
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:ANKRD13A-ATP2A2 (FusionGDB2 ID:4587) |
Fusion Gene Summary for ANKRD13A-ATP2A2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: ANKRD13A-ATP2A2 | Fusion gene ID: 4587 | Hgene | Tgene | Gene symbol | ANKRD13A | ATP2A2 | Gene ID | 88455 | 488 |
| Gene name | ankyrin repeat domain 13A | ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 2 | |
| Synonyms | ANKRD13|NY-REN-25 | ATP2B|DAR|DD|SERCA2 | |
| Cytomap | 12q24.11 | 12q24.11 | |
| Type of gene | protein-coding | protein-coding | |
| Description | ankyrin repeat domain-containing protein 13ANY-REN-25 antigen | sarcoplasmic/endoplasmic reticulum calcium ATPase 2ATPase Ca++ transporting cardiac muscle slow twitch 2ATPase, Ca++ dependent, slow-twitch, cardiac muscle-2SR Ca(2+)-ATPase 2calcium pump 2calcium-transporting ATPase sarcoplasmic reticulum type, slow | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | Q8IZ07 | P16615 | |
| Ensembl transtripts involved in fusion gene | ENST00000550404, ENST00000261739, | ENST00000308664, ENST00000395494, ENST00000539276, ENST00000550248, ENST00000552636, | |
| Fusion gene scores | * DoF score | 3 X 2 X 3=18 | 19 X 23 X 6=2622 |
| # samples | 3 | 23 | |
| ** MAII score | log2(3/18*10)=0.736965594166206 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | log2(23/2622*10)=-3.51096191927738 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: ANKRD13A [Title/Abstract] AND ATP2A2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | ANKRD13A(110437589)-ATP2A2(110734404), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | ANKRD13A | GO:1905667 | negative regulation of protein localization to endosome | 22298428 |
| Tgene | ATP2A2 | GO:0032469 | endoplasmic reticulum calcium ion homeostasis | 16402920 |
| Tgene | ATP2A2 | GO:0032470 | positive regulation of endoplasmic reticulum calcium ion concentration | 16402920 |
| Tgene | ATP2A2 | GO:0070588 | calcium ion transmembrane transport | 16402920 |
| Tgene | ATP2A2 | GO:1903515 | calcium ion transport from cytosol to endoplasmic reticulum | 16402920 |
 Fusion gene breakpoints across ANKRD13A (5'-gene) Fusion gene breakpoints across ANKRD13A (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
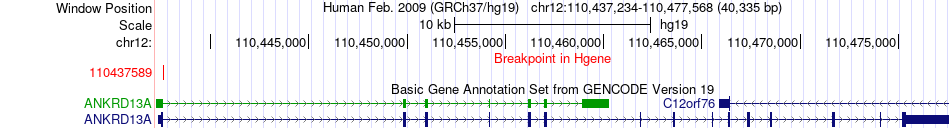 |
 Fusion gene breakpoints across ATP2A2 (3'-gene) Fusion gene breakpoints across ATP2A2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
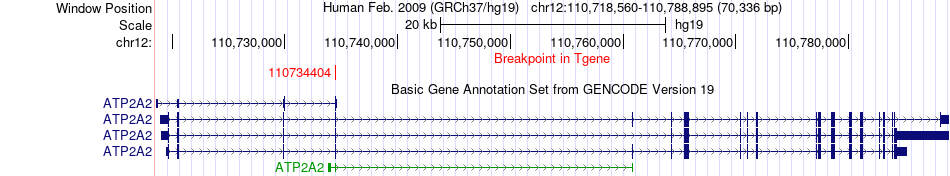 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | STAD | TCGA-D7-8574-01A | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
Top |
Fusion Gene ORF analysis for ANKRD13A-ATP2A2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 3UTR-3CDS | ENST00000550404 | ENST00000308664 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
| 3UTR-3CDS | ENST00000550404 | ENST00000395494 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
| 3UTR-3CDS | ENST00000550404 | ENST00000539276 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
| 3UTR-3UTR | ENST00000550404 | ENST00000550248 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
| 3UTR-3UTR | ENST00000550404 | ENST00000552636 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
| 5CDS-3UTR | ENST00000261739 | ENST00000550248 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
| 5CDS-3UTR | ENST00000261739 | ENST00000552636 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
| In-frame | ENST00000261739 | ENST00000308664 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
| In-frame | ENST00000261739 | ENST00000395494 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
| In-frame | ENST00000261739 | ENST00000539276 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000261739 | ANKRD13A | chr12 | 110437589 | + | ENST00000308664 | ATP2A2 | chr12 | 110734404 | + | 3717 | 262 | 166 | 2931 | 921 |
| ENST00000261739 | ANKRD13A | chr12 | 110437589 | + | ENST00000395494 | ATP2A2 | chr12 | 110734404 | + | 7604 | 262 | 166 | 2985 | 939 |
