
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:LONP1-RAD51 (FusionGDB2 ID:49423) |
Fusion Gene Summary for LONP1-RAD51 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: LONP1-RAD51 | Fusion gene ID: 49423 | Hgene | Tgene | Gene symbol | LONP1 | RAD51 | Gene ID | 9361 | 5888 |
| Gene name | lon peptidase 1, mitochondrial | RAD51 recombinase | |
| Synonyms | CODASS|LON|LONP|LonHS|PIM1|PRSS15|hLON | BRCC5|FANCR|HRAD51|HsRad51|HsT16930|MRMV2|RAD51A|RECA | |
| Cytomap | 19p13.3 | 15q15.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | lon protease homolog, mitochondrialhLON ATP-dependent proteaselon protease-like proteinmitochondrial ATP-dependent protease Lonmitochondrial lon protease-like proteinserine protease 15 | DNA repair protein RAD51 homolog 1BRCA1/BRCA2-containing complex, subunit 5RAD51 homolog ARecA, E. coli, homolog ofRecA-like proteinrecombination protein A | |
| Modification date | 20200313 | 20200322 | |
| UniProtAcc | P36776 | O15315 | |
| Ensembl transtripts involved in fusion gene | ENST00000360614, ENST00000540670, ENST00000585374, ENST00000590729, ENST00000593119, ENST00000590511, | ENST00000530766, ENST00000382643, ENST00000423169, ENST00000532743, ENST00000557850, ENST00000267868, | |
| Fusion gene scores | * DoF score | 18 X 14 X 12=3024 | 7 X 7 X 5=245 |
| # samples | 23 | 7 | |
| ** MAII score | log2(23/3024*10)=-3.7167523732767 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(7/245*10)=-1.8073549220576 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: LONP1 [Title/Abstract] AND RAD51 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | LONP1(5700800)-RAD51(41023253), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | LONP1-RAD51 seems lost the major protein functional domain in Hgene partner, which is a essential gene due to the frame-shifted ORF. LONP1-RAD51 seems lost the major protein functional domain in Tgene partner, which is a epigenetic factor due to the frame-shifted ORF. | ||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | LONP1 | GO:0034599 | cellular response to oxidative stress | 17420247 |
| Hgene | LONP1 | GO:0051603 | proteolysis involved in cellular protein catabolic process | 8248235|17420247 |
| Tgene | RAD51 | GO:0000724 | double-strand break repair via homologous recombination | 16428451 |
| Tgene | RAD51 | GO:0006268 | DNA unwinding involved in DNA replication | 7988572 |
| Tgene | RAD51 | GO:0006974 | cellular response to DNA damage stimulus | 23509288|25585578 |
| Tgene | RAD51 | GO:0010569 | regulation of double-strand break repair via homologous recombination | 23754376 |
| Tgene | RAD51 | GO:0031297 | replication fork processing | 25585578 |
| Tgene | RAD51 | GO:0051106 | positive regulation of DNA ligation | 8929543 |
| Tgene | RAD51 | GO:0071479 | cellular response to ionizing radiation | 23509288|23754376 |
| Tgene | RAD51 | GO:0072757 | cellular response to camptothecin | 23509288 |
 Fusion gene breakpoints across LONP1 (5'-gene) Fusion gene breakpoints across LONP1 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
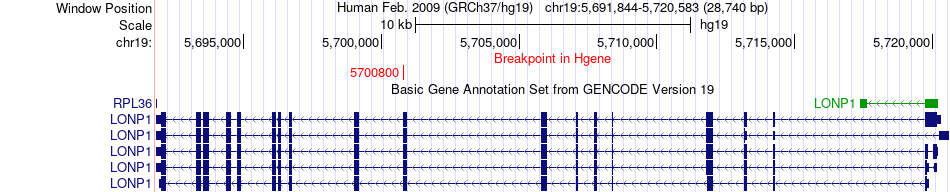 |
 Fusion gene breakpoints across RAD51 (3'-gene) Fusion gene breakpoints across RAD51 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
