
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:IQGAP1-CIB1 (FusionGDB2 ID:40157) |
Fusion Gene Summary for IQGAP1-CIB1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: IQGAP1-CIB1 | Fusion gene ID: 40157 | Hgene | Tgene | Gene symbol | IQGAP1 | CIB1 | Gene ID | 8826 | 10519 |
| Gene name | IQ motif containing GTPase activating protein 1 | calcium and integrin binding 1 | |
| Synonyms | HUMORFA01|SAR1|p195 | CIB|CIBP|KIP1|PRKDCIP|SIP2-28 | |
| Cytomap | 15q26.1 | 15q26.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ras GTPase-activating-like protein IQGAP1RasGAP-like with IQ motifs | calcium and integrin-binding protein 1DNA-PK interaction proteinDNA-PKcs-interacting proteinDNA-dependent protein kinase interacting proteinSNK-interacting protein 2-28calcium and integrin binding 1 (calmyrin)testicular secretory protein Li 9 | |
| Modification date | 20200327 | 20200313 | |
| UniProtAcc | P46940 | Q99828 | |
| Ensembl transtripts involved in fusion gene | ENST00000268182, ENST00000560020, ENST00000560738, | ENST00000328649, | |
| Fusion gene scores | * DoF score | 27 X 20 X 12=6480 | 4 X 5 X 2=40 |
| # samples | 32 | 6 | |
| ** MAII score | log2(32/6480*10)=-4.33985000288463 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(6/40*10)=0.584962500721156 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: IQGAP1 [Title/Abstract] AND CIB1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | IQGAP1(90969498)-CIB1(90774739), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | IQGAP1 | GO:0071277 | cellular response to calcium ion | 18567582 |
| Tgene | CIB1 | GO:0006974 | cellular response to DNA damage stimulus | 12011095 |
| Tgene | CIB1 | GO:0007026 | negative regulation of microtubule depolymerization | 21215777 |
| Tgene | CIB1 | GO:0007113 | endomitotic cell cycle | 21264284 |
| Tgene | CIB1 | GO:0010977 | negative regulation of neuron projection development | 21215777 |
| Tgene | CIB1 | GO:0033630 | positive regulation of cell adhesion mediated by integrin | 11756406 |
| Tgene | CIB1 | GO:0070886 | positive regulation of calcineurin-NFAT signaling cascade | 20639889 |
| Tgene | CIB1 | GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading | 11756406 |
| Tgene | CIB1 | GO:1990090 | cellular response to nerve growth factor stimulus | 21215777 |
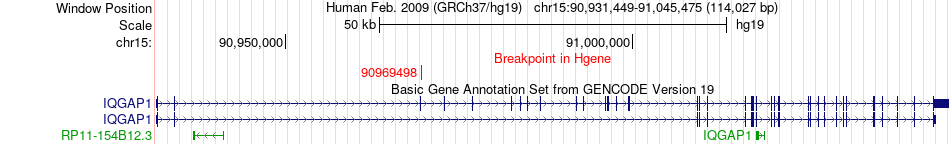
 Fusion gene breakpoints across IQGAP1 (5'-gene) Fusion gene breakpoints across IQGAP1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
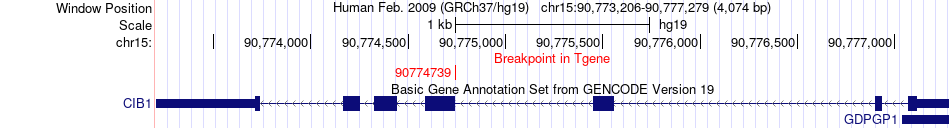
 Fusion gene breakpoints across CIB1 (3'-gene) Fusion gene breakpoints across CIB1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-E9-A1NF-01A | IQGAP1 | chr15 | 90969498 | + | CIB1 | chr15 | 90774739 | - |
