
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:LPCAT3-MYO15A (FusionGDB2 ID:49507) |
Fusion Gene Summary for LPCAT3-MYO15A |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: LPCAT3-MYO15A | Fusion gene ID: 49507 | Hgene | Tgene | Gene symbol | LPCAT3 | MYO15A | Gene ID | 10162 | 51168 |
| Gene name | lysophosphatidylcholine acyltransferase 3 | myosin XVA | |
| Synonyms | C3F|LPCAT|LPLAT 5|LPSAT|MBOAT5|OACT5|nessy | DFNB3|MYO15 | |
| Cytomap | 12p13.31 | 17p11.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | lysophospholipid acyltransferase 51-acylglycerophosphocholine O-acyltransferase1-acylglycerophosphoserine O-acyltransferaseO-acyltransferase (membrane bound) domain containing 5O-acyltransferase domain-containing protein 5lyso-PC acyltransferase 3ly | unconventional myosin-XVmyosin-XVunconventional myosin-15 | |
| Modification date | 20200313 | 20200322 | |
| UniProtAcc | Q6P1A2 | Q9UKN7 | |
| Ensembl transtripts involved in fusion gene | ENST00000261407, ENST00000535021, | ENST00000412324, ENST00000418233, ENST00000451725, ENST00000585180, ENST00000205890, | |
| Fusion gene scores | * DoF score | 7 X 5 X 6=210 | 4 X 5 X 3=60 |
| # samples | 6 | 5 | |
| ** MAII score | log2(6/210*10)=-1.8073549220576 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(5/60*10)=-0.263034405833794 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: LPCAT3 [Title/Abstract] AND MYO15A [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | LPCAT3(7090166)-MYO15A(18039025), # samples:2 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
 Fusion gene breakpoints across LPCAT3 (5'-gene) Fusion gene breakpoints across LPCAT3 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
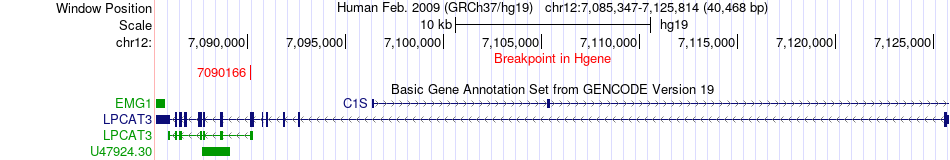 |
 Fusion gene breakpoints across MYO15A (3'-gene) Fusion gene breakpoints across MYO15A (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
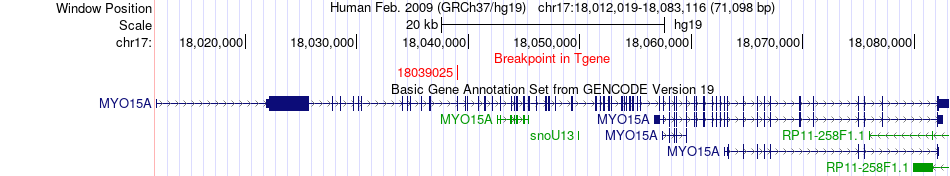 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-EW-A6SC-01A | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
Top |
Fusion Gene ORF analysis for LPCAT3-MYO15A |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000261407 | ENST00000412324 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
| 5CDS-intron | ENST00000261407 | ENST00000418233 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
| 5CDS-intron | ENST00000261407 | ENST00000451725 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
