
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ITFG1-LONP2 (FusionGDB2 ID:40375) |
Fusion Gene Summary for ITFG1-LONP2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ITFG1-LONP2 | Fusion gene ID: 40375 | Hgene | Tgene | Gene symbol | ITFG1 | LONP2 | Gene ID | 81533 | 83752 |
| Gene name | integrin alpha FG-GAP repeat containing 1 | lon peptidase 2, peroxisomal | |
| Synonyms | 2310047C21Rik|CDA08|LNKN-1|TIP | LONP|LONPL|PLON|PSLON | |
| Cytomap | 16q12.1 | 16q12.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | T-cell immunomodulatory proteinLINKINintegrin-alpha FG-GAP repeat-containing protein 1 | lon protease homolog 2, peroxisomallon protease 2lon protease-like protein 2peroxisomal LON protease likeperoxisomal Lon protease homolog 2 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q8TB96 | Q86WA8 | |
| Ensembl transtripts involved in fusion gene | ENST00000320640, ENST00000544001, ENST00000568047, | ENST00000564259, ENST00000285737, ENST00000535754, | |
| Fusion gene scores | * DoF score | 9 X 9 X 5=405 | 11 X 14 X 5=770 |
| # samples | 9 | 14 | |
| ** MAII score | log2(9/405*10)=-2.16992500144231 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(14/770*10)=-2.4594316186373 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: ITFG1 [Title/Abstract] AND LONP2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ITFG1(47399698)-LONP2(48368126), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | ITFG1-LONP2 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. ITFG1-LONP2 seems lost the major protein functional domain in Tgene partner, which is a cell metabolism gene due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across ITFG1 (5'-gene) Fusion gene breakpoints across ITFG1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
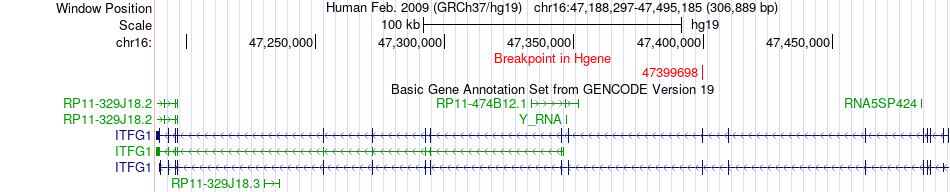 |
 Fusion gene breakpoints across LONP2 (3'-gene) Fusion gene breakpoints across LONP2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
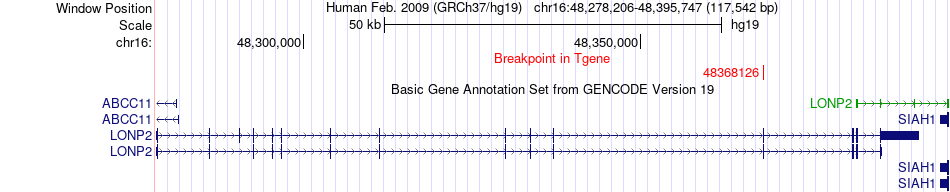 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-A8-A06Y | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
