
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:IBTK-GINM1 (FusionGDB2 ID:38175) |
Fusion Gene Summary for IBTK-GINM1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: IBTK-GINM1 | Fusion gene ID: 38175 | Hgene | Tgene | Gene symbol | IBTK | GINM1 | Gene ID | 25998 | 116254 |
| Gene name | inhibitor of Bruton tyrosine kinase | glycoprotein integral membrane 1 | |
| Synonyms | BTBD26|BTKI | C6orf72 | |
| Cytomap | 6q14.1 | 6q25.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | inhibitor of Bruton tyrosine kinaseBTK-binding proteinBruton agammaglobulinemia tyrosine kinase inhibitorinhibitor of Bruton's tyrosine kinase-alphainhibitor of Bruton's tyrosine kinase-betainhibitor of Bruton's tyrosine kinase-gamma | glycoprotein integral membrane protein 1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q9P2D0 | Q9NU53 | |
| Ensembl transtripts involved in fusion gene | ENST00000503631, ENST00000510291, ENST00000306270, | ENST00000367419, | |
| Fusion gene scores | * DoF score | 6 X 7 X 3=126 | 4 X 3 X 4=48 |
| # samples | 7 | 4 | |
| ** MAII score | log2(7/126*10)=-0.84799690655495 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(4/48*10)=-0.263034405833794 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: IBTK [Title/Abstract] AND GINM1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | IBTK(82900790)-GINM1(149899958), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | IBTK-GINM1 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. IBTK-GINM1 seems lost the major protein functional domain in Tgene partner, which is a essential gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | IBTK | GO:0001933 | negative regulation of protein phosphorylation | 11577348 |
| Hgene | IBTK | GO:0051209 | release of sequestered calcium ion into cytosol | 11577348 |
 Fusion gene breakpoints across IBTK (5'-gene) Fusion gene breakpoints across IBTK (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
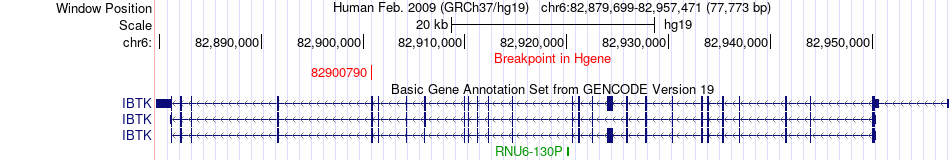 |
 Fusion gene breakpoints across GINM1 (3'-gene) Fusion gene breakpoints across GINM1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
