
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:HSP90B1-ALB (FusionGDB2 ID:37818) |
Fusion Gene Summary for HSP90B1-ALB |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: HSP90B1-ALB | Fusion gene ID: 37818 | Hgene | Tgene | Gene symbol | HSP90B1 | ALB | Gene ID | 7184 | 213 |
| Gene name | heat shock protein 90 beta family member 1 | albumin | |
| Synonyms | ECGP|GP96|GRP94|HEL-S-125m|HEL35|TRA1 | HSA|PRO0883|PRO0903|PRO1341 | |
| Cytomap | 12q23.3 | 4q13.3 | |
| Type of gene | protein-coding | protein-coding | |
| Description | endoplasmin94 kDa glucose-regulated proteinendothelial cell (HBMEC) glycoproteinepididymis luminal protein 35epididymis secretory sperm binding protein Li 125mheat shock protein 90 kDa beta member 1heat shock protein 90kDa beta (Grp94), member 1hea | serum albumin | |
| Modification date | 20200320 | 20200329 | |
| UniProtAcc | P14625 | P02768 | |
| Ensembl transtripts involved in fusion gene | ENST00000299767, | ENST00000505649, ENST00000401494, ENST00000415165, ENST00000503124, ENST00000509063, ENST00000295897, | |
| Fusion gene scores | * DoF score | 19 X 19 X 10=3610 | 49 X 62 X 5=15190 |
| # samples | 25 | 64 | |
| ** MAII score | log2(25/3610*10)=-3.85199883711245 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(64/15190*10)=-4.56890615450208 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: HSP90B1 [Title/Abstract] AND ALB [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | HSP90B1(104336987)-ALB(74274478), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | HSP90B1 | GO:0001666 | response to hypoxia | 15620698 |
| Hgene | HSP90B1 | GO:0031247 | actin rod assembly | 19000834 |
| Hgene | HSP90B1 | GO:0043666 | regulation of phosphoprotein phosphatase activity | 19000834 |
| Hgene | HSP90B1 | GO:0071318 | cellular response to ATP | 19000834 |
| Tgene | ALB | GO:0009267 | cellular response to starvation | 16245148 |
| Tgene | ALB | GO:0043066 | negative regulation of apoptotic process | 16153637 |
| Tgene | ALB | GO:0051659 | maintenance of mitochondrion location | 16153637 |
 Fusion gene breakpoints across HSP90B1 (5'-gene) Fusion gene breakpoints across HSP90B1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
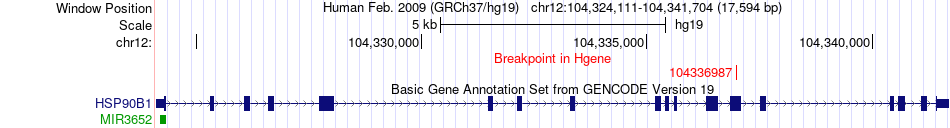 |
 Fusion gene breakpoints across ALB (3'-gene) Fusion gene breakpoints across ALB (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
