
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MAP4K2-FXR2 (FusionGDB2 ID:51387) |
Fusion Gene Summary for MAP4K2-FXR2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MAP4K2-FXR2 | Fusion gene ID: 51387 | Hgene | Tgene | Gene symbol | MAP4K2 | FXR2 | Gene ID | 5871 | 9513 |
| Gene name | mitogen-activated protein kinase kinase kinase kinase 2 | FMR1 autosomal homolog 2 | |
| Synonyms | BL44|GCK|RAB8IP | FMR1L2|FXR2P | |
| Cytomap | 11q13.1 | 17p13.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | mitogen-activated protein kinase kinase kinase kinase 2B lymphocyte serine/threonine protein kinaseMAPK/ERK kinase kinase kinase 2MEK kinase kinase 2MEKKK 2Rab8 interacting proteingerminal centre kinase (GC kinase) | fragile X mental retardation syndrome-related protein 2fragile X mental retardation, autosomal homolog 2fragile X-mental retardation 1-like 2 | |
| Modification date | 20200313 | 20200327 | |
| UniProtAcc | Q12851 | P51116 | |
| Ensembl transtripts involved in fusion gene | ENST00000294066, ENST00000377350, ENST00000468062, | ENST00000573057, ENST00000250113, | |
| Fusion gene scores | * DoF score | 3 X 4 X 3=36 | 10 X 9 X 6=540 |
| # samples | 3 | 11 | |
| ** MAII score | log2(3/36*10)=-0.263034405833794 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(11/540*10)=-2.29545588352617 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MAP4K2 [Title/Abstract] AND FXR2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MAP4K2(64563744)-FXR2(7506320), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | MAP4K2 | GO:0006468 | protein phosphorylation | 11784851 |
| Hgene | MAP4K2 | GO:0007257 | activation of JUN kinase activity | 17584736 |
| Hgene | MAP4K2 | GO:0035556 | intracellular signal transduction | 11784851 |
| Hgene | MAP4K2 | GO:0046330 | positive regulation of JNK cascade | 17584736 |
 Fusion gene breakpoints across MAP4K2 (5'-gene) Fusion gene breakpoints across MAP4K2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
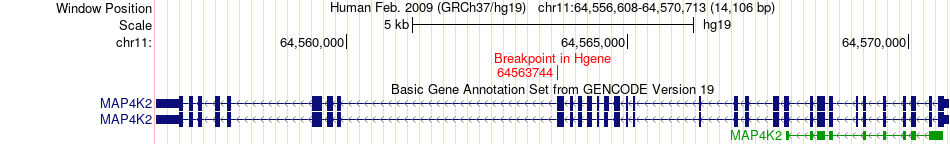 |
 Fusion gene breakpoints across FXR2 (3'-gene) Fusion gene breakpoints across FXR2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
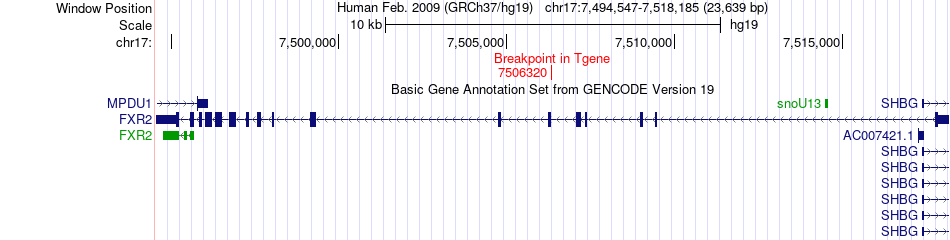 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | OV | TCGA-24-2033 | MAP4K2 | chr11 | 64563744 | - | FXR2 | chr17 | 7506320 | - |