| 5CDS-intron | ENST00000261407 | ENST00000585180 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
| 5UTR-3CDS | ENST00000535021 | ENST00000205890 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
| 5UTR-intron | ENST00000535021 | ENST00000412324 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
| 5UTR-intron | ENST00000535021 | ENST00000418233 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
| 5UTR-intron | ENST00000535021 | ENST00000451725 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
| 5UTR-intron | ENST00000535021 | ENST00000585180 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
| In-frame | ENST00000261407 | ENST00000205890 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000261407 | LPCAT3 | chr12 | 7090166 | - | ENST00000205890 | MYO15A | chr17 | 18039025 | + | 7806 | 763 | 823 | 6873 | 2016 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000261407 | ENST00000205890 | LPCAT3 | chr12 | 7090166 | - | MYO15A | chr17 | 18039025 | + | 0.007740744 | 0.9922592 |
Top |
Fusion Genomic Features for LPCAT3-MYO15A |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
| LPCAT3 | chr12 | 7090165 | - | MYO15A | chr17 | 18039024 | + | 3.76E-07 | 0.99999964 |
| LPCAT3 | chr12 | 7090165 | - | MYO15A | chr17 | 18039024 | + | 3.76E-07 | 0.99999964 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
 |
Top |
Fusion Protein Features for LPCAT3-MYO15A |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr12:7090166/chr17:18039025) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| LPCAT3 | MYO15A |
| FUNCTION: Lysophospholipid O-acyltransferase (LPLAT) that catalyzes the reacylation step of the phospholipid remodeling process also known as the Lands cycle (PubMed:18782225, PubMed:18195019, PubMed:18772128). Catalyzes transfer of the fatty acyl chain from fatty acyl-CoA to 1-acyl lysophospholipid to form various classes of phospholipids. Converts 1-acyl lysophosphatidylcholine (LPC) into phosphatidylcholine (PC) (LPCAT activity), 1-acyl lysophosphatidylserine (LPS) into phosphatidylserine (PS) (LPSAT activity) and 1-acyl lysophosphatidylethanolamine (LPE) into phosphatidylethanolamine (PE) (LPEAT activity) (PubMed:18782225, PubMed:18195019, PubMed:18772128). Favors polyunsaturated fatty acyl-CoAs as acyl donors compared to saturated fatty acyl-CoAs (PubMed:18195019, PubMed:18772128). Has higher activity for LPC acyl acceptors compared to LPEs and LPSs. Can also transfer the fatty acyl chain from fatty acyl-CoA to 1-O-alkyl lysophospholipid or 1-O-alkenyl lysophospholipid