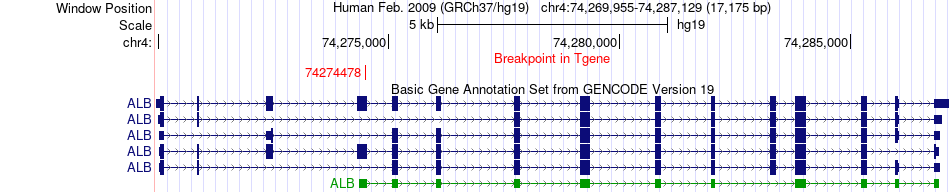 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | LIHC | TCGA-2Y-A9H0 | HSP90B1 | chr12 | 104336987 | + | ALB | chr4 | 74274478 | + |
Top |
Fusion Gene ORF analysis for HSP90B1-ALB |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000299767 | ENST00000505649 | HSP90B1 | chr12 | 104336987 | + | ALB | chr4 | 74274478 | + |
| 5CDS-intron | ENST00000299767 | ENST00000401494 | HSP90B1 | chr12 | 104336987 | + | ALB | chr4 | 74274478 | + |
| 5CDS-intron | ENST00000299767 | ENST00000415165 | HSP90B1 | chr12 | 104336987 | + | ALB | chr4 | 74274478 | + |
| 5CDS-intron | ENST00000299767 | ENST00000503124 | HSP90B1 | chr12 | 104336987 | + | ALB | chr4 | 74274478 | + |
| 5CDS-intron | ENST00000299767 | ENST00000509063 | HSP90B1 | chr12 | 104336987 | + | ALB | chr4 | 74274478 | + |
| In-frame | ENST00000299767 | ENST00000295897 | HSP90B1 | chr12 | 104336987 | + | ALB | chr4 | 74274478 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000299767 | HSP90B1 | chr12 | 104336987 | + | ENST00000295897 | ALB | chr4 | 74274478 | + | 3672 | 1936 | 182 | 1834 | 550 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000299767 | ENST00000295897 | HSP90B1 | chr12 | 104336987 | + | ALB | chr4 | 74274478 | + | 0.000454846 | 0.99954516 |
Top |
Fusion Genomic Features for HSP90B1-ALB |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
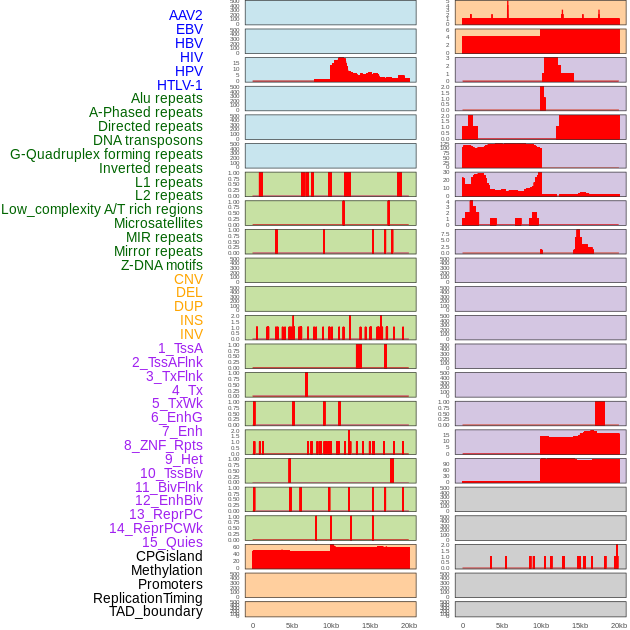 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for HSP90B1-ALB |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:104336987/chr4:74274478) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| HSP90B1 | ALB |
| FUNCTION: Molecular chaperone that functions in the processing and transport of secreted proteins (By similarity). When associated with CNPY3, required for proper folding of Toll-like receptors (By similarity). Functions in endoplasmic reticulum associated degradation (ERAD) (PubMed:18264092). Has ATPase activity (By similarity). May participate in the unfolding of cytosolic leaderless cargos (lacking the secretion signal sequence) such as the interleukin 1/IL-1 to facilitate their translocation into the ERGIC (endoplasmic reticulum-Golgi intermediate compartment) and secretion; the translocation process is mediated by the cargo receptor TMED10 (PubMed:32272059). {ECO:0000250|UniProtKB:P08113, ECO:0000269|PubMed:18264092, ECO:0000269|PubMed:32272059}. | FUNCTION: Binds water, Ca(2+), Na(+), K(+), fatty acids, hormones, bilirubin and drugs (Probable). Its main function is the regulation of the colloidal osmotic pressure of blood (Probable). Major zinc transporter in plasma, typically binds about 80% of all plasma zinc (PubMed:19021548). Major calcium and magnesium transporter in plasma, binds approximately 45% of circulating calcium and magnesium in plasma (By similarity). Potentially has more than two calcium-binding sites and might additionally bind calcium in a non-specific manner (By similarity). The shared binding site between zinc and calcium at residue Asp-273 suggests a crosstalk between zinc and calcium transport in the blood (By similarity). The rank order of affinity is zinc > calcium > magnesium (By similarity). Binds to the bacterial siderophore enterobactin and inhibits enterobactin-mediated iron uptake of E.coli from ferric transferrin, and may thereby limit the utilization of iron and growth of enteric bacteria such as E.coli (PubMed:6234017). Does not prevent iron uptake by the bacterial siderophore aerobactin (PubMed:6234017). {ECO:0000250|UniProtKB:P02769, ECO:0000269|PubMed:19021548, ECO:0000269|PubMed:6234017, ECO:0000305|PubMed:1630489}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | ALB | chr12:104336987 | chr4:74274478 | ENST00000295897 | 0 | 15 | 19_210 | 0 | 661.0 | Domain | Albumin 1 | |
| Tgene | ALB | chr12:104336987 | chr4:74274478 | ENST00000295897 | 0 | 15 | 211_403 | 0 | 661.0 | Domain | Albumin 2 | |
| Tgene | ALB | chr12:104336987 | chr4:74274478 | ENST00000295897 | 0 | 15 | 404_601 | 0 | 661.0 | Domain | Albumin 3 | |
| Tgene | ALB | chr12:104336987 | chr4:74274478 | ENST00000509063 | 0 | 14 | 19_210 | 0 | 605.0 | Domain | Albumin 1 | |
| Tgene | ALB | chr12:104336987 | chr4:74274478 | ENST00000509063 | 0 | 14 | 211_403 | 0 | 605.0 | Domain | Albumin 2 | |
| Tgene | ALB | chr12:104336987 | chr4:74274478 | ENST00000509063 | 0 | 14 | 404_601 | 0 | 605.0 | Domain | Albumin 3 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | HSP90B1 | chr12:104336987 | chr4:74274478 | ENST00000299767 | + | 1 | 18 | 800_803 | 0 | 804.0 | Motif | Note=Prevents secretion from ER |
Top |
