
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ANKIB1-GNAI1 (FusionGDB2 ID:4493) |
Fusion Gene Summary for ANKIB1-GNAI1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ANKIB1-GNAI1 | Fusion gene ID: 4493 | Hgene | Tgene | Gene symbol | ANKIB1 | GNAI1 | Gene ID | 54467 | 2770 |
| Gene name | ankyrin repeat and IBR domain containing 1 | G protein subunit alpha i1 | |
| Synonyms | - | Gi | |
| Cytomap | 7q21.2 | 7q21.11 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ankyrin repeat and IBR domain-containing protein 1 | guanine nucleotide-binding protein G(i) subunit alpha-1Gi1 protein alpha subunitadenylate cyclase-inhibiting G alpha proteinguanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 1 | |
| Modification date | 20200313 | 20200329 | |
| UniProtAcc | Q9P2G1 | P63096 | |
| Ensembl transtripts involved in fusion gene | ENST00000265742, ENST00000486698, | ENST00000490206, ENST00000351004, ENST00000457358, | |
| Fusion gene scores | * DoF score | 8 X 6 X 7=336 | 3 X 3 X 2=18 |
| # samples | 9 | 3 | |
| ** MAII score | log2(9/336*10)=-1.90046432644909 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(3/18*10)=0.736965594166206 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: ANKIB1 [Title/Abstract] AND GNAI1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ANKIB1(92021606)-GNAI1(79828541), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across ANKIB1 (5'-gene) Fusion gene breakpoints across ANKIB1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
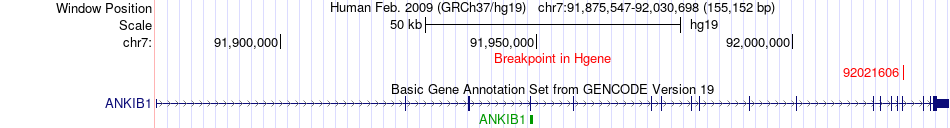 |
 Fusion gene breakpoints across GNAI1 (3'-gene) Fusion gene breakpoints across GNAI1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
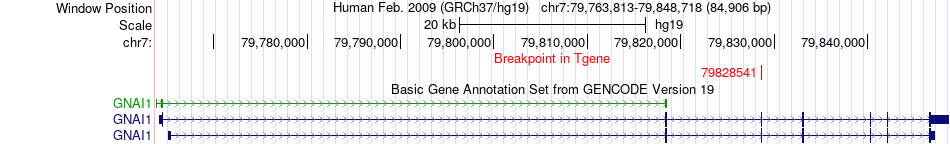 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | HNSC | TCGA-CV-7424-01A | ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828541 | + |
Top |
Fusion Gene ORF analysis for ANKIB1-GNAI1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000265742 | ENST00000490206 | ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828541 | + |
| In-frame | ENST00000265742 | ENST00000351004 | ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828541 | + |
| In-frame | ENST00000265742 | ENST00000457358 | ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828541 | + |
| intron-3CDS | ENST00000486698 | ENST00000351004 | ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828541 | + |
| intron-3CDS | ENST00000486698 | ENST00000457358 | ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828541 | + |
