
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:LMNB2-BMS1 (FusionGDB2 ID:45803) |
Fusion Gene Summary for LMNB2-BMS1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: LMNB2-BMS1 | Fusion gene ID: 45803 | Hgene | Tgene | Gene symbol | LMNB2 | BMS1 | Gene ID | 84823 | 9790 |
| Gene name | lamin B2 | BMS1 ribosome biogenesis factor | |
| Synonyms | EPM9|LAMB2|LMN2 | ACC|BMS1L | |
| Cytomap | 19p13.3 | 10q11.21 | |
| Type of gene | protein-coding | protein-coding | |
| Description | lamin-B2epididymis secretory sperm binding proteinlamin B3 | ribosome biogenesis protein BMS1 homologBMS1 homolog, ribosome assembly proteinBMS1-like, ribosome assembly proteinribosome assembly protein BMS1 homolog | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q03252 | Q14692 | |
| Ensembl transtripts involved in fusion gene | ENST00000325327, ENST00000582871, ENST00000475819, | ENST00000374518, | |
| Fusion gene scores | * DoF score | 4 X 5 X 3=60 | 6 X 6 X 3=108 |
| # samples | 5 | 6 | |
| ** MAII score | log2(5/60*10)=-0.263034405833794 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(6/108*10)=-0.84799690655495 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: LMNB2 [Title/Abstract] AND BMS1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | LMNB2(2434999)-BMS1(43297585), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across LMNB2 (5'-gene) Fusion gene breakpoints across LMNB2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
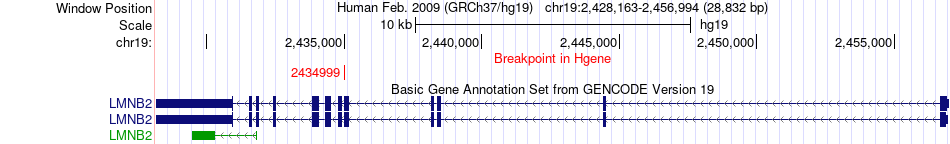 |
 Fusion gene breakpoints across BMS1 (3'-gene) Fusion gene breakpoints across BMS1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
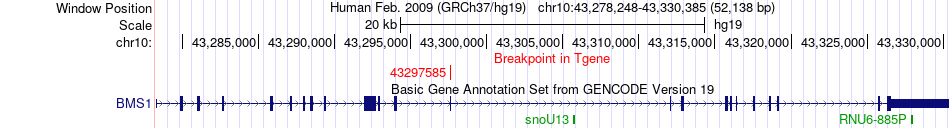 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-HU-A4GN-01A | LMNB2 | chr19 | 2434999 | - | BMS1 | chr10 | 43297585 | + |
Top |
Fusion Gene ORF analysis for LMNB2-BMS1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| In-frame | ENST00000325327 | ENST00000374518 | LMNB2 | chr19 | 2434999 | - | BMS1 | chr10 | 43297585 | + |
| In-frame | ENST00000582871 | ENST00000374518 | LMNB2 | chr19 | 2434999 | - | BMS1 | chr10 | 43297585 | + |
| intron-3CDS | ENST00000475819 | ENST00000374518 | LMNB2 | chr19 | 2434999 | - | BMS1 | chr10 | 43297585 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000325327 | LMNB2 | chr19 | 2434999 | - | ENST00000374518 | BMS1 | chr10 | 43297585 | + | 6361 | 918 | 63 | 2519 | 818 |
| ENST00000582871 | LMNB2 | chr19 | 2434999 | - | ENST00000374518 | BMS1 | chr10 | 43297585 | + | 6325 | 882 | 27 | 2483 | 818 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000325327 | ENST00000374518 | LMNB2 | chr19 | 2434999 | - | BMS1 | chr10 | 43297585 | + | 0.000687561 | 0.9993124 |