Top |
Fusion Gene ORF analysis for IQGAP1-CIB1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000268182 | ENST00000328649 | IQGAP1 | chr15 | 90969498 | + | CIB1 | chr15 | 90774739 | - |
| intron-3CDS | ENST00000560020 | ENST00000328649 | IQGAP1 | chr15 | 90969498 | + | CIB1 | chr15 | 90774739 | - |
| intron-3CDS | ENST00000560738 | ENST00000328649 | IQGAP1 | chr15 | 90969498 | + | CIB1 | chr15 | 90774739 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000268182 | IQGAP1 | chr15 | 90969498 | + | ENST00000328649 | CIB1 | chr15 | 90774739 | - | 1326 | 436 | 124 | 816 | 230 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000268182 | ENST00000328649 | IQGAP1 | chr15 | 90969498 | + | CIB1 | chr15 | 90774739 | - | 0.001027573 | 0.9989724 |
Top |
Fusion Genomic Features for IQGAP1-CIB1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
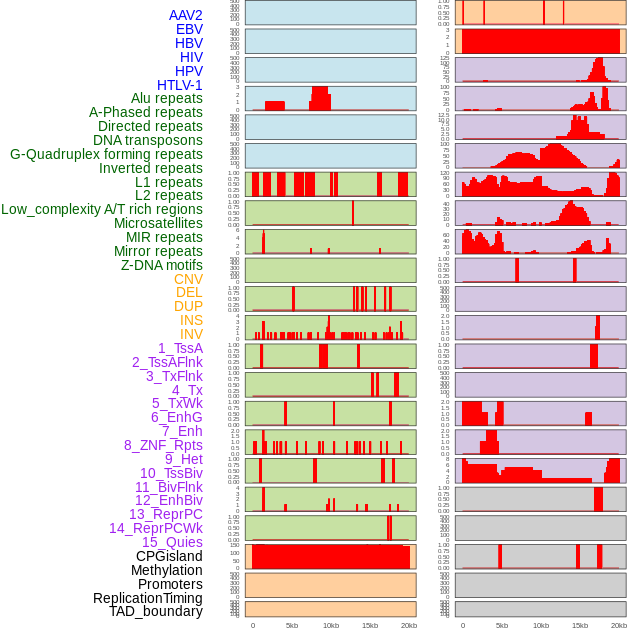 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for IQGAP1-CIB1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr15:90969498/chr15:90774739) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| IQGAP1 | CIB1 |
| FUNCTION: Plays a crucial role in regulating the dynamics and assembly of the actin cytoskeleton. Binds to activated CDC42 but does not stimulate its GTPase activity. It associates with calmodulin. Could serve as an assembly scaffold for the organization of a multimolecular complex that would interface incoming signals to the reorganization of the actin cytoskeleton at the plasma membrane. May promote neurite outgrowth (PubMed:15695813). May play a possible role in cell cycle regulation by contributing to cell cycle progression after DNA replication arrest (PubMed:20883816). {ECO:0000269|PubMed:15695813, ECO:0000269|PubMed:20883816}. | FUNCTION: Calcium-binding protein that plays a role in the regulation of numerous cellular processes, such as cell differentiation, cell division, cell proliferation, cell migration, thrombosis, angiogenesis, cardiac hypertrophy and apoptosis. Involved in bone marrow megakaryocyte differentiation by negatively regulating thrombopoietin-mediated signaling pathway. Participates in the endomitotic cell cycle of megakaryocyte, a form of mitosis in which both karyokinesis and cytokinesis are interrupted. Plays a role in integrin signaling by negatively regulating alpha-IIb/beta3 activation in thrombin-stimulated megakaryocytes preventing platelet aggregation. Up-regulates PTK2/FAK1 activity, and is also needed for the recruitment of PTK2/FAK1 to focal adhesions; it thus appears to play an important role in focal adhesion formation. Positively regulates cell migration on fibronectin in a CDC42-dependent