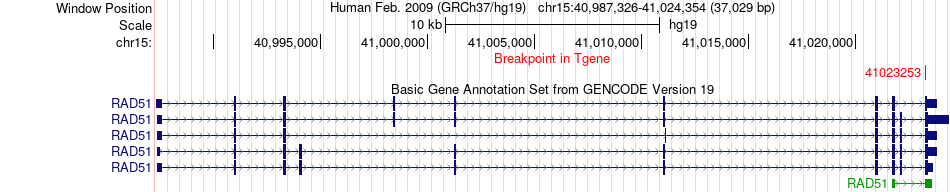 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | CESC | TCGA-EA-A50E-01A | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
Top |
Fusion Gene ORF analysis for LONP1-RAD51 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-3UTR | ENST00000360614 | ENST00000530766 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| 5CDS-3UTR | ENST00000540670 | ENST00000530766 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| 5CDS-3UTR | ENST00000585374 | ENST00000530766 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| 5CDS-3UTR | ENST00000590729 | ENST00000530766 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| 5CDS-3UTR | ENST00000593119 | ENST00000530766 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000360614 | ENST00000382643 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000360614 | ENST00000423169 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000360614 | ENST00000532743 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000360614 | ENST00000557850 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000540670 | ENST00000382643 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000540670 | ENST00000423169 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000540670 | ENST00000532743 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000540670 | ENST00000557850 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000585374 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000585374 | ENST00000382643 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000585374 | ENST00000423169 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000585374 | ENST00000532743 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000585374 | ENST00000557850 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000590729 | ENST00000382643 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000590729 | ENST00000423169 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000590729 | ENST00000532743 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000590729 | ENST00000557850 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000593119 | ENST00000382643 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000593119 | ENST00000423169 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000593119 | ENST00000532743 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| Frame-shift | ENST00000593119 | ENST00000557850 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| In-frame | ENST00000360614 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| In-frame | ENST00000540670 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| In-frame | ENST00000590729 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| In-frame | ENST00000593119 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| intron-3CDS | ENST00000590511 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| intron-3CDS | ENST00000590511 | ENST00000382643 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| intron-3CDS | ENST00000590511 | ENST00000423169 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| intron-3CDS | ENST00000590511 | ENST00000532743 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| intron-3CDS | ENST00000590511 | ENST00000557850 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
