
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MARK2-BATF2 (FusionGDB2 ID:51741) |
Fusion Gene Summary for MARK2-BATF2 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MARK2-BATF2 | Fusion gene ID: 51741 | Hgene | Tgene | Gene symbol | MARK2 | BATF2 | Gene ID | 2011 | 116071 |
| Gene name | microtubule affinity regulating kinase 2 | basic leucine zipper ATF-like transcription factor 2 | |
| Synonyms | EMK-1|EMK1|PAR-1|Par-1b|Par1b | SARI | |
| Cytomap | 11q13.1 | 11q13.1 | |
| Type of gene | protein-coding | protein-coding | |
| Description | serine/threonine-protein kinase MARK2ELKL motif kinase 1MAP/microtubule affinity-regulating kinase 2PAR1 homolog bSer/Thr protein kinase PAR-1Bserine/threonine protein kinase EMKtesticular tissue protein Li 117 | basic leucine zipper transcriptional factor ATF-like 2B-ATF-2basic leucine zipper transcription factor, ATF-like 2suppressor of AP-1 regulated by IFN | |
| Modification date | 20200329 | 20200313 | |
| UniProtAcc | Q7KZI7 | Q8N1L9 | |
| Ensembl transtripts involved in fusion gene | ENST00000315032, ENST00000350490, ENST00000361128, ENST00000377809, ENST00000402010, ENST00000413835, ENST00000502399, ENST00000508192, ENST00000377810, ENST00000408948, ENST00000425897, ENST00000509502, ENST00000513765, | ENST00000435842, ENST00000527716, ENST00000301887, | |
| Fusion gene scores | * DoF score | 13 X 9 X 11=1287 | 3 X 2 X 4=24 |
| # samples | 21 | 4 | |
| ** MAII score | log2(21/1287*10)=-2.61555082055458 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(4/24*10)=0.736965594166206 effective Gene in Pan-Cancer Fusion Genes (eGinPCFGs). DoF>8 and MAII>0 | |
| Context | PubMed: MARK2 [Title/Abstract] AND BATF2 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MARK2(63607032)-BATF2(64762021), # samples:3 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | MARK2 | GO:0006468 | protein phosphorylation | 14976552 |
| Hgene | MARK2 | GO:0010976 | positive regulation of neuron projection development | 12429843 |
| Hgene | MARK2 | GO:0018105 | peptidyl-serine phosphorylation | 10542369|16717194 |
| Hgene | MARK2 | GO:0030010 | establishment of cell polarity | 12429843 |
| Hgene | MARK2 | GO:0035556 | intracellular signal transduction | 14976552 |
| Hgene | MARK2 | GO:0045197 | establishment or maintenance of epithelial cell apical/basal polarity | 15324659 |
| Hgene | MARK2 | GO:0070507 | regulation of microtubule cytoskeleton organization | 10542369 |
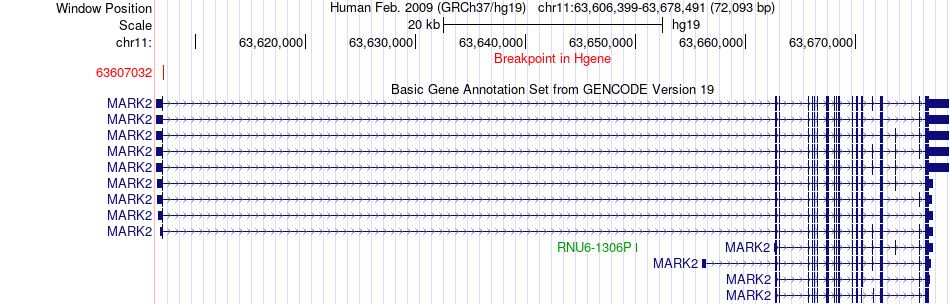
 Fusion gene breakpoints across MARK2 (5'-gene) Fusion gene breakpoints across MARK2 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
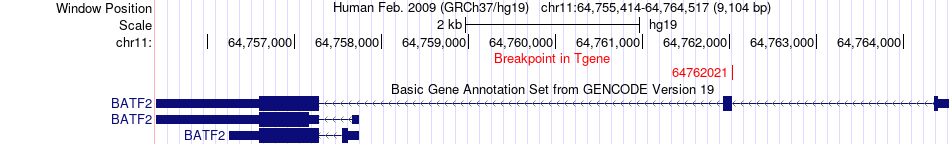
 Fusion gene breakpoints across BATF2 (3'-gene) Fusion gene breakpoints across BATF2 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | SKCM | TCGA-GN-A26D-06A | MARK2 | chr11 | 63607032 | - | BATF2 | chr11 | 64762021 | - |
| ChimerDB4 | SKCM | TCGA-GN-A26D-06A | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
Top |
Fusion Gene ORF analysis for MARK2-BATF2 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 5CDS-intron | ENST00000315032 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000315032 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000350490 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000350490 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000361128 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000361128 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000377809 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000377809 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000402010 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000402010 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000413835 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000413835 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000502399 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000502399 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000508192 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5CDS-intron | ENST00000508192 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5UTR-3CDS | ENST00000377810 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5UTR-intron | ENST00000377810 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| 5UTR-intron | ENST00000377810 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| In-frame | ENST00000315032 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| In-frame | ENST00000350490 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| In-frame | ENST00000361128 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| In-frame | ENST00000377809 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| In-frame | ENST00000402010 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| In-frame | ENST00000413835 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| In-frame | ENST00000502399 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| In-frame | ENST00000508192 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-3CDS | ENST00000408948 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-3CDS | ENST00000425897 