Top |
Fusion Gene ORF analysis for ITFG1-LONP2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000320640 | ENST00000564259 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
| 5CDS-intron | ENST00000544001 | ENST00000564259 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
| Frame-shift | ENST00000544001 | ENST00000285737 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
| Frame-shift | ENST00000544001 | ENST00000535754 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
| In-frame | ENST00000320640 | ENST00000285737 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
| In-frame | ENST00000320640 | ENST00000535754 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
| intron-3CDS | ENST00000568047 | ENST00000285737 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
| intron-3CDS | ENST00000568047 | ENST00000535754 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
| intron-intron | ENST00000568047 | ENST00000564259 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000320640 | ITFG1 | chr16 | 47399698 | - | ENST00000285737 | LONP2 | chr16 | 48368126 | + | 7342 | 1031 | 229 | 1794 | 521 |
| ENST00000320640 | ITFG1 | chr16 | 47399698 | - | ENST00000535754 | LONP2 | chr16 | 48368126 | + | 1892 | 1031 | 229 | 1794 | 521 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000320640 | ENST00000285737 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + | 0.000358663 | 0.9996413 |
| ENST00000320640 | ENST00000535754 | ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + | 0.000879014 | 0.999121 |
Top |
Fusion Genomic Features for ITFG1-LONP2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + | 0.022864684 | 0.9771353 |
| ITFG1 | chr16 | 47399698 | - | LONP2 | chr16 | 48368126 | + | 0.022864684 | 0.9771353 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
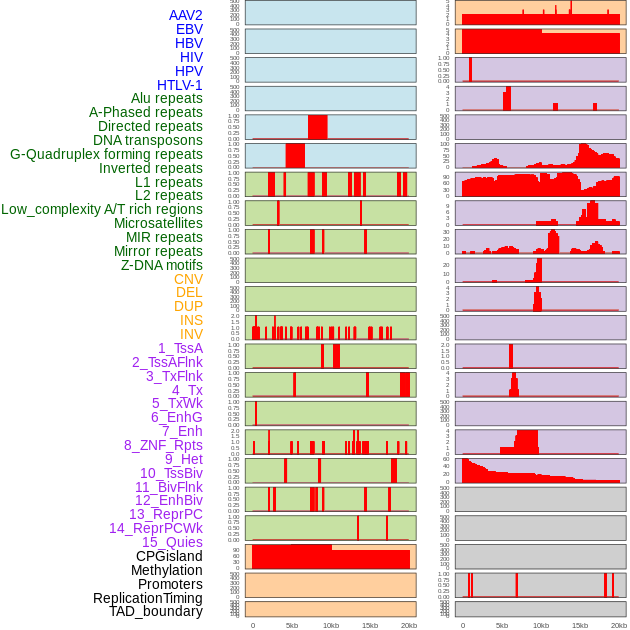 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
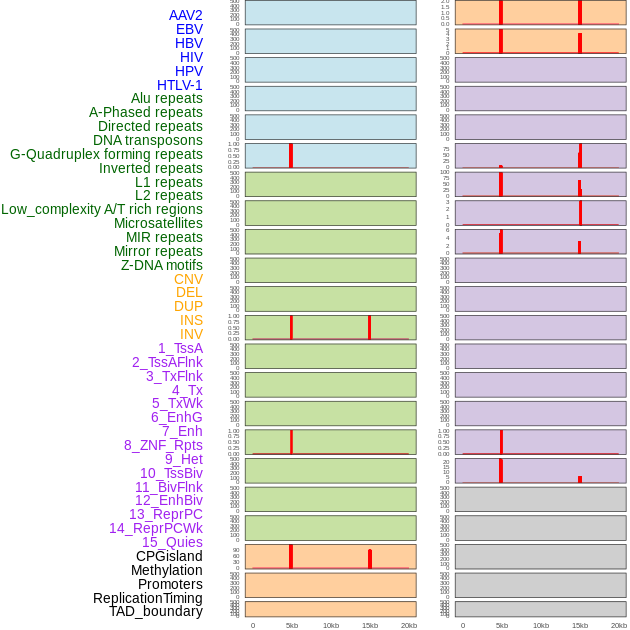 |
Top |
Fusion Protein Features for ITFG1-LONP2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr16:47399698/chr16:48368126) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| ITFG1 | LONP2 |
