
|
||||||
|
 | Fusion Gene Summary |
 | Fusion Gene ORF analysis |
 | Fusion Genomic Features |
 | Fusion Protein Features |
 | Fusion Gene Sequence |
 | Fusion Gene PPI analysis |
 | Related Drugs |
 | Related Diseases |
Fusion gene:MARK3-FRG1 (FusionGDB2 ID:51767) |
Fusion Gene Summary for MARK3-FRG1 |
 Fusion gene summary Fusion gene summary |
| Fusion gene information | Fusion gene name: MARK3-FRG1 | Fusion gene ID: 51767 | Hgene | Tgene | Gene symbol | MARK3 | FRG1 | Gene ID | 4140 | 2483 |
| Gene name | microtubule affinity regulating kinase 3 | FSHD region gene 1 | |
| Synonyms | CTAK1|KP78|PAR1A|Par-1a|VIPB | FRG1A|FSG1 | |
| Cytomap | 14q32.32-q32.33 | 4q35.2 | |
| Type of gene | protein-coding | protein-coding | |
| Description | MAP/microtubule affinity-regulating kinase 3C-TAK1ELKL motif kinase 2EMK-2cdc25C-associated protein kinase 1protein kinase STK10ser/Thr protein kinase PAR-1serine/threonine-protein kinase p78 | protein FRG1FSHD region gene 1 proteinfacioscapulohumeral muscular dystrophy region gene-1 | |
| Modification date | 20200313 | 20200313 | |
| UniProtAcc | P27448 | Q14331 | |
| Ensembl transtripts involved in fusion gene | ENST00000561071, ENST00000216288, ENST00000303622, ENST00000335102, ENST00000416682, ENST00000429436, ENST00000440884, ENST00000553942, | ENST00000226798, ENST00000514482, | |
| Fusion gene scores | * DoF score | 23 X 15 X 10=3450 | 5 X 3 X 5=75 |
| # samples | 25 | 5 | |
| ** MAII score | log2(25/3450*10)=-3.78659636189081 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | log2(5/75*10)=-0.584962500721156 possibly effective Gene in Pan-Cancer Fusion Genes (peGinPCFGs). DoF>8 and MAII<0 | |
| Context | PubMed: MARK3 [Title/Abstract] AND FRG1 [Title/Abstract] AND fusion [Title/Abstract] | ||
| Most frequent breakpoint | MARK3(103894777)-FRG1(190878553), # samples:1 | ||
| Anticipated loss of major functional domain due to fusion event. | |||
| * DoF score (Degree of Frequency) = # partners X # break points X # cancer types ** MAII score (Major Active Isofusion Index) = log2(# samples/DoF score*10) |
 Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez Gene ontology of each fusion partner gene with evidence of Inferred from Direct Assay (IDA) from Entrez |
| Partner | Gene | GO ID | GO term | PubMed ID |
| Hgene | MARK3 | GO:0018105 | peptidyl-serine phosphorylation | 9543386 |
| Hgene | MARK3 | GO:0032092 | positive regulation of protein binding | 9543386 |
| Hgene | MARK3 | GO:0035331 | negative regulation of hippo signaling | 28087714 |
| Hgene | MARK3 | GO:0036289 | peptidyl-serine autophosphorylation | 9543386 |
 Fusion gene breakpoints across MARK3 (5'-gene) Fusion gene breakpoints across MARK3 (5'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
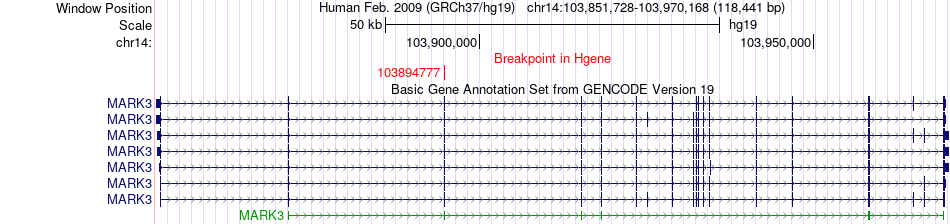 |
 Fusion gene breakpoints across FRG1 (3'-gene) Fusion gene breakpoints across FRG1 (3'-gene)* Click on the image to open the UCSC genome browser with custom track showing this image in a new window. |
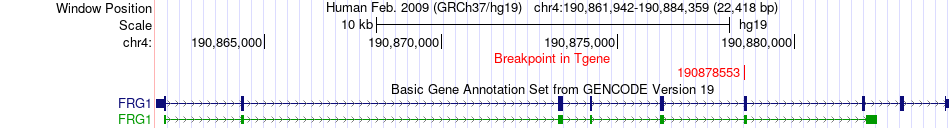 |
 Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0) Fusion gene information from two resources (ChiTars 5.0 and ChimerDB 4.0)* All genome coordinats were lifted-over on hg19. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| Source | Disease | Sample | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| ChimerDB4 | BRCA | TCGA-A8-A06U-01A | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
Top |
Fusion Gene ORF analysis for MARK3-FRG1 |
 Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. Open reading frame (ORF) analsis of fusion genes based on Ensembl gene isoform structure. * Click on the break point to see the gene structure around the break point region using the UCSC Genome Browser. |
| ORF | Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand |
| 3UTR-3CDS | ENST00000561071 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| 3UTR-3UTR | ENST00000561071 | ENST00000514482 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| 5CDS-3UTR | ENST00000216288 | ENST00000514482 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| 5CDS-3UTR | ENST00000303622 | ENST00000514482 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| 5CDS-3UTR | ENST00000335102 | ENST00000514482 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| 5CDS-3UTR | ENST00000416682 | ENST00000514482 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| 5CDS-3UTR | ENST00000429436 | ENST00000514482 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| 5CDS-3UTR | ENST00000440884 | ENST00000514482 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| 5CDS-3UTR | ENST00000553942 | ENST00000514482 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| In-frame | ENST00000216288 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| In-frame | ENST00000303622 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| In-frame | ENST00000335102 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| In-frame | ENST00000416682 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| In-frame | ENST00000429436 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| In-frame | ENST00000440884 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
| In-frame | ENST00000553942 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + |
 ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. ORFfinder result based on the fusion transcript sequence of in-frame fusion genes. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | Seq length (transcript) | BP loci (transcript) | Predicted start (transcript) | Predicted stop (transcript) | Seq length (amino acids) |
| ENST00000440884 | MARK3 | chr14 | 103894777 | + | ENST00000226798 | FRG1 | chr4 | 190878553 | + | 1355 | 935 | 638 | 1279 | 213 |
| ENST00000416682 | MARK3 | chr14 | 103894777 | + | ENST00000226798 | FRG1 | chr4 | 190878553 | + | 1294 | 874 | 577 | 1218 | 213 |
| ENST00000429436 | MARK3 | chr14 | 103894777 | + | ENST00000226798 | FRG1 | chr4 | 190878553 | + | 1227 | 807 | 510 | 1151 | 213 |
| ENST00000303622 | MARK3 | chr14 | 103894777 | + | ENST00000226798 | FRG1 | chr4 | 190878553 | + | 1206 | 786 | 489 | 1130 | 213 |
| ENST00000216288 | MARK3 | chr14 | 103894777 | + | ENST00000226798 | FRG1 | chr4 | 190878553 | + | 887 | 467 | 170 | 811 | 213 |
| ENST00000553942 | MARK3 | chr14 | 103894777 | + | ENST00000226798 | FRG1 | chr4 | 190878553 | + | 762 | 342 | 45 | 686 | 213 |
| ENST00000335102 | MARK3 | chr14 | 103894777 | + | ENST00000226798 | FRG1 | chr4 | 190878553 | + | 717 | 297 | 0 | 641 | 213 |
 DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. DeepORF prediction of the coding potential based on the fusion transcript sequence of in-frame fusion genes. DeepORF is a coding potential classifier based on convolutional neural network by comparing the real Ribo-seq data. If the no-coding score < 0.5 and coding score > 0.5, then the in-frame fusion transcript is predicted as being likely translated. |