| intron-3UTR | ENST00000590511 | ENST00000530766 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000360614 | LONP1 | chr19 | 5700800 | - | ENST00000267868 | RAD51 | chr15 | 41023253 | + | 2766 | 1664 | 38 | 1732 | 564 |
| ENST00000593119 | LONP1 | chr19 | 5700800 | - | ENST00000267868 | RAD51 | chr15 | 41023253 | + | 2449 | 1347 | 33 | 1415 | 460 |
| ENST00000540670 | LONP1 | chr19 | 5700800 | - | ENST00000267868 | RAD51 | chr15 | 41023253 | + | 2533 | 1431 | 369 | 1499 | 376 |
| ENST00000590729 | LONP1 | chr19 | 5700800 | - | ENST00000267868 | RAD51 | chr15 | 41023253 | + | 2277 | 1175 | 59 | 1243 | 394 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000360614 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + | 0.002372993 | 0.9976271 |
| ENST00000593119 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + | 0.001922995 | 0.99807703 |
| ENST00000540670 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + | 0.001983171 | 0.99801683 |
| ENST00000590729 | ENST00000267868 | LONP1 | chr19 | 5700800 | - | RAD51 | chr15 | 41023253 | + | 0.001130007 | 0.99886996 |
Top |
Fusion Genomic Features for LONP1-RAD51 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| LONP1 | chr19 | 5700799 | - | RAD51 | chr15 | 41023252 | + | 1.26E-06 | 0.9999987 |
| LONP1 | chr19 | 5700799 | - | RAD51 | chr15 | 41023252 | + | 1.26E-06 | 0.9999987 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
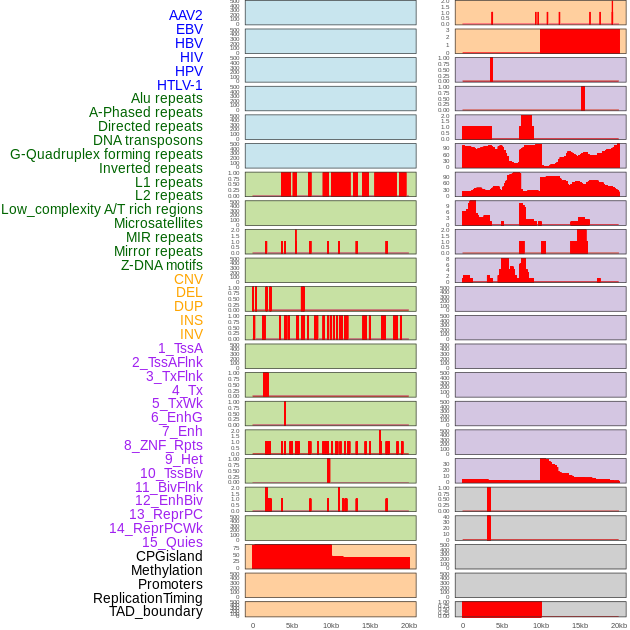 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
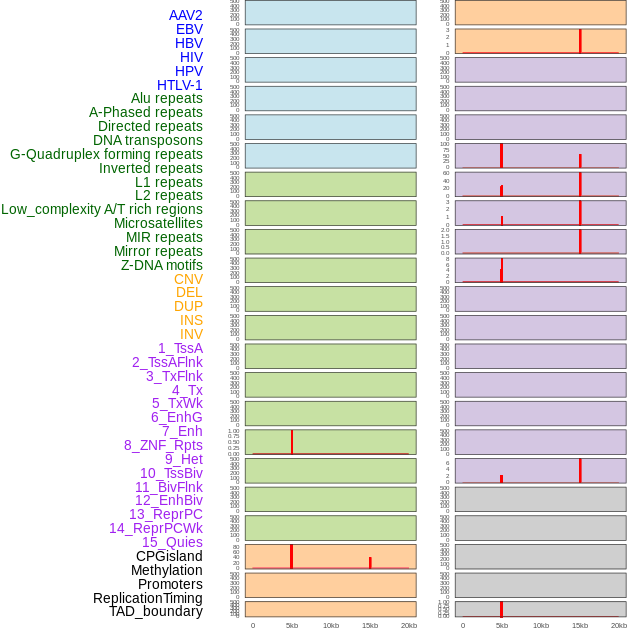 |
Top |
Fusion Protein Features for LONP1-RAD51 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr19:5700800/chr15:41023253) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| LONP1 | RAD51 |
| FUNCTION: ATP-dependent serine protease that mediates the selective degradation of misfolded, unassembled or oxidatively damaged polypeptides as well as certain short-lived regulatory proteins in the mitochondrial matrix. May also have a chaperone function in the assembly of inner membrane protein complexes. Participates in the regulation of mitochondrial gene expression and in the maintenance of the integrity of the mitochondrial genome. Binds to mitochondrial promoters and RNA in a single-stranded, site-specific, and strand-specific manner. May regulate mitochondrial DNA replication and/or gene expression using site-specific, single-stranded DNA binding to target the degradation of regulatory proteins binding to adjacent sites in mitochondrial promoters (PubMed:12198491, PubMed:15870080, PubMed:17420247, PubMed:8248235). Endogenous substrates include mitochondrial steroidogenic acute regulatory (StAR) protein, helicase Twinkle (TWNK) and the large ribosomal subunit protein bL32m. bL32m is protected from degradation by LONP1 when it is bound to a nucleic acid (RNA), but TWNK is not (PubMed:17579211, PubMed:28377575). {ECO:0000255|HAMAP-Rule:MF_03120, ECO:0000269|PubMed:12198491, ECO:0000269|PubMed:15870080, ECO:0000269|PubMed:17420247, ECO:0000269|PubMed:17579211, ECO:0000269|PubMed:28377575, ECO:0000269|PubMed:8248235}. | FUNCTION: Involved in the homologous recombination repair (HRR) pathway of double-stranded DNA breaks arising during DNA replication or induced by DNA-damaging agents. May promote the assembly of presynaptic RAD51 nucleoprotein filaments. Binds