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-3CDS | ENST00000509502 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-3CDS | ENST00000513765 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-intron | ENST00000408948 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-intron | ENST00000408948 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-intron | ENST00000425897 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-intron | ENST00000425897 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-intron | ENST00000509502 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-intron | ENST00000509502 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-intron | ENST00000513765 | ENST00000435842 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
| intron-intron | ENST00000513765 | ENST00000527716 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000377809 | MARK2 | chr11 | 63607032 | + | ENST00000301887 | BATF2 | chr11 | 64762021 | - | 2605 | 633 | 1008 | 55 | 317 |
| ENST00000402010 | MARK2 | chr11 | 63607032 | + | ENST00000301887 | BATF2 | chr11 | 64762021 | - | 2605 | 633 | 1008 | 55 | 317 |
| ENST00000413835 | MARK2 | chr11 | 63607032 | + | ENST00000301887 | BATF2 | chr11 | 64762021 | - | 2605 | 633 | 1008 | 55 | 317 |
| ENST00000315032 | MARK2 | chr11 | 63607032 | + | ENST00000301887 | BATF2 | chr11 | 64762021 | - | 2605 | 633 | 1008 | 55 | 317 |
| ENST00000508192 | MARK2 | chr11 | 63607032 | + | ENST00000301887 | BATF2 | chr11 | 64762021 | - | 2528 | 556 | 931 | 2 | 310 |
| ENST00000361128 | MARK2 | chr11 | 63607032 | + | ENST00000301887 | BATF2 | chr11 | 64762021 | - | 2528 | 556 | 931 | 2 | 310 |
| ENST00000350490 | MARK2 | chr11 | 63607032 | + | ENST00000301887 | BATF2 | chr11 | 64762021 | - | 2440 | 468 | 843 | 1 | 281 |
| ENST00000502399 | MARK2 | chr11 | 63607032 | + | ENST00000301887 | BATF2 | chr11 | 64762021 | - | 2238 | 266 | 212 | 1051 | 279 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000377809 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - | 0.30724803 | 0.692752 |
| ENST00000402010 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - | 0.30724803 | 0.692752 |
| ENST00000413835 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - | 0.30724803 | 0.692752 |
| ENST00000315032 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - | 0.30724803 | 0.692752 |
| ENST00000508192 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - | 0.25776607 | 0.742234 |
| ENST00000361128 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - | 0.25776607 | 0.742234 |
| ENST00000350490 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - | 0.27091786 | 0.7290821 |
| ENST00000502399 | ENST00000301887 | MARK2 | chr11 | 63607032 | + | BATF2 | chr11 | 64762021 | - | 0.368383 | 0.631617 |
Top |
Fusion Genomic Features for MARK2-BATF2 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
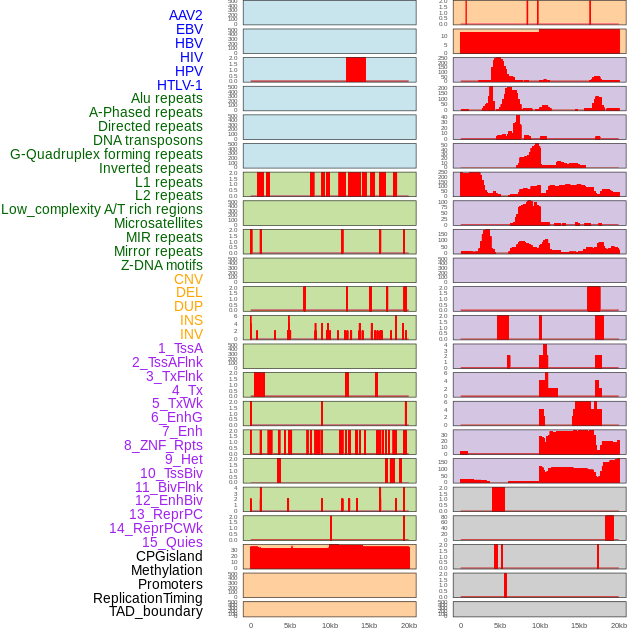 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
Top |
Fusion Protein Features for MARK2-BATF2 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr11:63607032/chr11:64762021) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MARK2 | BATF2 |
| FUNCTION: Serine/threonine-protein kinase (PubMed:23666762). Involved in cell polarity and microtubule dynamics regulation. Phosphorylates CRTC2/TORC2, DCX, HDAC7, KIF13B, MAP2, MAP4 and RAB11FIP2. Phosphorylates the microtubule-associated protein MAPT/TAU (PubMed:23666762). Plays a key role in cell polarity by phosphorylating the microtubule-associated proteins MAP2, MAP4 and MAPT/TAU at KXGS motifs, causing detachment from microtubules, and their disassembly. Regulates epithelial cell polarity by phosphorylating RAB11FIP2. Involved in the regulation of neuronal migration through its dual activities in regulating cellular polarity and microtubule dynamics, possibly by phosphorylating and regulating DCX. Regulates axogenesis by phosphorylating KIF13B, promoting interaction between KIF13B and 14-3-3 and inhibiting microtubule-dependent accumulation of KIF13B. Also required for neurite outgrowth and establishment of neuronal polarity. Regulates localization and activity of some histone deacetylases by mediating phosphorylation of HDAC7, promoting subsequent interaction between HDAC7 and 14-3-3 and export from the nucleus. Also acts as a positive regulator of the Wnt signaling pathway, probably by mediating phosphorylation of dishevelled proteins (DVL1, DVL2 and/or DVL3). Modulates the developmental decision to build a columnar versus a hepatic epithelial cell apparently by promoting a switch from a direct to a transcytotic mode of apical protein delivery. Essential for the asymmetric development of membrane domains of polarized epithelial cells. {ECO:0000269|PubMed:11433294, ECO:0000269|PubMed:12429843, ECO:0000269|PubMed:14976552, ECO:0000269|PubMed:15158914, ECO:0000269|PubMed:15324659, ECO:0000269|PubMed:15365179, ECO:0000269|PubMed:16775013, ECO:0000269|PubMed:16980613, ECO:0000269|PubMed:18626018, ECO:0000269|PubMed:20194617, ECO:0000269|PubMed:23666762}. | FUNCTION: AP-1 family transcription factor that controls the differentiation of lineage-specific