
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:IQGAP1-RASGRF1 (FusionGDB2 ID:40165) |
Fusion Gene Summary for IQGAP1-RASGRF1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: IQGAP1-RASGRF1 | Fusion gene ID: 40165 | Hgene | Tgene | Gene symbol | IQGAP1 | RASGRF1 | Gene ID | 8826 | 5923 |
| Gene name | IQ motif containing GTPase activating protein 1 | Ras protein specific guanine nucleotide releasing factor 1 | |
| Synonyms | HUMORFA01|SAR1|p195 | CDC25|CDC25L|GNRP|GRF1|GRF55|H-GRF55|PP13187|ras-GRF1 | |
| Cytomap | 15q26.1 | 15q25.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ras GTPase-activating-like protein IQGAP1RasGAP-like with IQ motifs | ras-specific guanine nucleotide-releasing factor 1Ras-specific guanine nucleotide-releasing factor, CDC25 homologRas-specific nucleotide exchange factor CDC25guanine nucleotide exchange factorguanine nucleotide-releasing factor 1guanine nucleotide-re | |
| Modification date | 20200327 | 20200313 | |
| UniProtAcc | P46940 | . | |
| Ensembl transtripts involved in fusion gene | ENST00000268182, ENST00000560020, ENST00000560738, | ENST00000560334, ENST00000394745, ENST00000419573, ENST00000558480, | |
| Fusion gene scores | * DoF score | 27 X 20 X 12=6480 | 24 X 19 X 4=1824 |
| # samples | 32 | 24 | |
| ** MAII score | log2(32/6480*10)=-4.33985000288463 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(24/1824*10)=-2.92599941855622 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: IQGAP1 [Title/Abstract] AND RASGRF1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | IQGAP1(90934105)-RASGRF1(79298776), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | IQGAP1 | GO:0071277 | cellular response to calcium ion | 18567582 |
 Fusion gene breakpoints across IQGAP1 (5'-gene) Fusion gene breakpoints across IQGAP1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
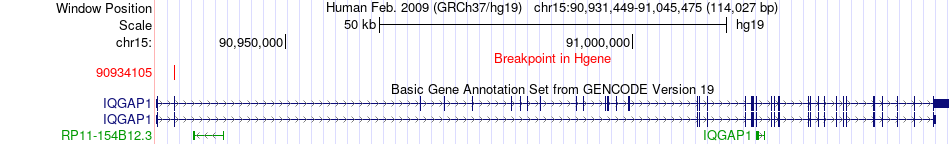 |
 Fusion gene breakpoints across RASGRF1 (3'-gene) Fusion gene breakpoints across RASGRF1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
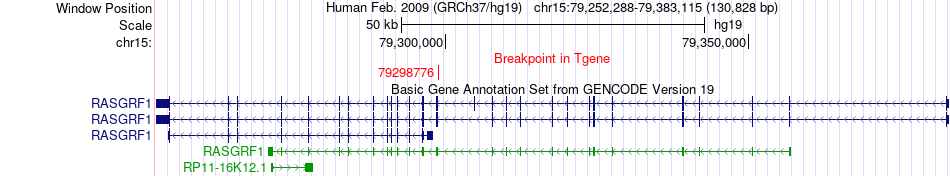 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SARC | TCGA-FX-A3TO-01A | IQGAP1 | chr15 | 90934105 | - | RASGRF1 | chr15 | 79298776 | - |
| ChimerDB4 | SARC | TCGA-FX-A3TO-01A | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
Top |