single-stranded DNA and double-stranded DNA and has DNA-dependent ATPase activity. Part of the RAD21 paralog protein complex BCDX2 which acts in the BRCA1-BRCA2-dependent HR pathway. Upon DNA damage, BCDX2 acts downstream of BRCA2 recruitment and upstream of RAD51 recruitment. BCDX2 binds predominantly to the intersection of the four duplex arms of the Holliday junction and to junction of replication forks. The BCDX2 complex was originally reported to bind single-stranded DNA, single-stranded gaps in duplex DNA and specifically to nicks in duplex DNA. The BCDX2 subcomplex RAD51B:RAD51C exhibits single-stranded DNA-dependent ATPase activity suggesting an involvement in early stages of the HR pathway. {ECO:0000269|PubMed:11751635, ECO:0000269|PubMed:11751636, ECO:0000269|PubMed:11842113, ECO:0000269|PubMed:12441335, ECO:0000269|PubMed:23108668, ECO:0000269|PubMed:23149936}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LONP1 | chr19:5700800 | chr15:41023253 | ENST00000360614 | - | 9 | 18 | 124_368 | 502 | 960.0 | Domain | Lon N-terminal |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LONP1 | chr19:5700800 | chr15:41023253 | ENST00000360614 | - | 9 | 18 | 759_949 | 502 | 960.0 | Domain | Lon proteolytic |
| Hgene | LONP1 | chr19:5700800 | chr15:41023253 | ENST00000360614 | - | 9 | 18 | 523_530 | 502 | 960.0 | Nucleotide binding | ATP |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000267868 | 8 | 10 | 48_77 | 298 | 340.0 | Domain | Note=HhH | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000382643 | 8 | 10 | 48_77 | 299 | 341.0 | Domain | Note=HhH | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000423169 | 7 | 9 | 48_77 | 258 | 281.0 | Domain | Note=HhH | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000532743 | 8 | 10 | 48_77 | 299 | 341.0 | Domain | Note=HhH | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000557850 | 6 | 8 | 48_77 | 201 | 243.0 | Domain | Note=HhH | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000267868 | 8 | 10 | 127_134 | 298 | 340.0 | Nucleotide binding | ATP | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000382643 | 8 | 10 | 127_134 | 299 | 341.0 | Nucleotide binding | ATP | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000423169 | 7 | 9 | 127_134 | 258 | 281.0 | Nucleotide binding | ATP | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000532743 | 8 | 10 | 127_134 | 299 | 341.0 | Nucleotide binding | ATP | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000557850 | 6 | 8 | 127_134 | 201 | 243.0 | Nucleotide binding | ATP |
Top |
Fusion Gene Sequence for LONP1-RAD51 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >49423_49423_1_LONP1-RAD51_LONP1_chr19_5700800_ENST00000360614_RAD51_chr15_41023253_ENST00000267868_length(transcript)=2766nt_BP=1664nt CGAACTCCGCCCCCTCTTCTCCGCGTAGGCCCAGCTCCCTGAAGCGGCTGTTTCGAGCCACGCGCCCATCGGGTACCGAGGCACGCGCCG GGCGTCACGTGCGTTTCGCGGCGAGCGGAAATGACGCGAGTTGTGTGAGCCGCCAGTATGGCCGGGCTATGGCGGCGAGCACTGGCTACG TGCGACTGTGGGGAGCGGCGCGGTGCTGGGTGCTGCGGCGGCCGATGCTGGCCGCCGCCGGGGGGCGGGTTCCCACTGCAGCAGGAGCGT GGTTGCTCCGAGGCCAGCGGACCTGCGACGCCTCTCCTCCTTGGGCACTGTGGGGCCGAGGCCCGGCAATTGGGGGCCAATGGCGGGGGT TTTGGGAAGCGAGCAGCCGCGGCGGAGGCGCATTCTCGGGGGGCGAGGACGCCTCCGAGGGCGGCGCGGAGGAAGGAGCCGGCGGCGCGG GGGGCAGCGCGGGCGCCGGGGAAGGCCCGGTCATAACGGCGCTCACGCCCATGACGATCCCCGATGTGTTTCCGCACCTGCCGCTCATCG