Top |
Fusion Gene ORF analysis for MAP4K2-FXR2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000294066 | ENST00000573057 | MAP4K2 | chr11 | 64563744 | - | FXR2 | chr17 | 7506320 | - |
| 5CDS-intron | ENST00000377350 | ENST00000573057 | MAP4K2 | chr11 | 64563744 | - | FXR2 | chr17 | 7506320 | - |
| In-frame | ENST00000294066 | ENST00000250113 | MAP4K2 | chr11 | 64563744 | - | FXR2 | chr17 | 7506320 | - |
| In-frame | ENST00000377350 | ENST00000250113 | MAP4K2 | chr11 | 64563744 | - | FXR2 | chr17 | 7506320 | - |
| intron-3CDS | ENST00000468062 | ENST00000250113 | MAP4K2 | chr11 | 64563744 | - | FXR2 | chr17 | 7506320 | - |
| intron-intron | ENST00000468062 | ENST00000573057 | MAP4K2 | chr11 | 64563744 | - | FXR2 | chr17 | 7506320 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000377350 | MAP4K2 | chr11 | 64563744 | - | ENST00000250113 | FXR2 | chr17 | 7506320 | - | 3992 | 1819 | 92 | 3391 | 1099 |
| ENST00000294066 | MAP4K2 | chr11 | 64563744 | - | ENST00000250113 | FXR2 | chr17 | 7506320 | - | 4016 | 1843 | 92 | 3415 | 1107 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000377350 | ENST00000250113 | MAP4K2 | chr11 | 64563744 | - | FXR2 | chr17 | 7506320 | - | 0.004174853 | 0.9958252 |
| ENST00000294066 | ENST00000250113 | MAP4K2 | chr11 | 64563744 | - | FXR2 | chr17 | 7506320 | - | 0.00312591 | 0.9968741 |
Top |
Fusion Genomic Features for MAP4K2-FXR2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
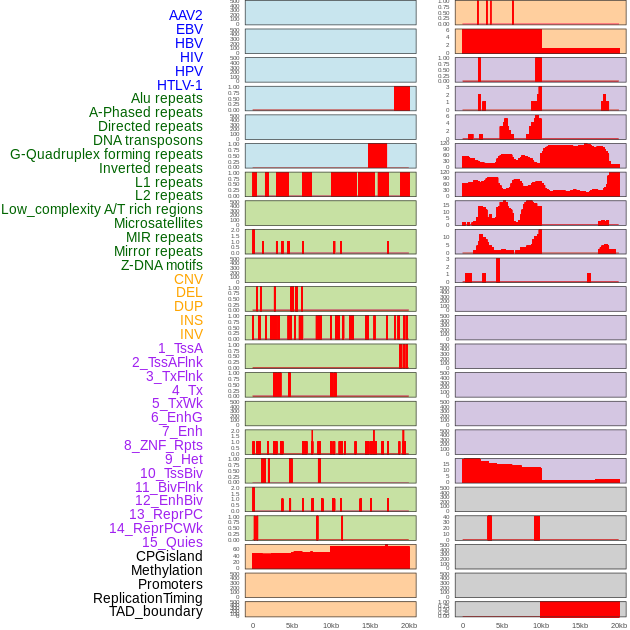 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for MAP4K2-FXR2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr11:64563744/chr17:7506320) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MAP4K2 | FXR2 |
| FUNCTION: Serine/threonine-protein kinase which acts as an essential component of the MAP kinase signal transduction pathway. Acts as a MAPK kinase kinase kinase (MAP4K) and is an upstream activator of the stress-activated protein kinase/c-Jun N-terminal kinase (SAP/JNK) signaling pathway and to a lesser extent of the p38 MAPKs signaling pathway. Required for the efficient activation of JNKs by TRAF6-dependent stimuli, including pathogen-associated molecular patterns (PAMPs) such as polyinosine-polycytidine (poly(IC)), lipopolysaccharides (LPS), lipid