| ENST00000261739 | ANKRD13A | chr12 | 110437589 | + | ENST00000539276 | ATP2A2 | chr12 | 110734404 | + | 3923 | 262 | 166 | 3066 | 966 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000261739 | ENST00000308664 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + | 0.000451152 | 0.99954885 |
| ENST00000261739 | ENST00000395494 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + | 0.000220331 | 0.99977964 |
| ENST00000261739 | ENST00000539276 | ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734404 | + | 0.000299179 | 0.9997009 |
Top |
Fusion Genomic Features for ANKRD13A-ATP2A2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734403 | + | 1.59E-14 | 1 |
| ANKRD13A | chr12 | 110437589 | + | ATP2A2 | chr12 | 110734403 | + | 1.59E-14 | 1 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
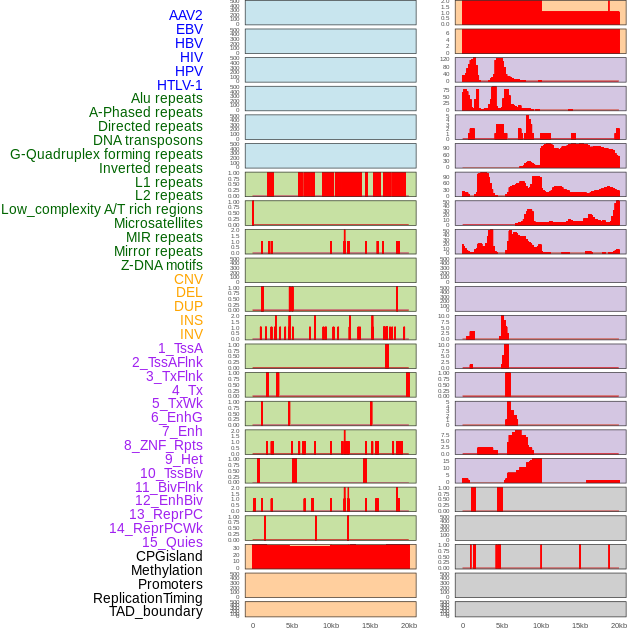 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for ANKRD13A-ATP2A2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:110437589/chr12:110734404) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| ANKRD13A | ATP2A2 |
| FUNCTION: Ubiquitin-binding protein that specifically recognizes and binds 'Lys-63'-linked ubiquitin. Does not bind 'Lys-48'-linked ubiquitin. Positively regulates the internalization of ligand-activated EGFR by binding to the Ub moiety of ubiquitinated EGFR at the cell membrane. {ECO:0000269|PubMed:22298428}. | FUNCTION: This magnesium-dependent enzyme catalyzes the hydrolysis of ATP coupled with the translocation of calcium from the cytosol to the sarcoplasmic reticulum lumen (PubMed:16402920). Involved in autophagy in response to starvation. Upon interaction with VMP1 and activation, controls ER-isolation membrane contacts for autophagosome formation (PubMed:28890335). Also modulates ER contacts with lipid droplets, mitochondria and endosomes (PubMed:28890335). {ECO:0000269|PubMed:16402920, ECO:0000269|PubMed:28890335}.; FUNCTION: [Isoform 2]: Involved in the regulation of the contraction/relaxation cycle. Acts as a regulator of TNFSF11-mediated Ca(2+) signaling pathways via its interaction with TMEM64 which is critical for the TNFSF11-induced CREB1 activation and mitochondrial ROS generation necessary for proper osteoclast generation. Association between TMEM64 and SERCA2 in the ER leads to cytosolic Ca(2+) spiking for activation of NFATC1 and production of mitochondrial ROS, thereby triggering Ca(2+) signaling cascades that promote osteoclast differentiation and activation. {ECO:0000250|UniProtKB:O55143}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 111_253 | 108 | 998.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 274_295 | 108 | 998.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 314_756 | 108 | 998.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 777_786 | 108 | 998.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 808_827 | 108 | 998.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 851_896 | 108 | 998.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 917_929 | 108 | 998.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 949_963 | 108 | 998.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 985_1042 | 108 | 998.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 111_253 | 108 | 1016.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 274_295 | 108 | 1016.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 314_756 | 108 | 1016.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 777_786 | 108 | 1016.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 808_827 | 108 | 1016.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 851_896 | 108 | 1016.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 917_929 | 108 | 1016.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 949_963 | 108 | 1016.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 985_1042 | 108 | 1016.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 111_253 | 108 | 1043.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 274_295 | 108 | 1043.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 314_756 | 108 | 1043.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 777_786 | 108 | 1043.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 808_827 | 108 | 1043.