manner, the effect being negatively regulated by PAK1. Functions as a negative regulator of stress activated MAP kinase (MAPK) signaling pathways. Down-regulates inositol 1,4,5-trisphosphate receptor-dependent calcium signaling. Involved in sphingosine kinase SPHK1 translocation to the plasma membrane in a N-myristoylation-dependent manner preventing TNF-alpha-induced apoptosis. Regulates serine/threonine-protein kinase PLK3 activity for proper completion of cell division progression. Plays a role in microtubule (MT) dynamics during neuronal development; disrupts the MT depolymerization activity of STMN2 attenuating NGF-induced neurite outgrowth and the MT reorganization at the edge of lamellipodia. Promotes cardiomyocyte hypertrophy via activation of the calcineurin/NFAT signaling pathway. Stimulates calcineurin PPP3R1 activity by mediating its anchoring to the sarcolemma. In ischemia-induced (pathological or adaptive) angiogenesis, stimulates endothelial cell proliferation, migration and microvessel formation by activating the PAK1 and ERK1/ERK2 signaling pathway. Promotes also cancer cell survival and proliferation. May regulate cell cycle and differentiation of spermatogenic germ cells, and/or differentiation of supporting Sertoli cells.; FUNCTION: [Isoform 2]: Plays a regulatory role in angiogenesis and tumor growth by mediating PKD/PRKD2-induced vascular endothelial growth factor A (VEGFA) secretion. {ECO:0000269|PubMed:23503467}.; FUNCTION: (Microbial infection) Involved in keratinocyte-intrinsic immunity to human beta-papillomaviruses (HPVs). {ECO:0000269|PubMed:30068544}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | CIB1 | chr15:90969498 | chr15:90774739 | ENST00000328649 | 2 | 7 | 116_127 | 65 | 192.0 | Calcium binding | Note=1 | |
| Tgene | CIB1 | chr15:90969498 | chr15:90774739 | ENST00000328649 | 2 | 7 | 161_172 | 65 | 192.0 | Calcium binding | Note=2 | |
| Tgene | CIB1 | chr15:90969498 | chr15:90774739 | ENST00000328649 | 2 | 7 | 103_138 | 65 | 192.0 | Domain | EF-hand 1 | |
| Tgene | CIB1 | chr15:90969498 | chr15:90774739 | ENST00000328649 | 2 | 7 | 148_183 | 65 | 192.0 | Domain | EF-hand 2 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | IQGAP1 | chr15:90969498 | chr15:90774739 | ENST00000268182 | + | 3 | 38 | 1004_1237 | 104 | 1658.0 | Domain | Ras-GAP |
| Hgene | IQGAP1 | chr15:90969498 | chr15:90774739 | ENST00000268182 | + | 3 | 38 | 44_159 | 104 | 1658.0 | Domain | Calponin-homology (CH) |
| Hgene | IQGAP1 | chr15:90969498 | chr15:90774739 | ENST00000268182 | + | 3 | 38 | 679_712 | 104 | 1658.0 | Domain | WW |
| Hgene | IQGAP1 | chr15:90969498 | chr15:90774739 | ENST00000268182 | + | 3 | 38 | 745_774 | 104 | 1658.0 | Domain | IQ 1 |