A, peptidoglycan (PGN), or bacterial flagellin. To a lesser degree, IL-1 and engagement of CD40 also stimulate MAP4K2-mediated JNKs activation. The requirement for MAP4K2/GCK is most pronounced for LPS signaling, and extends to LPS stimulation of c-Jun phosphorylation and induction of IL-8. Enhances MAP3K1 oligomerization, which may relieve N-terminal mediated MAP3K1 autoinhibition and lead to activation following autophosphorylation. Mediates also the SAP/JNK signaling pathway and the p38 MAPKs signaling pathway through activation of the MAP3Ks MAP3K10/MLK2 and MAP3K11/MLK3. May play a role in the regulation of vesicle targeting or fusion. regulation of vesicle targeting or fusion. {ECO:0000269|PubMed:11784851, ECO:0000269|PubMed:15456887, ECO:0000269|PubMed:17584736, ECO:0000269|PubMed:7477268, ECO:0000269|PubMed:7515885, ECO:0000269|PubMed:9712898}. | FUNCTION: RNA-binding protein. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MAP4K2 | chr11:64563744 | chr17:7506320 | ENST00000294066 | - | 24 | 32 | 16_273 | 583 | 821.0 | Domain | Protein kinase |
| Hgene | MAP4K2 | chr11:64563744 | chr17:7506320 | ENST00000294066 | - | 24 | 32 | 22_30 | 583 | 821.0 | Nucleotide binding | ATP |
| Hgene | MAP4K2 | chr11:64563744 | chr17:7506320 | ENST00000294066 | - | 24 | 32 | 294_314 | 583 | 821.0 | Region | Note=PEST1 |
| Hgene | MAP4K2 | chr11:64563744 | chr17:7506320 | ENST00000294066 | - | 24 | 32 | 344_360 | 583 | 821.0 | Region | Note=PEST2 |
| Hgene | MAP4K2 | chr11:64563744 | chr17:7506320 | ENST00000294066 | - | 24 | 32 | 405_448 | 583 | 821.0 | Region | Note=PEST3 |
| Tgene | FXR2 | chr11:64563744 | chr17:7506320 | ENST00000250113 | 4 | 17 | 414_418 | 149 | 674.0 | Compositional bias | Note=Poly-Ser | |
| Tgene | FXR2 | chr11:64563744 | chr17:7506320 | ENST00000250113 | 4 | 17 | 544_552 | 149 | 674.0 | Compositional bias | Note=Poly-Arg | |
| Tgene | FXR2 | chr11:64563744 | chr17:7506320 | ENST00000250113 | 4 | 17 | 584_594 | 149 | 674.0 | Compositional bias | Note=Poly-Arg | |
| Tgene | FXR2 | chr11:64563744 | chr17:7506320 | ENST00000250113 | 4 | 17 | 232_261 | 149 | 674.0 | Domain | KH 1 | |
| Tgene | FXR2 | chr11:64563744 | chr17:7506320 | ENST00000250113 | 4 | 17 | 295_324 | 149 | 674.0 | Domain | KH 2 |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MAP4K2 | chr11:64563744 | chr17:7506320 | ENST00000294066 | - | 24 | 32 | 482_793 | 583 | 821.0 | Domain | CNH |