| Henst | Tenst | Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | No-coding score | Coding score |
| ENST00000440884 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + | 0.00771416 | 0.99228585 |
| ENST00000416682 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + | 0.007659427 | 0.99234056 |
| ENST00000429436 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + | 0.005194881 | 0.99480516 |
| ENST00000303622 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + | 0.00615212 | 0.9938479 |
| ENST00000216288 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + | 0.005921984 | 0.99407804 |
| ENST00000553942 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + | 0.011285818 | 0.98871416 |
| ENST00000335102 | ENST00000226798 | MARK3 | chr14 | 103894777 | + | FRG1 | chr4 | 190878553 | + | 0.012265214 | 0.98773474 |
Top |
Fusion Genomic Features for MARK3-FRG1 |
 FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. FusionAI prediction of the potential fusion gene breakpoint based on the pre-mature RNA sequence context (+/- 5kb of individual partner genes, total 20kb length sequence). FusionAI is a fusion gene breakpoint classifier based on convolutional neural network by comparing the fusion positive and negative sequence context of ~ 20K fusion gene data. From here, we can have the relative potentency of the 20K genomic sequence how individual sequnce will be likely used as the gene fusion breakpoints. |
| Hgene | Hchr | Hbp | Hstrand | Tgene | Tchr | Tbp | Tstrand | 1-p | p (fusion gene breakpoint) |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions. We integrated a total of 44 different types of human genomic feature loci information across five big categories including virus integration sites, repeats, structural variants, chromatin states, and gene expression regulation. More details are in help page. |
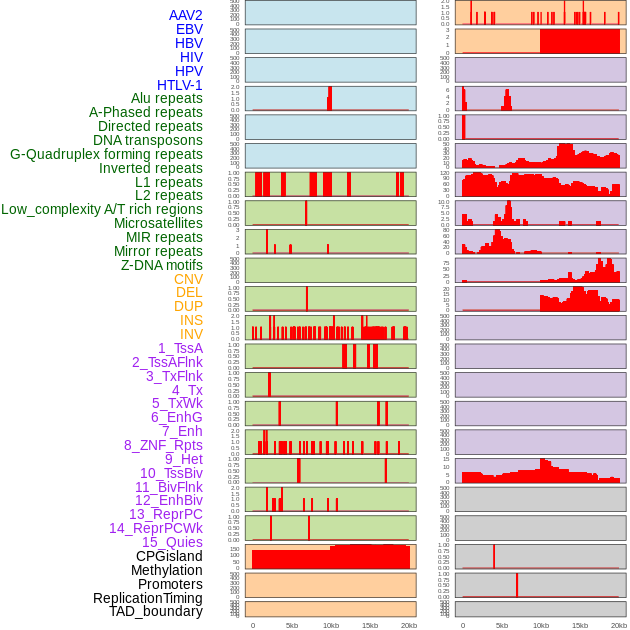 |
 Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. Distribution of 44 human genomic features loci across 20kb length fusion breakpoint regions that are ovelapped with the top 1% feature importance score regions. More details are in help page. |
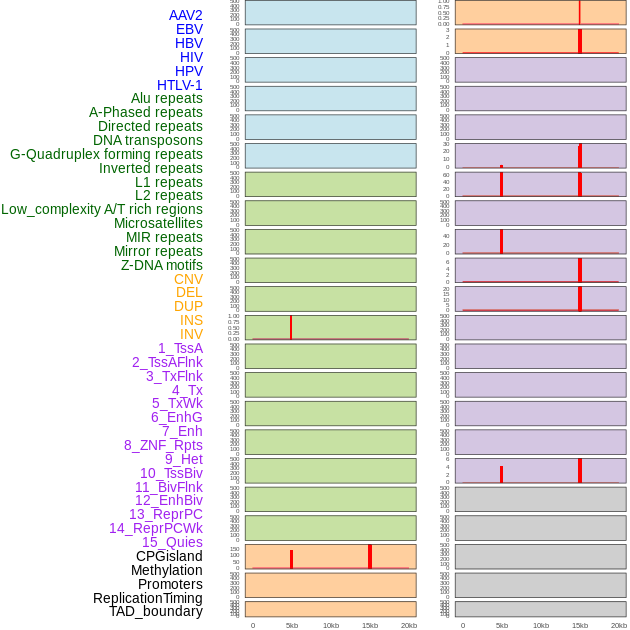 |
Top |
Fusion Protein Features for MARK3-FRG1 |
 Four levels of functional features of fusion genes Four levels of functional features of fusion genesGo to FGviewer search page for the most frequent breakpoint (https://ccsmweb.uth.edu/FGviewer/chr14:103894777/chr4:190878553) - FGviewer provides the online visualization of the retention search of the protein functional features across DNA, RNA, protein, and pathological levels. - How to search 1. Put your fusion gene symbol. 2. Press the tab key until there will be shown the breakpoint information filled. 4. Go down and press 'Search' tab twice. 4. Go down to have the hyperlink of the search result. 5. Click the hyperlink. 6. See the FGviewer result for your fusion gene. |
 |
 Main function of each fusion partner protein. (from UniProt) Main function of each fusion partner protein. (from UniProt) |
| Hgene | Tgene |
| MARK3 | FRG1 |