CCATCACCCGCAACCCGGTGTTCCCGCGCTTTATCAAGATTATCGAGGTTAAAAATAAGAAGTTGGTTGAGCTGCTGAGAAGGAAAGTTC GTCTCGCCCAGCCTTATGTCGGCGTCTTTCTAAAGAGAGATGACAGCAATGAGTCGGATGTGGTCGAGAGCCTGGATGAAATCTACCACA CGGGGACGTTTGCCCAGATCCATGAGATGCAGGACCTTGGGGACAAGCTGCGCATGATCGTCATGGGACACAGAAGAGTCCATATCAGCA GACAGCTGGAGGTGGAGCCCGAGGAGCCGGAGGCGGAGAACAAGCACAAGCCCCGCAGGAAGTCAAAGCGGGGCAAGAAGGAGGCGGAGG ACGAGCTGAGCGCCAGGCACCCGGCGGAGCTGGCGATGGAGCCCACCCCTGAGCTCCCGGCTGAGGTGCTCATGGTGGAGGTAGAGAACG TTGTCCACGAGGACTTCCAGGTCACGGAGGAGGTGAAAGCCCTGACTGCAGAGATCGTGAAGACCATCCGGGACATCATTGCCTTGAACC CTCTCTACAGGGAGTCAGTGCTGCAGATGATGCAGGCTGGCCAGCGGGTGGTGGACAACCCCATCTACCTGAGCGACATGGGCGCCGCGC TCACCGGGGCCGAGTCCCATGAGCTGCAGGACGTCCTGGAAGAGACCAATATTCCTAAGCGGCTGTACAAGGCCCTCTCCCTGCTGAAGA AGGAATTTGAACTGAGCAAGCTGCAGCAGCGCCTGGGGCGGGAGGTGGAGGAGAAGATCAAGCAGACCCACCGTAAGTACCTGCTGCAGG AGCAGCTAAAGATCATCAAGAAGGAGCTGGGCCTGGAGAAGGACGACAAGGATGCCATCGAGGAGAAGTTCCGGGAGCGCCTGAAGGAGC TCGTGGTCCCCAAGCACGTCATGGATGTTGTGGACGAGGAGCTGAGCAAGCTGGGCCTGCTGGACAACCACTCCTCGGAGTTCAATGTCA CCCGCAACTACCTAGACTGGCTCACGTCCATCCCTTGGGGCAAGTACAGCAACGAGAACCTGGACCTGGCGCGGGCACAGGCAGTGCTGG AGGAAGACCACTACGGCATGGAGGACGTCAAGAAACGCATCCTGATTGTATCTGAGGAAAGGAAGAGGGGAAACCAGAATCTGCAAAATC TACGACTCTCCCTGTCTTCCTGAAGCTGAAGCTATGTTCGCCATTAATGCAGATGGAGTGGGAGATGCCAAAGACTGAATCATTGGGTTT TTCCTCTGTTAAAAACCTTAAGTGCTGCAGCCTAATGAGAGTGCACTGCTCCCTGGGGTTCTCTACAGGCCTCTTCCTGTTGTGACTGCC AGGATAAAGCTTCCGGGAAAACAGCTATTATATCAGCTTTTCTGATGGTATAAACAGGAGACAGGTCAGTAGTCACAAACTGATCTAAAA TGTTTATTCCTTCTGTAGTGTATTAATCTCTGTGTGTTTTCTTTGGTTTTGGAGGAGGGGTATGAAGTATCTTTGACATGGTGCCTTAGG AATGACTTGGGTTTAACAAGCTGTCTACTGGACAATCTTATGTTTCCAAGAGAACTAAAGCTGGAGAGACCTGACCCTTCTCTCACTTCT AAATTAATGGTAAAATAAAATGCCTCAGCTATGTAGCAAAGGGAATGGGTCTGCACAGATTCTTTTTTTCTGTCAGTAAAACTCTCAAGC AGGTTTTTAAGTTGTCTGTCTGAATGATCTTGTGTAAGGTTTTGGTTATGGAGTCTTGTGCCAAACCTACTAGGCCATTAGCCCTTCACC ATCTACCTGCTTGGTCTTTCATTGCTAAGACTAACTCAAGATAATCCTAGAGTCTTAAAGCATTTCAGGCCAGTGTGGTGTCTTGCGCCT GTACTCCCAGCACTTTGGGAGGCCGAGGCAGGTGGATCGCTTGAGCCCAGGAGTTTTAAGTCCAGCTTGGCCAAGGTGGTGAAATCCCAT CTCTACAAAAAATGCAGAACTTAATCTGGACACACTGTTACACGTGCCTGTAGTCCCAGCTACTCGATAGCCTGAGGTGGGAGAATCACT TAAGCCTGGAAGGTGGAAGTTGCAGTGAGTCGAGATTGCACTGCTGCATTCCAGCCAGGGTGACAGAGTGAGACCATGTTTCAAACAAGA >49423_49423_1_LONP1-RAD51_LONP1_chr19_5700800_ENST00000360614_RAD51_chr15_41023253_ENST00000267868_length(amino acids)=564AA_BP=542 MKRLFRATRPSGTEARAGRHVRFAASGNDASCVSRQYGRAMAASTGYVRLWGAARCWVLRRPMLAAAGGRVPTAAGAWLLRGQRTCDASP PWALWGRGPAIGGQWRGFWEASSRGGGAFSGGEDASEGGAEEGAGGAGGSAGAGEGPVITALTPMTIPDVFPHLPLIAITRNPVFPRFIK IIEVKNKKLVELLRRKVRLAQPYVGVFLKRDDSNESDVVESLDEIYHTGTFAQIHEMQDLGDKLRMIVMGHRRVHISRQLEVEPEEPEAE NKHKPRRKSKRGKKEAEDELSARHPAELAMEPTPELPAEVLMVEVENVVHEDFQVTEEVKALTAEIVKTIRDIIALNPLYRESVLQMMQA GQRVVDNPIYLSDMGAALTGAESHELQDVLEETNIPKRLYKALSLLKKEFELSKLQQRLGREVEEKIKQTHRKYLLQEQLKIIKKELGLE KDDKDAIEEKFRERLKELVVPKHVMDVVDEELSKLGLLDNHSSEFNVTRNYLDWLTSIPWGKYSNENLDLARAQAVLEEDHYGMEDVKKR -------------------------------------------------------------- >49423_49423_2_LONP1-RAD51_LONP1_chr19_5700800_ENST00000540670_RAD51_chr15_41023253_ENST00000267868_length(transcript)=2533nt_BP=1431nt AGACGAATCCCGGGAAGCCCGAATCCGCGCACCCCAAAGTGCGCTCAGAGCCTGAGTATGCGCACCCGGCCACCCGTGGCCAGTCTAGAG TCTGAGCATGCGCATGTTGTCCGCGCGCCTAAGCATGTGCACAGGCTGCTCATGAGTTCCTAAACGTTGCGTTTGCAACACTACCAGTGT