC GATTCTCGGAATTCTTCCATCTTACCAAGGAGAGGTGCTTTGCTGAAAGTTAACCAGGAACTGGCAGGCTACACTGGCGGGGATGTGAGC TTCATCAAAGAAGATTTTGAACTTCAGTTGAACAAGCAACTCATATTTGATTCAGTTTTTTCAGCGTCTTTCTGGGGCGGAATGTTGGTA CCCATTGGTGATAAGCCGTCAAGCATTGCTGATAGGTTTTACCTTGGGGGACCCACAAGCATCCGCGGATTCAGCATGCACAGCATCGGG CCACAGAGCGAAGGAGACTACCTAGGTGGAGAAGCGTACTGGGCCGGCGGCCTGCACCTCTACACCCCATTACCTTTCCGGCCAGGCCAG GGTGGCTTTGGAGAACTTTTCCGAACACACTTCTTTCTCAACGCAGGAAACCTCTGCAACCTCAACTATGGGGAGGGCCCCAAAGCTCAT ATTCGTAAGCTGGCTGAGTGCATCCGCTGGTCGTACGGGGCCGGGATTGTCCTCAGGCTTGGCAACATCGCTCGGTTGGAACTTAATTAC TGCGTCCCCATGGGAGTACAGACAGGCGACAGGATATGTGATGGCGTCCAGTTTGGAGCTGGGATAAGGTTCCTGTAGCCGACACCCCTA CAGGAGAAGCTCTGGGACTGGGGCAGCAGCAAGGCGCCCATGCCACACACCGTCTCTCGAGGAAACGCGGTTCAGCGATTCTTTGACTGC >37121_37121_3_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000504898_SAMM50_chr22_44371935_ENST00000350028_length(amino acids)=490AA_BP=237 MSPLVAGAAAPRPSMSEAGEEQPMETTGATENGHEAVPEGESPAGAGTGAAAGAGGATAAPPSGNQNGAEGDQINASKNEEDAGKMFVGG LSWDTSKKDLKDYFTKFGEVVDCTIKMDPNTGRSRGFGFILFKDAASVEKVLDQKEHRLDGRVIDPKKAMAMKKDPVKKIFVGGLNPEAT EEKIREYFGEFGEIEAIELPMDPKLNKRRGFVFITFKEEEPVKKVLEKKFHTVSGSKFPIWKTSHTVKWEGVWRELGCLSRTASFAVRKE SGHSLKSSLSHAMVIDSRNSSILPRRGALLKVNQELAGYTGGDVSFIKEDFELQLNKQLIFDSVFSASFWGGMLVPIGDKPSSIADRFYL GGPTSIRGFSMHSIGPQSEGDYLGGEAYWAGGLHLYTPLPFRPGQGGFGELFRTHFFLNAGNLCNLNYGEGPKAHIRKLAECIRWSYGAG -------------------------------------------------------------- >37121_37121_4_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000506259_SAMM50_chr22_44371935_ENST00000350028_length(transcript)=1871nt_BP=959nt GCGGGACGGAGCTTGGCTGTTGGTCGGTGGGTTCCCGTGCGGCGGCGGCCAAGGAGGAGGAGACACAGTTGGAGCAGCTCCGTGGGCTGA CTGGGGCGAGGCCTCAGCAGCGCGAGCTTGAGTGCGGCCGAGCCTGCGGCGCCTTCCCCTGCGGGTGGGGACGAGCGGGCCCCGCGGCGT CATCGGCGGCGAGGTGAGCGGCGGCCGGCGGGCGGGAGCTCTGGCGGGAGCCGCGCCGGCCTGACGCGCTGTCGCCGCTGGTTGCAGGAG CCGCCGCGCCTCGGCCTAGCATGTCGGAAGCGGGCGAGGAGCAGCCCATGGAGACGACGGGCGCCACCGAGAACGGACATGAGGCCGTCC CCGAAGGCGAGTCGCCGGCCGGGGCTGGCACGGGCGCCGCGGCGGGGGCTGGAGGCGCGACCGCGGCGCCCCCGAGCGGGAATCAGAACG