Fusion Gene ORF analysis for IQGAP1-RASGRF1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-5UTR | ENST00000268182 | ENST00000560334 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| 5CDS-intron | ENST00000268182 | ENST00000394745 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| In-frame | ENST00000268182 | ENST00000419573 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| In-frame | ENST00000268182 | ENST00000558480 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| intron-3CDS | ENST00000560020 | ENST00000419573 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| intron-3CDS | ENST00000560020 | ENST00000558480 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| intron-3CDS | ENST00000560738 | ENST00000419573 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| intron-3CDS | ENST00000560738 | ENST00000558480 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| intron-5UTR | ENST00000560020 | ENST00000560334 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| intron-5UTR | ENST00000560738 | ENST00000560334 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| intron-intron | ENST00000560020 | ENST00000394745 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
| intron-intron | ENST00000560738 | ENST00000394745 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000268182 | IQGAP1 | chr15 | 90934105 | + | ENST00000558480 | RASGRF1 | chr15 | 79298776 | - | 4424 | 279 | 124 | 2226 | 700 |
| ENST00000268182 | IQGAP1 | chr15 | 90934105 | + | ENST00000419573 | RASGRF1 | chr15 | 79298776 | - | 4433 | 279 | 124 | 2235 | 703 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000268182 | ENST00000558480 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - | 0.003707179 | 0.99629277 |
| ENST00000268182 | ENST00000419573 | IQGAP1 | chr15 | 90934105 | + | RASGRF1 | chr15 | 79298776 | - | 0.0037512 | 0.9962488 |
Top |
Fusion Genomic Features for IQGAP1-RASGRF1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
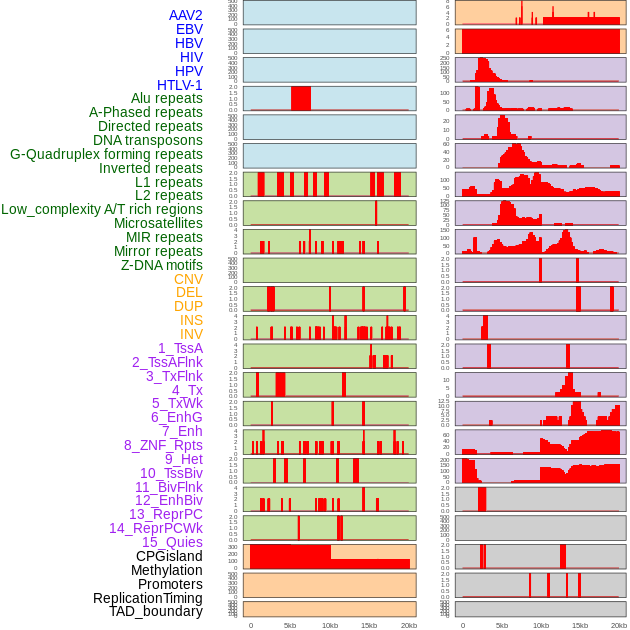 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for IQGAP1-RASGRF1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr15:90934105/chr15:79298776) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| IQGAP1 | . |