| ENST00000582871 | ENST00000374518 | LMNB2 | chr19 | 2434999 | - | BMS1 | chr10 | 43297585 | + | 0.000709822 | 0.9992901 |
Top |
Fusion Genomic Features for LMNB2-BMS1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| LMNB2 | chr19 | 2434998 | - | BMS1 | chr10 | 43297584 | + | 0.9496778 | 0.050322156 |
| LMNB2 | chr19 | 2434998 | - | BMS1 | chr10 | 43297584 | + | 0.9496778 | 0.050322156 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
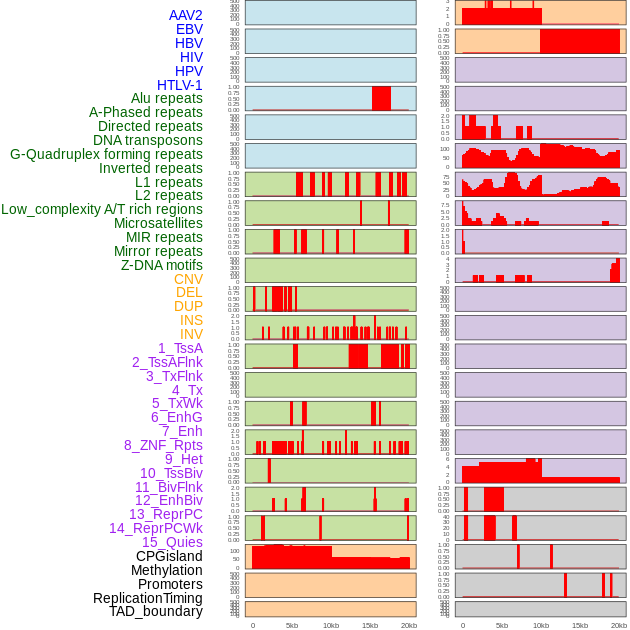 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
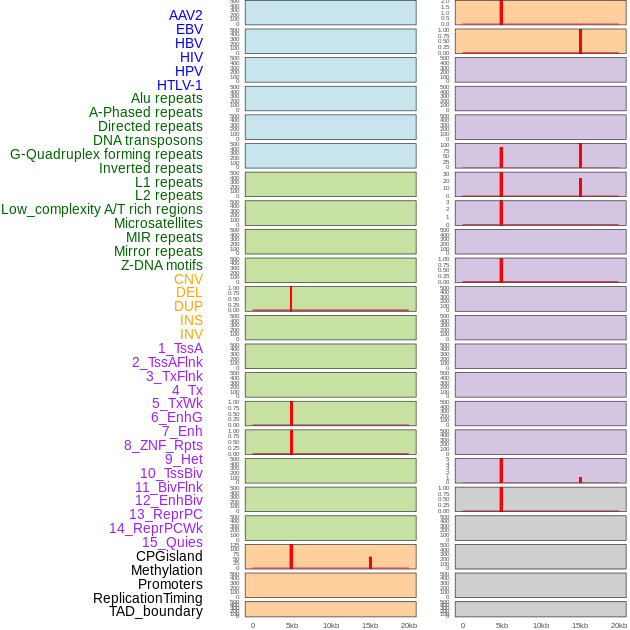 |
Top |
Fusion Protein Features for LMNB2-BMS1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:2434999/chr10:43297585) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| LMNB2 | BMS1 |
| FUNCTION: Lamins are components of the nuclear lamina, a fibrous layer on the nucleoplasmic side of the inner nuclear membrane, which is thought to provide a framework for the nuclear envelope and may also interact with chromatin. | FUNCTION: May act as a molecular switch during maturation of the 40S ribosomal subunit in the nucleolus. {ECO:0000250|UniProtKB:Q08965}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 1_48 | 265 | 601.0 | Region | Note=Head |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 230_256 | 265 | 601.0 | Region | Note=Linker 2 |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 49_83 | 265 | 601.0 | Region | Note=Coil 1A |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 84_95 | 265 | 601.0 | Region | Note=Linker 1 |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 96_229 | 265 | 601.0 | Region | Note=Coil 1B |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 590_602 | 265 | 601.0 | Compositional bias | Note=Asp/Glu-rich (highly acidic%3B could be involved in chromatin binding) |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 462_579 | 265 | 601.0 | Domain | LTD |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 46_402 | 265 | 601.0 | Domain | IF rod |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 435_440 | 265 | 601.0 | Motif | Nuclear localization signal |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 257_400 | 265 | 601.0 | Region | Note=Coil 2 |
| Hgene | LMNB2 | chr19:2434999 | chr10:43297585 | ENST00000582871 | - | 5 | 12 | 401_620 | 265 | 601.0 | Region | Note=Tail |
| Tgene | BMS1 | chr19:2434999 | chr10:43297585 | ENST00000374518 | 11 | 23 | 80_245 | 749 | 1283.0 | Domain | Bms1-type G | |
| Tgene | BMS1 | chr19:2434999 | chr10:43297585 | ENST00000374518 | 11 | 23 | 117_121 | 749 | 1283.0 | Region | G2 | |