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 851_896 | 108 | 1043.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 917_929 | 108 | 1043.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 949_963 | 108 | 1043.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 985_1042 | 108 | 1043.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 254_273 | 108 | 998.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 296_313 | 108 | 998.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 757_776 | 108 | 998.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 828_850 | 108 | 998.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 897_916 | 108 | 998.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 930_948 | 108 | 998.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 964_984 | 108 | 998.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 254_273 | 108 | 1016.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 296_313 | 108 | 1016.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 757_776 | 108 | 1016.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 828_850 | 108 | 1016.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 897_916 | 108 | 1016.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 930_948 | 108 | 1016.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 964_984 | 108 | 1016.0 | Transmembrane | Helical%3B Name%3D10 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 254_273 | 108 | 1043.0 | Transmembrane | Helical%3B Name%3D3 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 296_313 | 108 | 1043.0 | Transmembrane | Helical%3B Name%3D4 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 757_776 | 108 | 1043.0 | Transmembrane | Helical%3B Name%3D5 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 828_850 | 108 | 1043.0 | Transmembrane | Helical%3B Name%3D7 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 897_916 | 108 | 1043.0 | Transmembrane | Helical%3B Name%3D8 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 930_948 | 108 | 1043.0 | Transmembrane | Helical%3B Name%3D9 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 964_984 | 108 | 1043.0 | Transmembrane | Helical%3B Name%3D10 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | ANKRD13A | chr12:110437589 | chr12:110734404 | ENST00000261739 | + | 1 | 15 | 483_502 | 32 | 591.0 | Domain | UIM 1 |
| Hgene | ANKRD13A | chr12:110437589 | chr12:110734404 | ENST00000261739 | + | 1 | 15 | 519_538 | 32 | 591.0 | Domain | UIM 2 |
| Hgene | ANKRD13A | chr12:110437589 | chr12:110734404 | ENST00000261739 | + | 1 | 15 | 549_568 | 32 | 591.0 | Domain | UIM 3 |
| Hgene | ANKRD13A | chr12:110437589 | chr12:110734404 | ENST00000261739 | + | 1 | 15 | 574_590 | 32 | 591.0 | Domain | UIM 4 |
| Hgene | ANKRD13A | chr12:110437589 | chr12:110734404 | ENST00000261739 | + | 1 | 15 | 40_69 | 32 | 591.0 | Repeat | Note=ANK 1 |
| Hgene | ANKRD13A | chr12:110437589 | chr12:110734404 | ENST00000261739 | + | 1 | 15 | 73_102 | 32 | 591.0 | Repeat | Note=ANK 2 |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 1_48 | 108 | 998.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 70_89 | 108 | 998.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 1_48 | 108 | 1016.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 70_89 | 108 | 1016.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 1_48 | 108 | 1043.0 | Topological domain | Cytoplasmic | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 70_89 | 108 | 1043.0 | Topological domain | Lumenal | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 49_69 | 108 | 998.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 90_110 | 108 | 998.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 49_69 | 108 | 1016.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 90_110 | 108 | 1016.0 | Transmembrane | Helical%3B Name%3D2 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 49_69 | 108 | 1043.0 | Transmembrane | Helical%3B Name%3D1 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 90_110 | 108 | 1043.0 | Transmembrane | Helical%3B Name%3D2 |
Top |