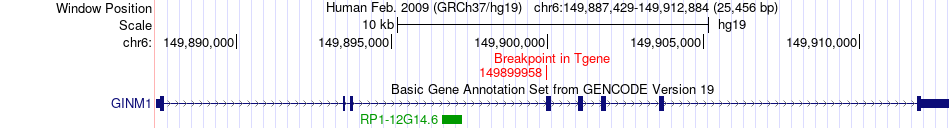 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | Non-Cancer | TCGA-IN-8462-11A | IBTK | chr6 | 82900790 | - | GINM1 | chr6 | 149899958 | + |
Top |
Fusion Gene ORF analysis for IBTK-GINM1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| Frame-shift | ENST00000503631 | ENST00000367419 | IBTK | chr6 | 82900790 | - | GINM1 | chr6 | 149899958 | + |
| Frame-shift | ENST00000510291 | ENST00000367419 | IBTK | chr6 | 82900790 | - | GINM1 | chr6 | 149899958 | + |
| In-frame | ENST00000306270 | ENST00000367419 | IBTK | chr6 | 82900790 | - | GINM1 | chr6 | 149899958 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000306270 | IBTK | chr6 | 82900790 | - | ENST00000367419 | GINM1 | chr6 | 149899958 | + | 5751 | 4125 | 484 | 4134 | 1216 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000306270 | ENST00000367419 | IBTK | chr6 | 82900790 | - | GINM1 | chr6 | 149899958 | + | 0.000108592 | 0.9998914 |
Top |
Fusion Genomic Features for IBTK-GINM1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| IBTK | chr6 | 82900789 | - | GINM1 | chr6 | 149899957 | + | 0.000156825 | 0.9998431 |
| IBTK | chr6 | 82900789 | - | GINM1 | chr6 | 149899957 | + | 0.000156825 | 0.9998431 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
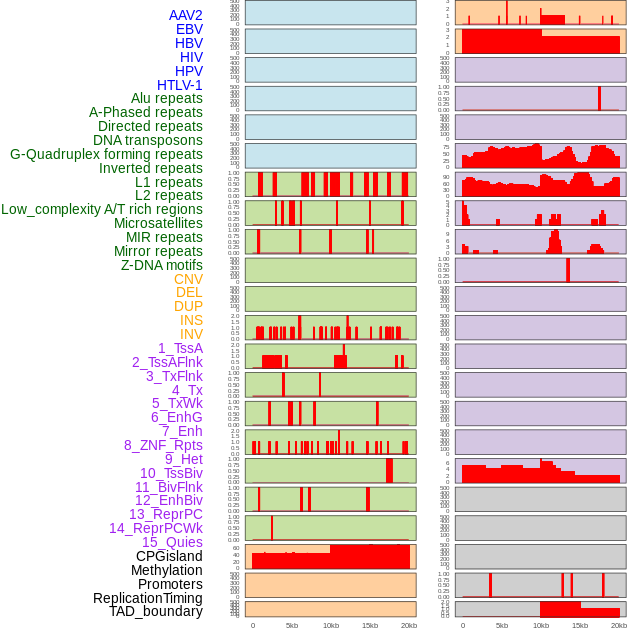 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
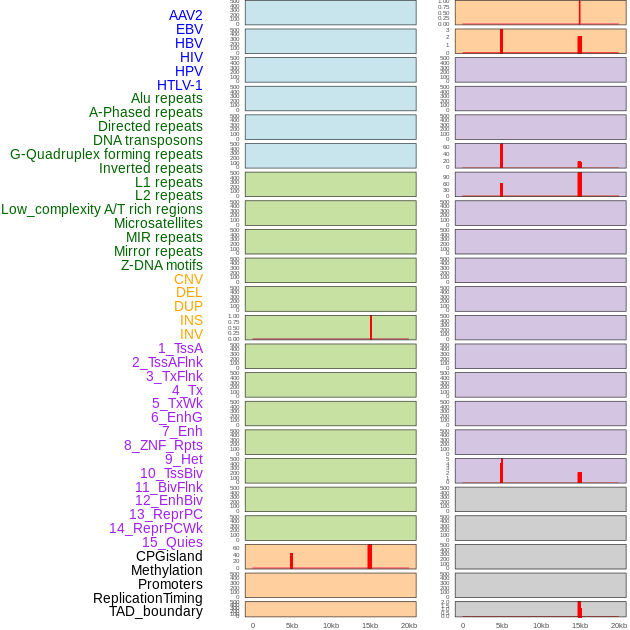 |
Top |
Fusion Protein Features for IBTK-GINM1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr6:82900790/chr6:149899958) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| IBTK | GINM1 |
| FUNCTION: Acts as an inhibitor of BTK tyrosine kinase activity, thereby playing a role in B-cell development. Down-regulates BTK kinase activity, leading to interference with BTK-mediated calcium mobilization and NF-kappa-B-driven transcription. {ECO:0000269|PubMed:11577348}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | IBTK | chr6:82900790 | chr6:149899958 | ENST00000306270 | - | 25 | 29 | 564_644 | 1191 | 1354.0 | Domain | BTB 1 |
| Hgene | IBTK | chr6:82900790 | chr6:149899958 | ENST00000306270 | - | 25 | 29 | 768_836 | 1191 | 1354.0 | Domain | BTB 2 |