| FUNCTION: Plays a crucial role in regulating the dynamics and assembly of the actin cytoskeleton. Binds to activated CDC42 but does not stimulate its GTPase activity. It associates with calmodulin. Could serve as an assembly scaffold for the organization of a multimolecular complex that would interface incoming signals to the reorganization of the actin cytoskeleton at the plasma membrane. May promote neurite outgrowth (PubMed:15695813). May play a possible role in cell cycle regulation by contributing to cell cycle progression after DNA replication arrest (PubMed:20883816). {ECO:0000269|PubMed:15695813, ECO:0000269|PubMed:20883816}. | FUNCTION: Transcriptional activator which is required for calcium-dependent dendritic growth and branching in cortical neurons. Recruits CREB-binding protein (CREBBP) to nuclear bodies. Component of the CREST-BRG1 complex, a multiprotein complex that regulates promoter activation by orchestrating a calcium-dependent release of a repressor complex and a recruitment of an activator complex. In resting neurons, transcription of the c-FOS promoter is inhibited by BRG1-dependent recruitment of a phospho-RB1-HDAC1 repressor complex. Upon calcium influx, RB1 is dephosphorylated by calcineurin, which leads to release of the repressor complex. At the same time, there is increased recruitment of CREBBP to the promoter by a CREST-dependent mechanism, which leads to transcriptional activation. The CREST-BRG1 complex also binds to the NR2B promoter, and activity-dependent induction of NR2B expression involves a release of HDAC1 and recruitment of CREBBP (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | RASGRF1 | chr15:90934105 | chr15:79298776 | ENST00000394745 | 0 | 14 | 1038_1270 | 0 | 490.0 | Domain | Ras-GEF | |
| Tgene | RASGRF1 | chr15:90934105 | chr15:79298776 | ENST00000394745 | 0 | 14 | 204_229 | 0 | 490.0 | Domain | IQ | |
| Tgene | RASGRF1 | chr15:90934105 | chr15:79298776 | ENST00000394745 | 0 | 14 | 22_129 | 0 | 490.0 | Domain | PH 1 | |
| Tgene | RASGRF1 | chr15:90934105 | chr15:79298776 | ENST00000394745 | 0 | 14 | 240_426 | 0 | 490.0 | Domain | DH | |
| Tgene | RASGRF1 | chr15:90934105 | chr15:79298776 | ENST00000394745 | 0 | 14 | 467_584 | 0 | 490.0 | Domain | PH 2 | |
| Tgene | RASGRF1 | chr15:90934105 | chr15:79298776 | ENST00000394745 | 0 | 14 | 644_761 | 0 | 490.0 | Domain | N-terminal Ras-GEF |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | IQGAP1 | chr15:90934105 | chr15:79298776 | ENST00000268182 | + | 2 | 38 | 1004_1237 | 51 | 1658.0 | Domain | Ras-GAP |
| Hgene | IQGAP1 | chr15:90934105 | chr15:79298776 | ENST00000268182 | + | 2 | 38 | 44_159 | 51 | 1658.0 | Domain | Calponin-homology (CH) |