| intron-intron | ENST00000486698 | ENST00000490206 | ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828541 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000265742 | ANKIB1 | chr7 | 92021606 | + | ENST00000351004 | GNAI1 | chr7 | 79828541 | + | 5330 | 2659 | 235 | 3420 | 1061 |
| ENST00000265742 | ANKIB1 | chr7 | 92021606 | + | ENST00000457358 | GNAI1 | chr7 | 79828541 | + | 3789 | 2659 | 235 | 3420 | 1061 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000265742 | ENST00000351004 | ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828541 | + | 0.000127793 | 0.9998722 |
| ENST00000265742 | ENST00000457358 | ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828541 | + | 0.000901144 | 0.9990989 |
Top |
Fusion Genomic Features for ANKIB1-GNAI1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828540 | + | 1.13E-05 | 0.9999887 |
| ANKIB1 | chr7 | 92021606 | + | GNAI1 | chr7 | 79828540 | + | 1.13E-05 | 0.9999887 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
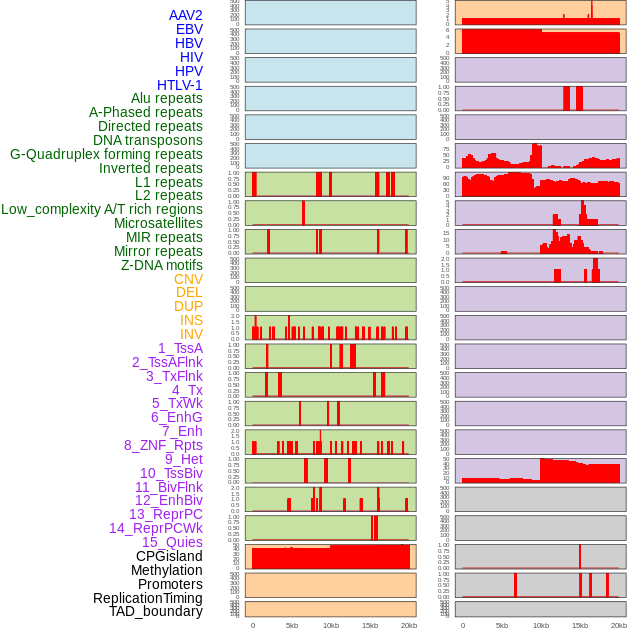 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
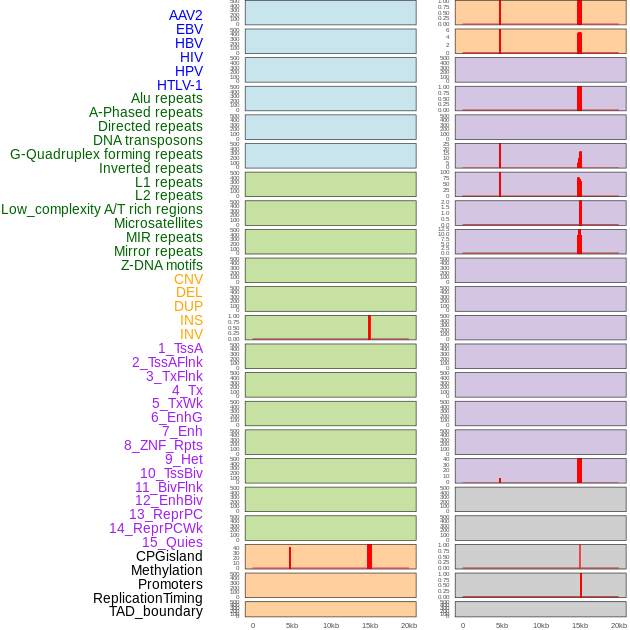 |
Top |
Fusion Protein Features for ANKIB1-GNAI1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr7:92021606/chr7:79828541) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| ANKIB1 | GNAI1 |
| FUNCTION: Might act as an E3 ubiquitin-protein ligase, or as part of E3 complex, which accepts ubiquitin from specific E2 ubiquitin-conjugating enzymes and then transfers it to substrates. {ECO:0000250}. | FUNCTION: Guanine nucleotide-binding proteins (G proteins) function as transducers downstream of G protein-coupled receptors (GPCRs) in numerous signaling cascades. The alpha chain contains the guanine nucleotide binding site and alternates between an active, GTP-bound state and an inactive, GDP-bound state. Signaling by an activated GPCR promotes GDP release and GTP binding. The alpha subunit has a low GTPase activity that converts bound GTP to GDP, thereby terminating the signal. Both GDP release and GTP hydrolysis are modulated by numerous regulatory proteins (PubMed:8774883, PubMed:18434541). Signaling is mediated via effector proteins, such as adenylate cyclase. Inhibits adenylate cyclase activity, leading to decreased intracellular cAMP levels (By similarity). The inactive GDP-bound form prevents the association of RGS14 with centrosomes and is required for the translocation of RGS14 from the cytoplasm to the plasma membrane. Required for normal cytokinesis during mitosis (PubMed:17635935). Required for cortical dynein-dynactin complex recruitment during metaphase (PubMed:22327364). {ECO:0000250|UniProtKB:P10824, ECO:0000269|PubMed:17635935, ECO:0000269|PubMed:18434541, ECO:0000269|PubMed:22327364, ECO:0000269|PubMed:8774883}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ANKIB1 | chr7:92021606 | chr7:79828541 | ENST00000265742 | + | 17 | 20 | 575_640 | 761 | 1090.0 | Coiled coil | Ontology_term=ECO:0000255 |