| Hgene | IBTK | chr6:82900790 | chr6:149899958 | ENST00000306270 | - | 25 | 29 | 141_194 | 1191 | 1354.0 | Repeat | Note=RCC1 1 |
| Hgene | IBTK | chr6:82900790 | chr6:149899958 | ENST00000306270 | - | 25 | 29 | 195_246 | 1191 | 1354.0 | Repeat | Note=RCC1 2 |
| Hgene | IBTK | chr6:82900790 | chr6:149899958 | ENST00000306270 | - | 25 | 29 | 248_301 | 1191 | 1354.0 | Repeat | Note=RCC1 3 |
| Hgene | IBTK | chr6:82900790 | chr6:149899958 | ENST00000306270 | - | 25 | 29 | 51_80 | 1191 | 1354.0 | Repeat | Note=ANK 1 |
| Hgene | IBTK | chr6:82900790 | chr6:149899958 | ENST00000306270 | - | 25 | 29 | 806_835 | 1191 | 1354.0 | Repeat | Note=ANK 3 |
| Hgene | IBTK | chr6:82900790 | chr6:149899958 | ENST00000306270 | - | 25 | 29 | 85_114 | 1191 | 1354.0 | Repeat | Note=ANK 2 |
| Tgene | GINM1 | chr6:82900790 | chr6:149899958 | ENST00000367419 | 2 | 8 | 290_330 | 92 | 331.0 | Topological domain | Cytoplasmic | |
| Tgene | GINM1 | chr6:82900790 | chr6:149899958 | ENST00000367419 | 2 | 8 | 269_289 | 92 | 331.0 | Transmembrane | Helical |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | GINM1 | chr6:82900790 | chr6:149899958 | ENST00000367419 | 2 | 8 | 24_268 | 92 | 331.0 | Topological domain | Extracellular |
Top |
Fusion Gene Sequence for IBTK-GINM1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >38175_38175_1_IBTK-GINM1_IBTK_chr6_82900790_ENST00000306270_GINM1_chr6_149899958_ENST00000367419_length(transcript)=5751nt_BP=4125nt CTTCCTCTACATCCCCGTTCCCGATTCCTGTAGTAGCGGCTGTATTGCAGCCGCCTGCCGAACTGACCCGGGTCTGGGGACTGGCCCCTC TGGCGCCGTTCGGTTTCTCTTATTGCCTTCACTGAGGATGAGTCCCTTTGTGGCTCTATGTGGACCCTGCGGAATCCACCGGCGCAGTTT CATCTAGCGACTGGTCACCCTTGCAATTATGGATATTTAAAAGGGTCAGACAGTGTGGAGGGGGAGTTCCCCTCCTCACTCCCCCTTGGT GCTTGACTCCAGGAATAATTTATAAACTGTGGAATTTTTTTAAATGAAGAACTTGTATTTGATATGAACTTTATAGAGCTATTTATAATT TTTTTGATTTAAGTGCCAAAAAATTGTATAAAGATATATAGTTTTATACTATTGTCAGGAGGATTTAAATTATCCTAAAAAGGTAATTTA TTCTCTGTAACTTCCTCAATAGCACCTTTGTGTCCTGGCTTTTTCATTTTTTAAAATTAGTTTTCACGATTCTGAAGTAAGTGGTATAAA AACAGTTAGGATGAGTTCACCCATGCCTGACTGCACATCAAAGTGTCGATCCCTGAAGCATGCTTTGGATGTCCTTTCTGTGGTAACAAA GGGGAGCGAAAACCAGATTAAGGCCTTTCTCTCCAGTCATTGTTACAATGCTGCAACTATCAAGGATGTTTTTGGCAGGAATGCCCTCCA