| Hgene | IQGAP1 | chr15:90934105 | chr15:79298776 | ENST00000268182 | + | 2 | 38 | 679_712 | 51 | 1658.0 | Domain | WW |
| Hgene | IQGAP1 | chr15:90934105 | chr15:79298776 | ENST00000268182 | + | 2 | 38 | 745_774 | 51 | 1658.0 | Domain | IQ 1 |
| Hgene | IQGAP1 | chr15:90934105 | chr15:79298776 | ENST00000268182 | + | 2 | 38 | 775_804 | 51 | 1658.0 | Domain | IQ 2 |
| Hgene | IQGAP1 | chr15:90934105 | chr15:79298776 | ENST00000268182 | + | 2 | 38 | 805_834 | 51 | 1658.0 | Domain | IQ 3 |
| Hgene | IQGAP1 | chr15:90934105 | chr15:79298776 | ENST00000268182 | + | 2 | 38 | 835_864 | 51 | 1658.0 | Domain | IQ 4 |
| Hgene | IQGAP1 | chr15:90934105 | chr15:79298776 | ENST00000268182 | + | 2 | 38 | 1276_1657 | 51 | 1658.0 | Region | Note=C2 |
| Hgene | IQGAP1 | chr15:90934105 | chr15:79298776 | ENST00000268182 | + | 2 | 38 | 956_1274 | 51 | 1658.0 | Region | Note=C1 |
Top |
Fusion Gene Sequence for IQGAP1-RASGRF1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >40165_40165_1_IQGAP1-RASGRF1_IQGAP1_chr15_90934105_ENST00000268182_RASGRF1_chr15_79298776_ENST00000419573_length(transcript)=4433nt_BP=279nt GACGGCACGGGGCGGGGCCTCGGGGACCCCGGCAAGCCCGCGCACTTGGCAGGAGCTGTAGCTACCGCCGTCCGCGCCTCCAAGGTTTCA CGGCTTCCTCAGCAGAGACTCGGGCTCGTCCGCCATGTCCGCCGCAGACGAGGTTGACGGGCTGGGCGTGGCCCGGCCGCACTATGGCTC TGTCCTGGATAATGAAAGACTTACTGCAGAGGAGATGGATGAAAGGAGACGTCAGAACGTGGCTTATGAGTACCTTTGTCATTTGGAAGA AGCGAAGAGGTCCGACGCCTCCTTATATTGTGATGATGTTGACATTCGCTTCAGCAAAACCATGAACTCCTGCAAAGTGCTGCAGATCCG CTACGCCAGTGTGGAGCGGCTGCTGGAGAGGCTGACGGACCTGCGCTTCCTGAGCATCGACTTCCTCAACACCTTCCTGCACTCCTACCG CGTCTTCACCACCGCCATCGTGGTCCTGGACAAGCTCATTACCATCTACAAGAAGCCTATCAGTGCCATTCCTGCCAGGTGGCTGAGGTC GCTGGAGCTCCTGTTTGCCAGTGGCCAGAACAATAAGCTCCTGTACGGTGAACCCCCCAAGTCCCCGCGCGCCACCCGCAAGTTCTCCTC GCCGCCACCTCTGTCCATCACCAAGACATCGTCACCGAGCCGCCGGCGGAAGCTCTCCCTGAACATCCCCATCATCACTGGCGGCAAGGC CCTGGACCTGGCCGCCCTCAGCTGCAACTCCAATGGCTACACCAGCATGTACTCGGCCATGTCACCCTTCAGCAAGGCCACGCTGGACAC CAGCAAGCTCTATGTGTCCAGCAGCTTCACCAACAAGATTCCAGATGAGGGCGATACGACCCCTGAGAAGCCCGAAGACCCTTCAGCGCT CAGCAAGCAGAGCTCAGAAGTCTCCATGAGAGAGGAGTCAGATATTGATCAAAACCAGAGTGATGATGGTGATACTGAAACATCACCAAC TAAATCTCCAACAACACCCAAATCAGTCAAAAACAAAAATTCTTCAGAGTTCCCACTCTTTTCCTATAACAATGGAGTCGTCATGACCTC CTGTCGTGAACTGGACAATAACCGCAGTGCCTTGTCGGCCGCCTCTGCCTTTGCCATAGCAACCGCCGGGGCCAACGAGGGCACCCCAAA CAAGGAGAAGTACCGGAGGATGTCCTTAGCCAGTGCAGGGTTTCCCCCAGACCAGAGGAATGGAGACAAGGAGTTTGTGATCCGCAGAGC