| Tgene | BMS1 | chr19:2434999 | chr10:43297585 | ENST00000374518 | 11 | 23 | 132_135 | 749 | 1283.0 | Region | G3 | |
| Tgene | BMS1 | chr19:2434999 | chr10:43297585 | ENST00000374518 | 11 | 23 | 184_187 | 749 | 1283.0 | Region | G4 | |
| Tgene | BMS1 | chr19:2434999 | chr10:43297585 | ENST00000374518 | 11 | 23 | 219_228 | 749 | 1283.0 | Region | G5 | |
| Tgene | BMS1 | chr19:2434999 | chr10:43297585 | ENST00000374518 | 11 | 23 | 89_96 | 749 | 1283.0 | Region | G1 |
Top |
Fusion Gene Sequence for LMNB2-BMS1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >45803_45803_1_LMNB2-BMS1_LMNB2_chr19_2434999_ENST00000325327_BMS1_chr10_43297585_ENST00000374518_length(transcript)=6361nt_BP=918nt GCCGCGGCGGGCGGCGGGCGCGGCGCGGCGCGGCGCCATCTTGCAGCCGGCGGCGCGGATTGAATGAGCCCGCCGAGCCCGGGCCGCCGT CGGGAGCAGCGCAGGCCGCGAGCCGCCGCCACCATGGCCACGCCGCTGCCCGGCCGCGCGGGCGGGCCCGCCACGCCGCTGTCGCCCACG CGCCTGTCGCGGCTGCAGGAGAAGGAGGAGCTGCGCGAGCTCAACGACCGCCTGGCGCACTACATCGACCGCGTCCGCGCGCTGGAGCTG GAGAACGACCGGCTCCTGCTCAAGATCTCAGAGAAGGAGGAGGTGACCACGCGCGAGGTGAGTGGCATCAAGGCGCTGTACGAGTCGGAG CTGGCCGATGCCCGGAGAGTCCTGGATGAGACGGCTCGAGAGCGTGCCCGGCTGCAGATAGAGATTGGGAAGCTGAGGGCAGAGTTGGAC GAGGTCAACAAGAGCGCCAAGAAGAGGGAGGGCGAGCTTACGGTGGCCCAGGGCCGTGTGAAGGACCTGGAGTCCCTGTTCCACCGGAGC GAGGTGGAGCTGGCAGCTGCCCTCAGCGACAAGCGCGGCCTGGAGAGTGACGTGGCTGAGCTGCGGGCCCAGCTGGCCAAGGCCGAGGAC GGTCATGCAGTGGCCAAAAAGCAGCTGGAGAAGGAGACGCTGATGCGTGTGGACCTGGAGAACCGCTGCCAGAGCCTGCAGGAGGAGCTG GACTTCCGGAAGAGTGTGTTCGAGGAGGAGGTGCGGGAGACGCGGCGGCGGCACGAGCGGCGCCTGGTGGAGGTGGACAGCAGCCGGCAG CAGGAGTACGACTTCAAGATGGCACAGGCGCTGGAGGAGCTGCGGAGCCAGCACGACGAGCAAGTGCGGCTCTACAAGCTGGAGCTGGAG CAGACCTACCAGGCCAAGGTTATGAACAGTATCAGAGATTGCTTCGTGACTGGAAAGTGGGAAGATGATAAAGATGCAGCCAAGGTCTTA GCAGAAGATGAGGAGCTCTACGGTGACTTTGAAGACTTGGAAACAGGGGACGTGCACAAGGGAAAATCAGGCCCCAATACTCAGAATGAA GATATAGAGAAAGAAGTTAAGGAAGAAATTGACCCCGACGAAGAAGAAAGTGCCAAGAAAAAGCATTTGGATAAGAAGAGAAAATTGAAG GAGATGTTTGATGCAGAATATGATGAAGGAGAAAGCACATATTTTGATGATCTTAAAGGAGAAATGCAGAAACAAGCACAGCTGAATCGC GCAGAATTTGAAGATCAAGATGATGAAGCCAGAGTTCAGTATGAGGGTTTTCGACCTGGGATGTACGTCCGCATTGAGATTGAAAATGTT CCCTGTGAATTTGTGCAGAACTTTGACCCCCATTACCCCATTATCCTGGGTGGCTTGGGCAACAGTGAGGGAAATGTTGGCTACGTGCAG ATGCGTCTGAAGAAACATCGCTGGTATAAGAAAATCCTCAAGTCCCGAGATCCAATCATATTTTCTGTAGGGTGGAGGAGGTTTCAGACC ATCCCACTGTATTATATCGAAGACCACAATGGAAGACAAAGGCTTCTAAAGTATACCCCACAGCACATGCATTGCGGAGCAGCCTTTTGG GGCCCTATCACTCCACAGGGAACTGGTTTCTTGGCAATACAGTCTGTCAGTGGCATAATGCCTGATTTTCGGATAGCTGCTACAGGAGTT GTCCTTGATCTGGATAAATCCATAAAAATTGTGAAGAAATTAAAGCTAACTGGTTTTCCATATAAAATTTTCAAGAACACTTCATTTATT AAGGGAATGTTTAATTCTGCCTTGGAAGTGGCCAAATTTGAAGGTGCTGTGATTCGAACAGTCAGTGGGATAAGGGGGCAGATCAAGAAA