| Hgene | ANKIB1 | chr7:92021606 | chr7:79828541 | ENST00000265742 | + | 17 | 20 | 329_569 | 761 | 1090.0 | Region | TRIAD supradomain |
| Hgene | ANKIB1 | chr7:92021606 | chr7:79828541 | ENST00000265742 | + | 17 | 20 | 144_173 | 761 | 1090.0 | Repeat | Note=ANK 2 |
| Hgene | ANKIB1 | chr7:92021606 | chr7:79828541 | ENST00000265742 | + | 17 | 20 | 45_74 | 761 | 1090.0 | Repeat | Note=ANK 1 |
| Hgene | ANKIB1 | chr7:92021606 | chr7:79828541 | ENST00000265742 | + | 17 | 20 | 333_383 | 761 | 1090.0 | Zinc finger | RING-type 1 |
| Hgene | ANKIB1 | chr7:92021606 | chr7:79828541 | ENST00000265742 | + | 17 | 20 | 401_478 | 761 | 1090.0 | Zinc finger | IBR-type |
| Hgene | ANKIB1 | chr7:92021606 | chr7:79828541 | ENST00000265742 | + | 17 | 20 | 519_548 | 761 | 1090.0 | Zinc finger | RING-type 2%3B atypical |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 175_181 | 101 | 355.0 | Nucleotide binding | GTP | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 200_204 | 101 | 355.0 | Nucleotide binding | GTP | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 269_272 | 101 | 355.0 | Nucleotide binding | GTP | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 175_181 | 49 | 303.0 | Nucleotide binding | GTP | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 200_204 | 49 | 303.0 | Nucleotide binding | GTP | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 269_272 | 49 | 303.0 | Nucleotide binding | GTP | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 173_181 | 101 | 355.0 | Region | G2 motif | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 196_205 | 101 | 355.0 | Region | G3 motif | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 265_272 | 101 | 355.0 | Region | G4 motif | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 324_329 | 101 | 355.0 | Region | G5 motif | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 173_181 | 49 | 303.0 | Region | G2 motif | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 196_205 | 49 | 303.0 | Region | G3 motif | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 265_272 | 49 | 303.0 | Region | G4 motif | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 324_329 | 49 | 303.0 | Region | G5 motif |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ANKIB1 | chr7:92021606 | chr7:79828541 | ENST00000265742 | + | 17 | 20 | 851_870 | 761 | 1090.0 | Domain | UIM |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 32_354 | 101 | 355.0 | Domain | G-alpha | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 32_354 | 49 | 303.0 | Domain | G-alpha | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 43_48 | 101 | 355.0 | Nucleotide binding | GTP | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 43_48 | 49 | 303.0 | Nucleotide binding | GTP | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000351004 | 2 | 8 | 35_48 | 101 | 355.0 | Region | G1 motif | |
| Tgene | GNAI1 | chr7:92021606 | chr7:79828541 | ENST00000457358 | 2 | 8 | 35_48 | 49 | 303.0 | Region | G1 motif |
Top |
Fusion Gene Sequence for ANKIB1-GNAI1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >4493_4493_1_ANKIB1-GNAI1_ANKIB1_chr7_92021606_ENST00000265742_GNAI1_chr7_79828541_ENST00000351004_length(transcript)=5330nt_BP=2659nt CGCAGCGGCCGGAGAGGGATGGGGGGCGCCCACCCAGTCTGAGCCTCGCCGCGGGCGCCTTCGGCTCACGCAGCGCTTCCCGCGGGCCGC CAGTGCGCACCCTCGGCCACATCAGCCTCCGCCTGGCGGGTGCACCCGGGCCGGCGAGGAAGGGGCAACCGTCCGGGTGGGTGGGGCGGG CTGGGTACCTGAGGTCACCAGCTCGGCTGTAGAGGCAGGGGCGGCGGAGGCGGAACTGCGGAGTTGCTGGGTCCACCGACCCTTACCCTC AGCGAGAGAAGTAACCAATGTGAAAGATGTTGCAGAAGTGTTCCAGAAGTGGCTGAAGATAGAAGGAAAAAAGTGCCACTGCCTATCAGA AAAAACAAAACAAAACATGGGAAATACAACCACCAAATTCCGTAAAGCACTCATCAATGGTGATGAAAACCTGGCCTGCCAAATATATGA AAACAATCCTCAGCTAAAAGAATCTCTTGATCCAAATACATCTTATGGGGAGCCCTACCAGCACAATACTCCATTACATTATGCTGCTAG ACATGGAATGAATAAAATATTAGGGACTTTTCTTGGTAGAGATGGAAATCCAAATAAACGGAATGTGCACAATGAAACATCTATGCATTT GTTGTGTATGGGACCTCAAATTATGATATCTGAAGGAGCCCTTCATCCTCGCTTGGCACGCCCCACAGAAGATGATTTCAGAAGAGCAGA TTGTCTGCAGATGATCTTAAAATGGAAAGGAGCAAAACTTGACCAGGGTGAATATGAGAGAGCAGCTATTGATGCTGTTGATAACAAAAA AAACACACCCTTGCACTATGCTGCTGCCTCAGGGATGAAAGCCTGTGTAGAGCTTTTAGTAAAACATGGAGGAGACTTGTTTGCTGAGAA TGAAAATAAAGATACTCCTTGTGATTGTGCTGAAAAGCAACACCACAAAGATTTGGCCCTCAATCTGGAATCTCAAATGGTATTCTCACG GGATCCCGAGGCTGAAGAAATAGAAGCTGAATATGCTGCATTAGACAAACGAGAGCCATATGAAGGATTAAGGCCTCAGGATCTTCGTAG ACTAAAAGATATGCTTATTGTGGAAACTGCAGACATGCTTCAGGCTCCTCTCTTTACTGCTGAAGCTTTACTTCGAGCTCATGACTGGGA CAGGGAGAAATTACTTGAAGCTTGGATGTCCAACCCGGAGAACTGCTGCCAACGATCAGGTGTTCAAATGCCAACTCCACCACCAAGTGG GTATAATGCCTGGGACACGCTCCCATCTCCAAGAACTCCAAGGACTACACGCTCTTCTGTCACCTCCCCAGATGAAATCAGCTTATCTCC TGGGGATTTAGACACCAGTTTGTGTGACATTTGTATGTGCAGTATCTCTGTATTTGAAGACCCTGTGGATATGCCCTGTGGACATGACTT TTGTAGAGGATGTTGGGAGTCGTTTTTGAATCTGAAAATTCAAGAAGGTGAAGCTCACAACATTTTTTGCCCTGCATATGATTGCTTCCA