| Tgene | FXR2 | chr11:64563744 | chr17:7506320 | ENST00000250113 | 4 | 17 | 14_60 | 149 | 674.0 | Domain | Agenet-like 1 | |
| Tgene | FXR2 | chr11:64563744 | chr17:7506320 | ENST00000250113 | 4 | 17 | 73_125 | 149 | 674.0 | Domain | Agenet-like 2 |
Top |
Fusion Gene Sequence for MAP4K2-FXR2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >51387_51387_1_MAP4K2-FXR2_MAP4K2_chr11_64563744_ENST00000294066_FXR2_chr17_7506320_ENST00000250113_length(transcript)=4016nt_BP=1843nt CAGAGCCACGGGCGCCCGCCCCGCCCCGCGCCGCCCCGCGCCGGCTCCGCAGCTCGCGCCCGCCCGCCTGCCGGCCCGCCCGGCGCCGGG CCATGGCGCTGCTGCGGGATGTGTCGCTGCAGGACCCGCGGGACCGCTTCGAGCTGCTGCAGCGCGTGGGGGCCGGGACCTATGGCGACG TCTACAAGGCCCGCGACACGGTCACGTCCGAACTGGCCGCCGTGAAGATAGTCAAGCTAGACCCAGGGGACGACATCAGCTCCCTCCAGC AGGAAATCACCATCCTGCGTGAGTGCCGCCACCCCAATGTGGTGGCCTACATTGGCAGCTACCTCAGGAATGACCGCTTGTGGATCTGCA TGGAGTTCTGCGGAGGGGGCTCCCTGCAGGAGATTTACCATGCCACTGGGCCCCTGGAGGAGCGGCAGATTGCCTACGTCTGCCGAGAGG CACTGAAGGGGCTCCACCACCTGCATTCTCAGGGGAAGATCCACAGAGACATCAAGGGAGCCAACCTTCTCCTCACTCTCCAGGGAGATG TCAAACTGGCTGACTTTGGGGTGTCAGGCGAGCTGACAGCGTCTGTGGCCAAGAGGAGGTCTTTCATTGGGACTCCCTACTGGATGGCTC CCGAGGTGGCTGCTGTGGAGCGCAAAGGTGGCTACAATGAGCTATGTGACGTCTGGGCCCTGGGCATCACTGCCATTGAGCTGGGCGAGC TGCAGCCCCCTCTGTTCCACCTGCACCCCATGAGGGCCCTGATGCTCATGTCGAAGAGCAGCTTCCAGCCGCCCAAACTGAGAGATAAGA CTCGCTGGACCCAGAATTTCCACCACTTTCTCAAACTGGCCCTGACCAAGAATCCTAAGAAGAGGCCGACAGCAGAGAAGCTCCTGCAGC ACCCGTTCACGACTCAGCAGCTCCCTCGGGCCCTCCTCACACAGCTGCTGGACAAAGCCAGTGACCCTCATCTGGGGACCCCCTCCCCTG AGGACTGTGAGCTGGAGACCTATGACATGTTTCCAGACACCATTCACTCCCGGGGGCAGCACGGCCCAGCCGAGAGGACCCCCTCGGAGA TCCAGTTTCACCAGGTGAAATTTGGCGCCCCACGCAGGAAGGAAACTGACCCACTGAATGAGCCGTGGGAGGAAGAGTGGACACTACTGG GAAAGGAAGAGTTGAGTGGGAGCCTGCTGCAGTCGGTCCAGGAGGCCCTGGAGGAAAGGAGTCTGACTATTCGGTCAGCCTCAGAATTCC AGGAGCTGGACTCCCCAGACGATACCATGGGAACCATCAAGCGGGCCCCGTTCCTAGGGCCACTCCCCACTGACCCTCCAGCAGAGGAGC CTCTGTCCAGTCCCCCAGGAACCCTGCCCCCACCTCCTTCAGGCCCCAACAGCTCCCCACTGCTGCCCACGGCCTGGGCCACCATGAAGC AGCGGGAGGATCCTGAGAGGTCATCCTGCCACGGGCTCCCCCCAACTCCCAAGGTGCATATGGGCGCCTGCTTCTCCAAGGTCTTCAATG GCTGCCCCCTGCGGATCCACGCTGCTGTCACCTGGATTCACCCTGTTACTCGGGACCAGTTCCTGGTGGTAGGGGCCGAGGAAGGCATCT ACACACTCAACCTGCATGAACTGCATGAGGATACGCTGGAGAAGCTGATTTCACATCGCTGCTCCTGGCTCTACTGCGTGAACAACGTGC