| Hgene | IQGAP1 | chr15:90969498 | chr15:90774739 | ENST00000268182 | + | 3 | 38 | 775_804 | 104 | 1658.0 | Domain | IQ 2 |
| Hgene | IQGAP1 | chr15:90969498 | chr15:90774739 | ENST00000268182 | + | 3 | 38 | 805_834 | 104 | 1658.0 | Domain | IQ 3 |
| Hgene | IQGAP1 | chr15:90969498 | chr15:90774739 | ENST00000268182 | + | 3 | 38 | 835_864 | 104 | 1658.0 | Domain | IQ 4 |
| Hgene | IQGAP1 | chr15:90969498 | chr15:90774739 | ENST00000268182 | + | 3 | 38 | 1276_1657 | 104 | 1658.0 | Region | Note=C2 |
| Hgene | IQGAP1 | chr15:90969498 | chr15:90774739 | ENST00000268182 | + | 3 | 38 | 956_1274 | 104 | 1658.0 | Region | Note=C1 |
Top |
Fusion Gene Sequence for IQGAP1-CIB1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >40157_40157_1_IQGAP1-CIB1_IQGAP1_chr15_90969498_ENST00000268182_CIB1_chr15_90774739_ENST00000328649_length(transcript)=1326nt_BP=436nt GACGGCACGGGGCGGGGCCTCGGGGACCCCGGCAAGCCCGCGCACTTGGCAGGAGCTGTAGCTACCGCCGTCCGCGCCTCCAAGGTTTCA CGGCTTCCTCAGCAGAGACTCGGGCTCGTCCGCCATGTCCGCCGCAGACGAGGTTGACGGGCTGGGCGTGGCCCGGCCGCACTATGGCTC TGTCCTGGATAATGAAAGACTTACTGCAGAGGAGATGGATGAAAGGAGACGTCAGAACGTGGCTTATGAGTACCTTTGTCATTTGGAAGA AGCGAAGAGGTGGATGGAAGCATGCCTAGGGGAAGATCTGCCTCCCACCACAGAACTGGAGGAGGGGCTTAGGAATGGGGTCTACCTTGC CAAACTGGGGAACTTCTTCTCTCCCAAAGTAGTGTCCCTGAAAAAAATCTATGATCGAGAACAGACCAGATACAAGGCCAACCCCTTCAA GGAGCGAATCTGCAGGGTCTTCTCCACATCCCCAGCCAAAGACAGCCTTAGCTTTGAGGACTTCCTGGATCTCCTCAGTGTGTTCAGTGA CACAGCCACGCCAGACATCAAGTCCCATTATGCCTTCCGCATCTTTGACTTTGATGATGACGGAACCTTGAACAGAGAAGACCTGAGCCG GCTGGTGAACTGCCTCACGGGAGAGGGCGAGGACACACGGCTTAGTGCGTCTGAGATGAAGCAGCTCATCGACAACATCCTGGAGGAGTC TGACATTGACAGGGATGGAACCATCAACCTCTCTGAGTTCCAGCACGTCATCTCCCGTTCTCCAGACTTTGCCAGCTCCTTTAAGATTGT CCTGTGACAGCAGCCCCAGCGTGTGTCCTGGCACCCTGTCCAAGAACCTTTCTACTGCTGAGCTGTGGCCAAGGTCAAGCCTGTGTTGCC AGTGCGGGCCAAGCTGGCCCAGCCTGGAGCTGGCGCTGTGCAGCCTCACCCCGGGCAGGGGCGGCCCTCGTTGTCAGGGCCTCTCCTCAC TGCTGTTGTCATTGCTCCGTTTGTGTTTGTACTAATCAGTAATAAAGGTTTAGAAGTTTGACCCTATGTGTGACATGAGATACACTGGTA TGAGGAAGGACTGGACTTCTCTTCTAAGAGCCTTCAATCATCCTAGGAATAAGCAGCATATCGAGCAGAGGCCAGCGGGAAGTCAAGCCG CACCCACGCTGGCTGTCCCCGTGGGAGGGTGTGAGGATGAAGGGGCAGCAGGAGGTGTGGGACTCGCCTGTATCCTGATTGGGGTGGTGG >40157_40157_1_IQGAP1-CIB1_IQGAP1_chr15_90969498_ENST00000268182_CIB1_chr15_90774739_ENST00000328649_length(amino acids)=230AA_BP=104 MSAADEVDGLGVARPHYGSVLDNERLTAEEMDERRRQNVAYEYLCHLEEAKRWMEACLGEDLPPTTELEEGLRNGVYLAKLGNFFSPKVV SLKKIYDREQTRYKANPFKERICRVFSTSPAKDSLSFEDFLDLLSVFSDTATPDIKSHYAFRIFDFDDDGTLNREDLSRLVNCLTGEGED -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for IQGAP1-CIB1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for IQGAP1-CIB1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for IQGAP1-CIB1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