cells in the immune system. Following infection, participates in the differentiation of CD8(+) thymic conventional dendritic cells in the immune system. Acts via the formation of a heterodimer with JUN family proteins that recognizes and binds DNA sequence 5'-TGA[CG]TCA-3' and regulates expression of target genes (By similarity). Selectively suppresses CCN1 transcription and hence blocks the downstream cell proliferation signals produced by CCN1 and inhibits CCN1-induced anchorage-independent growth and invasion in several cancer types, such as breast cancer, malignant glioma and metastatic melanoma. Possibly acts by interfering with AP-1 binding to CCN1 promoter. {ECO:0000250, ECO:0000269|PubMed:20531301}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Tgene | BATF2 | chr11:63607032 | chr11:64762021 | ENST00000301887 | 0 | 3 | 17_80 | 13 | 275.0 | Domain | bZIP | |
| Tgene | BATF2 | chr11:63607032 | chr11:64762021 | ENST00000435842 | 0 | 2 | 17_80 | 0 | 190.0 | Domain | bZIP | |
| Tgene | BATF2 | chr11:63607032 | chr11:64762021 | ENST00000301887 | 0 | 3 | 20_41 | 13 | 275.0 | Region | Basic motif | |
| Tgene | BATF2 | chr11:63607032 | chr11:64762021 | ENST00000301887 | 0 | 3 | 45_66 | 13 | 275.0 | Region | Leucine-zipper | |
| Tgene | BATF2 | chr11:63607032 | chr11:64762021 | ENST00000435842 | 0 | 2 | 20_41 | 0 | 190.0 | Region | Basic motif | |
| Tgene | BATF2 | chr11:63607032 | chr11:64762021 | ENST00000435842 | 0 | 2 | 45_66 | 0 | 190.0 | Region | Leucine-zipper |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000315032 | + | 1 | 18 | 323_362 | 18 | 780.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000315032 | + | 1 | 18 | 53_304 | 18 | 780.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000315032 | + | 1 | 18 | 739_788 | 18 | 780.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000350490 | + | 1 | 16 | 323_362 | 18 | 710.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000350490 | + | 1 | 16 | 53_304 | 18 | 710.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000350490 | + | 1 | 16 | 739_788 | 18 | 710.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000361128 | + | 1 | 17 | 323_362 | 18 | 720.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000361128 | + | 1 | 17 | 53_304 | 18 | 720.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000361128 | + | 1 | 17 | 739_788 | 18 | 720.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000377809 | + | 1 | 18 | 323_362 | 18 | 774.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000377809 | + | 1 | 18 | 53_304 | 18 | 774.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000377809 | + | 1 | 18 | 739_788 | 18 | 774.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000377810 | + | 1 | 17 | 323_362 | 0 | 692.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000377810 | + | 1 | 17 | 53_304 | 0 | 692.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000377810 | + | 1 | 17 | 739_788 | 0 | 692.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000402010 | + | 1 | 19 | 323_362 | 18 | 789.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000402010 | + | 1 | 19 | 53_304 | 18 | 789.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000402010 | + | 1 | 19 | 739_788 | 18 | 789.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000408948 | + | 1 | 16 | 323_362 | 0 | 692.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000408948 | + | 1 | 16 | 53_304 | 0 | 692.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000408948 | + | 1 | 16 | 739_788 | 0 | 692.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000413835 | + | 1 | 18 | 323_362 | 18 | 735.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000413835 | + | 1 | 18 | 53_304 | 18 | 735.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000413835 | + | 1 | 18 | 739_788 | 18 | 735.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000508192 | + | 1 | 17 | 323_362 | 18 | 725.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000508192 | + | 1 | 17 | 53_304 | 18 | 725.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000508192 | + | 1 | 17 | 739_788 | 18 | 725.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000509502 | + | 1 | 18 | 323_362 | 0 | 746.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000509502 | + | 1 | 18 | 53_304 | 0 | 746.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000509502 | + | 1 | 18 | 739_788 | 0 | 746.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000513765 | + | 1 | 18 | 323_362 | 0 | 756.0 | Domain | UBA |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000513765 | + | 1 | 18 | 53_304 | 0 | 756.0 | Domain | Protein kinase |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000513765 | + | 1 | 18 | 739_788 | 0 | 756.0 | Domain | KA1 |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000315032 | + | 1 | 18 | 59_67 | 18 | 780.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000350490 | + | 1 | 16 | 59_67 | 18 | 710.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000361128 | + | 1 | 17 | 59_67 | 18 | 720.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000377809 | + | 1 | 18 | 59_67 | 18 | 774.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000377810 | + | 1 | 17 | 59_67 | 0 | 692.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000402010 | + | 1 | 19 | 59_67 | 18 | 789.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000408948 | + | 1 | 16 | 59_67 | 0 | 692.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000413835 | + | 1 | 18 | 59_67 | 18 | 735.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000508192 | + | 1 | 17 | 59_67 | 18 | 725.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000509502 | + | 1 | 18 | 59_67 | 0 | 746.0 | Nucleotide binding | ATP |