CCTTGTTTCCTCCTGTGGAAAGAAAGGAGTGTTAGATTGGCTTATTCAGAAAGGAGTGGATCTGTTGGTGAAAGACAAAGAGTCTGGATG GACAGCATTGCACAGAAGCATTTTTTATGGACATATTGATTGTGTTTGGTCTCTATTGAAGCATGGTGTTAGTCTGTATATTCAAGATAA AGAAGGCTTGTCAGCTTTGGATCTTGTAATGAAGGATAGACCAACTCATGTAGTATTCAAGAATACTGATCCTACAGATGTTTATACTTG GGGCGATAATACAAATTTTACCCTGGGTCATGGAAGCCAGAATAGCAAACATCATCCAGAGTTGGTGGATCTGTTCTCCAGGAGTGGGAT TTATATCAAGCAGGTGGTGCTTTGTAAATTTCACTCCGTGTTTCTGTCTCAGAAAGGGCAGGTTTATACCTGTGGTCATGGTCCTGGAGG GCGATTAGGACATGGAGATGAACAGACATGCTTGGTCCCTCGGCTTGTGGAAGGACTGAATGGTCATAATTGTTCCCAAGTGGCAGCTGC TAAGGATCATACTGTTGTATTAACTGAAGATGGATGTGTTTATACATTTGGTCTAAACATTTTTCATCAATTAGGAATTATTCCACCGCC TTCCAGTTGTAATGTACCCAGACAGATACAGGCAAAATATCTGAAAGGAAGGACAATCATTGGCGTTGCAGCAGGCAGGTTTCATACAGT CCTATGGACTAGAGAAGCTGTTTACACTATGGGACTAAATGGTGGACAACTGGGTTGTTTGCTAGATCCCAATGGAGAAAAGTGTGTAAC TGCTCCTCGTCAGGTCTCTGCCCTTCACCATAAAGACATTGCTCTGTCTTTGGTTGCTGCAAGTGATGGAGCTACAGTCTGTGTTACCAC AAGGGGAGATATTTACTTACTTGCAGACTATCAGTGCAAGAAGATGGCTTCTAAACAGTTGAACTTGAAAAAAGTTCTTGTGTCTGGGGG TCATATGGAATACAAGGTTGATCCTGAACATTTGAAAGAAAATGGGGGTCAAAAAATTTGCATTCTTGCAATGGATGGAGCTGGAAGGGT GTTTTGCTGGAGATCAGTCAACAGTTCTCTGAAGCAGTGTCGATGGGCCTATCCACGTCAGGTCTTCATTTCTGATATTGCTTTAAATAG AAATGAAATTCTATTTGTTACGCAAGATGGAGAAGGATTTAGAGGGAGATGGTTTGAAGAGAAAAGAAAGAGTTCTGAAAAGAAAGAGAT TTTATCAAACCTTCACAATTCCTCATCAGATGTGTCTTATGTCTCTGATATAAATAGTGTGTATGAAAGAATTCGACTTGAGAAACTTAC CTTTGCACATAGAGCTGTTAGTGTCAGCACAGATCCAAGTGGATGCAACTTTGCAATCCTGCAGTCAGATCCTAAAACAAGCCTTTATGA AATTCCAGCTGTGTCCTCATCATCCTTTTTTGAAGAGTTTGGCAAACTGTTGAGGGAAGCAGATGAAATGGACAGCATTCATGATGTGAC ATTTCAAGTTGGCAATAGACTCTTCCCTGCACATAAATATATTTTGGCAGTGCATTCTGATTTTTTTCAGAAATTGTTTCTTTCAGATGG TAATACTTCAGAATTTACAGATATTTACCAGAAAGATGAAGATTCTGCAGGGTGCCATCTCTTTGTGGTAGAGAAGGTTCATCCTGACAT GTTTGAATACCTTTTACAATTTATATACACAGATACTTGTGACTTTTTAACTCATGGCTTCAAACCAAGAATACACTTAAACAAAAACCC AGAAGAATATCAGGGAACTCTGAATTCTCATTTGAATAAAGTGAATTTCCATGAAGATGATAACCAGAAGTCTGCATTTGAAGTTTACAA AAGTAATCAAGCTCAAACAGTTAGTGAGAGGCAGAAGAGCAAACCTAAATCTTGTAAAAAAGGAAAAAATATTAGGGAAGATGATCCTGT AAGAATGTTGCAAACTGTTGCAAAGAAATTCGACTTCAGTAATTTGAGTAGTAGGTTAGATGGAGTCAGATTTGAAAATGAAAAAATTAA TGTTATTGCCAAGAACACTGGTAATAAACTGAAGCTAAGTCAGAAAAAATGTTCATTTCTGTGTGACGTGACCATGAAATCAGTGGATGG