AGCCACCAATCGTGTCTTGAACGTGCTCCGCCACTGGGTGTCCAAGCACTCTCAGGACTTTGAGACCAACGATGAGCTCAAATGCAAGGT GATCGGCTTCCTGGAAGAAGTCATGCACGACCCGGAGCTCCTGACCCAGGAGCGGAAGGCTGCAGCCAACATCATCAGGACTCTGACCCA GGAGGACCCAGGTGACAACCAGATCACGCTGGAGGAGATCACGCAGATGGCTGAAGGCGTGAAGGCTGAGCCCTTTGAAAACCACTCAGC CCTGGAGATCGCGGAGCAGCTGACCCTGCTAGATCACCTCGTCTTCAAGAAGATTCCTTATGAGGAGTTCTTCGGACAAGGATGGATGAA ACTGGAAAAGAATGAAAGGACCCCTTATATCATGAAAACCACTAAGCACTTCAATGACATCAGTAACTTGATTGCTTCAGAAATCATCCG CAATGAGGACATCAACGCCAGGGTGAGCGCCATCGAGAAGTGGGTGGCCGTAGCTGACATATGCCGCTGCCTCCACAACTACAATGCCGT ACTGGAGATCACCTCGTCCATGAACCGCAGTGCAATCTTCCGGCTCAAAAAGACGTGGCTCAAAGTCTCTAAGCAGACTAAAGCTTTGAT TGATAAGCTCCAAAAGCTTGTGTCATCTGAGGGCAGATTTAAGAATCTCAGAGAAGCTCTGAAAAATTGTGACCCACCCTGTGTCCCTTA CCTGGGGATGTACCTCACCGACCTGGCCTTCATCGAGGAGGGGACGCCCAATTACACGGAAGACGGCCTGGTCAACTTCTCCAAGATGAG GATGATATCCCATATTATCCGAGAGATTCGCCAGTTTCAACAAACTGCCTACAAAATAGAGCACCAAGCAAAGGTAACGCAATATTTACT GGACCAATCTTTTGTAATGGATGAAGAAAGCCTCTACGAGTCTTCTCTCCGAATAGAACCAAAACTCCCCACCTGAAGCTGAGCCCAGCC CAGACCCAGCTGCTCCCGGGGACATGTGCTAGATGATACTGTACATATTCGTTTGGTTTCACTGGATTTTCTTCTTCAGTATGTGCTTCT CCAAGAATACAAATCGTCCTTGTTCTTAGATTCCTGTAGAACCGGAATATGAATTTCTGCACCGTTTCAGACTTCGCCCACCCATCCCTC CCCTCGTCTCCTGCAGTGCCTGTTTCTTTTAAACAGCTGTACTCTTGGTCCCTCTTCCCTAGACTCCTCACTCTTCTCAGAGGGGAAACA GCACCCTTGCATACAAGAATCTTAGACTCCAGTCCTGCTTCCGTCCCTCCCCAGCCCAGGCTCCTGGGTTGAGGTGGCCACCCAGGCATC CTCCCAGCATGTTCCATGTAGTTGTTATAACTGGGTGGGGGGTGGGGGGCGGCGGGAGGGGGAAACCATACCCTGAAGTCCAGTCATTGA CAAATCTTCCCGCTGATGGTGATGTCAATTTCCAAAGCAGCTTCTCCAGCCAAAATCGCAGGTCTCAAAACTCCCCATGTGTCTCCCAGT AACCTGGACGGAAGGGAGTCCTCTCTGCTCAGAACCACACCCCTCGTACCAATGCTGCCATCTCGTTCAGGCCCCACGTCGCTCCTTGCT CTAGGATTAACACCAGATAGCATGAATAGCAGTCTCAATACAACTGGATGAGGCCCTAGACTTCCCGAGGAAATGGAGTCACAAACACCA AGCCACGTCGACTCTTGCCGACCACTGGCCCAGCATCCACTGAAAGTCGGGAACACAAAAATGTCACTTTTCCTCTTTAAGCTGCCTAAC GGTCACTACAAGCTTACATTGAGATGCTCTAAGTCTGTATACCTTGTGATAATTAGTACCACACTGTAGAATTAGGAAATCAATAACCAA GTGTTTTTATTGCATATACTGTAGTCCTGTCTAAATGCCTTCTAGCAGGAATATAGTAATGTAGTGCTTATTCTGTGAATATTACCATGT ATTTTTTACACTGTACAGTATAGAAAACACAGGTTAACTCCGAAGTGGCAGGCATGCTCCACTACAACAGACAGACAACTTTTGCTGTAA GCAATACCTAGTGGTATGAGAATTCTATTTCAAGTCCTAAAATATTTCAACTTACAGCTTGTTTGGAAAAAAATTTCCATTTTTGTAAAA ATATATGTTTCCTGATTACCTCTTGCTAATAAATCTATTCTATCTCAGGGTGAACAGCGAGTTTTGATGGGTTTCATTCGTTCAGTCAAG GAATATTTGATGAGCACTTGCTCCTTACCCAGCCCAGCCTGGGACCGGAGATCCAGAGTGTCAAGGACACCATACCTGCTTTTTAGGCAC CAACAGTTTGGTGGGGAAGACTGTTTCAGTTGCCTATTGCACCAAGAACAAGCCACTCCAAAACATAATGGCTTAAAACCATTTTTGTTA