| FUNCTION: Modulator of T-cell function. Has a protective effect in graft versus host disease model (By similarity). {ECO:0000250}. | FUNCTION: ATP-dependent serine protease that mediates the selective degradation of misfolded and unassembled polypeptides in the peroxisomal matrix. Necessary for type 2 peroxisome targeting signal (PTS2)-containing protein processing and facilitates peroxisome matrix protein import (By similarity). May indirectly regulate peroxisomal fatty acid beta-oxidation through degradation of the self-processed forms of TYSND1. {ECO:0000255|HAMAP-Rule:MF_03121, ECO:0000269|PubMed:22002062}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | LONP2 | chr16:47399698 | chr16:48368126 | ENST00000285737 | 10 | 15 | 651_837 | 598 | 853.0 | Domain | Lon proteolytic |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ITFG1 | chr16:47399698 | chr16:48368126 | ENST00000320640 | - | 8 | 18 | 258_293 | 267 | 613.0 | Repeat | Note=FG-GAP%3B atypical |
| Hgene | ITFG1 | chr16:47399698 | chr16:48368126 | ENST00000320640 | - | 8 | 18 | 567_587 | 267 | 613.0 | Transmembrane | Helical |
| Tgene | LONP2 | chr16:47399698 | chr16:48368126 | ENST00000285737 | 10 | 15 | 13_220 | 598 | 853.0 | Domain | Lon N-terminal | |
| Tgene | LONP2 | chr16:47399698 | chr16:48368126 | ENST00000285737 | 10 | 15 | 375_382 | 598 | 853.0 | Nucleotide binding | ATP |
Top |
Fusion Gene Sequence for ITFG1-LONP2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >40375_40375_1_ITFG1-LONP2_ITFG1_chr16_47399698_ENST00000320640_LONP2_chr16_48368126_ENST00000285737_length(transcript)=7342nt_BP=1031nt ACTGTCGGGAGGCGCGCCTGCCGGAAGTGAGCGTGAGGGCGGCTTTCACCCTAGCTCCACTGCTTTTTTAGCGTCGGAAGGGCAGCCCTA GTGGGAGAGAGAAAAAGGAGCCGGCAGCGGCTCTTACGCGTCCCGGGGCTGCGCGCCACTCTCTCGGCCGGTAACGCGGTGCTTTGCGGC TGTCGTCAAGCGCGGCGTTGGGCCGGCGGGCGGGGGCTGAGGGGCTGCCATGGCGGCGGCGGGCCGGCTCCCGAGCTCCTGGGCCCTCTT CTCGCCGCTCCTCGCAGGGCTTGCACTACTGGGAGTCGGGCCGGTCCCAGCGCGGGCGCTGCACAACGTCACGGCCGAGCTCTTTGGGGC CGAGGCCTGGGGCACCCTTGCGGCTTTCGGGGACCTCAACTCCGACAAGCAGACGGATCTCTTCGTGCTGCGGGAAAGAAATGACTTAAT