Fusion Gene Sequence for HSP90B1-ALB |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >37818_37818_1_HSP90B1-ALB_HSP90B1_chr12_104336987_ENST00000299767_ALB_chr4_74274478_ENST00000295897_length(transcript)=3672nt_BP=1936nt GATTGGTGGGTTCATGTTTCCCGTCCCCCGCCCGCGGGAAGTGGGGGTGAAAAGCGGCCCGACCTGCTTGCGGTGTAGTGGGCGGACCGC GCGGCTGGAGGTGTGAGGATCCGAACCCAGGGGTGGGGGGTGGAGGCGGCTCCTGCGATCGAAGGGGACTTGAGACTCACCGGCCGCACG CCATGAGGGCCCTGTGGGTGCTGGGCCTCTGCTGCGTCCTGCTGACCTTCGGGTCGGTCAGAGCTGACGATGAAGTTGATGTGGATGGTA CAGTAGAAGAGGATCTGGGTAAAAGTAGAGAAGGATCAAGGACGGATGATGAAGTAGTACAGAGAGAGGAAGAAGCTATTCAGTTGGATG GATTAAATGCATCACAAATAAGAGAACTTAGAGAGAAGTCGGAAAAGTTTGCCTTCCAAGCCGAAGTTAACAGAATGATGAAACTTATCA TCAATTCATTGTATAAAAATAAAGAGATTTTCCTGAGAGAACTGATTTCAAATGCTTCTGATGCTTTAGATAAGATAAGGCTAATATCAC TGACTGATGAAAATGCTCTTTCTGGAAATGAGGAACTAACAGTCAAAATTAAGTGTGATAAGGAGAAGAACCTGCTGCATGTCACAGACA CCGGTGTAGGAATGACCAGAGAAGAGTTGGTTAAAAACCTTGGTACCATAGCCAAATCTGGGACAAGCGAGTTTTTAAACAAAATGACTG AAGCACAGGAAGATGGCCAGTCAACTTCTGAATTGATTGGCCAGTTTGGTGTCGGTTTCTATTCCGCCTTCCTTGTAGCAGATAAGGTTA TTGTCACTTCAAAACACAACAACGATACCCAGCACATCTGGGAGTCTGACTCCAATGAATTTTCTGTAATTGCTGACCCAAGAGGAAACA CTCTAGGACGGGGAACGACAATTACCCTTGTCTTAAAAGAAGAAGCATCTGATTACCTTGAATTGGATACAATTAAAAATCTCGTCAAAA AATATTCACAGTTCATAAACTTTCCTATTTATGTATGGAGCAGCAAGACTGAAACTGTTGAGGAGCCCATGGAGGAAGAAGAAGCAGCCA AAGAAGAGAAAGAAGAATCTGATGATGAAGCTGCAGTAGAGGAAGAAGAAGAAGAAAAGAAACCAAAGACTAAAAAAGTTGAAAAAACTG TCTGGGACTGGGAACTTATGAATGATATCAAACCAATATGGCAGAGACCATCAAAAGAAGTAGAAGAAGATGAATACAAAGCTTTCTACA AATCATTTTCAAAGGAAAGTGATGACCCCATGGCTTATATTCACTTTACTGCTGAAGGGGAAGTTACCTTCAAATCAATTTTATTTGTAC CCACATCTGCTCCACGTGGTCTGTTTGACGAATATGGATCTAAAAAGAGCGATTACATTAAGCTCTATGTGCGCCGTGTATTCATCACAG ACGACTTCCATGATATGATGCCTAAATACCTCAATTTTGTCAAGGGTGTGGTGGACTCAGATGATCTCCCCTTGAATGTTTCCCGCGAGA CTCTTCAGCAACATAAACTGCTTAAGGTGATTAGGAAGAAGCTTGTTCGTAAAACGCTGGACATGATCAAGAAGATTGCTGATGATAAAT ACAATGATACTTTTTGGAAAGAATTTGGTACCAACATCAAGCTTGGTGTGATTGAAGACCACTCGAATCGAACACGTCTTGCTAAACTTC TTAGGTTCCAGTCTTCTCATCATCCAACTGACATTACTAGCCTAGACCAGTATGTGGAAAGAATGAAGGAAAAACAAGACAAAATCTACT TCATGGCTGGGTCCAGCAGAAAAGAGAAGGAGTGAAGTTCGATGAAAGTGAGAAAACTAAGGAGAGTCGTGAAGCAGTTGAGAAAGAATT TGAGCCTCTGCTGAATTGGATGAAAGATAAAGCCCTTAAGGACAAGATGTGCACTGCTTTTCATGACAATGAAGAGACATTTTTGAAAAA ATACTTATATGAAATTGCCAGAAGACATCCTTACTTTTATGCCCCGGAACTCCTTTTCTTTGCTAAAAGGTATAAAGCTGCTTTTACAGA ATGTTGCCAAGCTGCTGATAAAGCTGCCTGCCTGTTGCCAAAGCTCGATGAACTTCGGGATGAAGGGAAGGCTTCGTCTGCCAAACAGAG