CCTTTCACAGTTCTATGGGTTGACTGGGTCCAGCTGTCAGTTCTTATTTGGAGTCTCTCATGAAGTTTCAGTCACTCAGCAACTGGGACT GGAGTCATCTGAAGGCTCAACTGGACCGGGTGGTGCTCTAGTTGGTGCACCAAGATGGCGCTGTCACATGGCTGGATGTTGATGTCTGCT GGGCTGCACTGGGGCTGTTCATGGGAGCATCCACACGTGGCCTCTCTGTGTGGCTTGGGCTCCTCACAGTCCGGAAGCTGGGTTCCAAGA GCGACTGTTCCAGCAGCCTCAGGCAGAAGCTACAAGGCTGCCCAGATGCCATCCTGCCCAGATGTGTCCTCACTGTCACCCAGGAAGGGA AGCCCCGGAACACCACCCTCAGTCACAGGCATGGGGCCCACTGCAAGGCACAACCCACTCCAGAGCTCCCAAGGCCGCCTCCCCTGGGAC CTGCAGCCAGCTGGCTCCATAACTGCTTTTCCGTTTCCTCCAGGCATTTTCATCCTAGCCCTACTTTGTGCTGGGATCTGAGGGGGATAG CAAGCGGAGAAAGAAGGGATCTAGAGGGAGAAGATTCTAGATCCTTCTTTTGACTGCCAGCTGATTTTGATTGCCAGCTTTACCTGCTTC ATGAGCTCAGCCAAGTCACTTAACCTCTCAGAGCTTCACTGTTCTCATCTGAAAAATGGGGATGAAAATGGTATCTCATTACAGAGTAGA >40165_40165_1_IQGAP1-RASGRF1_IQGAP1_chr15_90934105_ENST00000268182_RASGRF1_chr15_79298776_ENST00000419573_length(amino acids)=703AA_BP=51 MSAADEVDGLGVARPHYGSVLDNERLTAEEMDERRRQNVAYEYLCHLEEAKRSDASLYCDDVDIRFSKTMNSCKVLQIRYASVERLLERL TDLRFLSIDFLNTFLHSYRVFTTAIVVLDKLITIYKKPISAIPARWLRSLELLFASGQNNKLLYGEPPKSPRATRKFSSPPPLSITKTSS PSRRRKLSLNIPIITGGKALDLAALSCNSNGYTSMYSAMSPFSKATLDTSKLYVSSSFTNKIPDEGDTTPEKPEDPSALSKQSSEVSMRE ESDIDQNQSDDGDTETSPTKSPTTPKSVKNKNSSEFPLFSYNNGVVMTSCRELDNNRSALSAASAFAIATAGANEGTPNKEKYRRMSLAS AGFPPDQRNGDKEFVIRRAATNRVLNVLRHWVSKHSQDFETNDELKCKVIGFLEEVMHDPELLTQERKAAANIIRTLTQEDPGDNQITLE EITQMAEGVKAEPFENHSALEIAEQLTLLDHLVFKKIPYEEFFGQGWMKLEKNERTPYIMKTTKHFNDISNLIASEIIRNEDINARVSAI EKWVAVADICRCLHNYNAVLEITSSMNRSAIFRLKKTWLKVSKQTKALIDKLQKLVSSEGRFKNLREALKNCDPPCVPYLGMYLTDLAFI -------------------------------------------------------------- >40165_40165_2_IQGAP1-RASGRF1_IQGAP1_chr15_90934105_ENST00000268182_RASGRF1_chr15_79298776_ENST00000558480_length(transcript)=4424nt_BP=279nt GACGGCACGGGGCGGGGCCTCGGGGACCCCGGCAAGCCCGCGCACTTGGCAGGAGCTGTAGCTACCGCCGTCCGCGCCTCCAAGGTTTCA CGGCTTCCTCAGCAGAGACTCGGGCTCGTCCGCCATGTCCGCCGCAGACGAGGTTGACGGGCTGGGCGTGGCCCGGCCGCACTATGGCTC TGTCCTGGATAATGAAAGACTTACTGCAGAGGAGATGGATGAAAGGAGACGTCAGAACGTGGCTTATGAGTACCTTTGTCATTTGGAAGA AGCGAAGAGGTCCGACGCCTCCTTATATTGTGATGATGTTGACATTCGCTTCAGCAAAACCATGAACTCCTGCAAAGTGCTGCAGATCCG CTACGCCAGTGTGGAGCGGCTGCTGGAGAGGCTGACGGACCTGCGCTTCCTGAGCATCGACTTCCTCAACACCTTCCTGCACTCCTACCG CGTCTTCACCACCGCCATCGTGGTCCTGGACAAGCTCATTACCATCTACAAGAAGCCTATCAGTGCCATTCCTGCCAGGTCGCTGGAGCT CCTGTTTGCCAGTGGCCAGAACAATAAGCTCCTGTACGGTGAACCCCCCAAGTCCCCGCGCGCCACCCGCAAGTTCTCCTCGCCGCCACC