TTTGTACGGGCGCTGGCGCCTGCTCACAGCCTTCCACGTGACCGGAAAAGGCCTTCAGCGCATAAAGTGGGAGACCGGAAGAGGCGGAGA CGATCCGATGCTCGAACTCCGCCCCCTCTTCTCCGCGTAGGCCCAGCTCCCTGAAGCGGCTGTTTCGAGCCACGCGCCCATCGGGTTAAA AATAAGAAGTTGGTTGAGCTGCTGAGAAGGAAAGTTCGTCTCGCCCAGCCTTATGTCGGCGTCTTTCTAAAGAGAGATGACAGCAATGAG TCGGATGTGGTCGAGAGCCTGGATGAAATCTACCACACGGGGACGTTTGCCCAGATCCATGAGATGCAGGACCTTGGGGACAAGCTGCGC ATGATCGTCATGGGACACAGAAGAGTCCATATCAGCAGACAGCTGGAGGTGGAGCCCGAGGAGCCGGAGGCGGAGAACAAGCACAAGCCC CGCAGGAAGTCAAAGCGGGGCAAGAAGGAGGCGGAGGACGAGCTGAGCGCCAGGCACCCGGCGGAGCTGGCGATGGAGCCCACCCCTGAG CTCCCGGCTGAGGTGCTCATGGTGGAGGTAGAGAACGTTGTCCACGAGGACTTCCAGGTCACGGAGGAGGTGAAAGCCCTGACTGCAGAG ATCGTGAAGACCATCCGGGACATCATTGCCTTGAACCCTCTCTACAGGGAGTCAGTGCTGCAGATGATGCAGGCTGGCCAGCGGGTGGTG GACAACCCCATCTACCTGAGCGACATGGGCGCCGCGCTCACCGGGGCCGAGTCCCATGAGCTGCAGGACGTCCTGGAAGAGACCAATATT CCTAAGCGGCTGTACAAGGCCCTCTCCCTGCTGAAGAAGGAATTTGAACTGAGCAAGCTGCAGCAGCGCCTGGGGCGGGAGGTGGAGGAG AAGATCAAGCAGACCCACCGTAAGTACCTGCTGCAGGAGCAGCTAAAGATCATCAAGAAGGAGCTGGGCCTGGAGAAGGACGACAAGGAT GCCATCGAGGAGAAGTTCCGGGAGCGCCTGAAGGAGCTCGTGGTCCCCAAGCACGTCATGGATGTTGTGGACGAGGAGCTGAGCAAGCTG GGCCTGCTGGACAACCACTCCTCGGAGTTCAATGTCACCCGCAACTACCTAGACTGGCTCACGTCCATCCCTTGGGGCAAGTACAGCAAC GAGAACCTGGACCTGGCGCGGGCACAGGCAGTGCTGGAGGAAGACCACTACGGCATGGAGGACGTCAAGAAACGCATCCTGATTGTATCT GAGGAAAGGAAGAGGGGAAACCAGAATCTGCAAAATCTACGACTCTCCCTGTCTTCCTGAAGCTGAAGCTATGTTCGCCATTAATGCAGA TGGAGTGGGAGATGCCAAAGACTGAATCATTGGGTTTTTCCTCTGTTAAAAACCTTAAGTGCTGCAGCCTAATGAGAGTGCACTGCTCCC TGGGGTTCTCTACAGGCCTCTTCCTGTTGTGACTGCCAGGATAAAGCTTCCGGGAAAACAGCTATTATATCAGCTTTTCTGATGGTATAA ACAGGAGACAGGTCAGTAGTCACAAACTGATCTAAAATGTTTATTCCTTCTGTAGTGTATTAATCTCTGTGTGTTTTCTTTGGTTTTGGA GGAGGGGTATGAAGTATCTTTGACATGGTGCCTTAGGAATGACTTGGGTTTAACAAGCTGTCTACTGGACAATCTTATGTTTCCAAGAGA ACTAAAGCTGGAGAGACCTGACCCTTCTCTCACTTCTAAATTAATGGTAAAATAAAATGCCTCAGCTATGTAGCAAAGGGAATGGGTCTG CACAGATTCTTTTTTTCTGTCAGTAAAACTCTCAAGCAGGTTTTTAAGTTGTCTGTCTGAATGATCTTGTGTAAGGTTTTGGTTATGGAG TCTTGTGCCAAACCTACTAGGCCATTAGCCCTTCACCATCTACCTGCTTGGTCTTTCATTGCTAAGACTAACTCAAGATAATCCTAGAGT CTTAAAGCATTTCAGGCCAGTGTGGTGTCTTGCGCCTGTACTCCCAGCACTTTGGGAGGCCGAGGCAGGTGGATCGCTTGAGCCCAGGAG TTTTAAGTCCAGCTTGGCCAAGGTGGTGAAATCCCATCTCTACAAAAAATGCAGAACTTAATCTGGACACACTGTTACACGTGCCTGTAG TCCCAGCTACTCGATAGCCTGAGGTGGGAGAATCACTTAAGCCTGGAAGGTGGAAGTTGCAGTGAGTCGAGATTGCACTGCTGCATTCCA GCCAGGGTGACAGAGTGAGACCATGTTTCAAACAAGAAACATTTCAGAGGGTAAGTAAACAGATTTGATTGTGAGGCTTCTAATAAAGTA >49423_49423_2_LONP1-RAD51_LONP1_chr19_5700800_ENST00000540670_RAD51_chr15_41023253_ENST00000267868_length(amino acids)=376AA_BP=354 MVELLRRKVRLAQPYVGVFLKRDDSNESDVVESLDEIYHTGTFAQIHEMQDLGDKLRMIVMGHRRVHISRQLEVEPEEPEAENKHKPRRK SKRGKKEAEDELSARHPAELAMEPTPELPAEVLMVEVENVVHEDFQVTEEVKALTAEIVKTIRDIIALNPLYRESVLQMMQAGQRVVDNP IYLSDMGAALTGAESHELQDVLEETNIPKRLYKALSLLKKEFELSKLQQRLGREVEEKIKQTHRKYLLQEQLKIIKKELGLEKDDKDAIE EKFRERLKELVVPKHVMDVVDEELSKLGLLDNHSSEFNVTRNYLDWLTSIPWGKYSNENLDLARAQAVLEEDHYGMEDVKKRILIVSEER -------------------------------------------------------------- >49423_49423_3_LONP1-RAD51_LONP1_chr19_5700800_ENST00000590729_RAD51_chr15_41023253_ENST00000267868_length(transcript)=2277nt_BP=1175nt GCGGCGCGGGGGGCAGCGCGGGCGCCGGGGAAGGCCCGGTCATAACGGCGCTCACGCCCATGACGATCCCCGATGTGTTTCCGCACCTGC CGCTCATCGCCATCACCCGCAACCCGGTGTTCCCGCGCTTTATCAAGATTATCGAGGTTAAAAATAAGAAGTTGGTTGAGCTGCTGAGAA GGAAAGTTCGTCTCGCCCAGCCTTATGTCGGCGTCTTTCTAAAGAGAGATGACAGCAATGAGTCGGATGTGGTCGAGAGCCTGGATGAAA