Fusion Gene Sequence for ANKRD13A-ATP2A2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >4587_4587_1_ANKRD13A-ATP2A2_ANKRD13A_chr12_110437589_ENST00000261739_ATP2A2_chr12_110734404_ENST00000308664_length(transcript)=3717nt_BP=262nt CGGACCTCCCGCGCGCCCCGCACCCGACCGGCTCAGCCGGCCGGCAGCGTAACACGCCCTACGCTCGCTTGCTCGCCGGCCTCAGGGCAG GCAGGCGGGCGCGGGAGACCCCGCCGGGGCCGAGACTTGGGGCGGGCGACGAGGACCAGGTTACGGCCTCCTCGCCATGTCCTCGGCCTG CGACGCGGGCGACCACTACCCCCTGCACCTCCTAGTCTGGAAAAACGACTACCGGCAGCTCGAGAAGGAGCTGCAGGGCCAGGAAAGAAA TGCTGAAAATGCCATCGAAGCCCTTAAGGAATATGAGCCTGAAATGGGCAAAGTGTATCGACAGGACAGAAAGAGTGTGCAGCGGATTAA AGCTAAAGACATAGTTCCTGGTGATATTGTAGAAATTGCTGTTGGTGACAAAGTTCCTGCTGATATAAGGTTAACTTCCATCAAATCTAC CACACTAAGAGTTGACCAGTCAATTCTCACAGGTGAATCTGTCTCTGTCATCAAGCACACTGATCCCGTCCCTGACCCACGAGCTGTCAA CCAAGATAAAAAGAACATGCTGTTTTCTGGTACAAACATTGCTGCTGGGAAAGCTATGGGAGTGGTGGTAGCAACTGGAGTTAACACCGA AATTGGCAAGATCCGGGATGAAATGGTGGCAACAGAACAGGAGAGAACACCCCTTCAGCAAAAACTAGATGAATTTGGGGAACAGCTTTC CAAAGTCATCTCCCTTATTTGCATTGCAGTCTGGATCATAAATATTGGGCACTTCAATGACCCGGTTCATGGAGGGTCCTGGATCAGAGG TGCTATTTACTACTTTAAAATTGCAGTGGCCCTGGCTGTAGCAGCCATTCCTGAAGGTCTGCCTGCAGTCATCACCACCTGCCTGGCTCT TGGAACTCGCAGAATGGCAAAGAAAAATGCCATTGTTCGAAGCCTCCCGTCTGTGGAAACCCTTGGTTGTACTTCTGTTATCTGCTCAGA CAAGACTGGTACACTTACAACAAACCAGATGTCAGTCTGCAGGATGTTCATTCTGGACAGAGTGGAAGGTGATACTTGTTCCCTTAATGA GTTTACCATAACTGGATCAACTTATGCACCTATTGGAGAAGTGCATAAAGATGATAAACCAGTGAATTGTCACCAGTATGATGGTCTGGT AGAATTAGCAACAATTTGTGCTCTTTGTAATGACTCTGCTTTGGATTACAATGAGGCAAAGGGTGTGTATGAAAAAGTTGGAGAAGCTAC AGAGACTGCTCTCACTTGCCTAGTAGAGAAGATGAATGTATTTGATACCGAATTGAAGGGTCTTTCTAAAATAGAACGTGCAAATGCCTG CAACTCAGTCATTAAACAGCTGATGAAAAAGGAATTCACTCTAGAGTTTTCACGTGACAGAAAGTCAATGTCGGTTTACTGTACACCAAA TAAACCAAGCAGGACATCAATGAGCAAGATGTTTGTGAAGGGTGCTCCTGAAGGTGTCATTGACAGGTGCACCCACATTCGAGTTGGAAG TACTAAGGTTCCTATGACCTCTGGAGTCAAACAGAAGATCATGTCTGTCATTCGAGAGTGGGGTAGTGGCAGCGACACACTGCGATGCCT GGCCCTGGCCACTCATGACAACCCACTGAGAAGAGAAGAAATGCACCTTGAGGACTCTGCCAACTTTATTAAATATGAGACCAATCTGAC CTTCGTTGGCTGCGTGGGCATGCTGGATCCTCCGAGAATCGAGGTGGCCTCCTCCGTGAAGCTGTGCCGGCAAGCAGGCATCCGGGTCAT CATGATCACTGGGGACAACAAGGGCACTGCTGTGGCCATCTGTCGCCGCATCGGCATCTTCGGGCAGGATGAGGACGTGACGTCAAAAGC TTTCACAGGCCGGGAGTTTGATGAACTCAACCCCTCCGCCCAGCGAGACGCCTGCCTGAACGCCCGCTGTTTTGCTCGAGTTGAACCCTC CCACAAGTCTAAAATCGTAGAATTTCTTCAGTCTTTTGATGAGATTACAGCTATGACTGGCGATGGCGTGAACGATGCTCCTGCTCTGAA GAAAGCCGAGATTGGCATTGCTATGGGCTCTGGCACTGCGGTGGCTAAAACCGCCTCTGAGATGGTCCTGGCGGATGACAACTTCTCCAC CATTGTGGCTGCCGTTGAGGAGGGGCGGGCAATCTACAACAACATGAAACAGTTCATCCGCTACCTCATCTCGTCCAACGTCGGGGAAGT TGTCTGTATTTTCCTGACAGCAGCCCTTGGATTTCCCGAGGCTTTGATTCCTGTTCAGCTGCTCTGGGTCAATCTGGTGACAGATGGCCT GCCTGCCACTGCACTGGGGTTCAACCCTCCTGATCTGGACATCATGAATAAACCTCCCCGGAACCCAAAGGAACCATTGATCAGCGGGTG GCTCTTTTTCCGTTACTTGGCTATTGGCTGTTACGTCGGCGCTGCTACCGTGGGTGCTGCTGCATGGTGGTTCATTGCTGCTGACGGTGG TCCAAGAGTGTCCTTCTACCAGCTGAGTCATTTCCTACAGTGTAAAGAGGACAACCCGGACTTTGAAGGCGTGGATTGTGCAATCTTTGA ATCCCCATACCCGATGACAATGGCGCTCTCTGTTCTAGTAACTATAGAAATGTGTAACGCCCTCAACAGCTTGTCCGAAAACCAGTCCTT GCTGAGGATGCCCCCCTGGGAGAACATCTGGCTCGTGGGCTCCATCTGCCTGTCCATGTCACTCCACTTCCTGATCCTCTATGTCGAACC CTTGCCACTCATCTTCCAGATCACACCGCTGAACGTGACCCAGTGGCTGATGGTGCTGAAAATCTCCTTGCCCGTGATTCTCATGGATGA GACGCTCAAGTTTGTGGCCCGCAACTACCTGGAACCTGCAATACTGGAGTAACCGCTTCCTAAACCATTTTGCAGAAATGTAAGGGTGTT CGGTTGCGTGCATGTGCGTTTTTAGCAACACATCTACCAACCCTGTGCATGACTGATGTTGGGGAAAAAGAAAAGTAAAAAACTTCCCAA CTCACTTTGTGTTATGTGGAGGAAATGTGTATTACCAATGGGGTTGTTAGCTTTTAAATCAAAATACTGATTACAGATGTACAATTTAGC TTAATCAGAAAGCCTCTCCAGAGAAGTTTGGTTTCTTTGCTGCAAGAGGAATGAGGCTCTGTAACCTTATCTAAGAACTTGGAAGCCGTC AGCCAAGTCGCCACATTTCTCTGCAAAATGTCATAGCTTATATAAATGTACAGTATTCAATTGTAATGCATGCCTTCGGTTGTAAGTAGC CAGATCCCTCTCCAGTGACATTGGAACATGCTACTTTTTAATTGGCCCTGTACAGTTTGCTTATTTATAAATTCATTAAAAACACTACAG GTGTTGAATGGTTAAAATGTAGGCCTCCAGTTCATTTTCAGTTATTTTCTGAGTGTGCAGACAGCTATTTCGCACTGTATTAAATGTAAC TTATTTAATGAAATCAGAAGCAGTAGACAGATGTTGGTGCAATACAAATATTGTGATGCATTTATCTTAATAAAATGCTAAATGTCAATT TATCACTGCGCATGTTTGACTTTAGACTGTAAATAGAGATCAGTTTGTTTCTTTCTGTGCTGGTAACAATGAGCGTCGCACAGACATGGT >4587_4587_1_ANKRD13A-ATP2A2_ANKRD13A_chr12_110437589_ENST00000261739_ATP2A2_chr12_110734404_ENST00000308664_length(amino