TCTGTCCATCACCAAGACATCGTCACCGAGCCGCCGGCGGAAGCTCTCCCTGAACATCCCCATCATCACTGGCGGCAAGGCCCTGGACCT GGCCGCCCTCAGCTGCAACTCCAATGGCTACACCAGCATGTACTCGGCCATGTCACCCTTCAGCAAGGCCACGCTGGACACCAGCAAGCT CTATGTGTCCAGCAGCTTCACCAACAAGATTCCAGATGAGGGCGATACGACCCCTGAGAAGCCCGAAGACCCTTCAGCGCTCAGCAAGCA GAGCTCAGAAGTCTCCATGAGAGAGGAGTCAGATATTGATCAAAACCAGAGTGATGATGGTGATACTGAAACATCACCAACTAAATCTCC AACAACACCCAAATCAGTCAAAAACAAAAATTCTTCAGAGTTCCCACTCTTTTCCTATAACAATGGAGTCGTCATGACCTCCTGTCGTGA ACTGGACAATAACCGCAGTGCCTTGTCGGCCGCCTCTGCCTTTGCCATAGCAACCGCCGGGGCCAACGAGGGCACCCCAAACAAGGAGAA GTACCGGAGGATGTCCTTAGCCAGTGCAGGGTTTCCCCCAGACCAGAGGAATGGAGACAAGGAGTTTGTGATCCGCAGAGCAGCCACCAA TCGTGTCTTGAACGTGCTCCGCCACTGGGTGTCCAAGCACTCTCAGGACTTTGAGACCAACGATGAGCTCAAATGCAAGGTGATCGGCTT CCTGGAAGAAGTCATGCACGACCCGGAGCTCCTGACCCAGGAGCGGAAGGCTGCAGCCAACATCATCAGGACTCTGACCCAGGAGGACCC AGGTGACAACCAGATCACGCTGGAGGAGATCACGCAGATGGCTGAAGGCGTGAAGGCTGAGCCCTTTGAAAACCACTCAGCCCTGGAGAT CGCGGAGCAGCTGACCCTGCTAGATCACCTCGTCTTCAAGAAGATTCCTTATGAGGAGTTCTTCGGACAAGGATGGATGAAACTGGAAAA GAATGAAAGGACCCCTTATATCATGAAAACCACTAAGCACTTCAATGACATCAGTAACTTGATTGCTTCAGAAATCATCCGCAATGAGGA CATCAACGCCAGGGTGAGCGCCATCGAGAAGTGGGTGGCCGTAGCTGACATATGCCGCTGCCTCCACAACTACAATGCCGTACTGGAGAT CACCTCGTCCATGAACCGCAGTGCAATCTTCCGGCTCAAAAAGACGTGGCTCAAAGTCTCTAAGCAGACTAAAGCTTTGATTGATAAGCT CCAAAAGCTTGTGTCATCTGAGGGCAGATTTAAGAATCTCAGAGAAGCTCTGAAAAATTGTGACCCACCCTGTGTCCCTTACCTGGGGAT GTACCTCACCGACCTGGCCTTCATCGAGGAGGGGACGCCCAATTACACGGAAGACGGCCTGGTCAACTTCTCCAAGATGAGGATGATATC CCATATTATCCGAGAGATTCGCCAGTTTCAACAAACTGCCTACAAAATAGAGCACCAAGCAAAGGTAACGCAATATTTACTGGACCAATC TTTTGTAATGGATGAAGAAAGCCTCTACGAGTCTTCTCTCCGAATAGAACCAAAACTCCCCACCTGAAGCTGAGCCCAGCCCAGACCCAG CTGCTCCCGGGGACATGTGCTAGATGATACTGTACATATTCGTTTGGTTTCACTGGATTTTCTTCTTCAGTATGTGCTTCTCCAAGAATA CAAATCGTCCTTGTTCTTAGATTCCTGTAGAACCGGAATATGAATTTCTGCACCGTTTCAGACTTCGCCCACCCATCCCTCCCCTCGTCT CCTGCAGTGCCTGTTTCTTTTAAACAGCTGTACTCTTGGTCCCTCTTCCCTAGACTCCTCACTCTTCTCAGAGGGGAAACAGCACCCTTG CATACAAGAATCTTAGACTCCAGTCCTGCTTCCGTCCCTCCCCAGCCCAGGCTCCTGGGTTGAGGTGGCCACCCAGGCATCCTCCCAGCA TGTTCCATGTAGTTGTTATAACTGGGTGGGGGGTGGGGGGCGGCGGGAGGGGGAAACCATACCCTGAAGTCCAGTCATTGACAAATCTTC CCGCTGATGGTGATGTCAATTTCCAAAGCAGCTTCTCCAGCCAAAATCGCAGGTCTCAAAACTCCCCATGTGTCTCCCAGTAACCTGGAC GGAAGGGAGTCCTCTCTGCTCAGAACCACACCCCTCGTACCAATGCTGCCATCTCGTTCAGGCCCCACGTCGCTCCTTGCTCTAGGATTA ACACCAGATAGCATGAATAGCAGTCTCAATACAACTGGATGAGGCCCTAGACTTCCCGAGGAAATGGAGTCACAAACACCAAGCCACGTC GACTCTTGCCGACCACTGGCCCAGCATCCACTGAAAGTCGGGAACACAAAAATGTCACTTTTCCTCTTTAAGCTGCCTAACGGTCACTAC