GCACTCCGAGCTCCAGAAGGAGCTTTCAGGGCCAGCTTTGAGGATAAGCTGCTGATGAGCGATATTGTCTTCATGCGAACTTGGTATCCT GTTTCCATCCCAGCGTTCTATAACCCAGTAACATCTTTGTTGAAACCAGTGGGTGAGAAAGACACCTGGTCAGGAATGCGGACCACGGGC CAACTCAGGCTCGCCCATGGCGTCAGACTAAAGGCGAACAAGGACTCTCTGTATAAGCCAATCCTGAGGCAAAAGAAACATTTTAATTCA CTGCACATTCCAAAAGCCTTGCAGAAGGCCCTGCCATTTAAGAACAAGCCCAAGACCCAAGCAAAGGCAGGCAAGGTGCCAAAGGACAGG CGGAGACCGGCCGTCATACGCGAGCCTCATGAAAGAAAGATCCTTGCACTGCTGGATGCTCTGAGTACGGTGCATAGTCAGAAGATGAAG AAGGCCAAGGAGCAGCGGCACCTGCACAATAAAGAGCACTTCAGAGCCAAGCAGAAGGAGGAGGAGGAGAAGCTGAAGCGGCAGAAGGAC CTCAGGAAGAAGCTCTTCAGAATTCAGGGGCAGAAGGAAAGAAGAAACCAGAAGTCCAGTTTGAAGGGGGCTGAGGGCCAATTGCAGTGA GCCTTTGGACTGGAGGGACTGTCCCTGGATCTGCGGAGGTAGACAGTTTCAAACATCACAGTTTGAATGCCTGTGAATGACAAGTCAGTG GGAAAGAGCTCAAGAGATGTCTCTACTCAAACTGTGCCTGCAGGAGGAGGAACAGAGAAGCCTGGGCTGCTGGGACTGGGTTCATTCTCA TGACTTGGGGCTGTCGAGATTTAAAGTGATGTAAGCTGTGGTTATGTGGATTCTCTTACTTTCCTCTGCCTGCCTCAGTTTAATTATTTT GTCCTACAGAAATATCATTAAAATATTTTTTTGTTACTTTTGGCTTAGTAGTTTTCATTAGGGATGAATGCCTGACAATTCTTTGTGGAT AATTATTTATACTACCACCTTCATGAGGAGTCTTCCAGAAGAGAAAGAAATATTCACTTGAACTCTGAGCTCCCATGGAAGATTTTAGAG AATAGTGTGTAGTGTTTTTGTTTTGTTTTTTGTTTGAGTGACACACAAACCTAGATGGTACAGCCTACTGTGCACCCAGGATCTATGGGC TCCTTTGATCACTGGCCCTTGTGGACATTTTCTATCTTATTTCATATGTTCTGCACATTAGTATAGTGACAAGGTAACAGTAAGTACTTT TTTAAAGCTTTAGACTTAGGAGATTATTGTCTCAGGTGCAGAGACCAAGGAAATCGTATGTATCAGTGAGATAGGCTTCCCGTATTATGC TCATGCGTTATACCTGCCTAGCTACATTGCCTTTTTTTAAATTAAACATTATGTTGAAGTAATCATACTATTACATAGGCGATGTATGTA TTTTATGATTGTATGTTGATATAGTTATGGGGAATCTATTCCTATACGATGTAATCATAGATTCACATATAGTTGTATATTTTGCCCAAT TTCTCTCAGTGGTAATGTTTCACACATATAACAATCAGGATGTTGACATCAATACAGCCCGCTGATTTTATTCACATTTCCCCAGCTTTA CTTTCCCCAACGTATACTCCTCCATGTATGCATGGGTGTGTAGAAATATGGAACGCAGCTGGGCACAGTGACTCACGCCTGTAATCCCAG CACTTTGGGATGCTAAGGCAGGAGGATCACTTGAGCCCAGGAATTTGAGACCAACTTGGGCAACATGGCAAAACCTCATCGCTACAAAAA ATAAAAAATTCGCCAGGTGTGGTGGGTGCACACCTGTAGTTCCAGCTGCTTGGGAGGCTGAGGTAGGAGGATCACTTGAGCTTGGGAGGT TGAGGCTGCAGTGAGCCATGATAGCACCACTGCACTCCAGCCTGGGCGACAGGGCAAGACCCTGTTTCAAAAAAAAAAAATTATACATAT ATGTATGTAGAACACTTCATGAATTTGTGTGTCATCTGTGCATGGGGCCCATGCTGACCTCTGTGTTGTGCCAGTTATAGTATGTGTGCT GCTGATATGAGTGCCACATTGCCTTTACAACATCATAGGGGCCAAAGAGGGTTGGCAGGATTATGAGCAGTTGGCCTAGTCTGAGAGGCT CTCTAAAAGCCTAAAGTCTGTAGGCAGGTGCTCAGCTTTGTATCTCCATAAGAAGGGAAGAGGATGTGAGGATCCTGTCAGGGCTAGGTC CACCTGGGAAGTGACAATTGGAACACTCCCAGACCTCTCTGGGCAACTCTGGGTGTACATTTTCCTCTGTTCCTCAAGAGCTATTATGAA TTTCATCATCCATTATGCTGATAGGCCTGGGGACTGACACATAGTAAGCCCATAACAATTATAATTTAAAACTAATTTGTTATATAGCAT AATTGATATATTTTAAAATGCAAAACTATAATTCAACTAAAATATACAAGGCACTCAAGTCATACCATGTTTTAAGTAGCTTACAAGGTG ATCAGCAAGTGTCTTATACAACAAGCAAATTTGTTCAAAGAGAAGCATTGCAAATAAGTATTTATAATTGTATAGTTTTATATTTTAGCC