| Hgene | MARK2 | chr11:63607032 | chr11:64762021 | ENST00000513765 | + | 1 | 18 | 59_67 | 0 | 756.0 | Nucleotide binding | ATP |
Top |
Fusion Gene Sequence for MARK2-BATF2 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >51741_51741_1_MARK2-BATF2_MARK2_chr11_63607032_ENST00000315032_BATF2_chr11_64762021_ENST00000301887_length(transcript)=2605nt_BP=633nt AAACAGAGCCGAAGCTTCGTCTTCGGAATCGAGGAGGAGAGCGCGTCCCCGGGGTTTACCTTCTTCGGCTGTTTCCGGAATTCACCCGTA GTCTGTGGCCGGGAGGAGAAGGCGCGCCCCACCTCCCGTTTCTAGCCGCTTCGGATGTTTCCGGCTGCTGCCGGCGAAAAGAGCCGAGGG CCGGCGGTGGTGGCGGCCATGTTGGGAGCAGCAGGTCCGGCGGCGGCTGCCTGTGTGCCGGGCGCGGAGCAGTGCCGCTGAGGGCAGGGG AGGAGCGAGGCAGGCGGCCGGCTGCGGCGGCAGAGAGTAGGCGGAGCGGCGCGGCCCGGCCGAAAGGCGGCACAGCCCAGCCGGGGGTCG GGGGGGTGCGGTCCGGAGCCGCTCGGAGCCGGCGCGGCCTAGCCCGAGCGGCGCATCCCCGGGCTGGCGTGAGCGGCTGCCCGGCCTCCC CGCACCCCCGGCCGGGGCCCATGCGGCGGGTGCTCCTGCTGTGAGAAGCCCCGCCCGGCCGGGCTCCGCGCCTTCCCTTCCCTCCCTTCC TCCAAGCTTCTCGGTTCCCTCCCCCGAGATACCGGCGCCATGTCCAGCGCTCGGACCCCCCTACCCACGCTGAACGAGAGGGACACGGAG CAGGACCCCAAGGAGCAACAAAGGCAGCTGAAGAAGCAGAAGAACCGGGCAGCCGCCCAGCGAAGCCGGCAGAAGCACACAGACAAGGCA GACGCCCTGCACCAGCAGCACGAGTCTCTGGAAAAAGACAACCTCGCCCTGCGGAAGGAGATCCAGTCCCTGCAGGCCGAGCTGGCGTGG TGGAGCCGGACCCTGCACGTGCATGAGCGCCTGTGCCCCATGGATTGTGCCTCCTGCTCAGCTCCAGGGCTCCTGGGCTGCTGGGACCAG GCTGAGGGGCTCCTGGGCCCTGGCCCACAGGGACAACATGGCTGCCGGGAGCAGCTGGAGCTGTTCCAGACCCCGGGTTCCTGTTACCCA GCTCAGCCGCTCTCTCCAGGTCCACAGCCTCATGATTCTCCCAGCCTCCTCCAGTGCCCCCTGCCCTCACTGTCCCTTGGCCCCGCTGTG GTTGCTGAACCTCCTGTCCAGCTGTCCCCCAGCCCTCTCCTGTTTGCCTCGCACACTGGTTCCAGCCTGCAGGGGTCTTCCTCTAAGCTC AGTGCCCTCCAGCCCAGCCTCACGGCCCAAACTGCCCCTCCACAGCCCCTCGAGCTGGAGCATCCCACCAGAGGGAAGCTGGGGTCCTCT CCCGACAACCCTTCCTCTGCCCTGGGGCTTGCACGTCTGCAGAGCAGGGAGCACAAACCTGCTCTCTCAGCAGCCACTTGGCAAGGGCTG GTTGTGGATCCCAGCCCTCACCCTCTCCTGGCCTTTCCTCTGCTCTCCTCTGCTCAAGTCCACTTCTAACCTGGTCTTCGGAGCTGGGTT GGCCCCTTCTTTGGGCTCAGGAAGCAGCCTTAGCACACGGGCCTCTCCTCCCTCACTACTGGGTGCTGCCCTGCGTGGCTGACCAGCTGG CCCAGGATTTCACAGTCGAAAAGGAAGCCACCACTGATGCCTCCCACTGTGACAGGCCCTGTCACCACCAATATCTTATTTCAACCTCAC AGTTGACCTGAGAAATCGAGATTATCACTCCACTTTTTCAGACAAGGAAACTGAGGCTCAGGGAAGCCAAGTGACAAGTCCAAGGTCACG AAGACTTTCTTGGAGCCCGAAACACCACCCTCTGCTCCTCCTTCTCCTGTCCTGGCCCAGGCATCCTAGGGGCTGAAATCCTGGAAACCG TGGGCTGGTGTGAGAAGGTTTGCATGCTCAGAGCAGAGAAGGGCTCTCCCCACTGCTTCGTGATTCCAGGGCCAGAGCCATGCAGTCCCA GAAACCCCAACCTAGCTGGGGCAGGTCCAGAGTCCAAGCCCTGGTGGGTAGAGGCCAAGCAGAAGCCCTGAAGTGGACTCTTGCTTCCCC TAGTAGTGTTTTCAGTGCCAAGAAGCTGAAACTGTGAGCTGGAGTTGGGGAGAGGTCTGGAAGAGGACCATCTGGGATTTCTACAGCCTG GGTACCCATAGCCACACCAAGGCTTCTGGGAGATTCTGCAGGGTCAGCTTTCCAGGCTGTTCCCAAATAGCTCCCTGCCTCCCCACTGCC CCTAAAGCCACAGCAGAAGAGCCATTCATCTCATAAACAAAAAGGAAGAGGAAAGAATGAGGAAGGACCCTGTGCAAGGTTATTTGCAGG CAGGGATGGGCTTGTACCTGACAGCACCCACCCCTGTGTGGCCCCCAGGCCCTCATCACCCTCAGACCCCTCCTAAGCAGTTCCCTCATT GCTCTTTGGACTAGGCTGACAGCAGGAAGAGCAGGGCCCATGACCGGGTGGAAGTTCAGTTTTGGTGTCTGCTTCAAGAGGGGGTTTTAC ACTCTGATTCCAGGACAAGCACTCTGAGGCGGGTGGGGGAGAGAAACCCTGGCTCTTCACCCAGGTTTCACACACATGTAAATGAAACAC >51741_51741_1_MARK2-BATF2_MARK2_chr11_63607032_ENST00000315032_BATF2_chr11_64762021_ENST00000301887_length(amino acids)=317AA_BP=161 MERAAELGNRNPGSGTAPAAPGSHVVPVGQGPGAPQPGPSSPGALELSRRHNPWGTGAHARAGSGSTTPARPAGTGSPSAGRGCLFPETR AAGAGRLPCLCASAGFAGRLPGSSASSAAFVAPWGPAPCPSRSAWVGGSERWTWRRYLGGGNREAWRKGGKGRRGARPGGASHSRSTRRM GPGRGCGEAGQPLTPARGCAARARPRRLRAAPDRTPPTPGWAVPPFGRAAPLRLLSAAAAGRLPRSSPALSGTAPRPAHRQPPPDLLLPT -------------------------------------------------------------- >51741_51741_2_MARK2-BATF2_MARK2_chr11_63607032_ENST00000350490_BATF2_chr11_64762021_ENST00000301887_length(transcript)=2440nt_BP=468nt GAAAAGAGCCGAGGGCCGGCGGTGGTGGCGGCCATGTTGGGAGCAGCAGGTCCGGCGGCGGCTGCCTGTGTGCCGGGCGCGGAGCAGTGC CGCTGAGGGCAGGGGAGGAGCGAGGCAGGCGGCCGGCTGCGGCGGCAGAGAGTAGGCGGAGCGGCGCGGCCCGGCCGAAAGGCGGCACAG CCCAGCCGGGGGTCGGGGGGGTGCGGTCCGGAGCCGCTCGGAGCCGGCGCGGCCTAGCCCGAGCGGCGCATCCCCGGGCTGGCGTGAGCG GCTGCCCGGCCTCCCCGCACCCCCGGCCGGGGCCCATGCGGCGGGTGCTCCTGCTGTGAGAAGCCCCGCCCGGCCGGGCTCCGCGCCTTC CCTTCCCTCCCTTCCTCCAAGCTTCTCGGTTCCCTCCCCCGAGATACCGGCGCCATGTCCAGCGCTCGGACCCCCCTACCCACGCTGAAC GAGAGGGACACGGAGCAGGACCCCAAGGAGCAACAAAGGCAGCTGAAGAAGCAGAAGAACCGGGCAGCCGCCCAGCGAAGCCGGCAGAAG CACACAGACAAGGCAGACGCCCTGCACCAGCAGCACGAGTCTCTGGAAAAAGACAACCTCGCCCTGCGGAAGGAGATCCAGTCCCTGCAG GCCGAGCTGGCGTGGTGGAGCCGGACCCTGCACGTGCATGAGCGCCTGTGCCCCATGGATTGTGCCTCCTGCTCAGCTCCAGGGCTCCTG GGCTGCTGGGACCAGGCTGAGGGGCTCCTGGGCCCTGGCCCACAGGGACAACATGGCTGCCGGGAGCAGCTGGAGCTGTTCCAGACCCCG GGTTCCTGTTACCCAGCTCAGCCGCTCTCTCCAGGTCCACAGCCTCATGATTCTCCCAGCCTCCTCCAGTGCCCCCTGCCCTCACTGTCC CTTGGCCCCGCTGTGGTTGCTGAACCTCCTGTCCAGCTGTCCCCCAGCCCTCTCCTGTTTGCCTCGCACACTGGTTCCAGCCTGCAGGGG TCTTCCTCTAAGCTCAGTGCCCTCCAGCCCAGCCTCACGGCCCAAACTGCCCCTCCACAGCCCCTCGAGCTGGAGCATCCCACCAGAGGG