AAGCTTACATTGAGATGCTCTAAGTCTGTATACCTTGTGATAATTAGTACCACACTGTAGAATTAGGAAATCAATAACCAAGTGTTTTTA TTGCATATACTGTAGTCCTGTCTAAATGCCTTCTAGCAGGAATATAGTAATGTAGTGCTTATTCTGTGAATATTACCATGTATTTTTTAC ACTGTACAGTATAGAAAACACAGGTTAACTCCGAAGTGGCAGGCATGCTCCACTACAACAGACAGACAACTTTTGCTGTAAGCAATACCT AGTGGTATGAGAATTCTATTTCAAGTCCTAAAATATTTCAACTTACAGCTTGTTTGGAAAAAAATTTCCATTTTTGTAAAAATATATGTT TCCTGATTACCTCTTGCTAATAAATCTATTCTATCTCAGGGTGAACAGCGAGTTTTGATGGGTTTCATTCGTTCAGTCAAGGAATATTTG ATGAGCACTTGCTCCTTACCCAGCCCAGCCTGGGACCGGAGATCCAGAGTGTCAAGGACACCATACCTGCTTTTTAGGCACCAACAGTTT GGTGGGGAAGACTGTTTCAGTTGCCTATTGCACCAAGAACAAGCCACTCCAAAACATAATGGCTTAAAACCATTTTTGTTACCTTTCACA GTTCTATGGGTTGACTGGGTCCAGCTGTCAGTTCTTATTTGGAGTCTCTCATGAAGTTTCAGTCACTCAGCAACTGGGACTGGAGTCATC TGAAGGCTCAACTGGACCGGGTGGTGCTCTAGTTGGTGCACCAAGATGGCGCTGTCACATGGCTGGATGTTGATGTCTGCTGGGCTGCAC TGGGGCTGTTCATGGGAGCATCCACACGTGGCCTCTCTGTGTGGCTTGGGCTCCTCACAGTCCGGAAGCTGGGTTCCAAGAGCGACTGTT CCAGCAGCCTCAGGCAGAAGCTACAAGGCTGCCCAGATGCCATCCTGCCCAGATGTGTCCTCACTGTCACCCAGGAAGGGAAGCCCCGGA ACACCACCCTCAGTCACAGGCATGGGGCCCACTGCAAGGCACAACCCACTCCAGAGCTCCCAAGGCCGCCTCCCCTGGGACCTGCAGCCA GCTGGCTCCATAACTGCTTTTCCGTTTCCTCCAGGCATTTTCATCCTAGCCCTACTTTGTGCTGGGATCTGAGGGGGATAGCAAGCGGAG AAAGAAGGGATCTAGAGGGAGAAGATTCTAGATCCTTCTTTTGACTGCCAGCTGATTTTGATTGCCAGCTTTACCTGCTTCATGAGCTCA GCCAAGTCACTTAACCTCTCAGAGCTTCACTGTTCTCATCTGAAAAATGGGGATGAAAATGGTATCTCATTACAGAGTAGAATTAAATGA >40165_40165_2_IQGAP1-RASGRF1_IQGAP1_chr15_90934105_ENST00000268182_RASGRF1_chr15_79298776_ENST00000558480_length(amino acids)=700AA_BP=51 MSAADEVDGLGVARPHYGSVLDNERLTAEEMDERRRQNVAYEYLCHLEEAKRSDASLYCDDVDIRFSKTMNSCKVLQIRYASVERLLERL TDLRFLSIDFLNTFLHSYRVFTTAIVVLDKLITIYKKPISAIPARSLELLFASGQNNKLLYGEPPKSPRATRKFSSPPPLSITKTSSPSR RRKLSLNIPIITGGKALDLAALSCNSNGYTSMYSAMSPFSKATLDTSKLYVSSSFTNKIPDEGDTTPEKPEDPSALSKQSSEVSMREESD IDQNQSDDGDTETSPTKSPTTPKSVKNKNSSEFPLFSYNNGVVMTSCRELDNNRSALSAASAFAIATAGANEGTPNKEKYRRMSLASAGF PPDQRNGDKEFVIRRAATNRVLNVLRHWVSKHSQDFETNDELKCKVIGFLEEVMHDPELLTQERKAAANIIRTLTQEDPGDNQITLEEIT QMAEGVKAEPFENHSALEIAEQLTLLDHLVFKKIPYEEFFGQGWMKLEKNERTPYIMKTTKHFNDISNLIASEIIRNEDINARVSAIEKW VAVADICRCLHNYNAVLEITSSMNRSAIFRLKKTWLKVSKQTKALIDKLQKLVSSEGRFKNLREALKNCDPPCVPYLGMYLTDLAFIEEG -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for IQGAP1-RASGRF1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for IQGAP1-RASGRF1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for IQGAP1-RASGRF1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