AGAAGAGTGGAATCCTTTAGCAATGGGGGAAATAAAAGAATGTTTAATTACAGTTGCAGATAATTGCCTTTTCCAAAGACAATGTATGTA GGCAGAACTCATACCTGTTAATATTAAAAATCTTGGTTTATCATATTTGGAGAACAAGGGAAATAAAAAGTTTGCATATTAAGATTATTT AAACTAAAGTTAAGGAATCTCTAATGTCTGAAAGAAACAATAAATAGCATTTTTCATGTAGAATTTTTATTGGTGAAGGTTCCACCTGGC CAGGTTAGTAAAATACAGTCACACATCAGTTAAGGATGGGGATATATTCTGAGAAATGTATCATTAGGCAATTTTGTCATTGTGTAAACA TCATAGAGTGACACACAAACCTAGATGGTACAGCCTACTGTGCACCTGGGCTCTATGGTATGGCCTGTTGCTCCTAGGCTACAAACCTGG GCAGCATGTGACTGTACTGACTACTGTAGGCAGTTGTAACACAGTGGTAAACATTTGTGTATCGAAACATAGAAAAGGCGTAGCAAAAAT ACAGTATTGTGGAACCACTGCCATATACGTGGTCCATCCTTGACTGAACATCATGCGGCATGTGGCTGTATGTATACACATAGTTTTTGC TGTGAGTTTGCATCCATCTCATAGGAAGACATATATCAAAATAAGGACCAGACTGGAGAGTGTGTAAATTACTTTTCAATACTAATGGCT TATTAATTTGATATTGTCAGCAAGTATGTTGCAAAAACAAATTCCACCAGAATGTGAGTTCCATGAGAGCAGGAACTGGGTTGTTCACCT TGTTGGCACTTGGAGTAGAGCCACTGAGCTTCGTGCTGCATAAAATTGCTTGGAAGAAAGAAGGGAGGGCATTCCAGGTAACTGAAGCAT TGCTATGCAAAGTAAGCATTTGCCCTCTGAAATATTTTAACAATGTTTATAGAAACCTTTCTCAATTGTATTAAACTTTGTAAATTGTAA AAATTATTCTGTTATACCTTTTCTATTTACTAGGCACAGGCAGCCATAGGTGCTAGGTATGTTTTGAAAACATATTGAACCTACAGTGTA CATATCATTCCATGCCATTAAATATTCTTATTTAATCATAGATGTGTTCAGGTGTGTTTGGTTTCCTATTTGTGGATGTACTAGATTGTT TTCTTTTTTTAACAAAACCTGTGAACTTGCTTTAAGTATTATCATTTGGTGACATGCCCGCCATTTCAGAAATAAATGTTTGAGCAAAGA GGAAAGCCACCGGGTCTGATTCAGCTCTCTGGAGGCTCCAGAGCTTACCTCGCTGATGGGAAAAAGAGCATCGATTTAATTGCCCTTGGT ACCGACTTCTAGTATCTAGTATTTTGCCAGACTCAGGTATTTCTTAAAGCCATTATTCAGAGTCTTCAGCAAGGACATTGTATTTTACTT GTGTCTCCTTAGTTTTACCTAGTTTATTTCATGGTTTTAAAGTATTGTACATTTTTATAATCCTTTAACAAGACAAAAGGTTGAGATGTT GGACATTTAAATGCCAAAGGTGTTTTATTATGAGACTTAGCATTTCATATTTGGGGATGGTACCTTATCGAAAATCAGATAAACTTTGGT TGGATGGGTATTTGGAGGAATCTTTTAAGTCTTTTCTTAAGCAAAGTTTGGACACTCTAAAATAAGGTTGCCAACTGGCATGTAGAGAGA AAAGGATGTGTCATACATTTAGACCAGGTGGCCTGTCCTGTAAATTTTTTTGAGGGGAGGGGTGGGCTTGTTTTTTTCTGTGTTTCGGTT >45803_45803_1_LMNB2-BMS1_LMNB2_chr19_2434999_ENST00000325327_BMS1_chr10_43297585_ENST00000374518_length(amino acids)=818AA_BP=284 MSPPSPGRRREQRRPRAAATMATPLPGRAGGPATPLSPTRLSRLQEKEELRELNDRLAHYIDRVRALELENDRLLLKISEKEEVTTREVS GIKALYESELADARRVLDETARERARLQIEIGKLRAELDEVNKSAKKREGELTVAQGRVKDLESLFHRSEVELAAALSDKRGLESDVAEL RAQLAKAEDGHAVAKKQLEKETLMRVDLENRCQSLQEELDFRKSVFEEEVRETRRRHERRLVEVDSSRQQEYDFKMAQALEELRSQHDEQ VRLYKLELEQTYQAKVMNSIRDCFVTGKWEDDKDAAKVLAEDEELYGDFEDLETGDVHKGKSGPNTQNEDIEKEVKEEIDPDEEESAKKK HLDKKRKLKEMFDAEYDEGESTYFDDLKGEMQKQAQLNRAEFEDQDDEARVQYEGFRPGMYVRIEIENVPCEFVQNFDPHYPIILGGLGN SEGNVGYVQMRLKKHRWYKKILKSRDPIIFSVGWRRFQTIPLYYIEDHNGRQRLLKYTPQHMHCGAAFWGPITPQGTGFLAIQSVSGIMP DFRIAATGVVLDLDKSIKIVKKLKLTGFPYKIFKNTSFIKGMFNSALEVAKFEGAVIRTVSGIRGQIKKALRAPEGAFRASFEDKLLMSD