CGTCTTTTTGGCAGACCAGAATGCACCCTATTTTAAACCCAAAGTAAAGGTATCTTTCAAGAATCACAGTGCATTGATAACAAGTGTAGT CCCTGGGGATTATGATGGAGATTCTCAAATGGATGTCCTTCTGACATATCTTCCCAAAAATTATGCCAAGAGTGAATTAGGAGCTGTTAT CTTCTGGGGACAAAATCAAACATTAGATCCTAACAATATGACCATACTCAATAGGACTTTTCAAGATGAGCCACTAATTATGGATTTCAA TGGTGATCTAATTCCTGATATTTTTGGTATCACAAATGAATCCAACCAGCCACAGATACTATTAGGAGGGAATTTATCATGGCATCCAGC ATTGACCACTACAAGTAAAATGCGAATTCCACATTCTCATGCATTTATTGATCTGACTGAAGATTTTACAGCAGATTTATTCCTGACGAC ATTGAATGCCACCACTAGTACCTTCCAGTTTGAAATATGGGAAAATTTGGATGGAAACTTCTCTGTCAGTACTATATTGGAAAAACCTCA AAATATGATGGTGGTTGGACAGTCAGCATTTGCAGACTTTGGTTGCAGAGAACACATCTTAGAAGATGAAAAACCTGAATCTATCAGTGA CACTACTGACTTGGCTCTACCACCTGAAATGCCGATTTTGATTGATTTCCATGCTCTGAAAGACATCCTTGGGCCCCCGATGTATGAAAT GGAGGTATCTCAGCGTTTGAGTCAGCCAGGAGTAGCAATAGGTTTGGCTTGGACTCCCTTAGGTGGAGAAATCATGTTCGTGGAGGCGAG TCGAATGGATGGCGAGGGCCAGTTAACTCTGACCGGCCAGCTCGGGGACGTGATGAAGGAGTCCGCCCACCTCGCTATCAGCTGGCTCCG CAGCAACGCAAAGAAGTACCAGCTGACCAATGCTTTTGGAAGTTTTGATCTTCTTGACAACACAGACATCCATCTGCACTTCCCAGCTGG AGCTGTCACAAAAGATGGACCATCTGCTGGAGTTACCATAGTAACCTGTCTCGCCTCACTTTTTAGTGGGCGGCTGGTACGTTCAGATGT AGCCATGACTGGAGAAATTACACTGAGAGGTCTTGTTCTTCCAGTGGGTGGAATTAAAGACAAAGTGCTGGCGGCACACAGAGCGGGACT GAAGCAAGTCATTATTCCTCGGAGAAATGAAAAAGACCTTGAGGGAATCCCAGGCAACGTACGACAGGATTTAAGTTTTGTCACAGCAAG CTGCCTGGATGAGGTTCTTAATGCAGCTTTTGATGGTGGCTTTACTGTCAAGACCAGACCTGGTCTGTTAAATAGCAAACTGTAGGTCCA AATCTCAATTTTTTAGAATTTTAAGTTATGAAGTGCTCAAAGGTACTGACACAGTTGATTTTATTCACACCATTAGGGGTATGCAAGATG TCCCTGTTTTATAAACATAATCACAACAGTAATAAACCTCAAGTAGTGGCTAGTGTTTAGTATAGAAATATAAGATGTTGATTTAGTAAA CTGATAAAAATCGAATTCTTGTCTTTTTAGTGGGATCCTTACTGTCCCTGGAAAGATATAGCATAGTGGTTCTCAGCACAGTCTCCAGAA CAGAAGCATCTGTAGTACCTGGTAACTTGTTAGAAATGTACATTCTCAGGCTCCACAGCAGGCCGCCTGAATCAAATCCTGGGAGGTGGG GACAGAAATCTGTGTTTTAAGAAGCCTTCCAGGTAATTCTGCTGCACACTCAAGTTCAGGAACCACCGGTATAGACCATTACCTTAGTGG ATTTACCTGTAGAGTTTATTGGATCCTGAAACCAATCAATTACTTAGAACTAGGCAAAGATGAAAGTATAGCCAACTATTCTTGGCTATA TATATATATTCAAGTGGGCCGGGCGTGATGGCTCACACCTGTAATTCCAGCACTTTGGGAGGTCGAGGTAGGCAGATCACCGAGCCCAAG AGTTCAAGACAATCCTGGCCAACGGCGAAACTCTGTCTCTACAAAAAATATACAGGCGTGTTAGCATGTGCCTGTAATCCCAGCTTCTTG