AAAGGAATTTCCTTGTCATAAATGTGTTCTTTGTGCTAGACTTGAATATTTTCATAGTATGCTGAGTAGCTCATGGATTGAGGCTTCCAG TTGTGCAGCTCTGGAAATGCCAATACATTCTGATATATTAAAAGTTATTTTGGACTACCTCTATACTGATGAAGCTGTGGTGATAAAAGA ATCTCAAAATGTAGATTTTATTTGTAGTGTTCTTGTGGTGGCTGATCAACTTCTCATAACCCGGTTGAAAGAGATTTGTGAAGTAGCATT AACTGAAAAACTTACCCTGAAGAATGCTGCTATGCTACTGGAATTTGCAGCAATGTATAGTGCAAAACAGTTGAAACTGTCTTGTTTACA GTTTATAGGATTGAATATGGCAGCTTTACTTGAAGCAAGGTCTCTTGATGTTTTAAGCGATGGTGTTTTGAAGGATCTTTCTGAGTTTTA CCGGAAAATGATTCCAGCAATGGATAGAAGAGTCATTACACCATATCAAGATGGACCAGATATTAGCTATTTGGAAGTAGAAGATGGAGA TATCTTCTTGAAAGAAGAAATAAATATGGAACAAAATCATTCGGAAACTATGTTCAAGAAAGCAAAAACAAAAGCTAAAAAGAAGCCACG TAAACGTTCAGATAGTTCTGGAGGTTATAACCTTTCAGATATTATTCAGAGTCCATCATCTACAGGATTATTAAAGTCTGGTAAGACCAA TTCTGTGGAATCTCTTCCAGAACTGTTGACATCAGACTCTGAAGGAAGCTATGCAGGAGTGGGTAGTCCTAGAGATTTACAGTCCCCTGA TTTCACAACAGGATTTCATTCAGATAAGATTGAGGCGAAAGTCAAACCGTATGTTAATGGTACATCTCCTGTGTATTCTAGGGAAGATTT AAAACCATGGGAAAAGTCACCAATACTTAAAATATCTGCTCCACAGCCTATTCCCAGTAACAGAATTGATACTACCAGCTCTGCCAGTTG GGTTGCTGGTTCTTTCAGTCCTGTCAGCCCTCCTGTTGTGGATCTCAGAACTATCATGGAAATAGAAGAAAGTAGACAAAAATGTGGAGC TACACCAAAGTCACATTTAGGCAAAACAGTTTCTCATGGAGTTAAACTTTCTCAGAAGCAACGAAAAATGATTGCATTGACTACCAAGGA AAACAATTCAGGAATGAATAGCATGGAAACAGTTTTATTCACTCCTTCAAAAGCCCCCAAACCAGTGAATGCATGTGAAGAATGAAAATC TTGAAAATTTGGAGGAAAAAGAATATTTTGGAATTGTCAGTGTAAGGATTTTAGTTCATGAGTGGCCTATGACATCTGGTTCCAGTTTGC AACTAATTGTCATTCAAGAAGAGGTAGTAGAGATTGATGGAAAACAAGTTCAGCAAAAGGATGTCACTGAAATTGATATTTTAGTTAAGA ACCGGGGAGTACTCAGACATTCAAACTATACCCTCCCTTTGGAAGAAAGCATGCTCTACTCTATTTCTCGAGACAGTGACATTTTATTTA CCCTTCCTAACCTCTCCAAAAAAGAAAGTGTTAGTTCACTGCAAACCACTAGCCAGTATCTTATCAGGAATGTGGAAACCACTGTAGATG AAGATGTTTTACCTGGCAAGTTACCTGAAACTCCTCTCAGAGCAGAGCCGCCATCTTCATATAAGGTAATGTGTCAGTGGATGGAAAAGT TTAGAAAAGATCTGTGTAGGTTCTGGAGCAACGTTTTCCCAGTATTCTTTCAGTTTTTGAACATCATGGTGGTTGGAATTACAGGAGCAG CTGTGGTAATAACCATCTTAAAGGTGTTTTTCCCAGTTTCTGAATACAAAGGAATTCTTCAGTTGGATAAAGTGGACGTCATACCTGTGA CAGCTATCAACTTATATCCAGATGGTCCAGAGAAAAGAGCTGAAAACCTTGAAGATAAAACATGTATTTAAAACGCCATCTCATATCATG GACTCCGAAGTAGCCTGTTGCCTCCAAATTTGCCACTTGAATATAATTTTCTTTAAATCGTTAAGAATCAGTTTATACACTAGAGAAATT GCTAAACTCTAAGACTGCCTGAAAATTGACCTTTACAGTGCCAAGTTAAAGTTTACCTTATTCTCGGCCGGGTGCAGTGGCTCATGCCTG