IVFMRTWYPVSIPAFYNPVTSLLKPVGEKDTWSGMRTTGQLRLAHGVRLKANKDSLYKPILRQKKHFNSLHIPKALQKALPFKNKPKTQA KAGKVPKDRRRPAVIREPHERKILALLDALSTVHSQKMKKAKEQRHLHNKEHFRAKQKEEEEKLKRQKDLRKKLFRIQGQKERRNQKSSL -------------------------------------------------------------- >45803_45803_2_LMNB2-BMS1_LMNB2_chr19_2434999_ENST00000582871_BMS1_chr10_43297585_ENST00000374518_length(transcript)=6325nt_BP=882nt CATCTTGCAGCCGGCGGCGCGGATTGAATGAGCCCGCCGAGCCCGGGCCGCCGTCGGGAGCAGCGCAGGCCGCGAGCCGCCGCCACCATG GCCACGCCGCTGCCCGGCCGCGCGGGCGGGCCCGCCACGCCGCTGTCGCCCACGCGCCTGTCGCGGCTGCAGGAGAAGGAGGAGCTGCGC GAGCTCAACGACCGCCTGGCGCACTACATCGACCGCGTCCGCGCGCTGGAGCTGGAGAACGACCGGCTCCTGCTCAAGATCTCAGAGAAG GAGGAGGTGACCACGCGCGAGGTGAGTGGCATCAAGGCGCTGTACGAGTCGGAGCTGGCCGATGCCCGGAGAGTCCTGGATGAGACGGCT CGAGAGCGTGCCCGGCTGCAGATAGAGATTGGGAAGCTGAGGGCAGAGTTGGACGAGGTCAACAAGAGCGCCAAGAAGAGGGAGGGCGAG CTTACGGTGGCCCAGGGCCGTGTGAAGGACCTGGAGTCCCTGTTCCACCGGAGCGAGGTGGAGCTGGCAGCTGCCCTCAGCGACAAGCGC GGCCTGGAGAGTGACGTGGCTGAGCTGCGGGCCCAGCTGGCCAAGGCCGAGGACGGTCATGCAGTGGCCAAAAAGCAGCTGGAGAAGGAG ACGCTGATGCGTGTGGACCTGGAGAACCGCTGCCAGAGCCTGCAGGAGGAGCTGGACTTCCGGAAGAGTGTGTTCGAGGAGGAGGTGCGG GAGACGCGGCGGCGGCACGAGCGGCGCCTGGTGGAGGTGGACAGCAGCCGGCAGCAGGAGTACGACTTCAAGATGGCACAGGCGCTGGAG GAGCTGCGGAGCCAGCACGACGAGCAAGTGCGGCTCTACAAGCTGGAGCTGGAGCAGACCTACCAGGCCAAGGTTATGAACAGTATCAGA GATTGCTTCGTGACTGGAAAGTGGGAAGATGATAAAGATGCAGCCAAGGTCTTAGCAGAAGATGAGGAGCTCTACGGTGACTTTGAAGAC TTGGAAACAGGGGACGTGCACAAGGGAAAATCAGGCCCCAATACTCAGAATGAAGATATAGAGAAAGAAGTTAAGGAAGAAATTGACCCC GACGAAGAAGAAAGTGCCAAGAAAAAGCATTTGGATAAGAAGAGAAAATTGAAGGAGATGTTTGATGCAGAATATGATGAAGGAGAAAGC ACATATTTTGATGATCTTAAAGGAGAAATGCAGAAACAAGCACAGCTGAATCGCGCAGAATTTGAAGATCAAGATGATGAAGCCAGAGTT CAGTATGAGGGTTTTCGACCTGGGATGTACGTCCGCATTGAGATTGAAAATGTTCCCTGTGAATTTGTGCAGAACTTTGACCCCCATTAC CCCATTATCCTGGGTGGCTTGGGCAACAGTGAGGGAAATGTTGGCTACGTGCAGATGCGTCTGAAGAAACATCGCTGGTATAAGAAAATC CTCAAGTCCCGAGATCCAATCATATTTTCTGTAGGGTGGAGGAGGTTTCAGACCATCCCACTGTATTATATCGAAGACCACAATGGAAGA CAAAGGCTTCTAAAGTATACCCCACAGCACATGCATTGCGGAGCAGCCTTTTGGGGCCCTATCACTCCACAGGGAACTGGTTTCTTGGCA ATACAGTCTGTCAGTGGCATAATGCCTGATTTTCGGATAGCTGCTACAGGAGTTGTCCTTGATCTGGATAAATCCATAAAAATTGTGAAG AAATTAAAGCTAACTGGTTTTCCATATAAAATTTTCAAGAACACTTCATTTATTAAGGGAATGTTTAATTCTGCCTTGGAAGTGGCCAAA TTTGAAGGTGCTGTGATTCGAACAGTCAGTGGGATAAGGGGGCAGATCAAGAAAGCACTCCGAGCTCCAGAAGGAGCTTTCAGGGCCAGC TTTGAGGATAAGCTGCTGATGAGCGATATTGTCTTCATGCGAACTTGGTATCCTGTTTCCATCCCAGCGTTCTATAACCCAGTAACATCT TTGTTGAAACCAGTGGGTGAGAAAGACACCTGGTCAGGAATGCGGACCACGGGCCAACTCAGGCTCGCCCATGGCGTCAGACTAAAGGCG AACAAGGACTCTCTGTATAAGCCAATCCTGAGGCAAAAGAAACATTTTAATTCACTGCACATTCCAAAAGCCTTGCAGAAGGCCCTGCCA TTTAAGAACAAGCCCAAGACCCAAGCAAAGGCAGGCAAGGTGCCAAAGGACAGGCGGAGACCGGCCGTCATACGCGAGCCTCATGAAAGA