TCTACCACACGGGGACGTTTGCCCAGATCCATGAGATGCAGGACCTTGGGGACAAGCTGCGCATGATCGTCATGGGACACAGAAGAGTCC ATATCAGCAGACAGCTGGAGGTGGAGCCCGAGGAGCCGGAGGCGGAGAACAAGCACAAGCCCCGCAGGAAGTCAAAGCGGGGCAAGAAGG AGGCGGAGGACGAGCTGAGCGCCAGGCACCCGGCGGAGCTGGCGATGGAGCCCACCCCTGAGCTCCCGGCTGAGGTGCTCATGGTGGAGG CCCTGACTGCAGAGATCGTGAAGACCATCCGGGACATCATTGCCTTGAACCCTCTCTACAGGGAGTCAGTGCTGCAGATGATGCAGGCTG GCCAGCGGGTGGTGGACAACCCCATCTACCTGAGCGACATGGGCGCCGCGCTCACCGGGGCCGAGTCCCATGAGCTGCAGGACGTCCTGG AAGAGACCAATATTCCTAAGCGGCTGTACAAGGCCCTCTCCCTGCTGAAGAAGGAATTTGAACTGAGCAAGCTGCAGCAGCGCCTGGGGC GGGAGGTGGAGGAGAAGATCAAGCAGACCCACCGTAAGTACCTGCTGCAGGAGCAGCTAAAGATCATCAAGAAGGAGCTGGGCCTGGAGA AGGACGACAAGGATGCCATCGAGGAGAAGTTCCGGGAGCGCCTGAAGGAGCTCGTGGTCCCCAAGCACGTCATGGATGTTGTGGACGAGG AGCTGAGCAAGCTGGGCCTGCTGGACAACCACTCCTCGGAGTTCAATGTCACCCGCAACTACCTAGACTGGCTCACGTCCATCCCTTGGG GCAAGTACAGCAACGAGAACCTGGACCTGGCGCGGGCACAGGCAGTGCTGGAGGAAGACCACTACGGCATGGAGGACGTCAAGAAACGCA TCCTGATTGTATCTGAGGAAAGGAAGAGGGGAAACCAGAATCTGCAAAATCTACGACTCTCCCTGTCTTCCTGAAGCTGAAGCTATGTTC GCCATTAATGCAGATGGAGTGGGAGATGCCAAAGACTGAATCATTGGGTTTTTCCTCTGTTAAAAACCTTAAGTGCTGCAGCCTAATGAG AGTGCACTGCTCCCTGGGGTTCTCTACAGGCCTCTTCCTGTTGTGACTGCCAGGATAAAGCTTCCGGGAAAACAGCTATTATATCAGCTT TTCTGATGGTATAAACAGGAGACAGGTCAGTAGTCACAAACTGATCTAAAATGTTTATTCCTTCTGTAGTGTATTAATCTCTGTGTGTTT TCTTTGGTTTTGGAGGAGGGGTATGAAGTATCTTTGACATGGTGCCTTAGGAATGACTTGGGTTTAACAAGCTGTCTACTGGACAATCTT ATGTTTCCAAGAGAACTAAAGCTGGAGAGACCTGACCCTTCTCTCACTTCTAAATTAATGGTAAAATAAAATGCCTCAGCTATGTAGCAA AGGGAATGGGTCTGCACAGATTCTTTTTTTCTGTCAGTAAAACTCTCAAGCAGGTTTTTAAGTTGTCTGTCTGAATGATCTTGTGTAAGG TTTTGGTTATGGAGTCTTGTGCCAAACCTACTAGGCCATTAGCCCTTCACCATCTACCTGCTTGGTCTTTCATTGCTAAGACTAACTCAA GATAATCCTAGAGTCTTAAAGCATTTCAGGCCAGTGTGGTGTCTTGCGCCTGTACTCCCAGCACTTTGGGAGGCCGAGGCAGGTGGATCG CTTGAGCCCAGGAGTTTTAAGTCCAGCTTGGCCAAGGTGGTGAAATCCCATCTCTACAAAAAATGCAGAACTTAATCTGGACACACTGTT ACACGTGCCTGTAGTCCCAGCTACTCGATAGCCTGAGGTGGGAGAATCACTTAAGCCTGGAAGGTGGAAGTTGCAGTGAGTCGAGATTGC ACTGCTGCATTCCAGCCAGGGTGACAGAGTGAGACCATGTTTCAAACAAGAAACATTTCAGAGGGTAAGTAAACAGATTTGATTGTGAGG >49423_49423_3_LONP1-RAD51_LONP1_chr19_5700800_ENST00000590729_RAD51_chr15_41023253_ENST00000267868_length(amino acids)=394AA_BP=372 MTIPDVFPHLPLIAITRNPVFPRFIKIIEVKNKKLVELLRRKVRLAQPYVGVFLKRDDSNESDVVESLDEIYHTGTFAQIHEMQDLGDKL RMIVMGHRRVHISRQLEVEPEEPEAENKHKPRRKSKRGKKEAEDELSARHPAELAMEPTPELPAEVLMVEALTAEIVKTIRDIIALNPLY RESVLQMMQAGQRVVDNPIYLSDMGAALTGAESHELQDVLEETNIPKRLYKALSLLKKEFELSKLQQRLGREVEEKIKQTHRKYLLQEQL KIIKKELGLEKDDKDAIEEKFRERLKELVVPKHVMDVVDEELSKLGLLDNHSSEFNVTRNYLDWLTSIPWGKYSNENLDLARAQAVLEED -------------------------------------------------------------- >49423_49423_4_LONP1-RAD51_LONP1_chr19_5700800_ENST00000593119_RAD51_chr15_41023253_ENST00000267868_length(transcript)=2449nt_BP=1347nt GCGAGTTGTGTGAGCCGCCAGTATGGCCGGGCTATGGCGGCGAGCACTGGCTACGTGCGACTGTGGGGAGCGGCGCGGTGCTGGGTGCTG CGGCGGCCGATGCTGGCCGCCGCCGGGGGGCGGGTTCCCACTGCAGCAGGAGCGTGGTTGCTCCGAGGCCCGGTCATAACGGCGCTCACG CCCATGACGATCCCCGATGTGTTTCCGCACCTGCCGCTCATCGCCATCACCCGCAACCCGGTGTTCCCGCGCTTTATCAAGATTATCGAG GTTAAAAATAAGAAGTTGGTTGAGCTGCTGAGAAGGAAAGTTCGTCTCGCCCAGCCTTATGTCGGCGTCTTTCTAAAGAGAGATGACAGC AATGAGTCGGATGTGGTCGAGAGCCTGGATGAAATCTACCACACGGGGACGTTTGCCCAGATCCATGAGATGCAGGACCTTGGGGACAAG CTGCGCATGATCGTCATGGGACACAGAAGAGTCCATATCAGCAGACAGCTGGAGGTGGAGCCCGAGGAGCCGGAGGCGGAGAACAAGCAC AAGCCCCGCAGGAAGTCAAAGCGGGGCAAGAAGGAGGCGGAGGACGAGCTGAGCGCCAGGCACCCGGCGGAGCTGGCGATGGAGCCCACC