GGAAGCTGAGGCACAAGAATTGCCTGAACCCAGGAGGTGGAGGTTGCAGTGAGCTGGGATCGCGCCATTGCACTCCAGCCTGGCTGACAG AGCGAGACTGTCTCTAAAAAAAAAAGACTCAAGTGGACCCTACAATGAAGCCTACACATCCCAATAGAAGCCCCTTCTTATGCTGAGGGA AGCAGCCCTCAGAACATGATAGCTTGTATCCAGCAGAGTGGCACGTGCTGGCACACCTCACAGAAGCACCCTGGCCCTGGATGCCTGCAA CCTCAGAAGAGTGCAGCTCCCAGAGGGAGGCAGCCATCCATCTGGGATGGTCCTAAGCATGGAATCCTAACTCCTGATTCCGTCTCCTAT TTCTTGCTTGGCTACGCCAGTTCCCAAATCTGGTAGATGTCCATGCCCATGTGCTCCTGCTGGGACTCAATTCAGGCTATGTATGACTAT GAAGTCAGGCTCATCTGCTTACTGGCTGTGTGAACTTTTTGTATCTTGGTTTTCTTCATCCATGAAATCCAAGTAATACTACCTAATTGT TACTGTGGAGATTAAGTTCAAATGCAATGTATAGTAATATTAAGCAATTTCTAGTTATTATTCTAGCCAGTAATGGACTTCAGAATCTTT TATTACACAATATAAGAATATGTATGTAAAGACATTTTGGAATTTCCTGGATGAGAAGGAAGTCTGGGCTGGGCATGGTGGCTCACGCCT GTAACCCTAGCACTTTAGGAAATCGAGGCGAGTGGATCACTTAAGCTCAGGAGTTCAAGGCCAGCCTGGGCAACATGGCAAAACCCCATT TCTACAAAAAATACAAAAATTAGCTGGGCATGGTGGCACCCGCCTGTAGTCCAGCTACTTGAGGCTGAGATGGGAGGATGAGGGAGGTCG GGGCTGCAGTGAGCCAAGATCACGCCACTGCACTCCAGCACCCTGGGCGACAGAGTGAGACCCTGTCTCAAAAAAAAAAAAAAAAAAAAA GATTGGGCCAAAATACTGTGATAAAATAGCAGGCCTGCTGATAAAAGTTTATCTGAATGCATTGAGAGGAAAAGTCCAGACCTAGGACTA GTTATGGCAGTTGGAGAGAAAGAACATCGGGATGTTTGAAAATATGCCATTGACTATCTTAACTACTGTAATTTTATCATTTCCAACGTC ATCTAACTGGGGACTAGAACAAACTGTGAATTCACTTTCAGCAACCAGAGGGCGCTAATCCACACCCACATCGCTCTGCCCTGTTCCACC CAGCAGGGGCAACAAGGATATAACTTGGGGTTCTCGGTATTCTTCCTTTAGTCCTGACACAGGCAGCCTTGCACTTTGTAGCAGCAGGAG GGCACTTGCTTTAAGCATATCTTTCGAAAGGCATCCATTGAGAGAATAATGCATTCTCCCCTTGCTGTGTATGACATGGAACAGTATGAC CATTGCACTAGCTTTGTTTTTGGTTTGTTTTGTTTTTTTTTTAATTCCAGGGGTGGGAGGTTGGTTGGTTGAAGTCTAAGCTCTTCCTTT TTCACCTGGGTTTTTTTTTTTTGGGGGGGGTGGGGGGTTAGGGGTGGGGCGGGTGGGGAAGAGCCAAAATTTTAACCATTTTTAAGCATA TAATTTGGGGGTATCAGTTACTGGATCTAAGCATGTCCACTCTACACGCTTTTTTTTTTTTTTTTTTTTTTTTTTTTTGAGATGGAGTCT TGCTCTGTTACCCAGGCTGCACTGCAGTGGCATGATCTCGGCTCACTGCAACCTCTGCCTCCCCAGTTCAAGCAATTCTCATGCCTCAGC CTCCCAAGTAGCTGGGACCACAGGTGTGTGCCACCACGCCCAGCTAATTTTTGTATTTTTTAGTAGAGACAGGGTTTCACCATGTTGGCC AGGCTGGTCTCAAACTCCCAACCTCAAATGATCCGGCCACCTTGGCCTCCCAAAATGCTGGCATTACAGGCGTGAGCCACCACACCCGGC TTACATGCATTTTTTATCACCACAAACTGAAACTCTGTACCCACTGAAAAAGAGCTCCAAATTCCACCCTCACCCCAGACCCTGGTAACC