| FUNCTION: Serine/threonine-protein kinase (PubMed:23666762). Involved in the specific phosphorylation of microtubule-associated proteins for MAP2 and MAP4. Phosphorylates the microtubule-associated protein MAPT/TAU (PubMed:23666762). Phosphorylates CDC25C on 'Ser-216'. Regulates localization and activity of some histone deacetylases by mediating phosphorylation of HDAC7, promoting subsequent interaction between HDAC7 and 14-3-3 and export from the nucleus (PubMed:16980613). Negatively regulates the Hippo signaling pathway and antagonizes the phosphorylation of LATS1. Cooperates with DLG5 to inhibit the kinase activity of STK3/MST2 toward LATS1 (PubMed:28087714). {ECO:0000269|PubMed:16980613, ECO:0000269|PubMed:23666762, ECO:0000269|PubMed:28087714}. | FUNCTION: Binds to mRNA in a sequence-independent manner. May play a role in regulation of pre-mRNA splicing or in the assembly of rRNA into ribosomal subunits. May be involved in mRNA transport. May be involved in epigenetic regulation of muscle differentiation through regulation of activity of the histone-lysine N-methyltransferase KMT5B. {ECO:0000269|PubMed:11991638, ECO:0000269|PubMed:15060122, ECO:0000269|PubMed:20970242, ECO:0000269|PubMed:21699900, ECO:0000269|PubMed:23720823}. |
 Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at Retention analysis result of each fusion partner protein across 39 protein features of UniProt such as six molecule processing features, 13 region features, four site features, six amino acid modification features, two natural variation features, five experimental info features, and 3 secondary structure features. Here, because of limited space for viewing, we only show the protein feature retention information belong to the 13 regional features. All retention annotation result can be downloaded at * Minus value of BPloci means that the break pointn is located before the CDS. |
| - In-frame and retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000216288 | + | 3 | 16 | 62_70 | 99 | 714.0 | Nucleotide binding | ATP |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000303622 | + | 3 | 16 | 62_70 | 99 | 730.0 | Nucleotide binding | ATP |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000416682 | + | 3 | 17 | 62_70 | 99 | 753.0 | Nucleotide binding | ATP |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000429436 | + | 3 | 18 | 62_70 | 99 | 754.0 | Nucleotide binding | ATP |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000440884 | + | 3 | 16 | 62_70 | 99 | 660.0 | Nucleotide binding | ATP |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000553942 | + | 3 | 17 | 62_70 | 99 | 745.0 | Nucleotide binding | ATP |
| Tgene | FRG1 | chr14:103894777 | chr4:190878553 | ENST00000226798 | 4 | 9 | 235_251 | 144 | 259.0 | Motif | Bipartite nuclear localization signal |
| - In-frame and not-retained protein feature among the 13 regional features. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Protein feature | Protein feature note |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000216288 | + | 3 | 16 | 326_365 | 99 | 714.0 | Domain | UBA |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000216288 | + | 3 | 16 | 56_307 | 99 | 714.0 | Domain | Protein kinase |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000216288 | + | 3 | 16 | 704_753 | 99 | 714.0 | Domain | KA1 |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000303622 | + | 3 | 16 | 326_365 | 99 | 730.0 | Domain | UBA |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000303622 | + | 3 | 16 | 56_307 | 99 | 730.0 | Domain | Protein kinase |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000303622 | + | 3 | 16 | 704_753 | 99 | 730.0 | Domain | KA1 |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000416682 | + | 3 | 17 | 326_365 | 99 | 753.0 | Domain | UBA |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000416682 | + | 3 | 17 | 56_307 | 99 | 753.0 | Domain | Protein