TAATCCCAGGACTTTGGGAGGCCAATGCGGGCGGATCACGAGGTCAGATCAAGACCATCCTGCCAACATGGTGAAACCCTGTCTCTACTA AAAAAAATAAAAAAATTAGCTGGGTGTGGCGGTGCACGCCTGTAGTCCCAGCTACTTGGGAGGCTGAGGCAGGAGAATTGCTTGAACCCG GGAGGCGGAGGCTGCAGTGAGCCAAGATCACGCCACTGCACTCCAGCCTGGGTGACAGAGCGAGACTCTGTTTCAAAAAAAAAAAAGTTG ACCTTATTCTCTAAAAGGGCTGGCTATTCATATGATGAATTGTTAAGGAAAACTTAAAGTGGAAGAGAACACATGTGAAGAGACTTTGAA ATTATCAAAAGAAAAAAAAAAGACCAGACAAAATCTCATGTGCCAATAACTTTTCAAGGTGCCTTTGTTAAGGAAATTATATCCACTTAA TTACTATAATATATAAGACTTTATGAAAAGCACTTTATAAAATTCTAATTTAAAAGGTCAAGAACTTAACACTTATTTTTTTATTCAGTA CAGTGTGCCTTTATATTAAGTATGTGACTCAGTATAAAATCACTGAAGTGTATCATGATTTGATTGTATCCTGTACCAAGACTACTTACC >38175_38175_1_IBTK-GINM1_IBTK_chr6_82900790_ENST00000306270_GINM1_chr6_149899958_ENST00000367419_length(amino acids)=1216AA_BP= MAFSFFKISFHDSEVSGIKTVRMSSPMPDCTSKCRSLKHALDVLSVVTKGSENQIKAFLSSHCYNAATIKDVFGRNALHLVSSCGKKGVL DWLIQKGVDLLVKDKESGWTALHRSIFYGHIDCVWSLLKHGVSLYIQDKEGLSALDLVMKDRPTHVVFKNTDPTDVYTWGDNTNFTLGHG SQNSKHHPELVDLFSRSGIYIKQVVLCKFHSVFLSQKGQVYTCGHGPGGRLGHGDEQTCLVPRLVEGLNGHNCSQVAAAKDHTVVLTEDG CVYTFGLNIFHQLGIIPPPSSCNVPRQIQAKYLKGRTIIGVAAGRFHTVLWTREAVYTMGLNGGQLGCLLDPNGEKCVTAPRQVSALHHK DIALSLVAASDGATVCVTTRGDIYLLADYQCKKMASKQLNLKKVLVSGGHMEYKVDPEHLKENGGQKICILAMDGAGRVFCWRSVNSSLK QCRWAYPRQVFISDIALNRNEILFVTQDGEGFRGRWFEEKRKSSEKKEILSNLHNSSSDVSYVSDINSVYERIRLEKLTFAHRAVSVSTD PSGCNFAILQSDPKTSLYEIPAVSSSSFFEEFGKLLREADEMDSIHDVTFQVGNRLFPAHKYILAVHSDFFQKLFLSDGNTSEFTDIYQK DEDSAGCHLFVVEKVHPDMFEYLLQFIYTDTCDFLTHGFKPRIHLNKNPEEYQGTLNSHLNKVNFHEDDNQKSAFEVYKSNQAQTVSERQ KSKPKSCKKGKNIREDDPVRMLQTVAKKFDFSNLSSRLDGVRFENEKINVIAKNTGNKLKLSQKKCSFLCDVTMKSVDGKEFPCHKCVLC ARLEYFHSMLSSSWIEASSCAALEMPIHSDILKVILDYLYTDEAVVIKESQNVDFICSVLVVADQLLITRLKEICEVALTEKLTLKNAAM LLEFAAMYSAKQLKLSCLQFIGLNMAALLEARSLDVLSDGVLKDLSEFYRKMIPAMDRRVITPYQDGPDISYLEVEDGDIFLKEEINMEQ NHSETMFKKAKTKAKKKPRKRSDSSGGYNLSDIIQSPSSTGLLKSGKTNSVESLPELLTSDSEGSYAGVGSPRDLQSPDFTTGFHSDKIE AKVKPYVNGTSPVYSREDLKPWEKSPILKISAPQPIPSNRIDTTSSASWVAGSFSPVSPPVVDLRTIMEIEESRQKCGATPKSHLGKTVS -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for IBTK-GINM1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for IBTK-GINM1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for IBTK-GINM1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