ACTATTTCACTTTCTTTCTGAATTTCCCTTCATATGAGTGGGATCAAACAAGAGTTGTCCTTTTGTGTTTAGCATTCTTACTTAGCAAGC CATCAACTGCTCGAACAGTCACTGGGGCAACTACTTTTCGCCCAGTGTCCTCCAGAAAAGAAAGATCTGGAGGGTCAGGCCACTCTTTTT CTTCAAAATAATGGGGTGCAGAAGGGTAGGGATGAACCTCTTCCTCCTTTGCCACTTTAGCTTTAGCTCAAAGGAACTCCAACTTTTTAT TATTTGCAACAGTTAGCTTTCTTTGTTTCACATCTCTGCCTTGCAGAACCATGACCTATCAAGGTGTTTTGTCAAACAAAAACTAATACT TTCCCTGGGAAAAAGCCATGTGGTAGCAAGTACCAGAAATTTTTATGGCCAAATAATACACCACTATATCCATAGACCGCATTTGGTTTA TCCACTCATCTGCTGATTGTTTCCCCCTCTTGGGTATTATGAGTAATGCTATGAACATGGGTGTACACATCTCTCTTGAAGTCCCTTCTT AGGTCCTTTAGGGATACATATCCAGGAGCAGAACTACTAGATCACATATGCTAAGGTCTGAATGTCCCCCCACCAAATTCGTATGTTGAA GCAACCACCAACGTGATAGGTGAGGCTTTAGGAGGTGATTATGTCATGAGAGCTCCATCCTCATTAATGGATTTAGTGCCTTTATAAAGG GGCTCAAGGGAACCAATTACCTCCTTTTTGCCCTTCGATTCCCTTCCACCATATGAGGACACAGCAACAGGCCCCGTCCTAGAAGCTGAG ACTGGGCCCTCAATAGATGATACCAGTCTGCCAGCACGCTGATCGTGGGCTTCTCAGTCTTCACAACTATGAGAAATAAATTCGTACTTT TAAATTACCCAGTCTCAGGTATTCTGTTACGGTTAACACAAATGACTAAGACAACAGGATAATTCTGTTTAACTTTTTGAGAACTGCCAA ACTTTTCCACCGCAGCTGCACCATTTGACATTCCTTCCAGCAATGCACAAATGTTCCAGTTTCTCCACATCCTTGCCAGCACTCATTTTC TGGGTTTTTTATAATAGCCCTCCAAATGGGTGTGACGTATTTTGATTCTATGTCCCTAATGACTAGTGATGTTGAGCATTTTCTAGCATT GATTTTTAAGATGTTACCCAAAGACCCCTTGTATCAAAATAAGCTGGATTTTTTTATTGAAAATTATTAACTCTAGAAATTTTAGTTTAA ACTAGACTTAGGGATATGTGTATTTTACCGGTATTCCACGTTTTATGCATGGGTTTTTAAAACTTCTCAAGTATTAAAACTAAAAGCTTT AGGTGCTTTGCTTATCAAGAAATCCTACACTGTCCACTGGAGACATCCATGTTTTTACTTGGCTCTGCCCCTTTAGTGGTCCCTGTGAAC CTTACCTCAAACCATGCATCTGGGGCAGAGATCCTTACTTGCTTGGTGGTTACAAATGCAAATACAGTGAAGAATGTCATCTTTGTGATT GTTCCTGAAATAGTTCACGAGAAATCCATGACCGTAAAGTACTGTGATAGTGATGTCTACCACTGTGAGCTTCCAGTACTAGGTGATTGG TCTGCATTCACAGTGACCAAAATCAGCTATGTGGCCAGGTAATTCACTGCTGAGGGCTTTGGATTTTCCTTTATGAACTACTGAAATGAG GTCAACTTGACTATTACTAAGGGACATTTTGCTACAAAGAATGTTAGTTTTGCCAATTCCCTTTCCAAATCTAAAATTTATTTTAACCAG GATTTTAGATGTAAACATCAAGTAGTTTTGGTTGTTTCAATGAAGTAACATGTTTAAGCTCACATTATTTGAAGTACTTCAGTTCCTATT GCCATGAAAATTGTATCCAGCAGCTAAAAAAAAAAAAAAAAAAAAAGACTACAGTTAGTCATTATCCAATTTGATGATTTATGGTCCAAC ACTAATGCTCATTTTTTTTGTTTGTTTTACAAACATTTGGTGGATACCACAATGAAAACTGCACTTAAAAAACAAAAATGCTGAAAGAGG