kinase |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000416682 | + | 3 | 17 | 704_753 | 99 | 753.0 | Domain | KA1 |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000429436 | + | 3 | 18 | 326_365 | 99 | 754.0 | Domain | UBA |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000429436 | + | 3 | 18 | 56_307 | 99 | 754.0 | Domain | Protein kinase |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000429436 | + | 3 | 18 | 704_753 | 99 | 754.0 | Domain | KA1 |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000440884 | + | 3 | 16 | 326_365 | 99 | 660.0 | Domain | UBA |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000440884 | + | 3 | 16 | 56_307 | 99 | 660.0 | Domain | Protein kinase |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000440884 | + | 3 | 16 | 704_753 | 99 | 660.0 | Domain | KA1 |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000553942 | + | 3 | 17 | 326_365 | 99 | 745.0 | Domain | UBA |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000553942 | + | 3 | 17 | 56_307 | 99 | 745.0 | Domain | Protein kinase |
| Hgene | MARK3 | chr14:103894777 | chr4:190878553 | ENST00000553942 | + | 3 | 17 | 704_753 | 99 | 745.0 | Domain | KA1 |
| Tgene | FRG1 | chr14:103894777 | chr4:190878553 | ENST00000226798 | 4 | 9 | 8_32 | 144 | 259.0 | Compositional bias | Note=Lys-rich | |
| Tgene | FRG1 | chr14:103894777 | chr4:190878553 | ENST00000226798 | 4 | 9 | 22_32 | 144 | 259.0 | Motif | Nuclear localization signal |
Top |
Fusion Gene Sequence for MARK3-FRG1 |
 For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. For in-frame fusion transcripts, we provide the fusion transcript sequences and fusion amino acid sequences. To have fusion amino acid sequence, we ran ORFfinder and chose the longest ORF among the all predicted ones. |
| >51767_51767_1_MARK3-FRG1_MARK3_chr14_103894777_ENST00000216288_FRG1_chr4_190878553_ENST00000226798_length(transcript)=887nt_BP=467nt GACGGCCCGGGCCAGGCCCGGGATCTAGACGGCCGTAGGGGGAAGGGAGCCGCCCTCCCCACGGCGCCTTTTCGGAACTGCCGTGGACTC GAGGACGCTGGTCGCCGGCCTCCTAGGGCTGTGCTGTTTTGTTTTGACCCTCGCATTGTGCAGAATTAAAGTGCAGTAAAATGTCCACTA GGACCCCATTGCCAACGGTGAATGAACGAGACACTGAAAACCACACGTCACATGGAGATGGGCGTCAAGAAGTTACCTCTCGTACCAGCC GCTCAGGAGCTCGGTGTAGAAACTCTATAGCCTCCTGTGCAGATGAACAACCTCACATCGGAAACTACAGACTGTTGAAAACAATCGGCA AGGGGAATTTTGCAAAAGTAAAATTGGCAAGACATATCCTTACAGGCAGAGAGGTTGCAATAAAAATAATTGACAAAACTCAGTTGAATC CAACAAGTCTACAAAAGGGGAAAATGGCTTTGTTGGCCTCAAATAGCTGCTTTATTAGATGCAATGAAGCAGGGGACATAGAAGCAAAAA GTAAAACAGCAGGAGAAGAAGAAATGATCAAGATTAGATCCTGTGCTGAAAGAGAAACCAAGAAAAAAGATGACATTCCAGAAGAAGACA AAGGAAATGTAAAACAATGTGAAATCAATTATGTAAAGAAATTTCAGAGCTTCCAAGACCACAAACTTAAAATAAGTAAAGAAGACAGTA AAATTCTTAAAAAGGCTCGGAAAGATGGATTTTTGCATGAGACGCTTCTGGACAGGAGAGCCAAATTGAAAGCCGACAGATACTGCAAGT >51767_51767_1_MARK3-FRG1_MARK3_chr14_103894777_ENST00000216288_FRG1_chr4_190878553_ENST00000226798_length(amino acids)=213AA_BP=99 MSTRTPLPTVNERDTENHTSHGDGRQEVTSRTSRSGARCRNSIASCADEQPHIGNYRLLKTIGKGNFAKVKLARHILTGREVAIKIIDKT QLNPTSLQKGKMALLASNSCFIRCNEAGDIEAKSKTAGEEEMIKIRSCAERETKKKDDIPEEDKGNVKQCEINYVKKFQSFQDHKLKISK -------------------------------------------------------------- >51767_51767_2_MARK3-FRG1_MARK3_chr14_103894777_ENST00000303622_FRG1_chr4_190878553_ENST00000226798_length(transcript)=1206nt_BP=786nt GCTCCATGGTCGGCGCCCACAGCCCGCGGCGGCCTGTCTTGCGCTCCACTTCCTTCACATCCTCCTCCGCCTCCTCGTTTTCAGGCGCCG CCGGCGGCGCTGTGTGGAGGCCCGCGAGCTGAAATTCGCGGTGCGACGGGAGGGAGTGGAGAAGGAGGTGAGGGGGCCCAGGATCGCGGG GCGCCCTGAGGCAAGGGGACGCCGGCGGGCCGAAGCGCAGCCCGCCGCCCGCAGGCTCGGCTCCGCCACTGCCGCCCTCCCGGTCTCCTC