GCGCCGAGGGCGACCAGATCAACGCCAGCAAGAACGAGGAGGACGCGGGAAAAATGTTCGTTGGTGGCCTGAGCTGGGATACTAGCAAAA AAGATTTAAAAGACTATTTTACTAAATTTGGAGAGGTCGTTGACTGTACAATAAAAATGGATCCCAACACTGGACGGTCAAGAGGGTTTG GGTTTATCCTGTTCAAAGATGCAGCCAGTGTGGAGAAGGTCCTAGACCAGAAGGAGCACAGGCTGGATGGCCGTGTCATTGACCCTAAAA AGGCCATGGCTATGAAGAAGGACCCGGTGAAGAAAATCTTCGTTGGGGGTCTGAATCCTGAAGCCACTGAGGAAAAGATCAGGGAGTACT TTGGCGAGTTTGGGGAGATTGAGGCCATTGAATTGCCAATGGATCCAAAGTTGAACAAAAGACGAGGTTTTGTGTTTATCACCTTTAAAG AAGAAGAACCCGTGAAGAAGGTTCTGGAGAAAAAGTTCCATACTGTCAGTGGAAGCAAGTTTCCCATATGGAAGACCAGCCACACTGTCA AGTGGGAAGGCGTATGGCGAGAACTGGGCTGCCTCTCAAGGACGGCGTCATTTGCTGTTCGAAAAGAAAGCGGACATTCACTGAAATCAT CTCTTTCGCACGCCATGGTCATCGATTCTCGGAATTCTTCCATCTTACCAAGGAGAGGTGCTTTGCTGAAAGTTAACCAGGAACTGGCAG GCTACACTGGCGGGGATGTGAGCTTCATCAAAGAAGATTTTGAACTTCAGTTGAACAAGCAACTCATATTTGATTCAGTTTTTTCAGCGT CTTTCTGGGGCGGAATGTTGGTACCCATTGGTGATAAGCCGTCAAGCATTGCTGATAGGTTTTACCTTGGGGGACCCACAAGCATCCGCG GATTCAGCATGCACAGCATCGGGCCACAGAGCGAAGGAGACTACCTAGGTGGAGAAGCGTACTGGGCCGGCGGCCTGCACCTCTACACCC CATTACCTTTCCGGCCAGGCCAGGGTGGCTTTGGAGAACTTTTCCGAACACACTTCTTTCTCAACGCAGGAAACCTCTGCAACCTCAACT ATGGGGAGGGCCCCAAAGCTCATATTCGTAAGCTGGCTGAGTGCATCCGCTGGTCGTACGGGGCCGGGATTGTCCTCAGGCTTGGCAACA TCGCTCGGTTGGAACTTAATTACTGCGTCCCCATGGGAGTACAGACAGGCGACAGGATATGTGATGGCGTCCAGTTTGGAGCTGGGATAA GGTTCCTGTAGCCGACACCCCTACAGGAGAAGCTCTGGGACTGGGGCAGCAGCAAGGCGCCCATGCCACACACCGTCTCTCGAGGAAACG >37121_37121_4_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000506259_SAMM50_chr22_44371935_ENST00000350028_length(amino acids)=490AA_BP=237 MSPLVAGAAAPRPSMSEAGEEQPMETTGATENGHEAVPEGESPAGAGTGAAAGAGGATAAPPSGNQNGAEGDQINASKNEEDAGKMFVGG LSWDTSKKDLKDYFTKFGEVVDCTIKMDPNTGRSRGFGFILFKDAASVEKVLDQKEHRLDGRVIDPKKAMAMKKDPVKKIFVGGLNPEAT EEKIREYFGEFGEIEAIELPMDPKLNKRRGFVFITFKEEEPVKKVLEKKFHTVSGSKFPIWKTSHTVKWEGVWRELGCLSRTASFAVRKE SGHSLKSSLSHAMVIDSRNSSILPRRGALLKVNQELAGYTGGDVSFIKEDFELQLNKQLIFDSVFSASFWGGMLVPIGDKPSSIADRFYL GGPTSIRGFSMHSIGPQSEGDYLGGEAYWAGGLHLYTPLPFRPGQGGFGELFRTHFFLNAGNLCNLNYGEGPKAHIRKLAECIRWSYGAG -------------------------------------------------------------- >37121_37121_5_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000506339_SAMM50_chr22_44371935_ENST00000350028_length(transcript)=1812nt_BP=900nt AGGCGGCGGCACCGCGCGGGACGGAGCTTGGCTGTTGGTCGGTGGGTTCCCGTGCGGCGGCGGCCAAGGAGGAGGAGACACAGTTGGAGC AGCTCCGTGGGCTGACTGGGGCGAGGCCTCAGCAGCGCGAGCTTGAGTGCGGCCGAGCCTGCGGCGCCTTCCCCTGCGGGTGGGGACGAG CGGGCCCCGCGGCGTCATCGGCGGCGAGGAGCCGCCGCGCCTCGGCCTAGCATGTCGGAAGCGGGCGAGGAGCAGCCCATGGAGACGACG GGCGCCACCGAGAACGGACATGAGGCCGTCCCCGAAGGCGAGTCGCCGGCCGGGGCTGGCACGGGCGCCGCGGCGGGGGCTGGAGGCGCG ACCGCGGCGCCCCCGAGCGGGAATCAGAACGGCGCCGAGGGCGACCAGATCAACGCCAGCAAGAACGAGGAGGACGCGGGAAAAATGTTC GTTGGTGGCCTGAGCTGGGATACTAGCAAAAAAGATTTAAAAGACTATTTTACTAAATTTGGAGAGGTCGTTGACTGTACAATAAAAATG GATCCCAACACTGGACGGTCAAGAGGGTTTGGGTTTATCCTGTTCAAAGATGCAGCCAGTGTGGAGAAGGTCCTAGACCAGAAGGAGCAC AGGCTGGATGGCCGTGTCATTGACCCTAAAAAGGCCATGGCTATGAAGAAGGACCCGGTGAAGAAAATCTTCGTTGGGGGTCTGAATCCT GAAGCCACTGAGGAAAAGATCAGGGAGTACTTTGGCGAGTTTGGGGAGATTGAGGCCATTGAATTGCCAATGGATCCAAAGTTGAACAAA AGACGAGGTTTTGTGTTTATCACCTTTAAAGAAGAAGAACCCGTGAAGAAGGTTCTGGAGAAAAAGTTCCATACTGTCAGTGGAAGCAAG TTTCCCATATGGAAGACCAGCCACACTGTCAAGTGGGAAGGCGTATGGCGAGAACTGGGCTGCCTCTCAAGGACGGCGTCATTTGCTGTT CGAAAAGAAAGCGGACATTCACTGAAATCATCTCTTTCGCACGCCATGGTCATCGATTCTCGGAATTCTTCCATCTTACCAAGGAGAGGT GCTTTGCTGAAAGTTAACCAGGAACTGGCAGGCTACACTGGCGGGGATGTGAGCTTCATCAAAGAAGATTTTGAACTTCAGTTGAACAAG CAACTCATATTTGATTCAGTTTTTTCAGCGTCTTTCTGGGGCGGAATGTTGGTACCCATTGGTGATAAGCCGTCAAGCATTGCTGATAGG TTTTACCTTGGGGGACCCACAAGCATCCGCGGATTCAGCATGCACAGCATCGGGCCACAGAGCGAAGGAGACTACCTAGGTGGAGAAGCG TACTGGGCCGGCGGCCTGCACCTCTACACCCCATTACCTTTCCGGCCAGGCCAGGGTGGCTTTGGAGAACTTTTCCGAACACACTTCTTT CTCAACGCAGGAAACCTCTGCAACCTCAACTATGGGGAGGGCCCCAAAGCTCATATTCGTAAGCTGGCTGAGTGCATCCGCTGGTCGTAC GGGGCCGGGATTGTCCTCAGGCTTGGCAACATCGCTCGGTTGGAACTTAATTACTGCGTCCCCATGGGAGTACAGACAGGCGACAGGATA TGTGATGGCGTCCAGTTTGGAGCTGGGATAAGGTTCCTGTAGCCGACACCCCTACAGGAGAAGCTCTGGGACTGGGGCAGCAGCAAGGCG CCCATGCCACACACCGTCTCTCGAGGAAACGCGGTTCAGCGATTCTTTGACTGCGGACCCTGTGGGAAACCCCGTCAATAAATGTTAAAG >37121_37121_5_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000506339_SAMM50_chr22_44371935_ENST00000350028_length(amino