with lower efficiency (By similarity). Acts as a major LPC O-acyltransferase in liver and intestine. As a component of the liver X receptor/NR1H3 or NR1H2 signaling pathway, mainly catalyzes the incorporation of arachidonate into PCs of endoplasmic reticulum (ER) membranes, increasing membrane dynamics and enabling triacylglycerols transfer to nascent very low-density lipoprotein (VLDL) particles. Promotes processing of sterol regulatory protein SREBF1 in hepatocytes, likely by facilitating the translocation of SREBF1-SCAP complex from ER to the Golgi apparatus (By similarity). Participates in mechanisms by which the liver X receptor/NR1H3 or NR1H2 signaling pathway counteracts lipid-induced ER stress response and inflammation. Downregulates hepatic inflammation by limiting arachidonic acid availability for synthesis of inflammatory eicosanoids, such as prostaglandins (By similarity). In enterocytes, acts as a component of a gut-brain feedback loop that coordinates dietary lipid absorption and food intake. Regulates the abundance of PCs containing linoleate and arachidonate in enterocyte membranes, enabling passive diffusion of fatty acids and cholesterol across the membrane for efficient chylomicron assembly (By similarity). In the intestinal crypt, acts as a component of dietary-responsive phospholipid-cholesterol axis, regulating the biosynthesis of cholesterol and its mitogenic effects on intestinal stem cells (By similarity). {ECO:0000250|UniProtKB:Q91V01, ECO:0000269|PubMed:18195019, ECO:0000269|PubMed:18772128, ECO:0000269|PubMed:18782225}. | FUNCTION: Myosins are actin-based motor molecules with ATPase activity. Unconventional myosins serve in intracellular movements. Their highly divergent tails are presumed to bind to membranous compartments, which would be moved relative to actin filaments. Required for the arrangement of stereocilia in mature hair bundles (By similarity). {ECO:0000250}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 111_131 | 225 | 651.3333333333334 | Transmembrane | Helical |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 180_200 | 225 | 651.3333333333334 | Transmembrane | Helical |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 44_64 | 225 | 651.3333333333334 | Transmembrane | Helical |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 84_104 | 225 | 651.3333333333334 | Transmembrane | Helical |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 1902_1924 | 1494 | 3531.0 | Domain | IQ 1 | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 1925_1954 | 1494 | 3531.0 | Domain | IQ 2 | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 1955_1976 | 1494 | 3531.0 | Domain | IQ 3 | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 2065_2217 | 1494 | 3531.0 | Domain | MyTH4 1 | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 2867_2953 | 1494 | 3531.0 | Domain | SH3 | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 3050_3204 | 1494 | 3531.0 | Domain | MyTH4 2 | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 3209_3530 | 1494 | 3531.0 | Domain | FERM | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 1792_1799 | 1494 | 3531.0 | Region | Actin-binding | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 1888_2029 | 1494 | 3531.0 | Region | Note=Neck or regulatory domain | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 2030_3530 | 1494 | 3531.0 | Region | Note=Tail |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 484_487 | 225 | 651.3333333333334 | Motif | Note=Di-lysine motif |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 227_247 | 225 | 651.3333333333334 | Transmembrane | Helical |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 285_305 | 225 | 651.3333333333334 | Transmembrane | Helical |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 364_384 | 225 | 651.3333333333334 | Transmembrane | Helical |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 422_442 | 225 | 651.3333333333334 | Transmembrane | Helical |
| Hgene | LPCAT3 | chr12:7090166 | chr17:18039025 | ENST00000261407 | - | 6 | 13 | 453_473 | 225 | 651.3333333333334 | Transmembrane | Helical |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 1323_1350 | 1494 | 3531.0 | Coiled coil | Ontology_term=ECO:0000255 | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 1222_1899 | 1494 | 3531.0 | Domain | Myosin motor | |
| Tgene | MYO15A | chr12:7090166 | chr17:18039025 | ENST00000205890 | 11 | 66 | 1315_1322 | 1494 | 3531.0 | Nucleotide binding | ATP |
Top |
Fusion Gene Sequence for LPCAT3-MYO15A |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >49507_49507_1_LPCAT3-MYO15A_LPCAT3_chr12_7090166_ENST00000261407_MYO15A_chr17_18039025_ENST00000205890_length(transcript)=7806nt_BP=763nt CCGGAATTGGGGGTGAAGCGATAGCGTTTTGCCCGCATTCGGGGCGCGCGGAGCTGGGGGGTCCCTGTGGGGCTCCCGGAGTTAAGATGG CGTCCTCAGCGGAGGGGGACGAGGGGACTGTGGTGGCGCTGGCGGGGGTTCTGCAGTCGGGTTTCCAGGAGCTGAGCCTTAACAAGTTGG CGACGTCCCTGGGCGCGTCAGAACAGGCGCTGCGGCTGATCATCTCCATCTTCCTGGGTTACCCCTTTGCTTTGTTTTATCGGCATTACC