TGCTGTCACTCTCAGGGAAATCCACGCACATCTGGGCCCATGACCTCCCAGGCCTGTTTGAGCAGCGGAGGCTACAGCAACAGGTTCCCC TCTCCATCCCCACCAACCGCCTCACCCAGCGCATCATCCCCAGCTGCTCCAATGAAAACGTCCATAAAGAGTTCAAGAAAGCCCTGGGAG CCAACTGCATCTTTCTCAACATCACAAACAGTGAGCTCTTCATTCTGTCAACCACAGAAGCCCCTGTGAAGCGAGCATCTCTGCTGGGTG ATATGCATTTCCGAAGCCTGCGCACCAAACTGCTACTTATGTCCCGCAATGAAGAAGCTACCAAGCACCTAGAGACAAGCAAGCAGTTGG CAGCAGCCTTCCAAGAGGAGTTCACAGTGCGAGAGGACCTGATGGGACTGGCAATTGGGACTCACGGTGCCAACATCCAGCAGGCCCGAA AAGTACCTGGGGTGACCGCCATTGAGTTGGGTGAAGAGACCTGCACTTTCCGCATCTATGGGGAGACTCCCGAGGCTTGCCGACAGGCCC GAAGCTACCTTGAGTTTTCTGAGGACTCAGTGCAAGTGCCCAGGAACCTGGTTGGCAAAGTGATTGGAAAGAACGGGAAAGTGATCCAGG AGATTGTGGATAAATCTGGTGTGGTGAGGGTTCGAGTGGAAGGTGATAATGACAAGAAGAACCCCAGGGAGGAGGGAATGGTTCCCTTCA TTTTTGTTGGCACCCGAGAGAACATCAGCAATGCCCAGGCTTTGCTGGAGTATCACCTCTCCTACCTGCAGGAGGTAGAGCAGCTTCGCT TGGAGAGGCTACAAATTGATGAGCAGCTTCGGCAGATTGGGCTGGGCTTTCGCCCTCCTGGGAGTGGGCGGGGCAGCGGTGGCAGCGACA AGGCTGGATATAGCACTGATGAGAGCTCCTCCTCCTCCCTCCATGCGACTCGAACCTATGGGGGCAGCTATGGGGGCCGTGGCCGTGGCC GGAGGACAGGCGGTCCTGCCTATGGCCCCAGCTCAGATGTGTCTACAGCTTCAGAGACTGAGTCAGAGAAGAGAGAGGAGCCCAACCGAG CTGGGCCTGGCGACAGGGATCCCCCAACCCGAGGGGAAGAAAGCCGGAGGCGGCCGACTGGGGGCCGGGGTAGGGGACCCCCACCTGCCC CCCGGCCCACTTCGAGATACAATTCTTCATCTATTAGCTCAGTGCTGAAGGATCCAGACAGTAATCCCTACAGCCTATTGGACACGTCTG AACCAGAGCCCCCGGTTGATTCAGAACCTGGGGAACCCCCCCCAGCAAGTGCCAGGCGCCGCCGCTCCCGCCGCCGCCGCACTGATGAAG ACAGGACCGTCATGGATGGAGGCCTGGAATCAGATGGGCCCAACATGACAGAGAATGGCCTGGAAGATGAATCAAGACCTCAACGTCGTA ATCGCAGCCGCCGCCGCCGTAACCGTGGTAATCGGACTGATGGCTCTATCAGTGGAGACCGCCAGCCAGTGACTGTGGCTGACTATATCT CACGAGCAGAGTCTCAGAGCCGCCAGAGGCCACCCCTGGAACGCACTAAACCCTCAGAAGACTCTCTTTCAGGACAGAAGGGTGACTCTG TCAGCAAGCTTCCTAAGGGCCCCTCGGAGAATGGGGAGCTCTCCGCCCCCTTGGAGTTGGGTAGTATGGTGAATGGGGTTTCATAAAACC TCCAACCTGCACCCCTCCCTTCTCCATCTCGCTTGCTGCCCAACACCATGGCCCTCACAGGCCCAACTGACCTGCGCTGGAGCTGCTCTT ATCTAGGGGGGAGGGGGGTGGCACAGCAGCTTGGGTACCCCCCAACCTCCAGGAGCTAGTGGAGGGGTGTGTAACAGGGTCATACCCCCT CCCTCTTGTCCACCCTACCCCCAGGGTAAGGGGAGCCTCTCTCCTTCCCCATCAGACTGGATGTGCCTTTATCCTCTAATGCCCCAATCT CTCTCTGAACACCCCCATTCTCCACCTGTTGGTGGGGGGTGCTCCTCGACCCACCCAGATTTGACCGTTCAGGGGGCCTCCCCTGCTATC CCTCCTCCCATCCTGTACCCCCCATTTCTGGGGCCTCATCACTGTGGAAGACGGGGATAGTAAGAGATAAGTGGGTGGGAGGCACGGGGA AGGTTTTGGAGTAGAACCAGGGGTGTGTATGAAGGGGGGTGACAAGGTCCCCCTGGGGAGGGGACCAACCTTGTCTGGTGGATGAGAAGG >51387_51387_1_MAP4K2-FXR2_MAP4K2_chr11_64563744_ENST00000294066_FXR2_chr17_7506320_ENST00000250113_length(amino acids)=1107AA_BP=583 MALLRDVSLQDPRDRFELLQRVGAGTYGDVYKARDTVTSELAAVKIVKLDPGDDISSLQQEITILRECRHPNVVAYIGSYLRNDRLWICM EFCGGGSLQEIYHATGPLEERQIAYVCREALKGLHHLHSQGKIHRDIKGANLLLTLQGDVKLADFGVSGELTASVAKRRSFIGTPYWMAP EVAAVERKGGYNELCDVWALGITAIELGELQPPLFHLHPMRALMLMSKSSFQPPKLRDKTRWTQNFHHFLKLALTKNPKKRPTAEKLLQH PFTTQQLPRALLTQLLDKASDPHLGTPSPEDCELETYDMFPDTIHSRGQHGPAERTPSEIQFHQVKFGAPRRKETDPLNEPWEEEWTLLG KEELSGSLLQSVQEALEERSLTIRSASEFQELDSPDDTMGTIKRAPFLGPLPTDPPAEEPLSSPPGTLPPPPSGPNSSPLLPTAWATMKQ REDPERSSCHGLPPTPKVHMGACFSKVFNGCPLRIHAAVTWIHPVTRDQFLVVGAEEGIYTLNLHELHEDTLEKLISHRCSWLYCVNNVL LSLSGKSTHIWAHDLPGLFEQRRLQQQVPLSIPTNRLTQRIIPSCSNENVHKEFKKALGANCIFLNITNSELFILSTTEAPVKRASLLGD MHFRSLRTKLLLMSRNEEATKHLETSKQLAAAFQEEFTVREDLMGLAIGTHGANIQQARKVPGVTAIELGEETCTFRIYGETPEACRQAR SYLEFSEDSVQVPRNLVGKVIGKNGKVIQEIVDKSGVVRVRVEGDNDKKNPREEGMVPFIFVGTRENISNAQALLEYHLSYLQEVEQLRL ERLQIDEQLRQIGLGFRPPGSGRGSGGSDKAGYSTDESSSSSLHATRTYGGSYGGRGRGRRTGGPAYGPSSDVSTASETESEKREEPNRA GPGDRDPPTRGEESRRRPTGGRGRGPPPAPRPTSRYNSSSISSVLKDPDSNPYSLLDTSEPEPPVDSEPGEPPPASARRRRSRRRRTDED RTVMDGGLESDGPNMTENGLEDESRPQRRNRSRRRRNRGNRTDGSISGDRQPVTVADYISRAESQSRQRPPLERTKPSEDSLSGQKGDSV -------------------------------------------------------------- >51387_51387_2_MAP4K2-FXR2_MAP4K2_chr11_64563744_ENST00000377350_FXR2_chr17_7506320_ENST00000250113_length(transcript)=3992nt_BP=1819nt CAGAGCCACGGGCGCCCGCCCCGCCCCGCGCCGCCCCGCGCCGGCTCCGCAGCTCGCGCCCGCCCGCCTGCCGGCCCGCCCGGCGCCGGG CCATGGCGCTGCTGCGGGATGTGTCGCTGCAGGACCCGCGGGACCGCTTCGAGCTGCTGCAGCGCGTGGGGGCCGGGACCTATGGCGACG TCTACAAGGCCCGCGACACGGTCACGTCCGAACTGGCCGCCGTGAAGATAGTCAAGCTAGACCCAGGGGACGACATCAGCTCCCTCCAGC AGGAAATCACCATCCTGCGTGAGTGCCGCCACCCCAATGTGGTGGCCTACATTGGCAGCTACCTCAGGAATGACCGCTTGTGGATCTGCA TGGAGTTCTGCGGAGGGGGCTCCCTGCAGGAGATTTACCATGCCACTGGGCCCCTGGAGGAGCGGCAGATTGCCTACGTCTGCCGAGAGG CACTGAAGGGGCTCCACCACCTGCATTCTCAGGGGAAGATCCACAGAGACATCAAGGGAGCCAACCTTCTCCTCACTCTCCAGGGAGATG TCAAACTGGCTGACTTTGGGGTGTCAGGCGAGCTGACAGCGTCTGTGGCCAAGAGGAGGTCTTTCATTGGGACTCCCTACTGGATGGCTC CCGAGGTGGCTGCTGTGGAGCGCAAAGGTGGCTACAATGAGCTATGTGACGTCTGGGCCCTGGGCATCACTGCCATTGAGCTGGGCGAGC TGCAGCCCCCTCTGTTCCACCTGCACCCCATGAGGGCCCTGATGCTCATGTCGAAGAGCAGCTTCCAGCCGCCCAAACTGAGAGATAAGA CTCGCTGGACCCAGAATTTCCACCACTTTCTCAAACTGGCCCTGACCAAGAATCCTAAGAAGAGGCCGACAGCAGAGAAGCTCCTGCAGC