AAGATCCTTGCACTGCTGGATGCTCTGAGTACGGTGCATAGTCAGAAGATGAAGAAGGCCAAGGAGCAGCGGCACCTGCACAATAAAGAG CACTTCAGAGCCAAGCAGAAGGAGGAGGAGGAGAAGCTGAAGCGGCAGAAGGACCTCAGGAAGAAGCTCTTCAGAATTCAGGGGCAGAAG GAAAGAAGAAACCAGAAGTCCAGTTTGAAGGGGGCTGAGGGCCAATTGCAGTGAGCCTTTGGACTGGAGGGACTGTCCCTGGATCTGCGG AGGTAGACAGTTTCAAACATCACAGTTTGAATGCCTGTGAATGACAAGTCAGTGGGAAAGAGCTCAAGAGATGTCTCTACTCAAACTGTG CCTGCAGGAGGAGGAACAGAGAAGCCTGGGCTGCTGGGACTGGGTTCATTCTCATGACTTGGGGCTGTCGAGATTTAAAGTGATGTAAGC TGTGGTTATGTGGATTCTCTTACTTTCCTCTGCCTGCCTCAGTTTAATTATTTTGTCCTACAGAAATATCATTAAAATATTTTTTTGTTA CTTTTGGCTTAGTAGTTTTCATTAGGGATGAATGCCTGACAATTCTTTGTGGATAATTATTTATACTACCACCTTCATGAGGAGTCTTCC AGAAGAGAAAGAAATATTCACTTGAACTCTGAGCTCCCATGGAAGATTTTAGAGAATAGTGTGTAGTGTTTTTGTTTTGTTTTTTGTTTG AGTGACACACAAACCTAGATGGTACAGCCTACTGTGCACCCAGGATCTATGGGCTCCTTTGATCACTGGCCCTTGTGGACATTTTCTATC TTATTTCATATGTTCTGCACATTAGTATAGTGACAAGGTAACAGTAAGTACTTTTTTAAAGCTTTAGACTTAGGAGATTATTGTCTCAGG TGCAGAGACCAAGGAAATCGTATGTATCAGTGAGATAGGCTTCCCGTATTATGCTCATGCGTTATACCTGCCTAGCTACATTGCCTTTTT TTAAATTAAACATTATGTTGAAGTAATCATACTATTACATAGGCGATGTATGTATTTTATGATTGTATGTTGATATAGTTATGGGGAATC TATTCCTATACGATGTAATCATAGATTCACATATAGTTGTATATTTTGCCCAATTTCTCTCAGTGGTAATGTTTCACACATATAACAATC AGGATGTTGACATCAATACAGCCCGCTGATTTTATTCACATTTCCCCAGCTTTACTTTCCCCAACGTATACTCCTCCATGTATGCATGGG TGTGTAGAAATATGGAACGCAGCTGGGCACAGTGACTCACGCCTGTAATCCCAGCACTTTGGGATGCTAAGGCAGGAGGATCACTTGAGC CCAGGAATTTGAGACCAACTTGGGCAACATGGCAAAACCTCATCGCTACAAAAAATAAAAAATTCGCCAGGTGTGGTGGGTGCACACCTG TAGTTCCAGCTGCTTGGGAGGCTGAGGTAGGAGGATCACTTGAGCTTGGGAGGTTGAGGCTGCAGTGAGCCATGATAGCACCACTGCACT CCAGCCTGGGCGACAGGGCAAGACCCTGTTTCAAAAAAAAAAAATTATACATATATGTATGTAGAACACTTCATGAATTTGTGTGTCATC TGTGCATGGGGCCCATGCTGACCTCTGTGTTGTGCCAGTTATAGTATGTGTGCTGCTGATATGAGTGCCACATTGCCTTTACAACATCAT AGGGGCCAAAGAGGGTTGGCAGGATTATGAGCAGTTGGCCTAGTCTGAGAGGCTCTCTAAAAGCCTAAAGTCTGTAGGCAGGTGCTCAGC TTTGTATCTCCATAAGAAGGGAAGAGGATGTGAGGATCCTGTCAGGGCTAGGTCCACCTGGGAAGTGACAATTGGAACACTCCCAGACCT CTCTGGGCAACTCTGGGTGTACATTTTCCTCTGTTCCTCAAGAGCTATTATGAATTTCATCATCCATTATGCTGATAGGCCTGGGGACTG ACACATAGTAAGCCCATAACAATTATAATTTAAAACTAATTTGTTATATAGCATAATTGATATATTTTAAAATGCAAAACTATAATTCAA CTAAAATATACAAGGCACTCAAGTCATACCATGTTTTAAGTAGCTTACAAGGTGATCAGCAAGTGTCTTATACAACAAGCAAATTTGTTC AAAGAGAAGCATTGCAAATAAGTATTTATAATTGTATAGTTTTATATTTTAGCCAGAAGAGTGGAATCCTTTAGCAATGGGGGAAATAAA AGAATGTTTAATTACAGTTGCAGATAATTGCCTTTTCCAAAGACAATGTATGTAGGCAGAACTCATACCTGTTAATATTAAAAATCTTGG TTTATCATATTTGGAGAACAAGGGAAATAAAAAGTTTGCATATTAAGATTATTTAAACTAAAGTTAAGGAATCTCTAATGTCTGAAAGAA ACAATAAATAGCATTTTTCATGTAGAATTTTTATTGGTGAAGGTTCCACCTGGCCAGGTTAGTAAAATACAGTCACACATCAGTTAAGGA TGGGGATATATTCTGAGAAATGTATCATTAGGCAATTTTGTCATTGTGTAAACATCATAGAGTGACACACAAACCTAGATGGTACAGCCT