ACTTGTACCTGTGGATATCATAGAAAGTGTAGTTTCAAAGGAGATGGACAAACGATACCTACAGTTTGATATTAAGGCCTTTGTTGAAAA TAATCCTGCCATTAAATGGTGTCCTACTCCAGGCTGTGACAGAGCAGTAAGACTAACGAAACAAGGGTCAAATACATCTGGATCTGATAC ACTCAGCTTCCCATTGCTGAGAGCTCCTGCTGTTGATTGTGGAAAAGGACACCTCTTCTGCTGGGAGTGCCTTGGTGAAGCACATGAGCC TTGTGACTGCCAAACATGGAAGAATTGGCTGCAAAAAATAACCGAAATGAAACCAGAAGAACTTGTGGGAGTTAGTGAAGCCTACGAGGA TGCCGCCAATTGTCTCTGGTTATTAACTAACTCCAAGCCTTGTGCCAACTGTAAGTCTCCAATACAGAAGAATGAAGGCTGCAATCACAT GCAGTGTGCTAAGTGCAAGTATGACTTTTGCTGGATTTGCCTTGAAGAGTGGAAAAAACATAGTTCGTCCACTGGAGGTTATTACAGATG TACTCGCTATGAAGTCATTCAACACGTGGAGGAGCAATCCAAGGAAATGACTGTGGAGGCTGAGAAAAAACACAAACGATTTCAGGAACT TGACAGATTTATGCACTATTATACAAGATTTAAAAACCATGAGCATAGTTATCAGCTAGAACAACGCCTTCTTAAAACAGCCAAAGAAAA GATGGAGCAATTGAGCAGAGCTCTCAAAGAAACTGAAGGAGGCTGTCCAGATACCACTTTCATTGAAGATGCAGTTCATGTGCTCTTAAA AACTCGGCGCATTCTCAAGTGTTCTTATCCATATGGATTTTTCTTGGAACCTAAAAGCACAAAGAAAGAAATTTTTGAACTAATGCAAAC AGACCTAGAAATGGTCACTGAAGACCTTGCCCAGAAAGTCAATAGGCCTTACCTTCGCACACCCCGCCACAAGATCATCAAAGCAGCATG CCTTGTACAGCAGAAGAGGCAAGAATTCCTGGCATCTGTGGCTCGGGGAGTAGCTCCTGCAGACTCACCAGAAGCTCCAAGGCGCAGCTT TGCTGGTGGAACATGGGATTGGGAATATTTAGGATTTGCATCACCAGAGGATGATGCACGCCAACTCTTTGTGCTAGCTGGAGCTGCTGA AGAAGGCTTTATGACTGCAGAACTTGCTGGAGTTATAAAGAGATTGTGGAAAGATAGTGGTGTACAAGCCTGTTTCAACAGATCCCGAGA GTACCAGCTTAATGATTCTGCAGCATACTATTTGAATGACTTGGACAGAATAGCTCAACCAAATTACATCCCGACTCAACAAGATGTTCT CAGAACTAGAGTGAAAACTACAGGAATTGTTGAAACCCATTTTACTTTCAAAGATCTTCATTTTAAAATGTTTGATGTGGGAGGTCAGAG ATCTGAGCGGAAGAAGTGGATTCATTGCTTCGAAGGAGTGACGGCGATCATCTTCTGTGTAGCACTGAGTGACTACGACCTGGTTCTAGC TGAAGATGAAGAAATGAACCGAATGCATGAAAGCATGAAATTGTTTGACAGCATATGTAACAACAAGTGGTTTACAGATACATCCATTAT ACTTTTTCTAAACAAGAAGGATCTCTTTGAAGAAAAAATCAAAAAGAGCCCTCTCACTATATGCTATCCAGAATATGCAGGATCAAACAC ATATGAAGAGGCAGCTGCATATATTCAATGTCAGTTTGAAGACCTCAATAAAAGAAAGGACACAAAGGAAATATACACCCACTTCACATG TGCCACAGATACTAAGAATGTGCAGTTTGTTTTTGATGCTGTAACAGATGTCATCATAAAAAATAATCTAAAAGATTGTGGTCTCTTTTA AGTTTTGCAGTTCATGGTAAAATGCATTTTCAAACCAAATGAGTACTTATATATGGATCTCTGTAGACTAGAGTCTTGCAGCAACACAGA ATGTAATATAAGGCAAATGCATCTGGGACTTGACCAAAGTTGTTCTGTTTTGTTTTTTTAACTGAAAGTAACAGAAGGACCTTTCTTAAA TGTGACAGATGGTCCTGCAGTGTGAAACTGAAGGACAGTGTTAAAGCTGGGCTCTAGTATATTGATGATTTCTGCATAAGTGTAAATATG CAAATGTATGTATACATGTATTTATGACTTTAGTTTTCGACATTATTTTTAGGTTTTAAGAGTGGCAACTTAGGATTTTAGGGTGATGGC TTTGGAAATAACATAAATATACCTTGTACTGAATGACAGACTATTACTACGTTTGCCAGTTTTAAACAGCTTTATTTATGTTCATGTCCT GTAAATTTTTAAGTACAGTAATTAATATTAGGAAACATTACAGCCCTTATCTAGATTATATGTATACTTGTATTAATAAAAATGTTATTT GTACAAACATTGCACAGACTATTTTAATAACATGATTTGTTCTTTAAATTTTATGTGTTTTATTGAAATGTTCTTAAGATGAATACACCT GCCTTTGGATCAACTATTTAAACATTGTATGCATTTTGATTTTTCCTACTTTAAGAAAATAAAATAATTTAATTTTACATTAGATTCCAC