acids)=921AA_BP=0 MSSACDAGDHYPLHLLVWKNDYRQLEKELQGQERNAENAIEALKEYEPEMGKVYRQDRKSVQRIKAKDIVPGDIVEIAVGDKVPADIRLT SIKSTTLRVDQSILTGESVSVIKHTDPVPDPRAVNQDKKNMLFSGTNIAAGKAMGVVVATGVNTEIGKIRDEMVATEQERTPLQQKLDEF GEQLSKVISLICIAVWIINIGHFNDPVHGGSWIRGAIYYFKIAVALAVAAIPEGLPAVITTCLALGTRRMAKKNAIVRSLPSVETLGCTS VICSDKTGTLTTNQMSVCRMFILDRVEGDTCSLNEFTITGSTYAPIGEVHKDDKPVNCHQYDGLVELATICALCNDSALDYNEAKGVYEK VGEATETALTCLVEKMNVFDTELKGLSKIERANACNSVIKQLMKKEFTLEFSRDRKSMSVYCTPNKPSRTSMSKMFVKGAPEGVIDRCTH IRVGSTKVPMTSGVKQKIMSVIREWGSGSDTLRCLALATHDNPLRREEMHLEDSANFIKYETNLTFVGCVGMLDPPRIEVASSVKLCRQA GIRVIMITGDNKGTAVAICRRIGIFGQDEDVTSKAFTGREFDELNPSAQRDACLNARCFARVEPSHKSKIVEFLQSFDEITAMTGDGVND APALKKAEIGIAMGSGTAVAKTASEMVLADDNFSTIVAAVEEGRAIYNNMKQFIRYLISSNVGEVVCIFLTAALGFPEALIPVQLLWVNL VTDGLPATALGFNPPDLDIMNKPPRNPKEPLISGWLFFRYLAIGCYVGAATVGAAAWWFIAADGGPRVSFYQLSHFLQCKEDNPDFEGVD CAIFESPYPMTMALSVLVTIEMCNALNSLSENQSLLRMPPWENIWLVGSICLSMSLHFLILYVEPLPLIFQITPLNVTQWLMVLKISLPV -------------------------------------------------------------- >4587_4587_2_ANKRD13A-ATP2A2_ANKRD13A_chr12_110437589_ENST00000261739_ATP2A2_chr12_110734404_ENST00000395494_length(transcript)=7604nt_BP=262nt CGGACCTCCCGCGCGCCCCGCACCCGACCGGCTCAGCCGGCCGGCAGCGTAACACGCCCTACGCTCGCTTGCTCGCCGGCCTCAGGGCAG GCAGGCGGGCGCGGGAGACCCCGCCGGGGCCGAGACTTGGGGCGGGCGACGAGGACCAGGTTACGGCCTCCTCGCCATGTCCTCGGCCTG CGACGCGGGCGACCACTACCCCCTGCACCTCCTAGTCTGGAAAAACGACTACCGGCAGCTCGAGAAGGAGCTGCAGGGCCAGGAAAGAAA TGCTGAAAATGCCATCGAAGCCCTTAAGGAATATGAGCCTGAAATGGGCAAAGTGTATCGACAGGACAGAAAGAGTGTGCAGCGGATTAA AGCTAAAGACATAGTTCCTGGTGATATTGTAGAAATTGCTGGTGAATCTGTCTCTGTCATCAAGCACACTGATCCCGTCCCTGACCCACG AGCTGTCAACCAAGATAAAAAGAACATGCTGTTTTCTGGTACAAACATTGCTGCTGGGAAAGCTATGGGAGTGGTGGTAGCAACTGGAGT TAACACCGAAATTGGCAAGATCCGGGATGAAATGGTGGCAACAGAACAGGAGAGAACACCCCTTCAGCAAAAACTAGATGAATTTGGGGA ACAGCTTTCCAAAGTCATCTCCCTTATTTGCATTGCAGTCTGGATCATAAATATTGGGCACTTCAATGACCCGGTTCATGGAGGGTCCTG GATCAGAGGTGCTATTTACTACTTTAAAATTGCAGTGGCCCTGGCTGTAGCAGCCATTCCTGAAGGTCTGCCTGCAGTCATCACCACCTG CCTGGCTCTTGGAACTCGCAGAATGGCAAAGAAAAATGCCATTGTTCGAAGCCTCCCGTCTGTGGAAACCCTTGGTTGTACTTCTGTTAT CTGCTCAGACAAGACTGGTACACTTACAACAAACCAGATGTCAGTCTGCAGGATGTTCATTCTGGACAGAGTGGAAGGTGATACTTGTTC CCTTAATGAGTTTACCATAACTGGATCAACTTATGCACCTATTGGAGAAGTGCATAAAGATGATAAACCAGTGAATTGTCACCAGTATGA TGGTCTGGTAGAATTAGCAACAATTTGTGCTCTTTGTAATGACTCTGCTTTGGATTACAATGAGGCAAAGGGTGTGTATGAAAAAGTTGG AGAAGCTACAGAGACTGCTCTCACTTGCCTAGTAGAGAAGATGAATGTATTTGATACCGAATTGAAGGGTCTTTCTAAAATAGAACGTGC AAATGCCTGCAACTCAGTCATTAAACAGCTGATGAAAAAGGAATTCACTCTAGAGTTTTCACGTGACAGAAAGTCAATGTCGGTTTACTG TACACCAAATAAACCAAGCAGGACATCAATGAGCAAGATGTTTGTGAAGGGTGCTCCTGAAGGTGTCATTGACAGGTGCACCCACATTCG AGTTGGAAGTACTAAGGTTCCTATGACCTCTGGAGTCAAACAGAAGATCATGTCTGTCATTCGAGAGTGGGGTAGTGGCAGCGACACACT GCGATGCCTGGCCCTGGCCACTCATGACAACCCACTGAGAAGAGAAGAAATGCACCTTGAGGACTCTGCCAACTTTATTAAATATGAGAC CAATCTGACCTTCGTTGGCTGCGTGGGCATGCTGGATCCTCCGAGAATCGAGGTGGCCTCCTCCGTGAAGCTGTGCCGGCAAGCAGGCAT CCGGGTCATCATGATCACTGGGGACAACAAGGGCACTGCTGTGGCCATCTGTCGCCGCATCGGCATCTTCGGGCAGGATGAGGACGTGAC GTCAAAAGCTTTCACAGGCCGGGAGTTTGATGAACTCAACCCCTCCGCCCAGCGAGACGCCTGCCTGAACGCCCGCTGTTTTGCTCGAGT TGAACCCTCCCACAAGTCTAAAATCGTAGAATTTCTTCAGTCTTTTGATGAGATTACAGCTATGACTGGCGATGGCGTGAACGATGCTCC TGCTCTGAAGAAAGCCGAGATTGGCATTGCTATGGGCTCTGGCACTGCGGTGGCTAAAACCGCCTCTGAGATGGTCCTGGCGGATGACAA CTTCTCCACCATTGTGGCTGCCGTTGAGGAGGGGCGGGCAATCTACAACAACATGAAACAGTTCATCCGCTACCTCATCTCGTCCAACGT CGGGGAAGTTGTCTGTATTTTCCTGACAGCAGCCCTTGGATTTCCCGAGGCTTTGATTCCTGTTCAGCTGCTCTGGGTCAATCTGGTGAC AGATGGCCTGCCTGCCACTGCACTGGGGTTCAACCCTCCTGATCTGGACATCATGAATAAACCTCCCCGGAACCCAAAGGAACCATTGAT CAGCGGGTGGCTCTTTTTCCGTTACTTGGCTATTGGCTGTTACGTCGGCGCTGCTACCGTGGGTGCTGCTGCATGGTGGTTCATTGCTGC TGACGGTGGTCCAAGAGTGTCCTTCTACCAGCTGAGTCATTTCCTACAGTGTAAAGAGGACAACCCGGACTTTGAAGGCGTGGATTGTGC AATCTTTGAATCCCCATACCCGATGACAATGGCGCTCTCTGTTCTAGTAACTATAGAAATGTGTAACGCCCTCAACAGCTTGTCCGAAAA CCAGTCCTTGCTGAGGATGCCCCCCTGGGAGAACATCTGGCTCGTGGGCTCCATCTGCCTGTCCATGTCACTCCACTTCCTGATCCTCTA TGTCGAACCCTTGCCACTCATCTTCCAGATCACACCGCTGAACGTGACCCAGTGGCTGATGGTGCTGAAAATCTCCTTGCCCGTGATTCT CATGGATGAGACGCTCAAGTTTGTGGCCCGCAACTACCTGGAACCTGGTAAAGAGTGTGTGCAGCCTGCCACCAAATCCTGCTCGTTCTC GGCATGCACCGATGGGATTTCCTGGCCGTTTGTGCTGCTCATAATGCCCCTGGTGATCTGGGTCTATAGCACAGACACTAACTTTAGCGA