TTTTCTACAAGGAGACCTACCTCATCCACCTCTTCCATACCTTTACAGGCCTCTCAATTGCTTATTTTAACTTTGGAAACCAGCTCTACC ACTCCCTGCTGTGTATTGTGCTTCAGTTCCTCATCCTTCGACTAATGGGCCGCACCATCACTGCCGTCCTCACTACCTTTTGCTTCCAGA TGGCCTACCTTCTGGCTGGATACTATTACACTGCCACCGGCAACTACGATATCAAGTGGACAATGCCACATTGTGTTCTGACTTTGAAGC TGATTGGTTTGGCTGTTGACTACTTTGACGGAGGGAAAGATCAGAATTCCTTGTCCTCTGAGCAACAGAAATATGCCATACGTGGTGTTC CTTCCCTGCTGGAAGTTGCTGGTTTCTCCTACTTCTATGGGGCCTTCTTGGTAGGGCCCCAGTTCTCAATGAATCACTACATGAAGCTGG TGCAGGGAGAGCTGATTGACATACCAGGAAAGATACCAAACAGACGGATGCACAGGAGGTGGCCTCAGTGGTGAGTGCCCGAGAGATCCA GGCCGTGGCAGAGCTGCTGCAGATCTCCCCTGAGGGCCTGCAGAAGGCCATCACCTTCAAAGTGACCGAGACAATGCGAGAGAAGATCTT CACGCCCCTAACTGTGGAGAGCGCTGTGGATGCCAGGGACGCCATCGCCAAGGTCTTGTATGCACTGCTGTTCAGCTGGCTCATCACCAG GGTCAACGCGCTGGTGTCCCCAAGGCAGGACACACTGTCCATCGCCATCCTGGACATCTATGGTTTCGAGGACCTGAGCTTCAACAGCTT TGAGCAGCTGTGTATTAACTACGCAAACGAGAACCTTCAGTACCTTTTCAACAAGATCGTCTTCCAGGAGGAGCAGGAGGAGTACATCCG TGAGCAGATAGACTGGCAGGAGATCACCTTTGCTGACAACCAGCCCTGCATCAACCTCATCTCACTGAAGCCTTATGGCATCCTGCGGAT CCTTGACGACCAGTGTTGCTTTCCCCAGGCTACAGACCACACCTTCCTACAGAAGTGCCACTACCATCATGGCGCCAACCCGCTCTATTC CAAACCCAAGATGCCGCTGCCTGAGTTCACCATCAAGCACTATGCAGGCAAGGTCACCTACCAGGTGCACAAGTTCCTGGACAAGAACCA CGACCAAGTGCGCCAGGATGTGCTGGACCTGTTCGTACGGAGCCGGACACGGGTGGTGGCACACCTCTTCTCCAGCCATGCCCCACAGGC TGCCCCTCAGCGCCTGGGCAAGAGCAGCTCCGTCACTCGGCTCTACAAGGCGCACACTGTGGCCGCCAAGTTCCAGCAGTCACTCCTGGA TCTGGTGGAAAAGATGGAGAGGTGCAACCCCTTGTTCATGCGTTGCCTGAAGCCCAACCACAAGAAGGAGCCAGGTCTCTTTGAGCCAGA TGTGGTAATGGCACAATTACGCTATTCAGGGGTGCTGGAGACCGTGAGGATCCGCAAGGAGGGATTTCCAGTGCGCCTGCCTTTCCAGGG GTTCATCGACAGGTACTGCTGTCTAGTGGCCCTCAAGCATGACCTGCCGGCTAATGGGGACATGTGTGTGTCAGTGCTGAGTCGCCTGTG CAAAGTCATGCCAAACATGTACCGTGTTGGGGTCAGCAAGCTGTTCCTTAAGGAACACCTATACCAGCTGCTGGAGAGTATGCGAGAGCA TGTCCTGAATCTGGCAGCCCTCACTCTGCAGCGCTGCCTCCGTGGCTTCTTCATTAAGCGGCGATTCCGCTCTCTGCGCCACAAGATCAT CCTGCTGCAAAGCCGGGCCCGTGGCTACCTTGCCAGGCAACGCTATCAGCAGATGAGGAGGAGTCTGGTGAAGTTCCGGTCCCTGGTACA CGCATACGTGAGCCGCCGACGCTATCTCAAGCTGAGGGCAGAGTGGAGGTGCCAGGTGGAGGGGGCGCTGCTGTGGGAGCAGGAGGAGCT GAGCAAGCGGGAGGTAGTCGCTGTGGGGCACCTGGAGGTACCGGCTGAGCTGGCTGGGCTCTTGCAAGCAGTGGCAGGCCTCGGGCTGGC CCAGGTGCCTCAGGTGGCCCCTGTGAGGACTCCTCGACTCCAGGCTGAGCCCCGTGTCACACTGCCCCTGGACATCAACAACTATCCTAT GGCCAAGTTTGTCCAGTGCCACTTCAAGGAACCTGCCTTTGGGATGCTGACAGTGCCCCTGAGGACACCCCTCACGCAGCTGCCAGCCGA GCACCATGCAGAAGCCGTGAGCATCTTCAAGCTGATCCTGCGCTTCATGGGCGACCCCCACCTGCATGGTGCCCGGGAGAACATCTTCGG GAACTACATCGTGCAGAAGGGGCTGGCGGTGCCTGAGCTGCGGGATGAGATCCTGGCACAGCTGGCCAATCAGGTGTGGCACAATCACAA