AAGCTGGGGTCCTCTCCCGACAACCCTTCCTCTGCCCTGGGGCTTGCACGTCTGCAGAGCAGGGAGCACAAACCTGCTCTCTCAGCAGCC ACTTGGCAAGGGCTGGTTGTGGATCCCAGCCCTCACCCTCTCCTGGCCTTTCCTCTGCTCTCCTCTGCTCAAGTCCACTTCTAACCTGGT CTTCGGAGCTGGGTTGGCCCCTTCTTTGGGCTCAGGAAGCAGCCTTAGCACACGGGCCTCTCCTCCCTCACTACTGGGTGCTGCCCTGCG TGGCTGACCAGCTGGCCCAGGATTTCACAGTCGAAAAGGAAGCCACCACTGATGCCTCCCACTGTGACAGGCCCTGTCACCACCAATATC TTATTTCAACCTCACAGTTGACCTGAGAAATCGAGATTATCACTCCACTTTTTCAGACAAGGAAACTGAGGCTCAGGGAAGCCAAGTGAC AAGTCCAAGGTCACGAAGACTTTCTTGGAGCCCGAAACACCACCCTCTGCTCCTCCTTCTCCTGTCCTGGCCCAGGCATCCTAGGGGCTG AAATCCTGGAAACCGTGGGCTGGTGTGAGAAGGTTTGCATGCTCAGAGCAGAGAAGGGCTCTCCCCACTGCTTCGTGATTCCAGGGCCAG AGCCATGCAGTCCCAGAAACCCCAACCTAGCTGGGGCAGGTCCAGAGTCCAAGCCCTGGTGGGTAGAGGCCAAGCAGAAGCCCTGAAGTG GACTCTTGCTTCCCCTAGTAGTGTTTTCAGTGCCAAGAAGCTGAAACTGTGAGCTGGAGTTGGGGAGAGGTCTGGAAGAGGACCATCTGG GATTTCTACAGCCTGGGTACCCATAGCCACACCAAGGCTTCTGGGAGATTCTGCAGGGTCAGCTTTCCAGGCTGTTCCCAAATAGCTCCC TGCCTCCCCACTGCCCCTAAAGCCACAGCAGAAGAGCCATTCATCTCATAAACAAAAAGGAAGAGGAAAGAATGAGGAAGGACCCTGTGC AAGGTTATTTGCAGGCAGGGATGGGCTTGTACCTGACAGCACCCACCCCTGTGTGGCCCCCAGGCCCTCATCACCCTCAGACCCCTCCTA AGCAGTTCCCTCATTGCTCTTTGGACTAGGCTGACAGCAGGAAGAGCAGGGCCCATGACCGGGTGGAAGTTCAGTTTTGGTGTCTGCTTC AAGAGGGGGTTTTACACTCTGATTCCAGGACAAGCACTCTGAGGCGGGTGGGGGAGAGAAACCCTGGCTCTTCACCCAGGTTTCACACAC ATGTAAATGAAACACTATGTTAGTATCTAACACACTCCTGGATACAGAACACAAGTCTTGGCACATATGTGATGGAAATAAAGTGTTTTG >51741_51741_2_MARK2-BATF2_MARK2_chr11_63607032_ENST00000350490_BATF2_chr11_64762021_ENST00000301887_length(amino acids)=281AA_BP=161 MERAAELGNRNPGSGTAPAAPGSHVVPVGQGPGAPQPGPSSPGALELSRRHNPWGTGAHARAGSGSTTPARPAGTGSPSAGRGCLFPETR AAGAGRLPCLCASAGFAGRLPGSSASSAAFVAPWGPAPCPSRSAWVGGSERWTWRRYLGGGNREAWRKGGKGRRGARPGGASHSRSTRRM GPGRGCGEAGQPLTPARGCAARARPRRLRAAPDRTPPTPGWAVPPFGRAAPLRLLSAAAAGRLPRSSPALSGTAPRPAHRQPPPDLLLPT -------------------------------------------------------------- >51741_51741_3_MARK2-BATF2_MARK2_chr11_63607032_ENST00000361128_BATF2_chr11_64762021_ENST00000301887_length(transcript)=2528nt_BP=556nt GAATTCACCCGTAGTCTGTGGCCGGGAGGAGAAGGCGCGCCCCACCTCCCGTTTCTAGCCGCTTCGGATGTTTCCGGCTGCTGCCGGCGA AAAGAGCCGAGGGCCGGCGGTGGTGGCGGCCATGTTGGGAGCAGCAGGTCCGGCGGCGGCTGCCTGTGTGCCGGGCGCGGAGCAGTGCCG CTGAGGGCAGGGGAGGAGCGAGGCAGGCGGCCGGCTGCGGCGGCAGAGAGTAGGCGGAGCGGCGCGGCCCGGCCGAAAGGCGGCACAGCC CAGCCGGGGGTCGGGGGGGTGCGGTCCGGAGCCGCTCGGAGCCGGCGCGGCCTAGCCCGAGCGGCGCATCCCCGGGCTGGCGTGAGCGGC TGCCCGGCCTCCCCGCACCCCCGGCCGGGGCCCATGCGGCGGGTGCTCCTGCTGTGAGAAGCCCCGCCCGGCCGGGCTCCGCGCCTTCCC TTCCCTCCCTTCCTCCAAGCTTCTCGGTTCCCTCCCCCGAGATACCGGCGCCATGTCCAGCGCTCGGACCCCCCTACCCACGCTGAACGA GAGGGACACGGAGCAGGACCCCAAGGAGCAACAAAGGCAGCTGAAGAAGCAGAAGAACCGGGCAGCCGCCCAGCGAAGCCGGCAGAAGCA CACAGACAAGGCAGACGCCCTGCACCAGCAGCACGAGTCTCTGGAAAAAGACAACCTCGCCCTGCGGAAGGAGATCCAGTCCCTGCAGGC CGAGCTGGCGTGGTGGAGCCGGACCCTGCACGTGCATGAGCGCCTGTGCCCCATGGATTGTGCCTCCTGCTCAGCTCCAGGGCTCCTGGG CTGCTGGGACCAGGCTGAGGGGCTCCTGGGCCCTGGCCCACAGGGACAACATGGCTGCCGGGAGCAGCTGGAGCTGTTCCAGACCCCGGG TTCCTGTTACCCAGCTCAGCCGCTCTCTCCAGGTCCACAGCCTCATGATTCTCCCAGCCTCCTCCAGTGCCCCCTGCCCTCACTGTCCCT TGGCCCCGCTGTGGTTGCTGAACCTCCTGTCCAGCTGTCCCCCAGCCCTCTCCTGTTTGCCTCGCACACTGGTTCCAGCCTGCAGGGGTC TTCCTCTAAGCTCAGTGCCCTCCAGCCCAGCCTCACGGCCCAAACTGCCCCTCCACAGCCCCTCGAGCTGGAGCATCCCACCAGAGGGAA GCTGGGGTCCTCTCCCGACAACCCTTCCTCTGCCCTGGGGCTTGCACGTCTGCAGAGCAGGGAGCACAAACCTGCTCTCTCAGCAGCCAC TTGGCAAGGGCTGGTTGTGGATCCCAGCCCTCACCCTCTCCTGGCCTTTCCTCTGCTCTCCTCTGCTCAAGTCCACTTCTAACCTGGTCT TCGGAGCTGGGTTGGCCCCTTCTTTGGGCTCAGGAAGCAGCCTTAGCACACGGGCCTCTCCTCCCTCACTACTGGGTGCTGCCCTGCGTG GCTGACCAGCTGGCCCAGGATTTCACAGTCGAAAAGGAAGCCACCACTGATGCCTCCCACTGTGACAGGCCCTGTCACCACCAATATCTT ATTTCAACCTCACAGTTGACCTGAGAAATCGAGATTATCACTCCACTTTTTCAGACAAGGAAACTGAGGCTCAGGGAAGCCAAGTGACAA GTCCAAGGTCACGAAGACTTTCTTGGAGCCCGAAACACCACCCTCTGCTCCTCCTTCTCCTGTCCTGGCCCAGGCATCCTAGGGGCTGAA ATCCTGGAAACCGTGGGCTGGTGTGAGAAGGTTTGCATGCTCAGAGCAGAGAAGGGCTCTCCCCACTGCTTCGTGATTCCAGGGCCAGAG CCATGCAGTCCCAGAAACCCCAACCTAGCTGGGGCAGGTCCAGAGTCCAAGCCCTGGTGGGTAGAGGCCAAGCAGAAGCCCTGAAGTGGA CTCTTGCTTCCCCTAGTAGTGTTTTCAGTGCCAAGAAGCTGAAACTGTGAGCTGGAGTTGGGGAGAGGTCTGGAAGAGGACCATCTGGGA TTTCTACAGCCTGGGTACCCATAGCCACACCAAGGCTTCTGGGAGATTCTGCAGGGTCAGCTTTCCAGGCTGTTCCCAAATAGCTCCCTG CCTCCCCACTGCCCCTAAAGCCACAGCAGAAGAGCCATTCATCTCATAAACAAAAAGGAAGAGGAAAGAATGAGGAAGGACCCTGTGCAA GGTTATTTGCAGGCAGGGATGGGCTTGTACCTGACAGCACCCACCCCTGTGTGGCCCCCAGGCCCTCATCACCCTCAGACCCCTCCTAAG CAGTTCCCTCATTGCTCTTTGGACTAGGCTGACAGCAGGAAGAGCAGGGCCCATGACCGGGTGGAAGTTCAGTTTTGGTGTCTGCTTCAA GAGGGGGTTTTACACTCTGATTCCAGGACAAGCACTCTGAGGCGGGTGGGGGAGAGAAACCCTGGCTCTTCACCCAGGTTTCACACACAT GTAAATGAAACACTATGTTAGTATCTAACACACTCCTGGATACAGAACACAAGTCTTGGCACATATGTGATGGAAATAAAGTGTTTTGCA >51741_51741_3_MARK2-BATF2_MARK2_chr11_63607032_ENST00000361128_BATF2_chr11_64762021_ENST00000301887_length(amino acids)=310AA_BP=161 MERAAELGNRNPGSGTAPAAPGSHVVPVGQGPGAPQPGPSSPGALELSRRHNPWGTGAHARAGSGSTTPARPAGTGSPSAGRGCLFPETR AAGAGRLPCLCASAGFAGRLPGSSASSAAFVAPWGPAPCPSRSAWVGGSERWTWRRYLGGGNREAWRKGGKGRRGARPGGASHSRSTRRM GPGRGCGEAGQPLTPARGCAARARPRRLRAAPDRTPPTPGWAVPPFGRAAPLRLLSAAAAGRLPRSSPALSGTAPRPAHRQPPPDLLLPT -------------------------------------------------------------- >51741_51741_4_MARK2-BATF2_MARK2_chr11_63607032_ENST00000377809_BATF2_chr11_64762021_ENST00000301887_length(transcript)=2605nt_BP=633nt