ACCCGTTCACGACTCAGCAGCTCCCTCGGGCCCTCCTCACACAGCTGCTGGACAAAGCCAGTGACCCTCATCTGGGGACCCCCTCCCCTG AGGACTGTGAGCTGGAGACCTATGACATGTTTCCAGACACCATTCACTCCCGGGGGCAGCACGGCCCAGCCGAGAGGACCCCCTCGGAGA TCCAGTTTCACCAGGTGAAATTTGGCGCCCCACGCAGGAAGGAAACTGACCCACTGAATGAGCCGTGGGAGGAAGAGTGGACACTACTGG GAAAGGAAGAGTTGAGTGGGAGCCTGCTGCAGTCGGTCCAGGAGGCCCTGGAGGAAAGGAGTCTGACTATTCGGTCAGCCTCAGAATTCC AGGAGCTGGACTCCCCAGACGATACCATGGGAACCATCAAGCGGGCCCCGTTCCTAGGGCCACTCCCCACTGACCCTCCAGCAGAGGAGC CTCTGTCCAGTCCCCCAGGCCCCAACAGCTCCCCACTGCTGCCCACGGCCTGGGCCACCATGAAGCAGCGGGAGGATCCTGAGAGGTCAT CCTGCCACGGGCTCCCCCCAACTCCCAAGGTGCATATGGGCGCCTGCTTCTCCAAGGTCTTCAATGGCTGCCCCCTGCGGATCCACGCTG CTGTCACCTGGATTCACCCTGTTACTCGGGACCAGTTCCTGGTGGTAGGGGCCGAGGAAGGCATCTACACACTCAACCTGCATGAACTGC ATGAGGATACGCTGGAGAAGCTGATTTCACATCGCTGCTCCTGGCTCTACTGCGTGAACAACGTGCTGCTGTCACTCTCAGGGAAATCCA CGCACATCTGGGCCCATGACCTCCCAGGCCTGTTTGAGCAGCGGAGGCTACAGCAACAGGTTCCCCTCTCCATCCCCACCAACCGCCTCA CCCAGCGCATCATCCCCAGCTGCTCCAATGAAAACGTCCATAAAGAGTTCAAGAAAGCCCTGGGAGCCAACTGCATCTTTCTCAACATCA CAAACAGTGAGCTCTTCATTCTGTCAACCACAGAAGCCCCTGTGAAGCGAGCATCTCTGCTGGGTGATATGCATTTCCGAAGCCTGCGCA CCAAACTGCTACTTATGTCCCGCAATGAAGAAGCTACCAAGCACCTAGAGACAAGCAAGCAGTTGGCAGCAGCCTTCCAAGAGGAGTTCA CAGTGCGAGAGGACCTGATGGGACTGGCAATTGGGACTCACGGTGCCAACATCCAGCAGGCCCGAAAAGTACCTGGGGTGACCGCCATTG AGTTGGGTGAAGAGACCTGCACTTTCCGCATCTATGGGGAGACTCCCGAGGCTTGCCGACAGGCCCGAAGCTACCTTGAGTTTTCTGAGG ACTCAGTGCAAGTGCCCAGGAACCTGGTTGGCAAAGTGATTGGAAAGAACGGGAAAGTGATCCAGGAGATTGTGGATAAATCTGGTGTGG TGAGGGTTCGAGTGGAAGGTGATAATGACAAGAAGAACCCCAGGGAGGAGGGAATGGTTCCCTTCATTTTTGTTGGCACCCGAGAGAACA TCAGCAATGCCCAGGCTTTGCTGGAGTATCACCTCTCCTACCTGCAGGAGGTAGAGCAGCTTCGCTTGGAGAGGCTACAAATTGATGAGC AGCTTCGGCAGATTGGGCTGGGCTTTCGCCCTCCTGGGAGTGGGCGGGGCAGCGGTGGCAGCGACAAGGCTGGATATAGCACTGATGAGA GCTCCTCCTCCTCCCTCCATGCGACTCGAACCTATGGGGGCAGCTATGGGGGCCGTGGCCGTGGCCGGAGGACAGGCGGTCCTGCCTATG GCCCCAGCTCAGATGTGTCTACAGCTTCAGAGACTGAGTCAGAGAAGAGAGAGGAGCCCAACCGAGCTGGGCCTGGCGACAGGGATCCCC CAACCCGAGGGGAAGAAAGCCGGAGGCGGCCGACTGGGGGCCGGGGTAGGGGACCCCCACCTGCCCCCCGGCCCACTTCGAGATACAATT CTTCATCTATTAGCTCAGTGCTGAAGGATCCAGACAGTAATCCCTACAGCCTATTGGACACGTCTGAACCAGAGCCCCCGGTTGATTCAG AACCTGGGGAACCCCCCCCAGCAAGTGCCAGGCGCCGCCGCTCCCGCCGCCGCCGCACTGATGAAGACAGGACCGTCATGGATGGAGGCC TGGAATCAGATGGGCCCAACATGACAGAGAATGGCCTGGAAGATGAATCAAGACCTCAACGTCGTAATCGCAGCCGCCGCCGCCGTAACC