GCCTCGGCCGCCGAGGCAGGGAGAGAATGAGCCCCGGGACCCGCCGGGGGACGGCCCGGGCCAGGCCCGGGATCTAGACGGCCGTAGGGG GAAGGGAGCCGCCCTCCCCACGGCGCCTTTTCGGAACTGCCGTGGACTCGAGGACGCTGGTCGCCGGCCTCCTAGGGCTGTGCTGTTTTG TTTTGACCCTCGCATTGTGCAGAATTAAAGTGCAGTAAAATGTCCACTAGGACCCCATTGCCAACGGTGAATGAACGAGACACTGAAAAC CACACGTCACATGGAGATGGGCGTCAAGAAGTTACCTCTCGTACCAGCCGCTCAGGAGCTCGGTGTAGAAACTCTATAGCCTCCTGTGCA GATGAACAACCTCACATCGGAAACTACAGACTGTTGAAAACAATCGGCAAGGGGAATTTTGCAAAAGTAAAATTGGCAAGACATATCCTT ACAGGCAGAGAGGTTGCAATAAAAATAATTGACAAAACTCAGTTGAATCCAACAAGTCTACAAAAGGGGAAAATGGCTTTGTTGGCCTCA AATAGCTGCTTTATTAGATGCAATGAAGCAGGGGACATAGAAGCAAAAAGTAAAACAGCAGGAGAAGAAGAAATGATCAAGATTAGATCC TGTGCTGAAAGAGAAACCAAGAAAAAAGATGACATTCCAGAAGAAGACAAAGGAAATGTAAAACAATGTGAAATCAATTATGTAAAGAAA TTTCAGAGCTTCCAAGACCACAAACTTAAAATAAGTAAAGAAGACAGTAAAATTCTTAAAAAGGCTCGGAAAGATGGATTTTTGCATGAG ACGCTTCTGGACAGGAGAGCCAAATTGAAAGCCGACAGATACTGCAAGTGACTGGGATTTTTGTTTCTGCCTTATCTTTCTGTGTTTTTT >51767_51767_2_MARK3-FRG1_MARK3_chr14_103894777_ENST00000303622_FRG1_chr4_190878553_ENST00000226798_length(amino acids)=213AA_BP=99 MSTRTPLPTVNERDTENHTSHGDGRQEVTSRTSRSGARCRNSIASCADEQPHIGNYRLLKTIGKGNFAKVKLARHILTGREVAIKIIDKT QLNPTSLQKGKMALLASNSCFIRCNEAGDIEAKSKTAGEEEMIKIRSCAERETKKKDDIPEEDKGNVKQCEINYVKKFQSFQDHKLKISK -------------------------------------------------------------- >51767_51767_3_MARK3-FRG1_MARK3_chr14_103894777_ENST00000335102_FRG1_chr4_190878553_ENST00000226798_length(transcript)=717nt_BP=297nt ATGTCCACTAGGACCCCATTGCCAACGGTGAATGAACGAGACACTGAAAACCACACGTCACATGGAGATGGGCGTCAAGAAGTTACCTCT CGTACCAGCCGCTCAGGAGCTCGGTGTAGAAACTCTATAGCCTCCTGTGCAGATGAACAACCTCACATCGGAAACTACAGACTGTTGAAA ACAATCGGCAAGGGGAATTTTGCAAAAGTAAAATTGGCAAGACATATCCTTACAGGCAGAGAGGTTGCAATAAAAATAATTGACAAAACT CAGTTGAATCCAACAAGTCTACAAAAGGGGAAAATGGCTTTGTTGGCCTCAAATAGCTGCTTTATTAGATGCAATGAAGCAGGGGACATA GAAGCAAAAAGTAAAACAGCAGGAGAAGAAGAAATGATCAAGATTAGATCCTGTGCTGAAAGAGAAACCAAGAAAAAAGATGACATTCCA GAAGAAGACAAAGGAAATGTAAAACAATGTGAAATCAATTATGTAAAGAAATTTCAGAGCTTCCAAGACCACAAACTTAAAATAAGTAAA GAAGACAGTAAAATTCTTAAAAAGGCTCGGAAAGATGGATTTTTGCATGAGACGCTTCTGGACAGGAGAGCCAAATTGAAAGCCGACAGA >51767_51767_3_MARK3-FRG1_MARK3_chr14_103894777_ENST00000335102_FRG1_chr4_190878553_ENST00000226798_length(amino acids)=213AA_BP=99 MSTRTPLPTVNERDTENHTSHGDGRQEVTSRTSRSGARCRNSIASCADEQPHIGNYRLLKTIGKGNFAKVKLARHILTGREVAIKIIDKT QLNPTSLQKGKMALLASNSCFIRCNEAGDIEAKSKTAGEEEMIKIRSCAERETKKKDDIPEEDKGNVKQCEINYVKKFQSFQDHKLKISK -------------------------------------------------------------- >51767_51767_4_MARK3-FRG1_MARK3_chr14_103894777_ENST00000416682_FRG1_chr4_190878553_ENST00000226798_length(transcript)=1294nt_BP=874nt GCAGCGGCGGCCAGCAGGGCGGAGGCTGAGGCAGCAAGCTCGCTAGAGAGGGAGAAGCAGTCGGGCGCAGGCGCCTCCTCCGCAGCCCGC TCCATGGTCGGCGCCCACAGCCCGCGGCGGCCTGTCTTGCGCTCCACTTCCTTCACATCCTCCTCCGCCTCCTCGTTTTCAGGCGCCGCC GGCGGCGCTGTGTGGAGGCCCGCGAGCTGAAATTCGCGGTGCGACGGGAGGGAGTGGAGAAGGAGGTGAGGGGGCCCAGGATCGCGGGGC GCCCTGAGGCAAGGGGACGCCGGCGGGCCGAAGCGCAGCCCGCCGCCCGCAGGCTCGGCTCCGCCACTGCCGCCCTCCCGGTCTCCTCGC CTCGGCCGCCGAGGCAGGGAGAGAATGAGCCCCGGGACCCGCCGGGGGACGGCCCGGGCCAGGCCCGGGATCTAGACGGCCGTAGGGGGA AGGGAGCCGCCCTCCCCACGGCGCCTTTTCGGAACTGCCGTGGACTCGAGGACGCTGGTCGCCGGCCTCCTAGGGCTGTGCTGTTTTGTT TTGACCCTCGCATTGTGCAGAATTAAAGTGCAGTAAAATGTCCACTAGGACCCCATTGCCAACGGTGAATGAACGAGACACTGAAAACCA CACGTCACATGGAGATGGGCGTCAAGAAGTTACCTCTCGTACCAGCCGCTCAGGAGCTCGGTGTAGAAACTCTATAGCCTCCTGTGCAGA TGAACAACCTCACATCGGAAACTACAGACTGTTGAAAACAATCGGCAAGGGGAATTTTGCAAAAGTAAAATTGGCAAGACATATCCTTAC AGGCAGAGAGGTTGCAATAAAAATAATTGACAAAACTCAGTTGAATCCAACAAGTCTACAAAAGGGGAAAATGGCTTTGTTGGCCTCAAA