AAACAGAGCCGAAGCTTCGTCTTCGGAATCGAGGAGGAGAGCGCGTCCCCGGGGTTTACCTTCTTCGGCTGTTTCCGGAATTCACCCGTA GTCTGTGGCCGGGAGGAGAAGGCGCGCCCCACCTCCCGTTTCTAGCCGCTTCGGATGTTTCCGGCTGCTGCCGGCGAAAAGAGCCGAGGG CCGGCGGTGGTGGCGGCCATGTTGGGAGCAGCAGGTCCGGCGGCGGCTGCCTGTGTGCCGGGCGCGGAGCAGTGCCGCTGAGGGCAGGGG AGGAGCGAGGCAGGCGGCCGGCTGCGGCGGCAGAGAGTAGGCGGAGCGGCGCGGCCCGGCCGAAAGGCGGCACAGCCCAGCCGGGGGTCG GGGGGGTGCGGTCCGGAGCCGCTCGGAGCCGGCGCGGCCTAGCCCGAGCGGCGCATCCCCGGGCTGGCGTGAGCGGCTGCCCGGCCTCCC CGCACCCCCGGCCGGGGCCCATGCGGCGGGTGCTCCTGCTGTGAGAAGCCCCGCCCGGCCGGGCTCCGCGCCTTCCCTTCCCTCCCTTCC TCCAAGCTTCTCGGTTCCCTCCCCCGAGATACCGGCGCCATGTCCAGCGCTCGGACCCCCCTACCCACGCTGAACGAGAGGGACACGGAG CAGGACCCCAAGGAGCAACAAAGGCAGCTGAAGAAGCAGAAGAACCGGGCAGCCGCCCAGCGAAGCCGGCAGAAGCACACAGACAAGGCA GACGCCCTGCACCAGCAGCACGAGTCTCTGGAAAAAGACAACCTCGCCCTGCGGAAGGAGATCCAGTCCCTGCAGGCCGAGCTGGCGTGG TGGAGCCGGACCCTGCACGTGCATGAGCGCCTGTGCCCCATGGATTGTGCCTCCTGCTCAGCTCCAGGGCTCCTGGGCTGCTGGGACCAG GCTGAGGGGCTCCTGGGCCCTGGCCCACAGGGACAACATGGCTGCCGGGAGCAGCTGGAGCTGTTCCAGACCCCGGGTTCCTGTTACCCA GCTCAGCCGCTCTCTCCAGGTCCACAGCCTCATGATTCTCCCAGCCTCCTCCAGTGCCCCCTGCCCTCACTGTCCCTTGGCCCCGCTGTG GTTGCTGAACCTCCTGTCCAGCTGTCCCCCAGCCCTCTCCTGTTTGCCTCGCACACTGGTTCCAGCCTGCAGGGGTCTTCCTCTAAGCTC AGTGCCCTCCAGCCCAGCCTCACGGCCCAAACTGCCCCTCCACAGCCCCTCGAGCTGGAGCATCCCACCAGAGGGAAGCTGGGGTCCTCT CCCGACAACCCTTCCTCTGCCCTGGGGCTTGCACGTCTGCAGAGCAGGGAGCACAAACCTGCTCTCTCAGCAGCCACTTGGCAAGGGCTG GTTGTGGATCCCAGCCCTCACCCTCTCCTGGCCTTTCCTCTGCTCTCCTCTGCTCAAGTCCACTTCTAACCTGGTCTTCGGAGCTGGGTT GGCCCCTTCTTTGGGCTCAGGAAGCAGCCTTAGCACACGGGCCTCTCCTCCCTCACTACTGGGTGCTGCCCTGCGTGGCTGACCAGCTGG CCCAGGATTTCACAGTCGAAAAGGAAGCCACCACTGATGCCTCCCACTGTGACAGGCCCTGTCACCACCAATATCTTATTTCAACCTCAC AGTTGACCTGAGAAATCGAGATTATCACTCCACTTTTTCAGACAAGGAAACTGAGGCTCAGGGAAGCCAAGTGACAAGTCCAAGGTCACG AAGACTTTCTTGGAGCCCGAAACACCACCCTCTGCTCCTCCTTCTCCTGTCCTGGCCCAGGCATCCTAGGGGCTGAAATCCTGGAAACCG TGGGCTGGTGTGAGAAGGTTTGCATGCTCAGAGCAGAGAAGGGCTCTCCCCACTGCTTCGTGATTCCAGGGCCAGAGCCATGCAGTCCCA GAAACCCCAACCTAGCTGGGGCAGGTCCAGAGTCCAAGCCCTGGTGGGTAGAGGCCAAGCAGAAGCCCTGAAGTGGACTCTTGCTTCCCC TAGTAGTGTTTTCAGTGCCAAGAAGCTGAAACTGTGAGCTGGAGTTGGGGAGAGGTCTGGAAGAGGACCATCTGGGATTTCTACAGCCTG GGTACCCATAGCCACACCAAGGCTTCTGGGAGATTCTGCAGGGTCAGCTTTCCAGGCTGTTCCCAAATAGCTCCCTGCCTCCCCACTGCC CCTAAAGCCACAGCAGAAGAGCCATTCATCTCATAAACAAAAAGGAAGAGGAAAGAATGAGGAAGGACCCTGTGCAAGGTTATTTGCAGG CAGGGATGGGCTTGTACCTGACAGCACCCACCCCTGTGTGGCCCCCAGGCCCTCATCACCCTCAGACCCCTCCTAAGCAGTTCCCTCATT GCTCTTTGGACTAGGCTGACAGCAGGAAGAGCAGGGCCCATGACCGGGTGGAAGTTCAGTTTTGGTGTCTGCTTCAAGAGGGGGTTTTAC ACTCTGATTCCAGGACAAGCACTCTGAGGCGGGTGGGGGAGAGAAACCCTGGCTCTTCACCCAGGTTTCACACACATGTAAATGAAACAC >51741_51741_4_MARK2-BATF2_MARK2_chr11_63607032_ENST00000377809_BATF2_chr11_64762021_ENST00000301887_length(amino acids)=317AA_BP=161 MERAAELGNRNPGSGTAPAAPGSHVVPVGQGPGAPQPGPSSPGALELSRRHNPWGTGAHARAGSGSTTPARPAGTGSPSAGRGCLFPETR AAGAGRLPCLCASAGFAGRLPGSSASSAAFVAPWGPAPCPSRSAWVGGSERWTWRRYLGGGNREAWRKGGKGRRGARPGGASHSRSTRRM GPGRGCGEAGQPLTPARGCAARARPRRLRAAPDRTPPTPGWAVPPFGRAAPLRLLSAAAAGRLPRSSPALSGTAPRPAHRQPPPDLLLPT -------------------------------------------------------------- >51741_51741_5_MARK2-BATF2_MARK2_chr11_63607032_ENST00000402010_BATF2_chr11_64762021_ENST00000301887_length(transcript)=2605nt_BP=633nt AAACAGAGCCGAAGCTTCGTCTTCGGAATCGAGGAGGAGAGCGCGTCCCCGGGGTTTACCTTCTTCGGCTGTTTCCGGAATTCACCCGTA GTCTGTGGCCGGGAGGAGAAGGCGCGCCCCACCTCCCGTTTCTAGCCGCTTCGGATGTTTCCGGCTGCTGCCGGCGAAAAGAGCCGAGGG CCGGCGGTGGTGGCGGCCATGTTGGGAGCAGCAGGTCCGGCGGCGGCTGCCTGTGTGCCGGGCGCGGAGCAGTGCCGCTGAGGGCAGGGG AGGAGCGAGGCAGGCGGCCGGCTGCGGCGGCAGAGAGTAGGCGGAGCGGCGCGGCCCGGCCGAAAGGCGGCACAGCCCAGCCGGGGGTCG GGGGGGTGCGGTCCGGAGCCGCTCGGAGCCGGCGCGGCCTAGCCCGAGCGGCGCATCCCCGGGCTGGCGTGAGCGGCTGCCCGGCCTCCC CGCACCCCCGGCCGGGGCCCATGCGGCGGGTGCTCCTGCTGTGAGAAGCCCCGCCCGGCCGGGCTCCGCGCCTTCCCTTCCCTCCCTTCC TCCAAGCTTCTCGGTTCCCTCCCCCGAGATACCGGCGCCATGTCCAGCGCTCGGACCCCCCTACCCACGCTGAACGAGAGGGACACGGAG CAGGACCCCAAGGAGCAACAAAGGCAGCTGAAGAAGCAGAAGAACCGGGCAGCCGCCCAGCGAAGCCGGCAGAAGCACACAGACAAGGCA GACGCCCTGCACCAGCAGCACGAGTCTCTGGAAAAAGACAACCTCGCCCTGCGGAAGGAGATCCAGTCCCTGCAGGCCGAGCTGGCGTGG TGGAGCCGGACCCTGCACGTGCATGAGCGCCTGTGCCCCATGGATTGTGCCTCCTGCTCAGCTCCAGGGCTCCTGGGCTGCTGGGACCAG GCTGAGGGGCTCCTGGGCCCTGGCCCACAGGGACAACATGGCTGCCGGGAGCAGCTGGAGCTGTTCCAGACCCCGGGTTCCTGTTACCCA GCTCAGCCGCTCTCTCCAGGTCCACAGCCTCATGATTCTCCCAGCCTCCTCCAGTGCCCCCTGCCCTCACTGTCCCTTGGCCCCGCTGTG GTTGCTGAACCTCCTGTCCAGCTGTCCCCCAGCCCTCTCCTGTTTGCCTCGCACACTGGTTCCAGCCTGCAGGGGTCTTCCTCTAAGCTC AGTGCCCTCCAGCCCAGCCTCACGGCCCAAACTGCCCCTCCACAGCCCCTCGAGCTGGAGCATCCCACCAGAGGGAAGCTGGGGTCCTCT CCCGACAACCCTTCCTCTGCCCTGGGGCTTGCACGTCTGCAGAGCAGGGAGCACAAACCTGCTCTCTCAGCAGCCACTTGGCAAGGGCTG GTTGTGGATCCCAGCCCTCACCCTCTCCTGGCCTTTCCTCTGCTCTCCTCTGCTCAAGTCCACTTCTAACCTGGTCTTCGGAGCTGGGTT GGCCCCTTCTTTGGGCTCAGGAAGCAGCCTTAGCACACGGGCCTCTCCTCCCTCACTACTGGGTGCTGCCCTGCGTGGCTGACCAGCTGG CCCAGGATTTCACAGTCGAAAAGGAAGCCACCACTGATGCCTCCCACTGTGACAGGCCCTGTCACCACCAATATCTTATTTCAACCTCAC AGTTGACCTGAGAAATCGAGATTATCACTCCACTTTTTCAGACAAGGAAACTGAGGCTCAGGGAAGCCAAGTGACAAGTCCAAGGTCACG AAGACTTTCTTGGAGCCCGAAACACCACCCTCTGCTCCTCCTTCTCCTGTCCTGGCCCAGGCATCCTAGGGGCTGAAATCCTGGAAACCG TGGGCTGGTGTGAGAAGGTTTGCATGCTCAGAGCAGAGAAGGGCTCTCCCCACTGCTTCGTGATTCCAGGGCCAGAGCCATGCAGTCCCA GAAACCCCAACCTAGCTGGGGCAGGTCCAGAGTCCAAGCCCTGGTGGGTAGAGGCCAAGCAGAAGCCCTGAAGTGGACTCTTGCTTCCCC