TATGTTCTGGTCTTGACTGACAGTTTTCCATAAAGAAGATGTTTAACTTAATCAATTAATTTTTTTATTGTTTAAAGCAACTGTCTATTT CTGCTGAATTTTCACATGAACATACTGGCTGGTGATGGAGGTTTCATACTCTAGATTTTGTTTTGCTTTTTCTGACTCCAGTGGGGCAAG ATTTTCCTTTTTTATACACATAATTAAAGTGTCCATTGACATGTACAGAGAACTAACACTATTTTATGCAAATATTTTTTTGTAGATGAA AAAGCATGTACAGTGTTCTGTTTAATACTCATCCTTGTATAAAAAAAATAGTTGAGCCAGCAGACATTGTCAGCAAATTAATTGGCAGCA GATTTTAGGAAATGAATGTGTGTGGTTTTTTTTCTAAAACTAAATAGCATGTATTGTGTCTTTTGCATGATGATCCGGATTTAATTTGAT ATCACAGTCTAATTTTTATTCATAAGCCAATTTTTCTGCACTGAGCAGAGTCTTGCTACCTCAGTCAGTATTGTTTTGGTTTGCTACTTC CCTCACCCACTTTGGCCTCCGTTCACCCCACCCCACCCCACCTCTCCCCACCTTACCCCCGCCCCGCTTGGCTTCTTCTTTAGGATTGTG ATGGTTCGTTCTGTTTACATCAGTTTTAACGAGAGGTATGCCTGTACTCGCTTGTGCAGAAAACATTGTTCCAGATTCAATCGACTGGGT TTATGTCCCTTCACATAGTTTTTAAGGTTATTTATTTAAATGTCTAATGTATTTTATTGTAACAGACATTGTTTTGCCAACATTGCCTAT TTCAGTGGCACGTCATCTAGTTTTAAAAAAATAAAACATTTTAAATGGACAGAGAAAAATAACTGTCTTGTCTTTAACTCGTAAGTGGCT TACCTGGGACTAACAGACATGTCCAACTTTCTCTCCAGTTCTTAGCTCAGAACTTTAGTTGTACTCTGCTTGAGGGGAAGAAGGCTCCTG CTCTGCTGTGTAGGTAGTCATAGGAATTGTATTCTTAATGTACAGGCACTAATTGTCATCTGTGATGTACATTTTATGCAAGTTTCTGCT GGCCTGGTATAGAGAACATAAGGGCAAGTGTGTATGTGTGTGTATGTGTGTGTTTTGTAAAATCTGTAAATAGCACATGACCAAATGAAC ATATTGTATAGAACTATTTTTATTTGAATGTGGCACTAACCACCACCACCGTTACTACGATCAATGTTTGCGCATGTTCGAGATGAGTCT CACCAACAGTGTGTAAGTCATTAACAGTCCTAACTGTGGTGTTTTCCTCCAATGCCTTCCAACATCCATCAACTAACGTGAGTATTTTCT TCCTGGGATTTGGATGCTTTAGCCTAAAGGTGACTGCCACCAAGTGAGATAACTGTATGTCACTAACTTATAAGCCGCCTCCATGGCAGA TGCTGCTGTGCTCCCTGATGCCCTGTGAGCACCGGGGTTGCCTGTGGCGCCTGCCATGTGACTCGGGCGCAGCATCAGCTGGCTGGAGGT GTGGCTTTCATAGACCTCCACAGGCTGCCTAGGACAAGATGACCAGGAGGGCCCAAGCCAGACAAAAGCCGGAGTGGGGAGAGAGGCATT TCAGCCAGACCAACAGGCTGAAGGAGCTGTGCAGACTATTGCTAAAATGAGGGTTCGCAGCTGCCAGGAGTCATCCCAGAACATTGCTAC TTATTTATTAAAAAGCTAAAAACTAGTGAAAGCAAGTAACTGAACCAGTGAACTCTGGGTATCGATAGGTTCGTCTTAAATAGGCCACTT CCCCACTCCCCCACCCCCCCTTGCTTGGTCTTGTCCTTGGTGGCTAAGACTTAGCTCTGCAGGGGATGTTAAAGCACAGTTAGTAGGACG TGGCTCTGCACAGCCCAAAAACCAGCTTACTCCTTAGCCAGGGTGTGAGGCCTCGACTATATTCTTCAAAATGTACTTAGGCTCTGGTTA CTGGGATGGCCAGTAGATGTAATGCAGATGGTTGGAGTTTGGGGAGGGTTAGGAGGCATCAAGCAGGACAAGGTGCTGCTGAGTTCAGAG GGCCCACGTTCAAGGGATGGAGGTGGAACCTGGAGACCGACTCTTAAAAGCACAGTCCGTGGTTGGGTGTGGTGGCTCATGCCTGTAAGT AAGTCTCAGCCCTTTGGAGGCCACAGCAGAAGGATTGCTTGAGCCCAGGTACTCAAACCAGCCTGGGCAACAGAGTGAGGCCCTGTCAAA AAAAAATCAGCCTTACTGTGAAGCCTCCAAGGCTGCTCGCAGGCAGCTGTGGCTCTCGGGAAGGGAGGCTGCTTTGCCCAGCAGGGAACA TTTGGGGCAGGGGGTAAATTTTGCCAGTTTGAGCATCATGAGGTGTAACAAGAAATGGGTTGAATGGGCCAAATGCAAGGAGTGCATCTC TGGGCTGCAAACTGACTTGAGTGCTGCACTATTGCTATTCCGTGCAAACAAAACTCAGCTTTTCCTGACTCAGTTCCTTGACTTAGTGGC CTTTACAAAAAAAGTTGAGTAGTGTGTGGCCTGCTGTCGCACAGCCCCTAGTTAGCTTCATGGTTTCTCAGCTTCAGACCCCTCCAGCCC ACAGAGGAGCCCATGGAGGGACCCACTTCCCTTGGTCCAGACAGCTGGGAGTGGGTTAGGCCCACTGCTGTTTTGAGCAGGGCCACTTGC TCCATTTCACTGAAGGCTTTGCTGGGTGAAAACACTTCAGCATCTCCTCCTCAGGTCAACCCATAAAGACCAGGTCCAGCACCGTGGTCT TGGCACATCCCTGGCCTCAGGCCCTCACCTAACAGTGAGGCAGCAGCTGCCCAGCCCCGCAATGTGCCTGCTGTCAGGCAGCTCTTGCCT GAAACTTACTTCCACATTCTTTCCTGATGGGCAGGTGGCTGAAGGCCCAGCCATCAGTGTCGCTTGTTGCCACCCCGTGCCTCCCTTGGC CTCTCTGAGCTTTGCCCAGAAGACCAACAATCATACATACCCTAACTGGGACACCACTCTGCAGAATGCAGATGATCCATTCTGGAGGAA GCTGTCCCTTGAGCTCAGTGAGCTCCCAGGCAAGCAGGGCATCTGGCCGACTTCCCTCACAACAGCTGCTCCCACATCCCCTCGGACTGG AGCTTCAGCCCTGACTGAGGTGGGCAGACCTAAGACCTGAGACCACAAGATTAGCTCAGTGTCTACCAAGCATCTAGCCACTGTCCAGGG CCAGAGCATACCACGTCTGCAGTGCCTGTGAGCAGAGCCAGCAGTTGCCCTGTGACTGTAACCACCAAATTGTCCAAACACCCGCTGCAG TTAGCAAGAAGGGTAGGCTTCACCCTCCTTTACTGAGGAGAATGATGCGGAGGAGTTTCCTCTCCAGGGCTAGGCAAGGCAGGCGAGCAG CCAGAAGCCGGGTGCCCACAGGGCAGGGACAGGAAGGCTGTGCTGCTACTGGCTGCTCACTTCTCCATCAACCTCACCCTCTGCACCACT AACCAAGACCTTGTCCTCTTGCCTGTCTCGCTGCTTTCACAGCTGCAACGATTGTGTCTGCCTCATGGGGTTTTCCTCCAGAGCCTTTAT TCTGTAGCCAGACGACACGAGGAGTCTGTGTCACTGAGCCAGTGCTTCTAGATGCTACCCTGTGTGGGCGGCACCTCAGGGACAGTAAAT CAGAAATGCTGGTCTTGAAACCTTGAAAAGATCAAGCTGAATGTTCCTTTTCATCTGTCGCTGTTGATCTTCATCTATTTAAATAGGTAT TCTAACGTTTCCTCTCTGTATTTCATGAAGCTGATTTCCTCTCTCTTTCCTTTTCAGCAATACTGGAGTAACCGCTTCCTAAACCATTTT GCAGAAATGTAAGGGTGTTCGGTTGCGTGCATGTGCGTTTTTAGCAACACATCTACCAACCCTGTGCATGACTGATGTTGGGGAAAAAGA AAAGTAAAAAACTTCCCAACTCACTTTGTGTTATGTGGAGGAAATGTGTATTACCAATGGGGTTGTTAGCTTTTAAATCAAAATACTGAT