AAGGAAATATCAAAAAGGTCTGAATAGACAACAGGCAAATATGGTGAGTGGTCATTGAGATGTTTTAAAAGTTACAGCAAAAGGACTTCT AAAACAATTTTAGGAAAAGCTTTGTCATGAAAATTCATTTCTTTGGAAATACTGTAGTTTGTACTTAATGACACTGTCAGCACATTTCTA GGCAATACAAACTCTTGAGGCTAAATCCTCATCCTGACATGACAGTGCAAGCCTGTCAAAATGTGACCCCAAAACCAGGTGTCCTTTTGC TCCATTATTTATATCTGTAAACCTGTTATTATTTTCAGAATTCAGAAAGGCCTAAGAAAATTACATCTATGATAATAAAGTGTATTTCTT TGTCAAGTTTCTAGAGCTATATAAGGATAGCGAAAATATTTGGTTCAGCAAGACACTGGGGTATCAAGTCAGGCATAATAAGCACTTTAT AGCACATCTAGCCTTCCTCATTCCCTTACGCAGTGGACATCATACCCTTTTGTGGAGGAGGGACCAGGCAGAGAAGGTATTTAAATAAAA ACTGGTAGATACAGAGTCAGAAGTCAACCCAGGCAGTTAGAATCTACAACCCACACTGTTTGCCACCAAATCTTAGTAGATGCTTGATAA >40375_40375_1_ITFG1-LONP2_ITFG1_chr16_47399698_ENST00000320640_LONP2_chr16_48368126_ENST00000285737_length(amino acids)=521AA_BP=267 MAAAGRLPSSWALFSPLLAGLALLGVGPVPARALHNVTAELFGAEAWGTLAAFGDLNSDKQTDLFVLRERNDLIVFLADQNAPYFKPKVK VSFKNHSALITSVVPGDYDGDSQMDVLLTYLPKNYAKSELGAVIFWGQNQTLDPNNMTILNRTFQDEPLIMDFNGDLIPDIFGITNESNQ PQILLGGNLSWHPALTTTSKMRIPHSHAFIDLTEDFTADLFLTTLNATTSTFQFEIWENLDGNFSVSTILEKPQNMMVVGQSAFADFGCR EHILEDEKPESISDTTDLALPPEMPILIDFHALKDILGPPMYEMEVSQRLSQPGVAIGLAWTPLGGEIMFVEASRMDGEGQLTLTGQLGD VMKESAHLAISWLRSNAKKYQLTNAFGSFDLLDNTDIHLHFPAGAVTKDGPSAGVTIVTCLASLFSGRLVRSDVAMTGEITLRGLVLPVG -------------------------------------------------------------- >40375_40375_2_ITFG1-LONP2_ITFG1_chr16_47399698_ENST00000320640_LONP2_chr16_48368126_ENST00000535754_length(transcript)=1892nt_BP=1031nt ACTGTCGGGAGGCGCGCCTGCCGGAAGTGAGCGTGAGGGCGGCTTTCACCCTAGCTCCACTGCTTTTTTAGCGTCGGAAGGGCAGCCCTA GTGGGAGAGAGAAAAAGGAGCCGGCAGCGGCTCTTACGCGTCCCGGGGCTGCGCGCCACTCTCTCGGCCGGTAACGCGGTGCTTTGCGGC TGTCGTCAAGCGCGGCGTTGGGCCGGCGGGCGGGGGCTGAGGGGCTGCCATGGCGGCGGCGGGCCGGCTCCCGAGCTCCTGGGCCCTCTT CTCGCCGCTCCTCGCAGGGCTTGCACTACTGGGAGTCGGGCCGGTCCCAGCGCGGGCGCTGCACAACGTCACGGCCGAGCTCTTTGGGGC CGAGGCCTGGGGCACCCTTGCGGCTTTCGGGGACCTCAACTCCGACAAGCAGACGGATCTCTTCGTGCTGCGGGAAAGAAATGACTTAAT CGTCTTTTTGGCAGACCAGAATGCACCCTATTTTAAACCCAAAGTAAAGGTATCTTTCAAGAATCACAGTGCATTGATAACAAGTGTAGT CCCTGGGGATTATGATGGAGATTCTCAAATGGATGTCCTTCTGACATATCTTCCCAAAAATTATGCCAAGAGTGAATTAGGAGCTGTTAT