acids)=518AA_BP=265 MGRGLSSASLSAAEPAAPSPAGGDERAPRRHRRRGAAAPRPSMSEAGEEQPMETTGATENGHEAVPEGESPAGAGTGAAAGAGGATAAPP SGNQNGAEGDQINASKNEEDAGKMFVGGLSWDTSKKDLKDYFTKFGEVVDCTIKMDPNTGRSRGFGFILFKDAASVEKVLDQKEHRLDGR VIDPKKAMAMKKDPVKKIFVGGLNPEATEEKIREYFGEFGEIEAIELPMDPKLNKRRGFVFITFKEEEPVKKVLEKKFHTVSGSKFPIWK TSHTVKWEGVWRELGCLSRTASFAVRKESGHSLKSSLSHAMVIDSRNSSILPRRGALLKVNQELAGYTGGDVSFIKEDFELQLNKQLIFD SVFSASFWGGMLVPIGDKPSSIADRFYLGGPTSIRGFSMHSIGPQSEGDYLGGEAYWAGGLHLYTPLPFRPGQGGFGELFRTHFFLNAGN -------------------------------------------------------------- >37121_37121_6_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000514633_SAMM50_chr22_44371935_ENST00000350028_length(transcript)=1804nt_BP=892nt GCACCGCGCGGGACGGAGCTTGGCTGTTGGTCGGTGGGTTCCCGTGCGGCGGCGGCCAAGGAGGAGGAGACACAGTTGGAGCAGCTCCGT GGGCTGACTGGGGCGAGGCCTCAGCAGCGCGAGCTTGAGTGCGGCCGAGCCTGCGGCGCCTTCCCCTGCGGGTGGGGACGAGCGGGCCCC GCGGCGTCATCGGCGGCGAGGAGCCGCCGCGCCTCGGCCTAGCATGTCGGAAGCGGGCGAGGAGCAGCCCATGGAGACGACGGGCGCCAC CGAGAACGGACATGAGGCCGTCCCCGAAGGCGAGTCGCCGGCCGGGGCTGGCACGGGCGCCGCGGCGGGGGCTGGAGGCGCGACCGCGGC GCCCCCGAGCGGGAATCAGAACGGCGCCGAGGGCGACCAGATCAACGCCAGCAAGAACGAGGAGGACGCGGGAAAAATGTTCGTTGGTGG CCTGAGCTGGGATACTAGCAAAAAAGATTTAAAAGACTATTTTACTAAATTTGGAGAGGTCGTTGACTGTACAATAAAAATGGATCCCAA CACTGGACGGTCAAGAGGGTTTGGGTTTATCCTGTTCAAAGATGCAGCCAGTGTGGAGAAGGTCCTAGACCAGAAGGAGCACAGGCTGGA TGGCCGTGTCATTGACCCTAAAAAGGCCATGGCTATGAAGAAGGACCCGGTGAAGAAAATCTTCGTTGGGGGTCTGAATCCTGAAGCCAC TGAGGAAAAGATCAGGGAGTACTTTGGCGAGTTTGGGGAGATTGAGGCCATTGAATTGCCAATGGATCCAAAGTTGAACAAAAGACGAGG TTTTGTGTTTATCACCTTTAAAGAAGAAGAACCCGTGAAGAAGGTTCTGGAGAAAAAGTTCCATACTGTCAGTGGAAGCAAGTTTCCCAT ATGGAAGACCAGCCACACTGTCAAGTGGGAAGGCGTATGGCGAGAACTGGGCTGCCTCTCAAGGACGGCGTCATTTGCTGTTCGAAAAGA AAGCGGACATTCACTGAAATCATCTCTTTCGCACGCCATGGTCATCGATTCTCGGAATTCTTCCATCTTACCAAGGAGAGGTGCTTTGCT GAAAGTTAACCAGGAACTGGCAGGCTACACTGGCGGGGATGTGAGCTTCATCAAAGAAGATTTTGAACTTCAGTTGAACAAGCAACTCAT ATTTGATTCAGTTTTTTCAGCGTCTTTCTGGGGCGGAATGTTGGTACCCATTGGTGATAAGCCGTCAAGCATTGCTGATAGGTTTTACCT