GTTAGATTTGGTTTGAAAAACTAAAATTTCAGATTTCTGAGGATATACTGTCTTAGACTTATTGTACACACTTAGTTTTTATTCACTTGT TTTCACTCTGAATTTTAATATTTGGCTGATATGAATGCATTGCCTCAAAGGTGATGTCATCTTAATTTTTATTCACTTTAAATAACTACA TTTTTGTTTATAACTAAGTTTGGAGGGATCCTAAGAGCATTTTTGTGGGTAAAAAAAAAACCTGTGGACATAATGAATTTTGAGACATTG ATTGGTGAGGCTTTTATTTCCCTTGAGGAGTCTCTTGTACCTAGCATACATGATAGCTCCTTGTTGGGAAGATAACAAGAAGGATCTTTG AATACTCTATTGCTGATATAATGCAAGATTTAAATTTATACATATAACTAATTTCAAATGTAATTATCACACTATGTTAAAATTACTTTT TTCCCTTAGATAATTCAAATTTCTCCACTTGCTTGAGATTATATCATTTCTTTTTCAATTATACTATTATTTCTGAGAATGAAATGGACG ATTACACTTAGAAAATGAGTAATAGTGTTTAATAAGTCAGTGATTATATGTGTGCTCAAATAAGTGTTATGTATCAGCTAGATACTGAGC TTTGATAGAATAATTTTCTTTTGATTATTCATGATGTGTCATCTCTGACCTTGTTTCAGCAAAGTAAACAGCACTCCCCACCCCACCCTC CTTTTTTTACTCATCTTGGAAAAGGTTAGTCTTTCAGTACACGTTGCTGGTAAGTAGTTTCCAAGTTACGTGTTGTCACTGGGTTGAAGT ATATTTGTGTGTGTGTGTGTGTGTGTGTGTGTGTGTGTAACCATAAACTATATTCATATCTGTTTCATTTGGAGGATTTTCTTCTTTGTA ATGTAAAGAAATTCAAAGTTATCAAAGTTCCTTAAATGTGTTAGTTTAGATTCTTTATGTGCCTTTCATGAAAGATATGTTTTCATTAAT TTTACTGGTGGACCTGTAATATCCACATTGTGAAGCTGTGTATGAAATTCAACTATAATATGAATAAATTTGAATCATGAGAATTATGGG TTAAAAAGCCACAAAGAAGCACATATTGGTGACCATCATTAATGAAATCCTGAACTTTATTCTGTGTAATTGTGTTAATAAATCCTAATA >4493_4493_1_ANKIB1-GNAI1_ANKIB1_chr7_92021606_ENST00000265742_GNAI1_chr7_79828541_ENST00000351004_length(amino acids)=1061AA_BP=808 MRSCWVHRPLPSAREVTNVKDVAEVFQKWLKIEGKKCHCLSEKTKQNMGNTTTKFRKALINGDENLACQIYENNPQLKESLDPNTSYGEP YQHNTPLHYAARHGMNKILGTFLGRDGNPNKRNVHNETSMHLLCMGPQIMISEGALHPRLARPTEDDFRRADCLQMILKWKGAKLDQGEY ERAAIDAVDNKKNTPLHYAAASGMKACVELLVKHGGDLFAENENKDTPCDCAEKQHHKDLALNLESQMVFSRDPEAEEIEAEYAALDKRE PYEGLRPQDLRRLKDMLIVETADMLQAPLFTAEALLRAHDWDREKLLEAWMSNPENCCQRSGVQMPTPPPSGYNAWDTLPSPRTPRTTRS SVTSPDEISLSPGDLDTSLCDICMCSISVFEDPVDMPCGHDFCRGCWESFLNLKIQEGEAHNIFCPAYDCFQLVPVDIIESVVSKEMDKR YLQFDIKAFVENNPAIKWCPTPGCDRAVRLTKQGSNTSGSDTLSFPLLRAPAVDCGKGHLFCWECLGEAHEPCDCQTWKNWLQKITEMKP EELVGVSEAYEDAANCLWLLTNSKPCANCKSPIQKNEGCNHMQCAKCKYDFCWICLEEWKKHSSSTGGYYRCTRYEVIQHVEEQSKEMTV EAEKKHKRFQELDRFMHYYTRFKNHEHSYQLEQRLLKTAKEKMEQLSRALKETEGGCPDTTFIEDAVHVLLKTRRILKCSYPYGFFLEPK STKKEIFELMQTDLEMVTEDLAQKVNRPYLRTPRHKIIKAACLVQQKRQEFLASVARGVAPADSPEAPRRSFAGGTWDWEYLGFASPEDD ARQLFVLAGAAEEGFMTAELAGVIKRLWKDSGVQACFNRSREYQLNDSAAYYLNDLDRIAQPNYIPTQQDVLRTRVKTTGIVETHFTFKD LHFKMFDVGGQRSERKKWIHCFEGVTAIIFCVALSDYDLVLAEDEEMNRMHESMKLFDSICNNKWFTDTSIILFLNKKDLFEEKIKKSPL -------------------------------------------------------------- >4493_4493_2_ANKIB1-GNAI1_ANKIB1_chr7_92021606_ENST00000265742_GNAI1_chr7_79828541_ENST00000457358_length(transcript)=3789nt_BP=2659nt