TAGCTGCTTTATTAGATGCAATGAAGCAGGGGACATAGAAGCAAAAAGTAAAACAGCAGGAGAAGAAGAAATGATCAAGATTAGATCCTG TGCTGAAAGAGAAACCAAGAAAAAAGATGACATTCCAGAAGAAGACAAAGGAAATGTAAAACAATGTGAAATCAATTATGTAAAGAAATT TCAGAGCTTCCAAGACCACAAACTTAAAATAAGTAAAGAAGACAGTAAAATTCTTAAAAAGGCTCGGAAAGATGGATTTTTGCATGAGAC GCTTCTGGACAGGAGAGCCAAATTGAAAGCCGACAGATACTGCAAGTGACTGGGATTTTTGTTTCTGCCTTATCTTTCTGTGTTTTTTTC >51767_51767_4_MARK3-FRG1_MARK3_chr14_103894777_ENST00000416682_FRG1_chr4_190878553_ENST00000226798_length(amino acids)=213AA_BP=99 MSTRTPLPTVNERDTENHTSHGDGRQEVTSRTSRSGARCRNSIASCADEQPHIGNYRLLKTIGKGNFAKVKLARHILTGREVAIKIIDKT QLNPTSLQKGKMALLASNSCFIRCNEAGDIEAKSKTAGEEEMIKIRSCAERETKKKDDIPEEDKGNVKQCEINYVKKFQSFQDHKLKISK -------------------------------------------------------------- >51767_51767_5_MARK3-FRG1_MARK3_chr14_103894777_ENST00000429436_FRG1_chr4_190878553_ENST00000226798_length(transcript)=1227nt_BP=807nt CAGGCGCCTCCTCCGCAGCCCGCTCCATGGTCGGCGCCCACAGCCCGCGGCGGCCTGTCTTGCGCTCCACTTCCTTCACATCCTCCTCCG CCTCCTCGTTTTCAGGCGCCGCCGGCGGCGCTGTGTGGAGGCCCGCGAGCTGAAATTCGCGGTGCGACGGGAGGGAGTGGAGAAGGAGGT GAGGGGGCCCAGGATCGCGGGGCGCCCTGAGGCAAGGGGACGCCGGCGGGCCGAAGCGCAGCCCGCCGCCCGCAGGCTCGGCTCCGCCAC TGCCGCCCTCCCGGTCTCCTCGCCTCGGCCGCCGAGGCAGGGAGAGAATGAGCCCCGGGACCCGCCGGGGGACGGCCCGGGCCAGGCCCG GGATCTAGACGGCCGTAGGGGGAAGGGAGCCGCCCTCCCCACGGCGCCTTTTCGGAACTGCCGTGGACTCGAGGACGCTGGTCGCCGGCC TCCTAGGGCTGTGCTGTTTTGTTTTGACCCTCGCATTGTGCAGAATTAAAGTGCAGTAAAATGTCCACTAGGACCCCATTGCCAACGGTG AATGAACGAGACACTGAAAACCACACGTCACATGGAGATGGGCGTCAAGAAGTTACCTCTCGTACCAGCCGCTCAGGAGCTCGGTGTAGA AACTCTATAGCCTCCTGTGCAGATGAACAACCTCACATCGGAAACTACAGACTGTTGAAAACAATCGGCAAGGGGAATTTTGCAAAAGTA AAATTGGCAAGACATATCCTTACAGGCAGAGAGGTTGCAATAAAAATAATTGACAAAACTCAGTTGAATCCAACAAGTCTACAAAAGGGG AAAATGGCTTTGTTGGCCTCAAATAGCTGCTTTATTAGATGCAATGAAGCAGGGGACATAGAAGCAAAAAGTAAAACAGCAGGAGAAGAA GAAATGATCAAGATTAGATCCTGTGCTGAAAGAGAAACCAAGAAAAAAGATGACATTCCAGAAGAAGACAAAGGAAATGTAAAACAATGT GAAATCAATTATGTAAAGAAATTTCAGAGCTTCCAAGACCACAAACTTAAAATAAGTAAAGAAGACAGTAAAATTCTTAAAAAGGCTCGG AAAGATGGATTTTTGCATGAGACGCTTCTGGACAGGAGAGCCAAATTGAAAGCCGACAGATACTGCAAGTGACTGGGATTTTTGTTTCTG >51767_51767_5_MARK3-FRG1_MARK3_chr14_103894777_ENST00000429436_FRG1_chr4_190878553_ENST00000226798_length(amino acids)=213AA_BP=99 MSTRTPLPTVNERDTENHTSHGDGRQEVTSRTSRSGARCRNSIASCADEQPHIGNYRLLKTIGKGNFAKVKLARHILTGREVAIKIIDKT QLNPTSLQKGKMALLASNSCFIRCNEAGDIEAKSKTAGEEEMIKIRSCAERETKKKDDIPEEDKGNVKQCEINYVKKFQSFQDHKLKISK -------------------------------------------------------------- >51767_51767_6_MARK3-FRG1_MARK3_chr14_103894777_ENST00000440884_FRG1_chr4_190878553_ENST00000226798_length(transcript)=1355nt_BP=935nt GGAAGCCGAGGGGCGGCCATCTTGGCTCCGTGAGGCTCTGAGGTGCCGGGGTGCGGCGGCGGCAGCGGCGGCCAGCAGGGCGGAGGCTGA GGCAGCAAGCTCGCTAGAGAGGGAGAAGCAGTCGGGCGCAGGCGCCTCCTCCGCAGCCCGCTCCATGGTCGGCGCCCACAGCCCGCGGCG GCCTGTCTTGCGCTCCACTTCCTTCACATCCTCCTCCGCCTCCTCGTTTTCAGGCGCCGCCGGCGGCGCTGTGTGGAGGCCCGCGAGCTG AAATTCGCGGTGCGACGGGAGGGAGTGGAGAAGGAGGTGAGGGGGCCCAGGATCGCGGGGCGCCCTGAGGCAAGGGGACGCCGGCGGGCC GAAGCGCAGCCCGCCGCCCGCAGGCTCGGCTCCGCCACTGCCGCCCTCCCGGTCTCCTCGCCTCGGCCGCCGAGGCAGGGAGAGAATGAG CCCCGGGACCCGCCGGGGGACGGCCCGGGCCAGGCCCGGGATCTAGACGGCCGTAGGGGGAAGGGAGCCGCCCTCCCCACGGCGCCTTTT CGGAACTGCCGTGGACTCGAGGACGCTGGTCGCCGGCCTCCTAGGGCTGTGCTGTTTTGTTTTGACCCTCGCATTGTGCAGAATTAAAGT GCAGTAAAATGTCCACTAGGACCCCATTGCCAACGGTGAATGAACGAGACACTGAAAACCACACGTCACATGGAGATGGGCGTCAAGAAG TTACCTCTCGTACCAGCCGCTCAGGAGCTCGGTGTAGAAACTCTATAGCCTCCTGTGCAGATGAACAACCTCACATCGGAAACTACAGAC TGTTGAAAACAATCGGCAAGGGGAATTTTGCAAAAGTAAAATTGGCAAGACATATCCTTACAGGCAGAGAGGTTGCAATAAAAATAATTG