CCTGAGCTCCCGGCTGAGGTGCTCATGGTGGAGGTAGAGAACGTTGTCCACGAGGACTTCCAGGTCACGGAGGAGGTGAAAGCCCTGACT GCAGAGATCGTGAAGACCATCCGGGACATCATTGCCTTGAACCCTCTCTACAGGGAGTCAGTGCTGCAGATGATGCAGGCTGGCCAGCGG GTGGTGGACAACCCCATCTACCTGAGCGACATGGGCGCCGCGCTCACCGGGGCCGAGTCCCATGAGCTGCAGGACGTCCTGGAAGAGACC AATATTCCTAAGCGGCTGTACAAGGCCCTCTCCCTGCTGAAGAAGGAATTTGAACTGAGCAAGCTGCAGCAGCGCCTGGGGCGGGAGGTG GAGGAGAAGATCAAGCAGACCCACCGTAAGTACCTGCTGCAGGAGCAGCTAAAGATCATCAAGAAGGAGCTGGGCCTGGAGAAGGACGAC AAGGATGCCATCGAGGAGAAGTTCCGGGAGCGCCTGAAGGAGCTCGTGGTCCCCAAGCACGTCATGGATGTTGTGGACGAGGAGCTGAGC AAGCTGGGCCTGCTGGACAACCACTCCTCGGAGTTCAATGTCACCCGCAACTACCTAGACTGGCTCACGTCCATCCCTTGGGGCAAGTAC AGCAACGAGAACCTGGACCTGGCGCGGGCACAGGCAGTGCTGGAGGAAGACCACTACGGCATGGAGGACGTCAAGAAACGCATCCTGATT GTATCTGAGGAAAGGAAGAGGGGAAACCAGAATCTGCAAAATCTACGACTCTCCCTGTCTTCCTGAAGCTGAAGCTATGTTCGCCATTAA TGCAGATGGAGTGGGAGATGCCAAAGACTGAATCATTGGGTTTTTCCTCTGTTAAAAACCTTAAGTGCTGCAGCCTAATGAGAGTGCACT GCTCCCTGGGGTTCTCTACAGGCCTCTTCCTGTTGTGACTGCCAGGATAAAGCTTCCGGGAAAACAGCTATTATATCAGCTTTTCTGATG GTATAAACAGGAGACAGGTCAGTAGTCACAAACTGATCTAAAATGTTTATTCCTTCTGTAGTGTATTAATCTCTGTGTGTTTTCTTTGGT TTTGGAGGAGGGGTATGAAGTATCTTTGACATGGTGCCTTAGGAATGACTTGGGTTTAACAAGCTGTCTACTGGACAATCTTATGTTTCC AAGAGAACTAAAGCTGGAGAGACCTGACCCTTCTCTCACTTCTAAATTAATGGTAAAATAAAATGCCTCAGCTATGTAGCAAAGGGAATG GGTCTGCACAGATTCTTTTTTTCTGTCAGTAAAACTCTCAAGCAGGTTTTTAAGTTGTCTGTCTGAATGATCTTGTGTAAGGTTTTGGTT ATGGAGTCTTGTGCCAAACCTACTAGGCCATTAGCCCTTCACCATCTACCTGCTTGGTCTTTCATTGCTAAGACTAACTCAAGATAATCC TAGAGTCTTAAAGCATTTCAGGCCAGTGTGGTGTCTTGCGCCTGTACTCCCAGCACTTTGGGAGGCCGAGGCAGGTGGATCGCTTGAGCC CAGGAGTTTTAAGTCCAGCTTGGCCAAGGTGGTGAAATCCCATCTCTACAAAAAATGCAGAACTTAATCTGGACACACTGTTACACGTGC CTGTAGTCCCAGCTACTCGATAGCCTGAGGTGGGAGAATCACTTAAGCCTGGAAGGTGGAAGTTGCAGTGAGTCGAGATTGCACTGCTGC ATTCCAGCCAGGGTGACAGAGTGAGACCATGTTTCAAACAAGAAACATTTCAGAGGGTAAGTAAACAGATTTGATTGTGAGGCTTCTAAT >49423_49423_4_LONP1-RAD51_LONP1_chr19_5700800_ENST00000593119_RAD51_chr15_41023253_ENST00000267868_length(amino acids)=460AA_BP=438 MAASTGYVRLWGAARCWVLRRPMLAAAGGRVPTAAGAWLLRGPVITALTPMTIPDVFPHLPLIAITRNPVFPRFIKIIEVKNKKLVELLR RKVRLAQPYVGVFLKRDDSNESDVVESLDEIYHTGTFAQIHEMQDLGDKLRMIVMGHRRVHISRQLEVEPEEPEAENKHKPRRKSKRGKK EAEDELSARHPAELAMEPTPELPAEVLMVEVENVVHEDFQVTEEVKALTAEIVKTIRDIIALNPLYRESVLQMMQAGQRVVDNPIYLSDM GAALTGAESHELQDVLEETNIPKRLYKALSLLKKEFELSKLQQRLGREVEEKIKQTHRKYLLQEQLKIIKKELGLEKDDKDAIEEKFRER LKELVVPKHVMDVVDEELSKLGLLDNHSSEFNVTRNYLDWLTSIPWGKYSNENLDLARAQAVLEEDHYGMEDVKKRILIVSEERKRGNQN -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for LONP1-RAD51 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000267868 | 8 | 10 | 184_257 | 298.6666666666667 | 340.0 | PALB2 | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000382643 | 8 | 10 | 184_257 | 299.6666666666667 | 341.0 | PALB2 | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000423169 | 7 | 9 | 184_257 | 258.0 | 281.0 | PALB2 | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000532743 | 8 | 10 | 184_257 | 299.6666666666667 | 341.0 | PALB2 | |
| Tgene | RAD51 | chr19:5700800 | chr15:41023253 | ENST00000557850 | 6 | 8 | 184_257 | 201.66666666666666 | 243.0 | PALB2 |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for LONP1-RAD51 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for LONP1-RAD51 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