TAGTAGTGTTTTCAGTGCCAAGAAGCTGAAACTGTGAGCTGGAGTTGGGGAGAGGTCTGGAAGAGGACCATCTGGGATTTCTACAGCCTG GGTACCCATAGCCACACCAAGGCTTCTGGGAGATTCTGCAGGGTCAGCTTTCCAGGCTGTTCCCAAATAGCTCCCTGCCTCCCCACTGCC CCTAAAGCCACAGCAGAAGAGCCATTCATCTCATAAACAAAAAGGAAGAGGAAAGAATGAGGAAGGACCCTGTGCAAGGTTATTTGCAGG CAGGGATGGGCTTGTACCTGACAGCACCCACCCCTGTGTGGCCCCCAGGCCCTCATCACCCTCAGACCCCTCCTAAGCAGTTCCCTCATT GCTCTTTGGACTAGGCTGACAGCAGGAAGAGCAGGGCCCATGACCGGGTGGAAGTTCAGTTTTGGTGTCTGCTTCAAGAGGGGGTTTTAC ACTCTGATTCCAGGACAAGCACTCTGAGGCGGGTGGGGGAGAGAAACCCTGGCTCTTCACCCAGGTTTCACACACATGTAAATGAAACAC >51741_51741_5_MARK2-BATF2_MARK2_chr11_63607032_ENST00000402010_BATF2_chr11_64762021_ENST00000301887_length(amino acids)=317AA_BP=161 MERAAELGNRNPGSGTAPAAPGSHVVPVGQGPGAPQPGPSSPGALELSRRHNPWGTGAHARAGSGSTTPARPAGTGSPSAGRGCLFPETR AAGAGRLPCLCASAGFAGRLPGSSASSAAFVAPWGPAPCPSRSAWVGGSERWTWRRYLGGGNREAWRKGGKGRRGARPGGASHSRSTRRM GPGRGCGEAGQPLTPARGCAARARPRRLRAAPDRTPPTPGWAVPPFGRAAPLRLLSAAAAGRLPRSSPALSGTAPRPAHRQPPPDLLLPT -------------------------------------------------------------- >51741_51741_6_MARK2-BATF2_MARK2_chr11_63607032_ENST00000413835_BATF2_chr11_64762021_ENST00000301887_length(transcript)=2605nt_BP=633nt AAACAGAGCCGAAGCTTCGTCTTCGGAATCGAGGAGGAGAGCGCGTCCCCGGGGTTTACCTTCTTCGGCTGTTTCCGGAATTCACCCGTA GTCTGTGGCCGGGAGGAGAAGGCGCGCCCCACCTCCCGTTTCTAGCCGCTTCGGATGTTTCCGGCTGCTGCCGGCGAAAAGAGCCGAGGG CCGGCGGTGGTGGCGGCCATGTTGGGAGCAGCAGGTCCGGCGGCGGCTGCCTGTGTGCCGGGCGCGGAGCAGTGCCGCTGAGGGCAGGGG AGGAGCGAGGCAGGCGGCCGGCTGCGGCGGCAGAGAGTAGGCGGAGCGGCGCGGCCCGGCCGAAAGGCGGCACAGCCCAGCCGGGGGTCG GGGGGGTGCGGTCCGGAGCCGCTCGGAGCCGGCGCGGCCTAGCCCGAGCGGCGCATCCCCGGGCTGGCGTGAGCGGCTGCCCGGCCTCCC CGCACCCCCGGCCGGGGCCCATGCGGCGGGTGCTCCTGCTGTGAGAAGCCCCGCCCGGCCGGGCTCCGCGCCTTCCCTTCCCTCCCTTCC TCCAAGCTTCTCGGTTCCCTCCCCCGAGATACCGGCGCCATGTCCAGCGCTCGGACCCCCCTACCCACGCTGAACGAGAGGGACACGGAG CAGGACCCCAAGGAGCAACAAAGGCAGCTGAAGAAGCAGAAGAACCGGGCAGCCGCCCAGCGAAGCCGGCAGAAGCACACAGACAAGGCA GACGCCCTGCACCAGCAGCACGAGTCTCTGGAAAAAGACAACCTCGCCCTGCGGAAGGAGATCCAGTCCCTGCAGGCCGAGCTGGCGTGG TGGAGCCGGACCCTGCACGTGCATGAGCGCCTGTGCCCCATGGATTGTGCCTCCTGCTCAGCTCCAGGGCTCCTGGGCTGCTGGGACCAG GCTGAGGGGCTCCTGGGCCCTGGCCCACAGGGACAACATGGCTGCCGGGAGCAGCTGGAGCTGTTCCAGACCCCGGGTTCCTGTTACCCA GCTCAGCCGCTCTCTCCAGGTCCACAGCCTCATGATTCTCCCAGCCTCCTCCAGTGCCCCCTGCCCTCACTGTCCCTTGGCCCCGCTGTG GTTGCTGAACCTCCTGTCCAGCTGTCCCCCAGCCCTCTCCTGTTTGCCTCGCACACTGGTTCCAGCCTGCAGGGGTCTTCCTCTAAGCTC AGTGCCCTCCAGCCCAGCCTCACGGCCCAAACTGCCCCTCCACAGCCCCTCGAGCTGGAGCATCCCACCAGAGGGAAGCTGGGGTCCTCT CCCGACAACCCTTCCTCTGCCCTGGGGCTTGCACGTCTGCAGAGCAGGGAGCACAAACCTGCTCTCTCAGCAGCCACTTGGCAAGGGCTG GTTGTGGATCCCAGCCCTCACCCTCTCCTGGCCTTTCCTCTGCTCTCCTCTGCTCAAGTCCACTTCTAACCTGGTCTTCGGAGCTGGGTT GGCCCCTTCTTTGGGCTCAGGAAGCAGCCTTAGCACACGGGCCTCTCCTCCCTCACTACTGGGTGCTGCCCTGCGTGGCTGACCAGCTGG CCCAGGATTTCACAGTCGAAAAGGAAGCCACCACTGATGCCTCCCACTGTGACAGGCCCTGTCACCACCAATATCTTATTTCAACCTCAC AGTTGACCTGAGAAATCGAGATTATCACTCCACTTTTTCAGACAAGGAAACTGAGGCTCAGGGAAGCCAAGTGACAAGTCCAAGGTCACG AAGACTTTCTTGGAGCCCGAAACACCACCCTCTGCTCCTCCTTCTCCTGTCCTGGCCCAGGCATCCTAGGGGCTGAAATCCTGGAAACCG TGGGCTGGTGTGAGAAGGTTTGCATGCTCAGAGCAGAGAAGGGCTCTCCCCACTGCTTCGTGATTCCAGGGCCAGAGCCATGCAGTCCCA GAAACCCCAACCTAGCTGGGGCAGGTCCAGAGTCCAAGCCCTGGTGGGTAGAGGCCAAGCAGAAGCCCTGAAGTGGACTCTTGCTTCCCC TAGTAGTGTTTTCAGTGCCAAGAAGCTGAAACTGTGAGCTGGAGTTGGGGAGAGGTCTGGAAGAGGACCATCTGGGATTTCTACAGCCTG GGTACCCATAGCCACACCAAGGCTTCTGGGAGATTCTGCAGGGTCAGCTTTCCAGGCTGTTCCCAAATAGCTCCCTGCCTCCCCACTGCC CCTAAAGCCACAGCAGAAGAGCCATTCATCTCATAAACAAAAAGGAAGAGGAAAGAATGAGGAAGGACCCTGTGCAAGGTTATTTGCAGG CAGGGATGGGCTTGTACCTGACAGCACCCACCCCTGTGTGGCCCCCAGGCCCTCATCACCCTCAGACCCCTCCTAAGCAGTTCCCTCATT GCTCTTTGGACTAGGCTGACAGCAGGAAGAGCAGGGCCCATGACCGGGTGGAAGTTCAGTTTTGGTGTCTGCTTCAAGAGGGGGTTTTAC ACTCTGATTCCAGGACAAGCACTCTGAGGCGGGTGGGGGAGAGAAACCCTGGCTCTTCACCCAGGTTTCACACACATGTAAATGAAACAC >51741_51741_6_MARK2-BATF2_MARK2_chr11_63607032_ENST00000413835_BATF2_chr11_64762021_ENST00000301887_length(amino acids)=317AA_BP=161 MERAAELGNRNPGSGTAPAAPGSHVVPVGQGPGAPQPGPSSPGALELSRRHNPWGTGAHARAGSGSTTPARPAGTGSPSAGRGCLFPETR AAGAGRLPCLCASAGFAGRLPGSSASSAAFVAPWGPAPCPSRSAWVGGSERWTWRRYLGGGNREAWRKGGKGRRGARPGGASHSRSTRRM GPGRGCGEAGQPLTPARGCAARARPRRLRAAPDRTPPTPGWAVPPFGRAAPLRLLSAAAAGRLPRSSPALSGTAPRPAHRQPPPDLLLPT -------------------------------------------------------------- >51741_51741_7_MARK2-BATF2_MARK2_chr11_63607032_ENST00000502399_BATF2_chr11_64762021_ENST00000301887_length(transcript)=2238nt_BP=266nt GCGGTCCGGAGCCGCTCGGAGCCGGCGCGGCCTAGCCCGAGCGGCGCATCCCCGGGCTGGCGTGAGCGGCTGCCCGGCCTCCCCGCACCC CCGGCCGGGGCCCATGCGGCGGGTGCTCCTGCTGTGAGAAGCCCCGCCCGGCCGGGCTCCGCGCCTTCCCTTCCCTCCCTTCCTCCAAGC TTCTCGGTTCCCTCCCCCGAGATACCGGCGCCATGTCCAGCGCTCGGACCCCCCTACCCACGCTGAACGAGAGGGACACGGAGCAGGACC CCAAGGAGCAACAAAGGCAGCTGAAGAAGCAGAAGAACCGGGCAGCCGCCCAGCGAAGCCGGCAGAAGCACACAGACAAGGCAGACGCCC TGCACCAGCAGCACGAGTCTCTGGAAAAAGACAACCTCGCCCTGCGGAAGGAGATCCAGTCCCTGCAGGCCGAGCTGGCGTGGTGGAGCC GGACCCTGCACGTGCATGAGCGCCTGTGCCCCATGGATTGTGCCTCCTGCTCAGCTCCAGGGCTCCTGGGCTGCTGGGACCAGGCTGAGG GGCTCCTGGGCCCTGGCCCACAGGGACAACATGGCTGCCGGGAGCAGCTGGAGCTGTTCCAGACCCCGGGTTCCTGTTACCCAGCTCAGC CGCTCTCTCCAGGTCCACAGCCTCATGATTCTCCCAGCCTCCTCCAGTGCCCCCTGCCCTCACTGTCCCTTGGCCCCGCTGTGGTTGCTG AACCTCCTGTCCAGCTGTCCCCCAGCCCTCTCCTGTTTGCCTCGCACACTGGTTCCAGCCTGCAGGGGTCTTCCTCTAAGCTCAGTGCCC