CGCAGCGGCCGGAGAGGGATGGGGGGCGCCCACCCAGTCTGAGCCTCGCCGCGGGCGCCTTCGGCTCACGCAGCGCTTCCCGCGGGCCGC CAGTGCGCACCCTCGGCCACATCAGCCTCCGCCTGGCGGGTGCACCCGGGCCGGCGAGGAAGGGGCAACCGTCCGGGTGGGTGGGGCGGG CTGGGTACCTGAGGTCACCAGCTCGGCTGTAGAGGCAGGGGCGGCGGAGGCGGAACTGCGGAGTTGCTGGGTCCACCGACCCTTACCCTC AGCGAGAGAAGTAACCAATGTGAAAGATGTTGCAGAAGTGTTCCAGAAGTGGCTGAAGATAGAAGGAAAAAAGTGCCACTGCCTATCAGA AAAAACAAAACAAAACATGGGAAATACAACCACCAAATTCCGTAAAGCACTCATCAATGGTGATGAAAACCTGGCCTGCCAAATATATGA AAACAATCCTCAGCTAAAAGAATCTCTTGATCCAAATACATCTTATGGGGAGCCCTACCAGCACAATACTCCATTACATTATGCTGCTAG ACATGGAATGAATAAAATATTAGGGACTTTTCTTGGTAGAGATGGAAATCCAAATAAACGGAATGTGCACAATGAAACATCTATGCATTT GTTGTGTATGGGACCTCAAATTATGATATCTGAAGGAGCCCTTCATCCTCGCTTGGCACGCCCCACAGAAGATGATTTCAGAAGAGCAGA TTGTCTGCAGATGATCTTAAAATGGAAAGGAGCAAAACTTGACCAGGGTGAATATGAGAGAGCAGCTATTGATGCTGTTGATAACAAAAA AAACACACCCTTGCACTATGCTGCTGCCTCAGGGATGAAAGCCTGTGTAGAGCTTTTAGTAAAACATGGAGGAGACTTGTTTGCTGAGAA TGAAAATAAAGATACTCCTTGTGATTGTGCTGAAAAGCAACACCACAAAGATTTGGCCCTCAATCTGGAATCTCAAATGGTATTCTCACG GGATCCCGAGGCTGAAGAAATAGAAGCTGAATATGCTGCATTAGACAAACGAGAGCCATATGAAGGATTAAGGCCTCAGGATCTTCGTAG ACTAAAAGATATGCTTATTGTGGAAACTGCAGACATGCTTCAGGCTCCTCTCTTTACTGCTGAAGCTTTACTTCGAGCTCATGACTGGGA CAGGGAGAAATTACTTGAAGCTTGGATGTCCAACCCGGAGAACTGCTGCCAACGATCAGGTGTTCAAATGCCAACTCCACCACCAAGTGG GTATAATGCCTGGGACACGCTCCCATCTCCAAGAACTCCAAGGACTACACGCTCTTCTGTCACCTCCCCAGATGAAATCAGCTTATCTCC TGGGGATTTAGACACCAGTTTGTGTGACATTTGTATGTGCAGTATCTCTGTATTTGAAGACCCTGTGGATATGCCCTGTGGACATGACTT TTGTAGAGGATGTTGGGAGTCGTTTTTGAATCTGAAAATTCAAGAAGGTGAAGCTCACAACATTTTTTGCCCTGCATATGATTGCTTCCA ACTTGTACCTGTGGATATCATAGAAAGTGTAGTTTCAAAGGAGATGGACAAACGATACCTACAGTTTGATATTAAGGCCTTTGTTGAAAA TAATCCTGCCATTAAATGGTGTCCTACTCCAGGCTGTGACAGAGCAGTAAGACTAACGAAACAAGGGTCAAATACATCTGGATCTGATAC ACTCAGCTTCCCATTGCTGAGAGCTCCTGCTGTTGATTGTGGAAAAGGACACCTCTTCTGCTGGGAGTGCCTTGGTGAAGCACATGAGCC TTGTGACTGCCAAACATGGAAGAATTGGCTGCAAAAAATAACCGAAATGAAACCAGAAGAACTTGTGGGAGTTAGTGAAGCCTACGAGGA TGCCGCCAATTGTCTCTGGTTATTAACTAACTCCAAGCCTTGTGCCAACTGTAAGTCTCCAATACAGAAGAATGAAGGCTGCAATCACAT GCAGTGTGCTAAGTGCAAGTATGACTTTTGCTGGATTTGCCTTGAAGAGTGGAAAAAACATAGTTCGTCCACTGGAGGTTATTACAGATG TACTCGCTATGAAGTCATTCAACACGTGGAGGAGCAATCCAAGGAAATGACTGTGGAGGCTGAGAAAAAACACAAACGATTTCAGGAACT TGACAGATTTATGCACTATTATACAAGATTTAAAAACCATGAGCATAGTTATCAGCTAGAACAACGCCTTCTTAAAACAGCCAAAGAAAA GATGGAGCAATTGAGCAGAGCTCTCAAAGAAACTGAAGGAGGCTGTCCAGATACCACTTTCATTGAAGATGCAGTTCATGTGCTCTTAAA AACTCGGCGCATTCTCAAGTGTTCTTATCCATATGGATTTTTCTTGGAACCTAAAAGCACAAAGAAAGAAATTTTTGAACTAATGCAAAC AGACCTAGAAATGGTCACTGAAGACCTTGCCCAGAAAGTCAATAGGCCTTACCTTCGCACACCCCGCCACAAGATCATCAAAGCAGCATG CCTTGTACAGCAGAAGAGGCAAGAATTCCTGGCATCTGTGGCTCGGGGAGTAGCTCCTGCAGACTCACCAGAAGCTCCAAGGCGCAGCTT