TGCCCACAATGCTGAGCGGGGCTGGCTGCTGCTGGCCGCCTGCCTCAGTGGCTTTGCACCTTCCCCGTGCTTCAACAAGTACCTTCTCAA GTTTGTGTCTGATTATGGGCGGAATGGCTTCCAGGCTGTGTGTCAGCACCGCCTCATGCAGGCCATGGGCCGGGCCCAACAGCAGGGCTC GGGGGCTGCCCGCACCTTACCCCCGACCCAGCTCGAGTGGACAGCGACCTATGAGAAGGCCAGCATGGCGCTGGACGTGGGCTGCTTCAA TGGTGACCAGTTCTCCTGCCCGGTGCACTCCTGGAGTACGGGGGAAGAGGTGGCTGGAGACATTCTGAGGCACAGGGGGCTGGCAGATGG CTGGCGCGGCTGGACCGTGGCCATGAAGAATGGTGTCCAGTGGGCAGAGCTGGCTGGCCACGACTACGTGTTAGACCTGGTGTCGGACCT GGAGCTGCTCAGGGACTTCCCTCGACAGAAGTCCTACTTCATTGTGGGCACAGAGGGGCCTGCAGCCAGCAGGGGAGGCCCCAAAGTGGT GTTTGGGAACAGCTGGGACTCGGATGAGGACATGTCCACTAGACCCCAGCCCCAGGAGCACATGCCCAAAGTACTTGACTCTGATGGGTA CAGCAGCCACAATCAGGACGGTACAAATGGGGAGACTGAGGCCCAAAGAGGGACAGCAACCCACCAAGAGTCAGACAGTCTTGGAGAGCC TGCTGTGCCCCACAAGGGGCTGGACTGCTACCTGGATAGCCTCTTTGACCCTGTGCTGTCCTACGGGGATGCGGACCTGGAGAAGCCAAC AGCCATTGCCTACCGCATGAAAGGGGGAGGCCAGCCCGGTGGAGGCAGCAGTAGTGGTACTGAAGACACCCCCAGGAGACCCCCAGAGCC AAAGCCAATCCCAGGCCTGGATGCCTCCACATTGGCTCTGCAGCAAGCCTTCATCCACAAACAGGCCGTGCTGCTGGCCCGGGAGATGAC CCTGCAGGCCACGGCACTCCAGCAGCAGCCCCTGAGTGCTGCCCTGAGATCCTTGCCCGCAGAGAAACCCCCAGCACCAGAGGCACAGCC GACGTCTGTAGGCACCGGTCCCCCTGCCAAACCCGTGCTCCTGCGTGCCACTCCAAAGCCCTTGGCCCCAGCCCCTCTGGCCAAGGCTCC AAGGCTCCCCATCAAGCCTGTGGCTGCCCCTGTTCTAGCTCAGGATCAGGCTTCTCCAGAAACCACTTCACCCTCCCCAGAGCTGGTCCG GTACTCTACGCTCAACTCTGAGCACTTCCCACAGCCCACACAGCAGATCAAGAATATTGTCAGGCAGTACCAGCAGCCGTTCCGGGGAGG CCGGCCTGAGGCCCTCAGGAAGGATGGCGGGAAAGTGTTCATGAAGCGGCCAGACCCTCATGAGGAGGCCCTGATGATCCTGAAAGGGCA GATGACCCACCTGGCAGCTGCACCTGGCACCCAGGTGTCCAGAGAGGCCGTGGCCCTGGTGAAGCCGGTGACCAGTGCACCAAGGCCATC CATGGCACCCACTTCAGCTCTGCCCTCGCGATCGCTGGAGCCCCCTGAGGAACTCACGCAGACGCGGCTGCACCGCCTCATCAATCCCAA CTTCTACGGCTATCAGGACGCCCCCTGGAAGATCTTCCTGCGCAAAGAGGTGTTTTACCCCAAGGACAGCTACAGCCATCCTGTGCAGCT TGACCTCCTGTTCCGGCAGATCCTGCACGACACGCTCTCCGAGGCCTGCCTTCGCATCTCTGAGGATGAGAGGCTCAGGATGAAGGCCTT GTTTGCCCAGAACCAGCTGGACACACAGAAGCCTCTGGTAACGGAAAGCGTGAAGCGGGCCGTGGTCAGCACTGCACGAGACACCTGGGA GGTCTACTTCTCCCGCATCTTCCCCGCCACGGGCAGCGTGGGCACTGGTGTGCAGCTCCTAGCTGTGTCCCACGTGGGCATCAAACTCCT GAGGATGGTCAAGGGTGGCCAGGAGGCCGGCGGGCAGCTGCGGGTCCTGCGTGCATACAGCTTTGCAGATATCCTGTTTGTGACCATGCC CTCCCAGAACATGCTGGAGTTCAACCTGGCCAGTGAGAAGGTCATCCTCTTCTCAGCCCGAGCGCACCAGGTCAAGACCCTGGTAGATGA CTTCATCTTGGAGCTGAAGAAGGACTCTGACTACGTGGTCGCTGTGAGGAACTTCCTGCCTGAGGACCCTGCGCTGCTGGCTTTCCACAA GGGTGACATCATACACCTGCAGCCCCTAGAGCCACCTCGAGTGGGCTACAGTGCTGGCTGCGTGGTTCGCAGGAAGGTGGTGTACCTGGA GGAGCTGCGACGTAGAGGCCCCGACTTTGGCTGGAGGTTCGGGACCATCCACGGGCGCGTGGGCCGCTTCCCTTCGGAGCTGGTGCAGCC CGCTGCTGCCCCCGACTTCCTGCAGCTGCCAACGGAGCCAGGCCGCGGCCGAGCAGCCGCCGTGGCCGCTGCTGTGGCCTCTGCAGCCGC