TACAGATGTACAATTTAGCTTAATCAGAAAGCCTCTCCAGAGAAGTTTGGTTTCTTTGCTGCAAGAGGAATGAGGCTCTGTAACCTTATC TAAGAACTTGGAAGCCGTCAGCCAAGTCGCCACATTTCTCTGCAAAATGTCATAGCTTATATAAATGTACAGTATTCAATTGTAATGCAT GCCTTCGGTTGTAAGTAGCCAGATCCCTCTCCAGTGACATTGGAACATGCTACTTTTTAATTGGCCCTGTACAGTTTGCTTATTTATAAA TTCATTAAAAACACTACAGGTGTTGAATGGTTAAAATGTAGGCCTCCAGTTCATTTTCAGTTATTTTCTGAGTGTGCAGACAGCTATTTC GCACTGTATTAAATGTAACTTATTTAATGAAATCAGAAGCAGTAGACAGATGTTGGTGCAATACAAATATTGTGATGCATTTATCTTAAT AAAATGCTAAATGTCAATTTATCACTGCGCATGTTTGACTTTAGACTGTAAATAGAGATCAGTTTGTTTCTTTCTGTGCTGGTAACAATG >4587_4587_2_ANKRD13A-ATP2A2_ANKRD13A_chr12_110437589_ENST00000261739_ATP2A2_chr12_110734404_ENST00000395494_length(amino acids)=939AA_BP=0 MSSACDAGDHYPLHLLVWKNDYRQLEKELQGQERNAENAIEALKEYEPEMGKVYRQDRKSVQRIKAKDIVPGDIVEIAGESVSVIKHTDP VPDPRAVNQDKKNMLFSGTNIAAGKAMGVVVATGVNTEIGKIRDEMVATEQERTPLQQKLDEFGEQLSKVISLICIAVWIINIGHFNDPV HGGSWIRGAIYYFKIAVALAVAAIPEGLPAVITTCLALGTRRMAKKNAIVRSLPSVETLGCTSVICSDKTGTLTTNQMSVCRMFILDRVE GDTCSLNEFTITGSTYAPIGEVHKDDKPVNCHQYDGLVELATICALCNDSALDYNEAKGVYEKVGEATETALTCLVEKMNVFDTELKGLS KIERANACNSVIKQLMKKEFTLEFSRDRKSMSVYCTPNKPSRTSMSKMFVKGAPEGVIDRCTHIRVGSTKVPMTSGVKQKIMSVIREWGS GSDTLRCLALATHDNPLRREEMHLEDSANFIKYETNLTFVGCVGMLDPPRIEVASSVKLCRQAGIRVIMITGDNKGTAVAICRRIGIFGQ DEDVTSKAFTGREFDELNPSAQRDACLNARCFARVEPSHKSKIVEFLQSFDEITAMTGDGVNDAPALKKAEIGIAMGSGTAVAKTASEMV LADDNFSTIVAAVEEGRAIYNNMKQFIRYLISSNVGEVVCIFLTAALGFPEALIPVQLLWVNLVTDGLPATALGFNPPDLDIMNKPPRNP KEPLISGWLFFRYLAIGCYVGAATVGAAAWWFIAADGGPRVSFYQLSHFLQCKEDNPDFEGVDCAIFESPYPMTMALSVLVTIEMCNALN SLSENQSLLRMPPWENIWLVGSICLSMSLHFLILYVEPLPLIFQITPLNVTQWLMVLKISLPVILMDETLKFVARNYLEPGKECVQPATK -------------------------------------------------------------- >4587_4587_3_ANKRD13A-ATP2A2_ANKRD13A_chr12_110437589_ENST00000261739_ATP2A2_chr12_110734404_ENST00000539276_length(transcript)=3923nt_BP=262nt CGGACCTCCCGCGCGCCCCGCACCCGACCGGCTCAGCCGGCCGGCAGCGTAACACGCCCTACGCTCGCTTGCTCGCCGGCCTCAGGGCAG GCAGGCGGGCGCGGGAGACCCCGCCGGGGCCGAGACTTGGGGCGGGCGACGAGGACCAGGTTACGGCCTCCTCGCCATGTCCTCGGCCTG CGACGCGGGCGACCACTACCCCCTGCACCTCCTAGTCTGGAAAAACGACTACCGGCAGCTCGAGAAGGAGCTGCAGGGCCAGGAAAGAAA TGCTGAAAATGCCATCGAAGCCCTTAAGGAATATGAGCCTGAAATGGGCAAAGTGTATCGACAGGACAGAAAGAGTGTGCAGCGGATTAA AGCTAAAGACATAGTTCCTGGTGATATTGTAGAAATTGCTGTTGGTGACAAAGTTCCTGCTGATATAAGGTTAACTTCCATCAAATCTAC CACACTAAGAGTTGACCAGTCAATTCTCACAGGTGAATCTGTCTCTGTCATCAAGCACACTGATCCCGTCCCTGACCCACGAGCTGTCAA CCAAGATAAAAAGAACATGCTGTTTTCTGGTACAAACATTGCTGCTGGGAAAGCTATGGGAGTGGTGGTAGCAACTGGAGTTAACACCGA AATTGGCAAGATCCGGGATGAAATGGTGGCAACAGAACAGGAGAGAACACCCCTTCAGCAAAAACTAGATGAATTTGGGGAACAGCTTTC CAAAGTCATCTCCCTTATTTGCATTGCAGTCTGGATCATAAATATTGGGCACTTCAATGACCCGGTTCATGGAGGGTCCTGGATCAGAGG TGCTATTTACTACTTTAAAATTGCAGTGGCCCTGGCTGTAGCAGCCATTCCTGAAGGTCTGCCTGCAGTCATCACCACCTGCCTGGCTCT TGGAACTCGCAGAATGGCAAAGAAAAATGCCATTGTTCGAAGCCTCCCGTCTGTGGAAACCCTTGGTTGTACTTCTGTTATCTGCTCAGA CAAGACTGGTACACTTACAACAAACCAGATGTCAGTCTGCAGGATGTTCATTCTGGACAGAGTGGAAGGTGATACTTGTTCCCTTAATGA GTTTACCATAACTGGATCAACTTATGCACCTATTGGAGAAGTGCATAAAGATGATAAACCAGTGAATTGTCACCAGTATGATGGTCTGGT AGAATTAGCAACAATTTGTGCTCTTTGTAATGACTCTGCTTTGGATTACAATGAGGCAAAGGGTGTGTATGAAAAAGTTGGAGAAGCTAC AGAGACTGCTCTCACTTGCCTAGTAGAGAAGATGAATGTATTTGATACCGAATTGAAGGGTCTTTCTAAAATAGAACGTGCAAATGCCTG CAACTCAGTCATTAAACAGCTGATGAAAAAGGAATTCACTCTAGAGTTTTCACGTGACAGAAAGTCAATGTCGGTTTACTGTACACCAAA TAAACCAAGCAGGACATCAATGAGCAAGATGTTTGTGAAGGGTGCTCCTGAAGGTGTCATTGACAGGTGCACCCACATTCGAGTTGGAAG TACTAAGGTTCCTATGACCTCTGGAGTCAAACAGAAGATCATGTCTGTCATTCGAGAGTGGGGTAGTGGCAGCGACACACTGCGATGCCT GGCCCTGGCCACTCATGACAACCCACTGAGAAGAGAAGAAATGCACCTTGAGGACTCTGCCAACTTTATTAAATATGAGACCAATCTGAC CTTCGTTGGCTGCGTGGGCATGCTGGATCCTCCGAGAATCGAGGTGGCCTCCTCCGTGAAGCTGTGCCGGCAAGCAGGCATCCGGGTCAT CATGATCACTGGGGACAACAAGGGCACTGCTGTGGCCATCTGTCGCCGCATCGGCATCTTCGGGCAGGATGAGGACGTGACGTCAAAAGC TTTCACAGGCCGGGAGTTTGATGAACTCAACCCCTCCGCCCAGCGAGACGCCTGCCTGAACGCCCGCTGTTTTGCTCGAGTTGAACCCTC CCACAAGTCTAAAATCGTAGAATTTCTTCAGTCTTTTGATGAGATTACAGCTATGACTGGCGATGGCGTGAACGATGCTCCTGCTCTGAA GAAAGCCGAGATTGGCATTGCTATGGGCTCTGGCACTGCGGTGGCTAAAACCGCCTCTGAGATGGTCCTGGCGGATGACAACTTCTCCAC CATTGTGGCTGCCGTTGAGGAGGGGCGGGCAATCTACAACAACATGAAACAGTTCATCCGCTACCTCATCTCGTCCAACGTCGGGGAAGT