GTGGTAATCGGACTGATGGCTCTATCAGTGGAGACCGCCAGCCAGTGACTGTGGCTGACTATATCTCACGAGCAGAGTCTCAGAGCCGCC AGAGGCCACCCCTGGAACGCACTAAACCCTCAGAAGACTCTCTTTCAGGACAGAAGGGTGACTCTGTCAGCAAGCTTCCTAAGGGCCCCT CGGAGAATGGGGAGCTCTCCGCCCCCTTGGAGTTGGGTAGTATGGTGAATGGGGTTTCATAAAACCTCCAACCTGCACCCCTCCCTTCTC CATCTCGCTTGCTGCCCAACACCATGGCCCTCACAGGCCCAACTGACCTGCGCTGGAGCTGCTCTTATCTAGGGGGGAGGGGGGTGGCAC AGCAGCTTGGGTACCCCCCAACCTCCAGGAGCTAGTGGAGGGGTGTGTAACAGGGTCATACCCCCTCCCTCTTGTCCACCCTACCCCCAG GGTAAGGGGAGCCTCTCTCCTTCCCCATCAGACTGGATGTGCCTTTATCCTCTAATGCCCCAATCTCTCTCTGAACACCCCCATTCTCCA CCTGTTGGTGGGGGGTGCTCCTCGACCCACCCAGATTTGACCGTTCAGGGGGCCTCCCCTGCTATCCCTCCTCCCATCCTGTACCCCCCA TTTCTGGGGCCTCATCACTGTGGAAGACGGGGATAGTAAGAGATAAGTGGGTGGGAGGCACGGGGAAGGTTTTGGAGTAGAACCAGGGGT GTGTATGAAGGGGGGTGACAAGGTCCCCCTGGGGAGGGGACCAACCTTGTCTGGTGGATGAGAAGGCGTATTTATTTTTCACTGTACAGT >51387_51387_2_MAP4K2-FXR2_MAP4K2_chr11_64563744_ENST00000377350_FXR2_chr17_7506320_ENST00000250113_length(amino acids)=1099AA_BP=575 MALLRDVSLQDPRDRFELLQRVGAGTYGDVYKARDTVTSELAAVKIVKLDPGDDISSLQQEITILRECRHPNVVAYIGSYLRNDRLWICM EFCGGGSLQEIYHATGPLEERQIAYVCREALKGLHHLHSQGKIHRDIKGANLLLTLQGDVKLADFGVSGELTASVAKRRSFIGTPYWMAP EVAAVERKGGYNELCDVWALGITAIELGELQPPLFHLHPMRALMLMSKSSFQPPKLRDKTRWTQNFHHFLKLALTKNPKKRPTAEKLLQH PFTTQQLPRALLTQLLDKASDPHLGTPSPEDCELETYDMFPDTIHSRGQHGPAERTPSEIQFHQVKFGAPRRKETDPLNEPWEEEWTLLG KEELSGSLLQSVQEALEERSLTIRSASEFQELDSPDDTMGTIKRAPFLGPLPTDPPAEEPLSSPPGPNSSPLLPTAWATMKQREDPERSS CHGLPPTPKVHMGACFSKVFNGCPLRIHAAVTWIHPVTRDQFLVVGAEEGIYTLNLHELHEDTLEKLISHRCSWLYCVNNVLLSLSGKST HIWAHDLPGLFEQRRLQQQVPLSIPTNRLTQRIIPSCSNENVHKEFKKALGANCIFLNITNSELFILSTTEAPVKRASLLGDMHFRSLRT KLLLMSRNEEATKHLETSKQLAAAFQEEFTVREDLMGLAIGTHGANIQQARKVPGVTAIELGEETCTFRIYGETPEACRQARSYLEFSED SVQVPRNLVGKVIGKNGKVIQEIVDKSGVVRVRVEGDNDKKNPREEGMVPFIFVGTRENISNAQALLEYHLSYLQEVEQLRLERLQIDEQ LRQIGLGFRPPGSGRGSGGSDKAGYSTDESSSSSLHATRTYGGSYGGRGRGRRTGGPAYGPSSDVSTASETESEKREEPNRAGPGDRDPP TRGEESRRRPTGGRGRGPPPAPRPTSRYNSSSISSVLKDPDSNPYSLLDTSEPEPPVDSEPGEPPPASARRRRSRRRRTDEDRTVMDGGL ESDGPNMTENGLEDESRPQRRNRSRRRRNRGNRTDGSISGDRQPVTVADYISRAESQSRQRPPLERTKPSEDSLSGQKGDSVSKLPKGPS -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MAP4K2-FXR2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MAP4K2-FXR2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MAP4K2-FXR2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