TGCACAGGAGGTGGGCCGCAGGAGAGAGGGTCCCCCAGTCAGGGCCCGCTCTGCTGACCATGGGGAGGACGCCCTGGCGCTCCCACCCTA CACAATGCTCGAGTTTGCCCAGAAGTATTTCCGAGACCCTCAGAGGAGACCCCAGGATGGCCTCAGGCTGAAATCCAAGGAGCCTCGGGA GTCCAGAACCTTGGAGGACATGCTTTGCTTCACCAAGACTCCCCTCCAGGAATCCCTCATCGAACTCAGCGACAGCAGCCTCAGCAAGAT GGCCACCGACATGTTCCTAGCTGTAATGAGGTTCATGGGGGATGCCCCACTGAAGGGCCAGAGTGACCTGGACGTGCTTTGTAACCTCCT GAAGCTGTGCGGGGACCATGAGGTCATGCGGGATGAATGTTACTGCCAAGTTGTGAAGCAGATCACAGACAATACCAGCTCCAAGCAGGA CAGCTGCCAGCGAGGCTGGAGGCTGCTGTATATCGTGACCGCCTACCACAGCTGCTCTGAGGTCCTCCACCCACACCTCACTCGCTTCCT CCAAGACGTGAGCCGGACCCCAGGCCTGCCCTTTCAGGGGATCGCCAAGGCCTGCGAGCAGAACCTGCAGAAAACCTTGCGCTTCGGAGG TCGTCTGGAGCTCCCCAGCAGCATAGAGCTTCGGGCCATGTTGGCAGGCCGCAGTTCCAAGAGGCAACTCTTTCTTCTTCCTGGAGGCCT TGAACGCCATCTCAAAATCAAAACATGCACTGTGGCCCTGGACGTGGTGGAAGAGATATGTGCTGAGATGGCTCTGACACGCCCTGAGGC CTTCAATGAATATGTTATCTTCGTTGTCACCAACCGTGGCCAGCATGTGTGCCCACTCAGTCGCCGTGCTTACATCCTGGATGTGGCCTC AGAGATGGAGCAGGTGGACGGCGGCTACATGCTCTGGTTCCGGCGTGTGCTCTGGGATCAGCCACTCAAGTTCGAGAATGAGCTATATGT GACCATGCACTACAACCAGGTCCTGCCTGACTACCTGAAGGGACTCTTCAGCAGTGTGCCGGCCAGCCGGCCCAGCGAGCAGCTGCTGCA GCAGGTGTCCAAGCTGGCTTCACTGCAGCATCGCGCCAAGGACCACTTCTACCTGCCGAGCGTGCGGGAAGTCCAGGAGTACATCCCAGC CCAGCTCTACCGTACAACGGCAGGCTCGACCTGGCTCAACCTGGTCAGCCAGCACCGGCAGCAGACACAGGCGCTCAGCCCCCACCAGGC CCGTGCCCAGTTTCTGGGCCTCCTCAGCGCCTTACCTATGTTCGGCTCCTCCTTCTTCTTCATCCAGAGCTGCAGCAACATTGCTGTGCC AGCCCCTTGCATCCTTGCCATCAACCACAATGGCCTCAACTTTCTCAGCACAGAGACTCATGAATTGATGGTGAAGTTCCCCCTGAAGGA GATCCAGTCGACGCGGACCCAGCGGCCCACGGCCAACTCCAGCTACCCCTATGTGGAGATTGCGCTGGGGGACGTGGCGGCCCAGCGCAC CTTGCAGCTGCAGCTGGAGCAGGGACTGGAACTGTGTCGTGTGGTGGCCGTGCACGTGGAGAACCTGCTCAGTGCCCATGAGAAGCGGCT CACATTGCCCCCCAGCGAGATCACCCTGCTCTGACCCAGCCCCCAGCCCTCCAGTACCTTCTGCCAGAAGACTCACTGTGTGGCCTCAGA GAAATCACTGAACCTCTCAGGATCAATGACCCCTGTAAGGGGCCAGAGCCTTGGAGGACACTAAGAGGAGGCAGGAGGAGCAACTCAAAT CCCCAAGAACACAAGAAGACCCATCCTGAACTGGGATGGAATGGCAGCATGCAAACTTGGATCAGATAGCAGGAGGAACTTTCAAAAGTC TGGCCCACTGTGCAGTGGAGCAGAAGGCAGGACCATGAGGCCTCCTGCCATGTACCCATTGCAGACCCTGCCCCTAACTCCTGCCTATGA CACAGAAGCCCCACACCAGTTGCCCAGATGAACTGGCCTCTGCCTTTGGTTTACTCAGGGTCTGATGTTGGAATCTGCTCCAACTCCACA CCCTAGCCCTTACATGTCCTCCTAAGGGGCCCCTCCTTGTGCTGCCAGTCAGCCTGGATTTCTGGTCTTTGGTTATTTCTGTGCAAACAA AAGGTGTGCCTGGCAGCCATTTCTCCATGGAGTTGCTAAGTGGCCGGAAAACAAGCCTGAGGGAGGAGGCAGGAGTTGGAGTTACCTTAG GCCCCTGATTCACTGCCTATGAACAGACCATCCCCCACTCCTTGGGTATCCCCAACCCCAGACCCCCATCACTTGATGGGCCACACAAGT TTGAGAGTGGTACAAGGGAGAAGTTTGGGAAAAGCCTTCTTGGAAAATGGGACATTAGCATTGAGTTTTGAAAGATGAGTAGGAGTTTGC