ACAAAACTCAGTTGAATCCAACAAGTCTACAAAAGGGGAAAATGGCTTTGTTGGCCTCAAATAGCTGCTTTATTAGATGCAATGAAGCAG GGGACATAGAAGCAAAAAGTAAAACAGCAGGAGAAGAAGAAATGATCAAGATTAGATCCTGTGCTGAAAGAGAAACCAAGAAAAAAGATG ACATTCCAGAAGAAGACAAAGGAAATGTAAAACAATGTGAAATCAATTATGTAAAGAAATTTCAGAGCTTCCAAGACCACAAACTTAAAA TAAGTAAAGAAGACAGTAAAATTCTTAAAAAGGCTCGGAAAGATGGATTTTTGCATGAGACGCTTCTGGACAGGAGAGCCAAATTGAAAG CCGACAGATACTGCAAGTGACTGGGATTTTTGTTTCTGCCTTATCTTTCTGTGTTTTTTTCTGAATAAAATATTCAGAGGAAATGCTTTT >51767_51767_6_MARK3-FRG1_MARK3_chr14_103894777_ENST00000440884_FRG1_chr4_190878553_ENST00000226798_length(amino acids)=213AA_BP=99 MSTRTPLPTVNERDTENHTSHGDGRQEVTSRTSRSGARCRNSIASCADEQPHIGNYRLLKTIGKGNFAKVKLARHILTGREVAIKIIDKT QLNPTSLQKGKMALLASNSCFIRCNEAGDIEAKSKTAGEEEMIKIRSCAERETKKKDDIPEEDKGNVKQCEINYVKKFQSFQDHKLKISK -------------------------------------------------------------- >51767_51767_7_MARK3-FRG1_MARK3_chr14_103894777_ENST00000553942_FRG1_chr4_190878553_ENST00000226798_length(transcript)=762nt_BP=342nt GTTTTGTTTTGACCCTCGCATTGTGCAGAATTAAAGTGCAGTAAAATGTCCACTAGGACCCCATTGCCAACGGTGAATGAACGAGACACT GAAAACCACACGTCACATGGAGATGGGCGTCAAGAAGTTACCTCTCGTACCAGCCGCTCAGGAGCTCGGTGTAGAAACTCTATAGCCTCC TGTGCAGATGAACAACCTCACATCGGAAACTACAGACTGTTGAAAACAATCGGCAAGGGGAATTTTGCAAAAGTAAAATTGGCAAGACAT ATCCTTACAGGCAGAGAGGTTGCAATAAAAATAATTGACAAAACTCAGTTGAATCCAACAAGTCTACAAAAGGGGAAAATGGCTTTGTTG GCCTCAAATAGCTGCTTTATTAGATGCAATGAAGCAGGGGACATAGAAGCAAAAAGTAAAACAGCAGGAGAAGAAGAAATGATCAAGATT AGATCCTGTGCTGAAAGAGAAACCAAGAAAAAAGATGACATTCCAGAAGAAGACAAAGGAAATGTAAAACAATGTGAAATCAATTATGTA AAGAAATTTCAGAGCTTCCAAGACCACAAACTTAAAATAAGTAAAGAAGACAGTAAAATTCTTAAAAAGGCTCGGAAAGATGGATTTTTG CATGAGACGCTTCTGGACAGGAGAGCCAAATTGAAAGCCGACAGATACTGCAAGTGACTGGGATTTTTGTTTCTGCCTTATCTTTCTGTG >51767_51767_7_MARK3-FRG1_MARK3_chr14_103894777_ENST00000553942_FRG1_chr4_190878553_ENST00000226798_length(amino acids)=213AA_BP=99 MSTRTPLPTVNERDTENHTSHGDGRQEVTSRTSRSGARCRNSIASCADEQPHIGNYRLLKTIGKGNFAKVKLARHILTGREVAIKIIDKT QLNPTSLQKGKMALLASNSCFIRCNEAGDIEAKSKTAGEEEMIKIRSCAERETKKKDDIPEEDKGNVKQCEINYVKKFQSFQDHKLKISK -------------------------------------------------------------- |
Top |
Fusion Gene PPI Analysis for MARK3-FRG1 |
 Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in Go to ChiPPI (Chimeric Protein-Protein interactions) to see the chimeric PPI interaction in |
 Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) Protein-protein interactors with each fusion partner protein in wild-type (BIOGRID-3.4.160) |
| Hgene | Hgene's interactors | Tgene | Tgene's interactors |
 - Retained PPIs in in-frame fusion. - Retained PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Still interaction with |
 - Lost PPIs in in-frame fusion. - Lost PPIs in in-frame fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
 - Retained PPIs, but lost function due to frame-shift fusion. - Retained PPIs, but lost function due to frame-shift fusion. |
| Partner | Gene | Hbp | Tbp | ENST | Strand | BPexon | TotalExon | Protein feature loci | *BPloci | TotalLen | Interaction lost with |
Top |
Related Drugs for MARK3-FRG1 |
 Drugs targeting genes involved in this fusion gene. Drugs targeting genes involved in this fusion gene. (DrugBank Version 5.1.8 2021-05-08) |
| Partner | Gene | UniProtAcc | DrugBank ID | Drug name | Drug activity | Drug type | Drug status |
Top |
Related Diseases for MARK3-FRG1 |
 Diseases associated with fusion partners. Diseases associated with fusion partners. (DisGeNet 4.0) |
| Partner | Gene | Disease ID | Disease name | # pubmeds | Source |