TGGGGGACCCACAAGCATCCGCGGATTCAGCATGCACAGCATCGGGCCACAGAGCGAAGGAGACTACCTAGGTGGAGAAGCGTACTGGGC CGGCGGCCTGCACCTCTACACCCCATTACCTTTCCGGCCAGGCCAGGGTGGCTTTGGAGAACTTTTCCGAACACACTTCTTTCTCAACGC AGGAAACCTCTGCAACCTCAACTATGGGGAGGGCCCCAAAGCTCATATTCGTAAGCTGGCTGAGTGCATCCGCTGGTCGTACGGGGCCGG GATTGTCCTCAGGCTTGGCAACATCGCTCGGTTGGAACTTAATTACTGCGTCCCCATGGGAGTACAGACAGGCGACAGGATATGTGATGG CGTCCAGTTTGGAGCTGGGATAAGGTTCCTGTAGCCGACACCCCTACAGGAGAAGCTCTGGGACTGGGGCAGCAGCAAGGCGCCCATGCC ACACACCGTCTCTCGAGGAAACGCGGTTCAGCGATTCTTTGACTGCGGACCCTGTGGGAAACCCCGTCAATAAATGTTAAAGACACACTC >37121_37121_6_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000514633_SAMM50_chr22_44371935_ENST00000350028_length(amino acids)=518AA_BP=265 MGRGLSSASLSAAEPAAPSPAGGDERAPRRHRRRGAAAPRPSMSEAGEEQPMETTGATENGHEAVPEGESPAGAGTGAAAGAGGATAAPP SGNQNGAEGDQINASKNEEDAGKMFVGGLSWDTSKKDLKDYFTKFGEVVDCTIKMDPNTGRSRGFGFILFKDAASVEKVLDQKEHRLDGR VIDPKKAMAMKKDPVKKIFVGGLNPEATEEKIREYFGEFGEIEAIELPMDPKLNKRRGFVFITFKEEEPVKKVLEKKFHTVSGSKFPIWK TSHTVKWEGVWRELGCLSRTASFAVRKESGHSLKSSLSHAMVIDSRNSSILPRRGALLKVNQELAGYTGGDVSFIKEDFELQLNKQLIFD SVFSASFWGGMLVPIGDKPSSIADRFYLGGPTSIRGFSMHSIGPQSEGDYLGGEAYWAGGLHLYTPLPFRPGQGGFGELFRTHFFLNAGN -------------------------------------------------------------- >37121_37121_7_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000515193_SAMM50_chr22_44371935_ENST00000350028_length(transcript)=1801nt_BP=889nt CCGCGCGGGACGGAGCTTGGCTGTTGGTCGGTGGGTTCCCGTGCGGCGGCGGCCAAGGAGGAGGAGACACAGTTGGAGCAGCTCCGTGGG CTGACTGGGGCGAGGCCTCAGCAGCGCGAGCTTGAGTGCGGCCGAGCCTGCGGCGCCTTCCCCTGCGGGTGGGGACGAGCGGGCCCCGCG GCGTCATCGGCGGCGAGGAGCCGCCGCGCCTCGGCCTAGCATGTCGGAAGCGGGCGAGGAGCAGCCCATGGAGACGACGGGCGCCACCGA GAACGGACATGAGGCCGTCCCCGAAGGCGAGTCGCCGGCCGGGGCTGGCACGGGCGCCGCGGCGGGGGCTGGAGGCGCGACCGCGGCGCC CCCGAGCGGGAATCAGAACGGCGCCGAGGGCGACCAGATCAACGCCAGCAAGAACGAGGAGGACGCGGGAAAAATGTTCGTTGGTGGCCT GAGCTGGGATACTAGCAAAAAAGATTTAAAAGACTATTTTACTAAATTTGGAGAGGTCGTTGACTGTACAATAAAAATGGATCCCAACAC TGGACGGTCAAGAGGGTTTGGGTTTATCCTGTTCAAAGATGCAGCCAGTGTGGAGAAGGTCCTAGACCAGAAGGAGCACAGGCTGGATGG