ACTCAAGTGTGCCAGTCTCCAAAAATTTGGAGAAAGAGCTTTCAAAGCATGGGCAGTAGCTCGCCTGAGCCAGAGATTTCCCAAAGCTGA GTTTGCAGAAGTTTCCAAGTTAGTGACAGATCTTACCAAAGTCCACACGGAATGCTGCCATGGAGATCTGCTTGAATGTGCTGATGACAG GGCGGACCTTGCCAAGTATATCTGTGAAAATCAAGATTCGATCTCCAGTAAACTGAAGGAATGCTGTGAAAAACCTCTGTTGGAAAAATC CCACTGCATTGCCGAAGTGGAAAATGATGAGATGCCTGCTGACTTGCCTTCATTAGCTGCTGATTTTGTTGAAAGTAAGGATGTTTGCAA AAACTATGCTGAGGCAAAGGATGTCTTCCTGGGCATGTTTTTGTATGAATATGCAAGAAGGCATCCTGATTACTCTGTCGTGCTGCTGCT GAGACTTGCCAAGACATATGAAACCACTCTAGAGAAGTGCTGTGCCGCTGCAGATCCTCATGAATGCTATGCCAAAGTGTTCGATGAATT TAAACCTCTTGTGGAAGAGCCTCAGAATTTAATCAAACAAAATTGTGAGCTTTTTGAGCAGCTTGGAGAGTACAAATTCCAGAATGCGCT ATTAGTTCGTTACACCAAGAAAGTACCCCAAGTGTCAACTCCAACTCTTGTAGAGGTCTCAAGAAACCTAGGAAAAGTGGGCAGCAAATG TTGTAAACATCCTGAAGCAAAAAGAATGCCCTGTGCAGAAGACTATCTATCCGTGGTCCTGAACCAGTTATGTGTGTTGCATGAGAAAAC GCCAGTAAGTGACAGAGTCACCAAATGCTGCACAGAATCCTTGGTGAACAGGCGACCATGCTTTTCAGCTCTGGAAGTCGATGAAACATA CGTTCCCAAAGAGTTTAATGCTGAAACATTCACCTTCCATGCAGATATATGCACACTTTCTGAGAAGGAGAGACAAATCAAGAAACAAAC TGCACTTGTTGAGCTCGTGAAACACAAGCCCAAGGCAACAAAAGAGCAACTGAAAGCTGTTATGGATGATTTCGCAGCTTTTGTAGAGAA GTGCTGCAAGGCTGACGATAAGGAGACCTGCTTTGCCGAGGAGGGTAAAAAACTTGTTGCTGCAAGTCAAGCTGCCTTAGGCTTATAACA TCACATTTAAAAGCATCTCAGCCTACCATGAGAATAAGAGAAAGAAAATGAAGATCAAAAGCTTATTCATCTGTTTTTCTTTTTCGTTGG TGTAAAGCCAACACCCTGTCTAAAAAACATAAATTTCTTTAATCATTTTGCCTCTTTTCTCTGTGCTTCAATTAATAAAAAATGGAAAGA ATCTAATAGAGTGGTACAGCACTGTTATTTTTCAAAGATGTGTTGCTATCCTGAAAATTCTGTAGGTTCTGTGGAAGTTCCAGTGTTCTC >37818_37818_1_HSP90B1-ALB_HSP90B1_chr12_104336987_ENST00000299767_ALB_chr4_74274478_ENST00000295897_length(amino acids)=550AA_BP= MRALWVLGLCCVLLTFGSVRADDEVDVDGTVEEDLGKSREGSRTDDEVVQREEEAIQLDGLNASQIRELREKSEKFAFQAEVNRMMKLII NSLYKNKEIFLRELISNASDALDKIRLISLTDENALSGNEELTVKIKCDKEKNLLHVTDTGVGMTREELVKNLGTIAKSGTSEFLNKMTE AQEDGQSTSELIGQFGVGFYSAFLVADKVIVTSKHNNDTQHIWESDSNEFSVIADPRGNTLGRGTTITLVLKEEASDYLELDTIKNLVKK YSQFINFPIYVWSSKTETVEEPMEEEEAAKEEKEESDDEAAVEEEEEEKKPKTKKVEKTVWDWELMNDIKPIWQRPSKEVEEDEYKAFYK SFSKESDDPMAYIHFTAEGEVTFKSILFVPTSAPRGLFDEYGSKKSDYIKLYVRRVFITDDFHDMMPKYLNFVKGVVDSDDLPLNVSRET LQQHKLLKVIRKKLVRKTLDMIKKIADDKYNDTFWKEFGTNIKLGVIEDHSNRTRLAKLLRFQSSHHPTDITSLDQYVERMKEKQDKIYF -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for HSP90B1-ALB |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for HSP90B1-ALB |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for HSP90B1-ALB |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