TGTCTGTATTTTCCTGACAGCAGCCCTTGGATTTCCCGAGGCTTTGATTCCTGTTCAGCTGCTCTGGGTCAATCTGGTGACAGATGGCCT GCCTGCCACTGCACTGGGGTTCAACCCTCCTGATCTGGACATCATGAATAAACCTCCCCGGAACCCAAAGGAACCATTGATCAGCGGGTG GCTCTTTTTCCGTTACTTGGCTATTGGCTGTTACGTCGGCGCTGCTACCGTGGGTGCTGCTGCATGGTGGTTCATTGCTGCTGACGGTGG TCCAAGAGTGTCCTTCTACCAGCTGAGTCATTTCCTACAGTGTAAAGAGGACAACCCGGACTTTGAAGGCGTGGATTGTGCAATCTTTGA ATCCCCATACCCGATGACAATGGCGCTCTCTGTTCTAGTAACTATAGAAATGTGTAACGCCCTCAACAGCTTGTCCGAAAACCAGTCCTT GCTGAGGATGCCCCCCTGGGAGAACATCTGGCTCGTGGGCTCCATCTGCCTGTCCATGTCACTCCACTTCCTGATCCTCTATGTCGAACC CTTGCCACTCATCTTCCAGATCACACCGCTGAACGTGACCCAGTGGCTGATGGTGCTGAAAATCTCCTTGCCCGTGATTCTCATGGATGA GACGCTCAAGTTTGTGGCCCGCAACTACCTGGAACCTGGTAAAGAGTGTGTGCAGCCTGCCACCAAATCCTGCTCGTTCTCGGCATGCAC CGATGGGATTTCCTGGCCGTTTGTGCTGCTCATAATGCCCCTGGTGATCTGGGTCTATAGCACAGACACTAACTTTAGCGATATGTTCTG GTCTTGACTGACAGTTTTCCATAAAGAAGATGTTTAACTTAATCAATTAATTTTTTTATTGTTTAAAGCAACTGTCTATTTCTGCTGAAT TTTCACATGAACATACTGGCTGGTGATGGAGGTTTCATACTCTAGATTTTGTTTTGCTTTTTCTGACTCCAGTGGGGCAAGATTTTCCTT TTTTATACACATAATTAAAGTGTCCATTGACATGTACAGAGAACTAACACTATTTTATGCAAATATTTTTTTGTAGATGAAAAAGCATGT ACAGTGTTCTGTTTAATACTCATCCTTGTATAAAAAAAATAGTTGAGCCAGCAGACATTGTCAGCAAATTAATTGGCAGCAGATTTTAGG AAATGAATGTGTGTGGTTTTTTTTCTAAAACTAAATAGCATGTATTGTGTCTTTTGCATGATGATCCGGATTTAATTTGATATCACAGTC TAATTTTTATTCATAAGCCAATTTTTCTGCACTGAGCAGAGTCTTGCTACCTCAGTCAGTATTGTTTTGGTTTGCTACTTCCCTCACCCA CTTTGGCCTCCGTTCACCCCACCCCACCCCACCTCTCCCCACCTTACCCCCGCCCCGCTTGGCTTCTTCTTTAGGATTGTGATGGTTCGT TCTGTTTACATCAGTTTTAACGAGAGGTATGCCTGTACTCGCTTGTGCAGAAAACATTGTTCCAGATTCAATCGACTGGGTTTATGTCCC TTCACATAGTTTTTAAGGTTATTTATTTAAATGTCTAATGTATTTTATTGTAACAGACATTGTTTTGCCAACATTGCCTATTTCAGTGGC >4587_4587_3_ANKRD13A-ATP2A2_ANKRD13A_chr12_110437589_ENST00000261739_ATP2A2_chr12_110734404_ENST00000539276_length(amino acids)=966AA_BP=0 MSSACDAGDHYPLHLLVWKNDYRQLEKELQGQERNAENAIEALKEYEPEMGKVYRQDRKSVQRIKAKDIVPGDIVEIAVGDKVPADIRLT SIKSTTLRVDQSILTGESVSVIKHTDPVPDPRAVNQDKKNMLFSGTNIAAGKAMGVVVATGVNTEIGKIRDEMVATEQERTPLQQKLDEF GEQLSKVISLICIAVWIINIGHFNDPVHGGSWIRGAIYYFKIAVALAVAAIPEGLPAVITTCLALGTRRMAKKNAIVRSLPSVETLGCTS VICSDKTGTLTTNQMSVCRMFILDRVEGDTCSLNEFTITGSTYAPIGEVHKDDKPVNCHQYDGLVELATICALCNDSALDYNEAKGVYEK VGEATETALTCLVEKMNVFDTELKGLSKIERANACNSVIKQLMKKEFTLEFSRDRKSMSVYCTPNKPSRTSMSKMFVKGAPEGVIDRCTH IRVGSTKVPMTSGVKQKIMSVIREWGSGSDTLRCLALATHDNPLRREEMHLEDSANFIKYETNLTFVGCVGMLDPPRIEVASSVKLCRQA GIRVIMITGDNKGTAVAICRRIGIFGQDEDVTSKAFTGREFDELNPSAQRDACLNARCFARVEPSHKSKIVEFLQSFDEITAMTGDGVND APALKKAEIGIAMGSGTAVAKTASEMVLADDNFSTIVAAVEEGRAIYNNMKQFIRYLISSNVGEVVCIFLTAALGFPEALIPVQLLWVNL VTDGLPATALGFNPPDLDIMNKPPRNPKEPLISGWLFFRYLAIGCYVGAATVGAAAWWFIAADGGPRVSFYQLSHFLQCKEDNPDFEGVD CAIFESPYPMTMALSVLVTIEMCNALNSLSENQSLLRMPPWENIWLVGSICLSMSLHFLILYVEPLPLIFQITPLNVTQWLMVLKISLPV -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for ANKRD13A-ATP2A2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 575_594 | 108.0 | 998.0 | HAX1 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 575_594 | 108.0 | 1016.0 | HAX1 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 575_594 | 108.0 | 1043.0 | HAX1 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 787_807 | 108.0 | 998.0 | PLN | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 931_942 | 108.0 | 998.0 | PLN | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 787_807 | 108.0 | 1016.0 | PLN | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 931_942 | 108.0 | 1016.0 | PLN | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 787_807 | 108.0 | 1043.0 | PLN | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 931_942 | 108.0 | 1043.0 | PLN | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000308664 | 3 | 21 | 788_1042 | 108.0 | 998.0 | TMEM64 and PDIA3 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000395494 | 3 | 19 | 788_1042 | 108.0 | 1016.0 | TMEM64 and PDIA3 | |
| Tgene | ATP2A2 | chr12:110437589 | chr12:110734404 | ENST00000539276 | 3 | 20 | 788_1042 | 108.0 | 1043.0 | TMEM64 and PDIA3 |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for ANKRD13A-ATP2A2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for ANKRD13A-ATP2A2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