CCGTGTCATTGACCCTAAAAAGGCCATGGCTATGAAGAAGGACCCGGTGAAGAAAATCTTCGTTGGGGGTCTGAATCCTGAAGCCACTGA GGAAAAGATCAGGGAGTACTTTGGCGAGTTTGGGGAGATTGAGGCCATTGAATTGCCAATGGATCCAAAGTTGAACAAAAGACGAGGTTT TGTGTTTATCACCTTTAAAGAAGAAGAACCCGTGAAGAAGGTTCTGGAGAAAAAGTTCCATACTGTCAGTGGAAGCAAGTTTCCCATATG GAAGACCAGCCACACTGTCAAGTGGGAAGGCGTATGGCGAGAACTGGGCTGCCTCTCAAGGACGGCGTCATTTGCTGTTCGAAAAGAAAG CGGACATTCACTGAAATCATCTCTTTCGCACGCCATGGTCATCGATTCTCGGAATTCTTCCATCTTACCAAGGAGAGGTGCTTTGCTGAA AGTTAACCAGGAACTGGCAGGCTACACTGGCGGGGATGTGAGCTTCATCAAAGAAGATTTTGAACTTCAGTTGAACAAGCAACTCATATT TGATTCAGTTTTTTCAGCGTCTTTCTGGGGCGGAATGTTGGTACCCATTGGTGATAAGCCGTCAAGCATTGCTGATAGGTTTTACCTTGG GGGACCCACAAGCATCCGCGGATTCAGCATGCACAGCATCGGGCCACAGAGCGAAGGAGACTACCTAGGTGGAGAAGCGTACTGGGCCGG CGGCCTGCACCTCTACACCCCATTACCTTTCCGGCCAGGCCAGGGTGGCTTTGGAGAACTTTTCCGAACACACTTCTTTCTCAACGCAGG AAACCTCTGCAACCTCAACTATGGGGAGGGCCCCAAAGCTCATATTCGTAAGCTGGCTGAGTGCATCCGCTGGTCGTACGGGGCCGGGAT TGTCCTCAGGCTTGGCAACATCGCTCGGTTGGAACTTAATTACTGCGTCCCCATGGGAGTACAGACAGGCGACAGGATATGTGATGGCGT CCAGTTTGGAGCTGGGATAAGGTTCCTGTAGCCGACACCCCTACAGGAGAAGCTCTGGGACTGGGGCAGCAGCAAGGCGCCCATGCCACA CACCGTCTCTCGAGGAAACGCGGTTCAGCGATTCTTTGACTGCGGACCCTGTGGGAAACCCCGTCAATAAATGTTAAAGACACACTCCGA >37121_37121_7_HNRNPAB-SAMM50_HNRNPAB_chr5_177634226_ENST00000515193_SAMM50_chr22_44371935_ENST00000350028_length(amino acids)=518AA_BP=265 MGRGLSSASLSAAEPAAPSPAGGDERAPRRHRRRGAAAPRPSMSEAGEEQPMETTGATENGHEAVPEGESPAGAGTGAAAGAGGATAAPP SGNQNGAEGDQINASKNEEDAGKMFVGGLSWDTSKKDLKDYFTKFGEVVDCTIKMDPNTGRSRGFGFILFKDAASVEKVLDQKEHRLDGR VIDPKKAMAMKKDPVKKIFVGGLNPEATEEKIREYFGEFGEIEAIELPMDPKLNKRRGFVFITFKEEEPVKKVLEKKFHTVSGSKFPIWK TSHTVKWEGVWRELGCLSRTASFAVRKESGHSLKSSLSHAMVIDSRNSSILPRRGALLKVNQELAGYTGGDVSFIKEDFELQLNKQLIFD SVFSASFWGGMLVPIGDKPSSIADRFYLGGPTSIRGFSMHSIGPQSEGDYLGGEAYWAGGLHLYTPLPFRPGQGGFGELFRTHFFLNAGN -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for HNRNPAB-SAMM50 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for HNRNPAB-SAMM50 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for HNRNPAB-SAMM50 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