TGCTGGTGGAACATGGGATTGGGAATATTTAGGATTTGCATCACCAGAGGATGATGCACGCCAACTCTTTGTGCTAGCTGGAGCTGCTGA AGAAGGCTTTATGACTGCAGAACTTGCTGGAGTTATAAAGAGATTGTGGAAAGATAGTGGTGTACAAGCCTGTTTCAACAGATCCCGAGA GTACCAGCTTAATGATTCTGCAGCATACTATTTGAATGACTTGGACAGAATAGCTCAACCAAATTACATCCCGACTCAACAAGATGTTCT CAGAACTAGAGTGAAAACTACAGGAATTGTTGAAACCCATTTTACTTTCAAAGATCTTCATTTTAAAATGTTTGATGTGGGAGGTCAGAG ATCTGAGCGGAAGAAGTGGATTCATTGCTTCGAAGGAGTGACGGCGATCATCTTCTGTGTAGCACTGAGTGACTACGACCTGGTTCTAGC TGAAGATGAAGAAATGAACCGAATGCATGAAAGCATGAAATTGTTTGACAGCATATGTAACAACAAGTGGTTTACAGATACATCCATTAT ACTTTTTCTAAACAAGAAGGATCTCTTTGAAGAAAAAATCAAAAAGAGCCCTCTCACTATATGCTATCCAGAATATGCAGGATCAAACAC ATATGAAGAGGCAGCTGCATATATTCAATGTCAGTTTGAAGACCTCAATAAAAGAAAGGACACAAAGGAAATATACACCCACTTCACATG TGCCACAGATACTAAGAATGTGCAGTTTGTTTTTGATGCTGTAACAGATGTCATCATAAAAAATAATCTAAAAGATTGTGGTCTCTTTTA AGTTTTGCAGTTCATGGTAAAATGCATTTTCAAACCAAATGAGTACTTATATATGGATCTCTGTAGACTAGAGTCTTGCAGCAACACAGA ATGTAATATAAGGCAAATGCATCTGGGACTTGACCAAAGTTGTTCTGTTTTGTTTTTTTAACTGAAAGTAACAGAAGGACCTTTCTTAAA TGTGACAGATGGTCCTGCAGTGTGAAACTGAAGGACAGTGTTAAAGCTGGGCTCTAGTATATTGATGATTTCTGCATAAGTGTAAATATG CAAATGTATGTATACATGTATTTATGACTTTAGTTTTCGACATTATTTTTAGGTTTTAAGAGTGGCAACTTAGGATTTTAGGGTGATGGC >4493_4493_2_ANKIB1-GNAI1_ANKIB1_chr7_92021606_ENST00000265742_GNAI1_chr7_79828541_ENST00000457358_length(amino acids)=1061AA_BP=808 MRSCWVHRPLPSAREVTNVKDVAEVFQKWLKIEGKKCHCLSEKTKQNMGNTTTKFRKALINGDENLACQIYENNPQLKESLDPNTSYGEP YQHNTPLHYAARHGMNKILGTFLGRDGNPNKRNVHNETSMHLLCMGPQIMISEGALHPRLARPTEDDFRRADCLQMILKWKGAKLDQGEY ERAAIDAVDNKKNTPLHYAAASGMKACVELLVKHGGDLFAENENKDTPCDCAEKQHHKDLALNLESQMVFSRDPEAEEIEAEYAALDKRE PYEGLRPQDLRRLKDMLIVETADMLQAPLFTAEALLRAHDWDREKLLEAWMSNPENCCQRSGVQMPTPPPSGYNAWDTLPSPRTPRTTRS SVTSPDEISLSPGDLDTSLCDICMCSISVFEDPVDMPCGHDFCRGCWESFLNLKIQEGEAHNIFCPAYDCFQLVPVDIIESVVSKEMDKR YLQFDIKAFVENNPAIKWCPTPGCDRAVRLTKQGSNTSGSDTLSFPLLRAPAVDCGKGHLFCWECLGEAHEPCDCQTWKNWLQKITEMKP EELVGVSEAYEDAANCLWLLTNSKPCANCKSPIQKNEGCNHMQCAKCKYDFCWICLEEWKKHSSSTGGYYRCTRYEVIQHVEEQSKEMTV EAEKKHKRFQELDRFMHYYTRFKNHEHSYQLEQRLLKTAKEKMEQLSRALKETEGGCPDTTFIEDAVHVLLKTRRILKCSYPYGFFLEPK STKKEIFELMQTDLEMVTEDLAQKVNRPYLRTPRHKIIKAACLVQQKRQEFLASVARGVAPADSPEAPRRSFAGGTWDWEYLGFASPEDD ARQLFVLAGAAEEGFMTAELAGVIKRLWKDSGVQACFNRSREYQLNDSAAYYLNDLDRIAQPNYIPTQQDVLRTRVKTTGIVETHFTFKD LHFKMFDVGGQRSERKKWIHCFEGVTAIIFCVALSDYDLVLAEDEEMNRMHESMKLFDSICNNKWFTDTSIILFLNKKDLFEEKIKKSPL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for ANKIB1-GNAI1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ANKIB1-GNAI1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ANKIB1-GNAI1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