TCCAGCCCAGCCTCACGGCCCAAACTGCCCCTCCACAGCCCCTCGAGCTGGAGCATCCCACCAGAGGGAAGCTGGGGTCCTCTCCCGACA ACCCTTCCTCTGCCCTGGGGCTTGCACGTCTGCAGAGCAGGGAGCACAAACCTGCTCTCTCAGCAGCCACTTGGCAAGGGCTGGTTGTGG ATCCCAGCCCTCACCCTCTCCTGGCCTTTCCTCTGCTCTCCTCTGCTCAAGTCCACTTCTAACCTGGTCTTCGGAGCTGGGTTGGCCCCT TCTTTGGGCTCAGGAAGCAGCCTTAGCACACGGGCCTCTCCTCCCTCACTACTGGGTGCTGCCCTGCGTGGCTGACCAGCTGGCCCAGGA TTTCACAGTCGAAAAGGAAGCCACCACTGATGCCTCCCACTGTGACAGGCCCTGTCACCACCAATATCTTATTTCAACCTCACAGTTGAC CTGAGAAATCGAGATTATCACTCCACTTTTTCAGACAAGGAAACTGAGGCTCAGGGAAGCCAAGTGACAAGTCCAAGGTCACGAAGACTT TCTTGGAGCCCGAAACACCACCCTCTGCTCCTCCTTCTCCTGTCCTGGCCCAGGCATCCTAGGGGCTGAAATCCTGGAAACCGTGGGCTG GTGTGAGAAGGTTTGCATGCTCAGAGCAGAGAAGGGCTCTCCCCACTGCTTCGTGATTCCAGGGCCAGAGCCATGCAGTCCCAGAAACCC CAACCTAGCTGGGGCAGGTCCAGAGTCCAAGCCCTGGTGGGTAGAGGCCAAGCAGAAGCCCTGAAGTGGACTCTTGCTTCCCCTAGTAGT GTTTTCAGTGCCAAGAAGCTGAAACTGTGAGCTGGAGTTGGGGAGAGGTCTGGAAGAGGACCATCTGGGATTTCTACAGCCTGGGTACCC ATAGCCACACCAAGGCTTCTGGGAGATTCTGCAGGGTCAGCTTTCCAGGCTGTTCCCAAATAGCTCCCTGCCTCCCCACTGCCCCTAAAG CCACAGCAGAAGAGCCATTCATCTCATAAACAAAAAGGAAGAGGAAAGAATGAGGAAGGACCCTGTGCAAGGTTATTTGCAGGCAGGGAT GGGCTTGTACCTGACAGCACCCACCCCTGTGTGGCCCCCAGGCCCTCATCACCCTCAGACCCCTCCTAAGCAGTTCCCTCATTGCTCTTT GGACTAGGCTGACAGCAGGAAGAGCAGGGCCCATGACCGGGTGGAAGTTCAGTTTTGGTGTCTGCTTCAAGAGGGGGTTTTACACTCTGA TTCCAGGACAAGCACTCTGAGGCGGGTGGGGGAGAGAAACCCTGGCTCTTCACCCAGGTTTCACACACATGTAAATGAAACACTATGTTA >51741_51741_7_MARK2-BATF2_MARK2_chr11_63607032_ENST00000502399_BATF2_chr11_64762021_ENST00000301887_length(amino acids)=279AA_BP=17 MSSARTPLPTLNERDTEQDPKEQQRQLKKQKNRAAAQRSRQKHTDKADALHQQHESLEKDNLALRKEIQSLQAELAWWSRTLHVHERLCP MDCASCSAPGLLGCWDQAEGLLGPGPQGQHGCREQLELFQTPGSCYPAQPLSPGPQPHDSPSLLQCPLPSLSLGPAVVAEPPVQLSPSPL LFASHTGSSLQGSSSKLSALQPSLTAQTAPPQPLELEHPTRGKLGSSPDNPSSALGLARLQSREHKPALSAATWQGLVVDPSPHPLLAFP -------------------------------------------------------------- >51741_51741_8_MARK2-BATF2_MARK2_chr11_63607032_ENST00000508192_BATF2_chr11_64762021_ENST00000301887_length(transcript)=2528nt_BP=556nt GAATTCACCCGTAGTCTGTGGCCGGGAGGAGAAGGCGCGCCCCACCTCCCGTTTCTAGCCGCTTCGGATGTTTCCGGCTGCTGCCGGCGA AAAGAGCCGAGGGCCGGCGGTGGTGGCGGCCATGTTGGGAGCAGCAGGTCCGGCGGCGGCTGCCTGTGTGCCGGGCGCGGAGCAGTGCCG CTGAGGGCAGGGGAGGAGCGAGGCAGGCGGCCGGCTGCGGCGGCAGAGAGTAGGCGGAGCGGCGCGGCCCGGCCGAAAGGCGGCACAGCC CAGCCGGGGGTCGGGGGGGTGCGGTCCGGAGCCGCTCGGAGCCGGCGCGGCCTAGCCCGAGCGGCGCATCCCCGGGCTGGCGTGAGCGGC TGCCCGGCCTCCCCGCACCCCCGGCCGGGGCCCATGCGGCGGGTGCTCCTGCTGTGAGAAGCCCCGCCCGGCCGGGCTCCGCGCCTTCCC TTCCCTCCCTTCCTCCAAGCTTCTCGGTTCCCTCCCCCGAGATACCGGCGCCATGTCCAGCGCTCGGACCCCCCTACCCACGCTGAACGA GAGGGACACGGAGCAGGACCCCAAGGAGCAACAAAGGCAGCTGAAGAAGCAGAAGAACCGGGCAGCCGCCCAGCGAAGCCGGCAGAAGCA CACAGACAAGGCAGACGCCCTGCACCAGCAGCACGAGTCTCTGGAAAAAGACAACCTCGCCCTGCGGAAGGAGATCCAGTCCCTGCAGGC CGAGCTGGCGTGGTGGAGCCGGACCCTGCACGTGCATGAGCGCCTGTGCCCCATGGATTGTGCCTCCTGCTCAGCTCCAGGGCTCCTGGG CTGCTGGGACCAGGCTGAGGGGCTCCTGGGCCCTGGCCCACAGGGACAACATGGCTGCCGGGAGCAGCTGGAGCTGTTCCAGACCCCGGG TTCCTGTTACCCAGCTCAGCCGCTCTCTCCAGGTCCACAGCCTCATGATTCTCCCAGCCTCCTCCAGTGCCCCCTGCCCTCACTGTCCCT TGGCCCCGCTGTGGTTGCTGAACCTCCTGTCCAGCTGTCCCCCAGCCCTCTCCTGTTTGCCTCGCACACTGGTTCCAGCCTGCAGGGGTC TTCCTCTAAGCTCAGTGCCCTCCAGCCCAGCCTCACGGCCCAAACTGCCCCTCCACAGCCCCTCGAGCTGGAGCATCCCACCAGAGGGAA GCTGGGGTCCTCTCCCGACAACCCTTCCTCTGCCCTGGGGCTTGCACGTCTGCAGAGCAGGGAGCACAAACCTGCTCTCTCAGCAGCCAC TTGGCAAGGGCTGGTTGTGGATCCCAGCCCTCACCCTCTCCTGGCCTTTCCTCTGCTCTCCTCTGCTCAAGTCCACTTCTAACCTGGTCT TCGGAGCTGGGTTGGCCCCTTCTTTGGGCTCAGGAAGCAGCCTTAGCACACGGGCCTCTCCTCCCTCACTACTGGGTGCTGCCCTGCGTG GCTGACCAGCTGGCCCAGGATTTCACAGTCGAAAAGGAAGCCACCACTGATGCCTCCCACTGTGACAGGCCCTGTCACCACCAATATCTT ATTTCAACCTCACAGTTGACCTGAGAAATCGAGATTATCACTCCACTTTTTCAGACAAGGAAACTGAGGCTCAGGGAAGCCAAGTGACAA GTCCAAGGTCACGAAGACTTTCTTGGAGCCCGAAACACCACCCTCTGCTCCTCCTTCTCCTGTCCTGGCCCAGGCATCCTAGGGGCTGAA ATCCTGGAAACCGTGGGCTGGTGTGAGAAGGTTTGCATGCTCAGAGCAGAGAAGGGCTCTCCCCACTGCTTCGTGATTCCAGGGCCAGAG CCATGCAGTCCCAGAAACCCCAACCTAGCTGGGGCAGGTCCAGAGTCCAAGCCCTGGTGGGTAGAGGCCAAGCAGAAGCCCTGAAGTGGA CTCTTGCTTCCCCTAGTAGTGTTTTCAGTGCCAAGAAGCTGAAACTGTGAGCTGGAGTTGGGGAGAGGTCTGGAAGAGGACCATCTGGGA TTTCTACAGCCTGGGTACCCATAGCCACACCAAGGCTTCTGGGAGATTCTGCAGGGTCAGCTTTCCAGGCTGTTCCCAAATAGCTCCCTG CCTCCCCACTGCCCCTAAAGCCACAGCAGAAGAGCCATTCATCTCATAAACAAAAAGGAAGAGGAAAGAATGAGGAAGGACCCTGTGCAA GGTTATTTGCAGGCAGGGATGGGCTTGTACCTGACAGCACCCACCCCTGTGTGGCCCCCAGGCCCTCATCACCCTCAGACCCCTCCTAAG CAGTTCCCTCATTGCTCTTTGGACTAGGCTGACAGCAGGAAGAGCAGGGCCCATGACCGGGTGGAAGTTCAGTTTTGGTGTCTGCTTCAA GAGGGGGTTTTACACTCTGATTCCAGGACAAGCACTCTGAGGCGGGTGGGGGAGAGAAACCCTGGCTCTTCACCCAGGTTTCACACACAT GTAAATGAAACACTATGTTAGTATCTAACACACTCCTGGATACAGAACACAAGTCTTGGCACATATGTGATGGAAATAAAGTGTTTTGCA >51741_51741_8_MARK2-BATF2_MARK2_chr11_63607032_ENST00000508192_BATF2_chr11_64762021_ENST00000301887_length(amino acids)=310AA_BP=161 MERAAELGNRNPGSGTAPAAPGSHVVPVGQGPGAPQPGPSSPGALELSRRHNPWGTGAHARAGSGSTTPARPAGTGSPSAGRGCLFPETR AAGAGRLPCLCASAGFAGRLPGSSASSAAFVAPWGPAPCPSRSAWVGGSERWTWRRYLGGGNREAWRKGGKGRRGARPGGASHSRSTRRM GPGRGCGEAGQPLTPARGCAARARPRRLRAAPDRTPPTPGWAVPPFGRAAPLRLLSAAAAGRLPRSSPALSGTAPRPAHRQPPPDLLLPT -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MARK2-BATF2 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MARK2-BATF2 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MARK2-BATF2 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