CTTCTGGGGACAAAATCAAACATTAGATCCTAACAATATGACCATACTCAATAGGACTTTTCAAGATGAGCCACTAATTATGGATTTCAA TGGTGATCTAATTCCTGATATTTTTGGTATCACAAATGAATCCAACCAGCCACAGATACTATTAGGAGGGAATTTATCATGGCATCCAGC ATTGACCACTACAAGTAAAATGCGAATTCCACATTCTCATGCATTTATTGATCTGACTGAAGATTTTACAGCAGATTTATTCCTGACGAC ATTGAATGCCACCACTAGTACCTTCCAGTTTGAAATATGGGAAAATTTGGATGGAAACTTCTCTGTCAGTACTATATTGGAAAAACCTCA AAATATGATGGTGGTTGGACAGTCAGCATTTGCAGACTTTGGTTGCAGAGAACACATCTTAGAAGATGAAAAACCTGAATCTATCAGTGA CACTACTGACTTGGCTCTACCACCTGAAATGCCGATTTTGATTGATTTCCATGCTCTGAAAGACATCCTTGGGCCCCCGATGTATGAAAT GGAGGTATCTCAGCGTTTGAGTCAGCCAGGAGTAGCAATAGGTTTGGCTTGGACTCCCTTAGGTGGAGAAATCATGTTCGTGGAGGCGAG TCGAATGGATGGCGAGGGCCAGTTAACTCTGACCGGCCAGCTCGGGGACGTGATGAAGGAGTCCGCCCACCTCGCTATCAGCTGGCTCCG CAGCAACGCAAAGAAGTACCAGCTGACCAATGCTTTTGGAAGTTTTGATCTTCTTGACAACACAGACATCCATCTGCACTTCCCAGCTGG AGCTGTCACAAAAGATGGACCATCTGCTGGAGTTACCATAGTAACCTGTCTCGCCTCACTTTTTAGTGGGCGGCTGGTACGTTCAGATGT AGCCATGACTGGAGAAATTACACTGAGAGGTCTTGTTCTTCCAGTGGGTGGAATTAAAGACAAAGTGCTGGCGGCACACAGAGCGGGACT GAAGCAAGTCATTATTCCTCGGAGAAATGAAAAAGACCTTGAGGGAATCCCAGGCAACGTACGACAGGATTTAAGTTTTGTCACAGCAAG CTGCCTGGATGAGGTTCTTAATGCAGCTTTTGATGGTGGCTTTACTGTCAAGACCAGACCTGGTCTGTTAAATAGCAAACTGTAGGTCCA AATCTCAATTTTTTAGAATTTTAAGTTATGAAGTGCTCAAAGGTACTGACACAGTTGATTTTATTCACACCATTAGGGGTATGCAAGATG >40375_40375_2_ITFG1-LONP2_ITFG1_chr16_47399698_ENST00000320640_LONP2_chr16_48368126_ENST00000535754_length(amino acids)=521AA_BP=267 MAAAGRLPSSWALFSPLLAGLALLGVGPVPARALHNVTAELFGAEAWGTLAAFGDLNSDKQTDLFVLRERNDLIVFLADQNAPYFKPKVK VSFKNHSALITSVVPGDYDGDSQMDVLLTYLPKNYAKSELGAVIFWGQNQTLDPNNMTILNRTFQDEPLIMDFNGDLIPDIFGITNESNQ PQILLGGNLSWHPALTTTSKMRIPHSHAFIDLTEDFTADLFLTTLNATTSTFQFEIWENLDGNFSVSTILEKPQNMMVVGQSAFADFGCR EHILEDEKPESISDTTDLALPPEMPILIDFHALKDILGPPMYEMEVSQRLSQPGVAIGLAWTPLGGEIMFVEASRMDGEGQLTLTGQLGD VMKESAHLAISWLRSNAKKYQLTNAFGSFDLLDNTDIHLHFPAGAVTKDGPSAGVTIVTCLASLFSGRLVRSDVAMTGEITLRGLVLPVG -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for ITFG1-LONP2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ITFG1-LONP2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ITFG1-LONP2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