TAAGAATAGATGGAAGACAGCAGGATAAACATTCCAGAGAAAATCATGTTTATTCCCTGCTGTATCTTCCAGAACCTAGGAGGATGCCTA >49507_49507_1_LPCAT3-MYO15A_LPCAT3_chr12_7090166_ENST00000261407_MYO15A_chr17_18039025_ENST00000205890_length(amino acids)=2016AA_BP= MLQISPEGLQKAITFKVTETMREKIFTPLTVESAVDARDAIAKVLYALLFSWLITRVNALVSPRQDTLSIAILDIYGFEDLSFNSFEQLC INYANENLQYLFNKIVFQEEQEEYIREQIDWQEITFADNQPCINLISLKPYGILRILDDQCCFPQATDHTFLQKCHYHHGANPLYSKPKM PLPEFTIKHYAGKVTYQVHKFLDKNHDQVRQDVLDLFVRSRTRVVAHLFSSHAPQAAPQRLGKSSSVTRLYKAHTVAAKFQQSLLDLVEK MERCNPLFMRCLKPNHKKEPGLFEPDVVMAQLRYSGVLETVRIRKEGFPVRLPFQGFIDRYCCLVALKHDLPANGDMCVSVLSRLCKVMP NMYRVGVSKLFLKEHLYQLLESMREHVLNLAALTLQRCLRGFFIKRRFRSLRHKIILLQSRARGYLARQRYQQMRRSLVKFRSLVHAYVS RRRYLKLRAEWRCQVEGALLWEQEELSKREVVAVGHLEVPAELAGLLQAVAGLGLAQVPQVAPVRTPRLQAEPRVTLPLDINNYPMAKFV QCHFKEPAFGMLTVPLRTPLTQLPAEHHAEAVSIFKLILRFMGDPHLHGARENIFGNYIVQKGLAVPELRDEILAQLANQVWHNHNAHNA ERGWLLLAACLSGFAPSPCFNKYLLKFVSDYGRNGFQAVCQHRLMQAMGRAQQQGSGAARTLPPTQLEWTATYEKASMALDVGCFNGDQF SCPVHSWSTGEEVAGDILRHRGLADGWRGWTVAMKNGVQWAELAGHDYVLDLVSDLELLRDFPRQKSYFIVGTEGPAASRGGPKVVFGNS WDSDEDMSTRPQPQEHMPKVLDSDGYSSHNQDGTNGETEAQRGTATHQESDSLGEPAVPHKGLDCYLDSLFDPVLSYGDADLEKPTAIAY RMKGGGQPGGGSSSGTEDTPRRPPEPKPIPGLDASTLALQQAFIHKQAVLLAREMTLQATALQQQPLSAALRSLPAEKPPAPEAQPTSVG TGPPAKPVLLRATPKPLAPAPLAKAPRLPIKPVAAPVLAQDQASPETTSPSPELVRYSTLNSEHFPQPTQQIKNIVRQYQQPFRGGRPEA LRKDGGKVFMKRPDPHEEALMILKGQMTHLAAAPGTQVSREAVALVKPVTSAPRPSMAPTSALPSRSLEPPEELTQTRLHRLINPNFYGY QDAPWKIFLRKEVFYPKDSYSHPVQLDLLFRQILHDTLSEACLRISEDERLRMKALFAQNQLDTQKPLVTESVKRAVVSTARDTWEVYFS RIFPATGSVGTGVQLLAVSHVGIKLLRMVKGGQEAGGQLRVLRAYSFADILFVTMPSQNMLEFNLASEKVILFSARAHQVKTLVDDFILE LKKDSDYVVAVRNFLPEDPALLAFHKGDIIHLQPLEPPRVGYSAGCVVRRKVVYLEELRRRGPDFGWRFGTIHGRVGRFPSELVQPAAAP DFLQLPTEPGRGRAAAVAAAVASAAAAQEVGRRREGPPVRARSADHGEDALALPPYTMLEFAQKYFRDPQRRPQDGLRLKSKEPRESRTL EDMLCFTKTPLQESLIELSDSSLSKMATDMFLAVMRFMGDAPLKGQSDLDVLCNLLKLCGDHEVMRDECYCQVVKQITDNTSSKQDSCQR GWRLLYIVTAYHSCSEVLHPHLTRFLQDVSRTPGLPFQGIAKACEQNLQKTLRFGGRLELPSSIELRAMLAGRSSKRQLFLLPGGLERHL KIKTCTVALDVVEEICAEMALTRPEAFNEYVIFVVTNRGQHVCPLSRRAYILDVASEMEQVDGGYMLWFRRVLWDQPLKFENELYVTMHY NQVLPDYLKGLFSSVPASRPSEQLLQQVSKLASLQHRAKDHFYLPSVREVQEYIPAQLYRTTAGSTWLNLVSQHRQQTQALSPHQARAQF LGLLSALPMFGSSFFFIQSCSNIAVPAPCILAINHNGLNFLSTETHELMVKFPLKEIQSTRTQRPTANSSYPYVEIALGDVAAQRTLQLQ -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for LPCAT3-MYO15A |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for LPCAT3-MYO15A |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for LPCAT3-MYO15A |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