ACTGTGCACCTGGGCTCTATGGTATGGCCTGTTGCTCCTAGGCTACAAACCTGGGCAGCATGTGACTGTACTGACTACTGTAGGCAGTTG TAACACAGTGGTAAACATTTGTGTATCGAAACATAGAAAAGGCGTAGCAAAAATACAGTATTGTGGAACCACTGCCATATACGTGGTCCA TCCTTGACTGAACATCATGCGGCATGTGGCTGTATGTATACACATAGTTTTTGCTGTGAGTTTGCATCCATCTCATAGGAAGACATATAT CAAAATAAGGACCAGACTGGAGAGTGTGTAAATTACTTTTCAATACTAATGGCTTATTAATTTGATATTGTCAGCAAGTATGTTGCAAAA ACAAATTCCACCAGAATGTGAGTTCCATGAGAGCAGGAACTGGGTTGTTCACCTTGTTGGCACTTGGAGTAGAGCCACTGAGCTTCGTGC TGCATAAAATTGCTTGGAAGAAAGAAGGGAGGGCATTCCAGGTAACTGAAGCATTGCTATGCAAAGTAAGCATTTGCCCTCTGAAATATT TTAACAATGTTTATAGAAACCTTTCTCAATTGTATTAAACTTTGTAAATTGTAAAAATTATTCTGTTATACCTTTTCTATTTACTAGGCA CAGGCAGCCATAGGTGCTAGGTATGTTTTGAAAACATATTGAACCTACAGTGTACATATCATTCCATGCCATTAAATATTCTTATTTAAT CATAGATGTGTTCAGGTGTGTTTGGTTTCCTATTTGTGGATGTACTAGATTGTTTTCTTTTTTTAACAAAACCTGTGAACTTGCTTTAAG TATTATCATTTGGTGACATGCCCGCCATTTCAGAAATAAATGTTTGAGCAAAGAGGAAAGCCACCGGGTCTGATTCAGCTCTCTGGAGGC TCCAGAGCTTACCTCGCTGATGGGAAAAAGAGCATCGATTTAATTGCCCTTGGTACCGACTTCTAGTATCTAGTATTTTGCCAGACTCAG GTATTTCTTAAAGCCATTATTCAGAGTCTTCAGCAAGGACATTGTATTTTACTTGTGTCTCCTTAGTTTTACCTAGTTTATTTCATGGTT TTAAAGTATTGTACATTTTTATAATCCTTTAACAAGACAAAAGGTTGAGATGTTGGACATTTAAATGCCAAAGGTGTTTTATTATGAGAC TTAGCATTTCATATTTGGGGATGGTACCTTATCGAAAATCAGATAAACTTTGGTTGGATGGGTATTTGGAGGAATCTTTTAAGTCTTTTC TTAAGCAAAGTTTGGACACTCTAAAATAAGGTTGCCAACTGGCATGTAGAGAGAAAAGGATGTGTCATACATTTAGACCAGGTGGCCTGT CCTGTAAATTTTTTTGAGGGGAGGGGTGGGCTTGTTTTTTTCTGTGTTTCGGTTTTTTCCAATTTTTGTTTTGTGATAAAATATGCATAA >45803_45803_2_LMNB2-BMS1_LMNB2_chr19_2434999_ENST00000582871_BMS1_chr10_43297585_ENST00000374518_length(amino acids)=818AA_BP=284 MSPPSPGRRREQRRPRAAATMATPLPGRAGGPATPLSPTRLSRLQEKEELRELNDRLAHYIDRVRALELENDRLLLKISEKEEVTTREVS GIKALYESELADARRVLDETARERARLQIEIGKLRAELDEVNKSAKKREGELTVAQGRVKDLESLFHRSEVELAAALSDKRGLESDVAEL RAQLAKAEDGHAVAKKQLEKETLMRVDLENRCQSLQEELDFRKSVFEEEVRETRRRHERRLVEVDSSRQQEYDFKMAQALEELRSQHDEQ VRLYKLELEQTYQAKVMNSIRDCFVTGKWEDDKDAAKVLAEDEELYGDFEDLETGDVHKGKSGPNTQNEDIEKEVKEEIDPDEEESAKKK HLDKKRKLKEMFDAEYDEGESTYFDDLKGEMQKQAQLNRAEFEDQDDEARVQYEGFRPGMYVRIEIENVPCEFVQNFDPHYPIILGGLGN SEGNVGYVQMRLKKHRWYKKILKSRDPIIFSVGWRRFQTIPLYYIEDHNGRQRLLKYTPQHMHCGAAFWGPITPQGTGFLAIQSVSGIMP DFRIAATGVVLDLDKSIKIVKKLKLTGFPYKIFKNTSFIKGMFNSALEVAKFEGAVIRTVSGIRGQIKKALRAPEGAFRASFEDKLLMSD IVFMRTWYPVSIPAFYNPVTSLLKPVGEKDTWSGMRTTGQLRLAHGVRLKANKDSLYKPILRQKKHFNSLHIPKALQKALPFKNKPKTQA KAGKVPKDRRRPAVIREPHERKILALLDALSTVHSQKMKKAKEQRHLHNKEHFRAKQKEEEEKLKRQKDLRKKLFRIQGQKERRNQKSSL -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for LMNB2-BMS1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for LMNB2-BMS1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for LMNB2-BMS1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
